Abstract
Pyruvate (PYR) dehydrogenase complex (PDC) is an enzymatic system that plays a crucial role in cellular metabolism as it controls the entry of carbon into the Krebs cycle. From a structural point of view, PDC is formed by three different subunits (E1, E2 and E3) capable of catalyzing the three reaction steps necessary for the full conversion of pyruvate to acetyl-CoA. Recent investigations pointed out the crucial role of this enzyme in the replication and survival of specific cancer cell lines, renewing the interest of the scientific community. Here, we report the results of our molecular dynamics studies on the mechanism by which posttranslational modifications, in particular the phosphorylation of three serine residues (Ser-264-α, Ser-271-α, and Ser-203-α), influence the enzymatic function of the protein. Our results support the hypothesis that the phosphorylation of Ser-264-α and Ser-271-α leads to (1) a perturbation of the catalytic site structure and dynamics and, especially in the case of Ser-264-α, to (2) a reduction in the affinity of E1 for the substrate. Additionally, an analysis of the channels connecting the external environment with the catalytic site indicates that the inhibitory effect should not be due to the occlusion of the access/egress pathways to/from the active site.
Similar content being viewed by others
Introduction
Pyruvate (PYR) dehydrogenase complex (PDC) is a multisubunit molecular machine responsible for the conversion of PYR into acetyl-CoA through a process known as pyruvate decarboxylation1.
From a biochemical point of view, PDC is the gatekeeper controlling the entry of carbon into the TCA cycle (also known as Krebs cycle) from two main sources: carbohydrates and gluconeogenic amino acids.
Given its central role in cellular metabolism2,3, PDC has been considered a promising target for the development of antibacterial4,5 and anticancer drugs6,7. In fact, recent experimental investigations pointed out that an increase in PDC activity can sustain the progression of some cancer types (for example prostate cancer and glioblastoma)8 as well as the survival of cancer stem cells7,9.
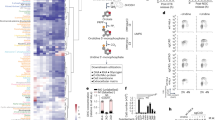
From the structural point of view, PDC is composed of multiple copies of three different subunits, usually called E1 (pyruvate dehydrogenase), E2 (dihydrolipoyl trans-acetylase) and E3 (dihydrolipoyl dehydrogenase)1,10. While the first two subunits are involved in the synthesis of acetyl-coenzyme A (Ac-CoA), the third subunit (E3) is a FAD/NAD+ dependent system performing redox recycling (Fig. 1).
(I) Schematic view of the three PDC subunits with the corresponding reaction schemes. E1 (pink, pyruvate dehydrogenase) catalyzes pyruvate decarboxylation. E2 (blue, dihydrolipoyl trans-acetylase) catalyzes the acetyl transfer to CoA. E3 (white, dihydrolipoyl dehydrogenase) catalyzes the regeneration of the lipoamide moiety. (II) General PDC catalyzed reaction.
Structural studies1 have shown that the E1 PDC subunit is formed by two different chains (α and β) and is characterized by the presence of a magnesium ion coordinated by three amino acid residues (Asp167-α, Αsn196-α, Τyr198-α) and a thiamine diphosphate (ThDP) molecule acting as a cofactor. ThDP is directly involved in the enzymatic reaction where after activation by Glu59 and the consequent formation of a dipolar molecule bearing a formally negatively charged atom (Fig. 1)1,11 called ylide, it carries out a nucleophile attack on the PYR molecule to form a covalent ThDP-acetyl adduct (step A, Fig. 1). Then, the reaction proceeds with the involvement of the E2 subunit. This second enzymatic subunit bears a long amino acidic arm ending with a lipoamide group, which is able to swing between the E1 and E3 active sites. In particular, the acetyl group from the ThDP-acetyl in the E1 subunit is first transferred to the lipoamide. Then, the swinging arm moves out of the E1 subunit and enters into the E2 catalytic site, where the acetyl group is transferred to the CoA molecule to form the Ac-CoA. Finally, the arm moves to the E3 catalytic site, where the initial oxidation state of the lipoamide group is restored.
Given its importance in cellular metabolism, the PDC complex has been at the center of numerous structural and functional studies1. However, several questions regarding (1) the PDC global stoichiometry, (2) the influence of posttranslational modifications on the function of PDC and (3) a full characterization of the enzymatic reaction at an atomistic level are still unanswered.
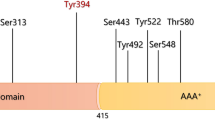
In regards to question (2), it has been extensively reported that the activity of PDC is regulated by the phosphorylation of three serine residues1 and that the level of phosphorylation is controlled by four pyruvate dehydrogenase kinases (PDKs1-4) and two pyruvate dehydrogenase phosphatases (PDP1-2).
In particular, experimental data clearly indicate that the phosphorylation of specific serine residues (Ser-264-α, Ser-271-α, and Ser-203-α in the human sequence, subsequently indicated as Ser-264-α-P, Ser-271-α-P and Ser-203-α-P when phosphorylated) results in different degrees of enzyme inhibition, with the strongest effect observed with Ser-264-α-P1,6.
In vitro studies demonstrated12 that PDK1 is the only kinase that can phosphorylate all the three sites, even if with different preferences and kinetics, while PDK2, PDK3 and PDK4 only act on Ser-264-α and Ser-271-α. In contrast, a similar action on all the three sites was observed for the two phosphatases13.
Furthermore, other investigations have shown that these posttranslational modifications are independent and not sequential13,14 and that the phosphorylation of Ser-264-α drastically reduces the enzymatic activity, while the phosphorylation of Ser-271-α or Ser-203-α have less clear inhibitory effects13,15,16.
Finally, experimental investigations6,17,18 indicated that the levels of PDC phosphorylation are significant for several widely diffused diseases such as cancer6,17,18 diabetes and neuro-metabolic disorders19,20. Understanding, at an atomistic level of detail, how posttranslational modifications alter enzymatic activity could open the way to the development of innovative therapeutic tools based on PDC inhibition/activation.
Two different hypotheses to explain the PDC inhibition induced by phosphorylation have been proposed in the scientific literature21,22. Kato et al.21 suggested that a perturbation of protein dynamics, in particular of two phosphorylation loops (loop A: from 259-α to 282-α, loop B: from 195-α to 205-α), as origin of the inhibition while Seifert et al.22 hypothesized that Ser-264-α-P could hinder the substrates access to the active site22.
Over the years, molecular simulations have become a valuable tool to answer these types of questions23,24,25,26,27,28,29,30,31,32,33,34 as they provide a detailed atomistic picture of the biomolecular dynamics. In particular, they have been recently used to obtain a low resolution model of the entire PCD structure35 and to investigate other aspects of this enzymatic complex36,37.
Thus, we carried out an MD simulation study in which the PDC-E1 system in its unphosphorylated and phosphorylated forms (Ser-264-α-P, Ser-271-α-P and Ser-203-α-P) was simulated for 1 μsec (MD). Finally, the trajectories were analyzed to investigate the effects of the posttranslational modifications on the protein structure, dynamics and on its ability to bind the substrate.
Results and Discussion
Analysis of molecular dynamics simulations
The molecular dynamics trajectories for the different systems were compared by considering different quantities such as RMSD, RMSF, Lindemann coefficient and pyruvate binding energy, as well as, the number, position, length and diameter of the tunnels leading to the active site.
A detailed analysis of our simulations (Figs 2–4 and Table 1) clearly shows that Ser-264-α-P and Ser-271-α-P have a different influence on the protein structure and dynamics when compared to Ser-203-α-P. In particular, RMSD and RMSF analyses of the protein backbone carried out on the entire 1 μs MD simulations showed that Ser-203-α-P stabilizes the protein in a conformation that is more similar to that observed in X-ray experiments (PDB ID: 1NI4) and reduces atomic fluctuations (lower RMSF). In contrast, higher RMSD and RMSF values were observed for Ser-264-α-P and Ser-271-α-P.
Differences between the RMSF values measured for the three phosphorylated species, Ser-264-α-P (light blue), Ser-271-α-P (orange), Ser-203-α-P (violet), and the WT. Positive values correspond to an increment of RMSF values with respect to WT, while negative values correspond to a reduction. The data for the α (A) and β (B) subunits are reported in separate diagrams for clarity.
(A) Backbone RMSD calculated with respect to the starting coordinates of the WT protein over the entire MD simulations. WT (violet) and the three phosphorylated species, Ser-264-α-P (light blue), Ser-271-α-P (orange), and Ser-203-α-P (green)). (B) Binding site RMSD (His63-α, Met200-α, His263-α, Phe61-α, Tyr89-α, Met82-β) calculated for the entire residues (backbone + sidechains) over the entire simulation time. WT (violet) and the three phosphorylated species Ser-264-α-P (light blue), Ser-271-α-P (orange), and Ser-203-α-P (green).
Next, we aimed to quantify the observed differences in protein dynamics by computing the Lindemann coefficient38, which gives an estimation of the solid-liquid behavior of a protein. The values of the coefficients (Table 1) computed for the entire protein as well as for different portions, confirm a stiffening of the entire molecule induced by Ser-203-α-P. Moreover, the values indicate that phosphorylation of Ser-264-α and Ser-271-α increases the flexibility of the region from Asp-92-β to Pro-110β, while that on Ser-203-α rigidifies the same region.
A different trend was observed for the two phosphorylation loops (for loop definitions, see introduction). In fact, while Ser-203-α-P stiffens both loops, Ser-264-α-P and Ser-271-α-P significantly increase the flexibility of loop B but not that of loop A. Concerning this point, it should be noted that Ser-264-α and Ser-271-α are inside loop A, while Ser-203-α is placed in loop B.
To better understand the origins of these differences, we visually inspected the trajectories of all the simulations.
This analysis showed that during the Ser-203-α-P simulation, this residue gets surrounded by three arginine residues (Arg273-α, Arg206-α, Αrg197-α) forming a cationic-hole (Fig. SI1). Interestingly, Arg273-α is located in the terminal region of loop A; thus, the interaction with this residue could reasonably be at the origin of the greater rigidity of this loop during the MD simulations with the phosphorylated Ser-203-α. Α similar, but less stable, cationic hole is formed around Ser-271-α-P that interacts with Lys39−β, Arg273-α and Arg275-α (Fig. SI2). In contrast to Ser-271-α-P and Ser-203-α-P, similar interactions were not visible for Ser-264-α.
Several experimental and theoretical investigations21,39,40 have pointed out the role of specific amino acids residues in the catalytic activity of E1-PCD. We, therefore, analyzed our simulations focusing on the consequences of phosphorylation on the catalytic site structure and dynamics.
The RMSD values (Fig. 3B), computed taking into account all atoms of the amino acids residues forming the catalytic site (His63-α, Met200-α, His263-α, Phe61-α, Tyr89-α, Met82-β), showed that Ser-264-α-P and Ser-271-α-P perturb the starting structure more significantly than Ser-203-α-P.
In summary, the analysis of our simulations shows that phosphorylation of the three investigated serine residues have marked structural and dynamics effects on the PDC-E1-α subunit. In particular, both Ser-264-α-P and Ser-271-α-P increase the flexibility of loop B, the loop that directly coordinates the magnesium ion interacting with the ThDP cofactor. These findings are in line with the conclusions of Kato et al.21 that also pointed out an increased disorder in the two phosphorylation loops (loop A and B).
At the same time, the stiffening of the protein structure observed in the presence of Ser-203-α-P, suggests that this posttranslational modification should not inhibit PDC or, alternatively, that the inhibition could have a different explanation.
Phosphorylation of single residues can influence protein properties such as hydrophobicity, lipophilicity41 and conformational entropy42. We, therefore, used specific computational tools to analyze our simulations with respect to these properties.
The entropic contribution (conformational entropy) to the total free energy (−ΤΔS) was estimated to be −3.4 ± 0.1, −5.5 ± 0.1 and 26.2 ± 0.1 kcal/mol for the systems bearing Ser-264-α-P, Ser-271-α-P and Ser-203-α-P, respectively. These values confirm a stabilizing effect of Ser-203-α-P and a slight destabilizing action of Ser-264-α-P and Ser-271-α-P.
Following the suggestion of Polyansky and Zagrovic41, we also investigated the effects of phosphorylation on the protein hydrophobicity/lipophilicity by computing the molecular hydrophobicity potential (MHP) on multiple snapshots extracted from our MD simulations.
These calculations resulted in ΔMHP values of −994, −835 and 258 log P units for the systems with Ser-264-α-P, Ser-271-α-P and Ser-203-α-P, respectively. These results indicate a decrease in the hydrophobicity of the protein in the first two cases and an opposite effect in the last one.
Influence of phosphorylation on pyruvate binding affinity
Our previous calculations, as well as experimental studies14, suggest that changes in the enzyme affinity for the substrate and/or the cofactors could be one of the consequences of the phosphorylation. Because of technical problems in the purification of phosphorylated proteins, experimental investigations have been carried out using glutamate and aspartate mutations of the serine amino acids to mimic the effects of phosphorylation. However, while these mutations effectively reproduce the negative net charge of the phosphate group, they less efficiently mimic other properties, in particular the amino acid size. Therefore, to better clarify this point, we estimated the affinity between PDC-E1 and PYR or ThDP in all the investigated systems by MM-GBSA (Table 2).
For PYR, our calculations indicate that Ser-264-α-P significantly reduces the enzyme affinity. In contrast, Ser-203-α-P increases the affinity for PYR, while Ser-271-α-P has no significant effect. Next, we performed a per-residue decomposition of the binding energy to better understand which residues facilitate or antagonize the binding of the substrate (Fig. 5).
(Left column), the results of per-residue Energy decomposition carried out for the WT (A) and the systems bearing a Ser-264-α-P, Ser-271-α-P, or Ser-203-α-P. For sake of clarity, only contributions higher the 0.2 kcal/mol (absolute value) were reported. (Right column) representative structure of the catalytic site in the four investigated systems. The residues with the higher contributions to the binding and the phosphorylated residues are depicted as sticks; the nonpolar hydrogens are not shown for the sake of clarity.
From the analysis of these results, we noticed that the network of residues with the strongest contribution to the binding energy is generally conserved among all the investigated systems. However, in the case of the Ser-264-α-P mutant, we observed a large (~2 kcal/mol) positive, i.e. repulsive, contribution from Ser-264-α-P and a reduction (~1.5 kcal/mol with respect to the unphosphorylated protein) in the contribution to the binding energy given by His63-α and His263-α. In fact, in the WT protein, these two residues are close to each other on the interior of the active site and are able to form stable H-bond interactions with PYR. In the system with Ser-264-α-P, the phosphate group interacts with His263-α, inducing its reorientation to the exterior of the active site and resulting in the breaking of the H-bonds with the substrate.
In the case of Ser-203-α-P, the increase in the predicted affinity, with respect to the unphosphorylated protein, is due to a larger contribution of Phe61-α and an additional interaction with Gln128-β.
The calculations carried out to estimate the protein affinity for ThDP did not show any negative effects of the phosphorylation of the three serine residues. In contrast, the computed affinities for the proteins bearing Ser-264-α-P and Ser-203-α-P were higher than for the unphosphorylated ones.
In summary, MM-GBSA calculations indicated that Ser-264-α-P has a repulsive effect on PYR reducing its affinity for the catalytic site, while a similar effect was not observed for Ser-271-α-P and Ser-203-α-P. Moreover, the results indicated that serine phosphorylations do not affect the affinity for ThDP.
Analysis of the protein channels by Caver3.0
Seifert et al.22, studying the structure of the a pseudophosphorylated variant in which Ser-264-α was mutated to glutamate, concluded that the Ser-264-α-P hinders the substrate access to the active site. However, their conclusions were drawn by structural investigation on a system bearing a phosphomimetic mutation and using an experimental technique, X-ray crystallography, that provides only limited information about the dynamics of the system. Furthermore, the study22 focused on only one of the three possible phosphorylations.
Therefore, aiming to determine if this hypothesis could be confirmed by simulations of the different systems bearing an explicit phosphate group, we analyzed the 1 μs MD simulations outputs with Caver3.043, a software specifically designed to identify tunnels in protein and previously used in similar investigations25,44,45.
In the case of the WT simulation, the tunnel with the best score identified in the 65% of the analyzed snapshots and corresponding to that described by Tittmann and coworkers22 is topologically located near the protein region occupied by Ser-264-α. Moreover, a second channel (present in the 20% of the snapshots, Fig. 6 and Table 3) is detected on the other site of the protein. For both tunnels, the average bottleneck diameter is larger than 3.0 Å, which is enough to allow the transit of a PYR molecule.
Graphical representation of the Caver3.0 results. The first and the second tunnels are represented in blue and green, respectively. The three phosphorylated serine residues are always depicted in magenta as spatial references. In the cases of Ser-264-α-P and Ser-271-α-P, in which the tunnels have a common pathway, only one color is used to depict the common part.
In addition, in the case of the Ser-264-α-P system, the best scoring channel still corresponds to that which was proposed from the observation of the X-ray structure and passes near Ser-264-α-P22. This channel is characterized by an average bottleneck radius larger than that measured for the same channel in the WT and it is present in the 74% of the analyzed snapshots. A second channel, starting from a slightly different position but converging in tunnel 1, was also detected in the 75% of the snapshots.
A similar behavior was observed for Ser-271-α-P. In fact, in this case, the first two channels (detected in the 69% and 48% of the snapshots) start from different positions but in their terminal portion, converge on the same pathway passing near the regions of the protein occupied by Ser-264-α and Ser-271-α. It is also worth noting that in these cases, the channel diameters are larger than those detected for the main channels of the WT system.
Finally, in the case of Ser-203-α-P, the two best scoring channels detected by Caver were essentially identical to those observed in the WT, only with a reverse ranking.
Taken together, the results of our analysis strongly suggest that the channel passing near Ser-264-α and Ser-271-α, corresponding to what is already described in the literature22, should represent the main pathway connecting the protein surface with the active site. In all the phosphorylated systems, this channel was demonstrated to be as viable as in the WT system. We can therefore conclude that the origin of the inhibitory effect of phosphorylations15,16,21 is not due to the closure of the access route to the active site.
Summary and Conclusions
In this work, we focused on the effects of specific serine phosphorylations on the enzymatic activity of the PDC E1 subunit.
Our calculations highlighted an increase in the protein flexibility induced by the phosphorylation of Ser-264-α and Ser-271-α at both the global (entire protein) and local (catalytic site) levels, while the phosphorylation of Ser-203-α was associated with an increase in protein rigidity. In particular, in the Ser-264-α-P and Ser-271-α-P systems, we observed a significant increase in the flexibility of the phosphorylation loop B, which is directly involved in the binding of both the ThDP cofactor and of the ThDP coordinated Mg2+ ion.
Binding energy calculations helped to shed light on the consequences of the increased flexibility induced by the presence Ser-264-α-P and Ser-271-α-P. In fact, while the affinity for the ThDP cofactor is unaltered, Ser-264-α-P reduces the affinity of PYR for the catalytic site. A similar effect was not observed for Ser-271-α-P, while the affinity in the Ser-203-α-P system was significantly increased by the posttranslational modification.
Finally, Caver calculations showed that the presence of the phosphate groups in the three considered serine residues does not induce a closure of the access/release pathway to/from the catalytic site.
Taken together, our findings suggest that the strong inhibitory effect observed for Ser-264-α-P is due to both a perturbation of the active site structure and dynamics and to a repulsive interaction between the modified residue and PYR that leads to a reduced protein-ligand affinity.
In the cases of Ser-271-α and Ser-203-α, the picture emerging from our studies is less clear. In fact, the phosphorylation of Ser-271-α, seems to show only a small increase in protein flexibility that could however be the cause of its experimentally observed milder inhibitory effect13,15,16.
Regarding Ser-203-α-P, the effects of phosphorylation are completely different with respect to Ser-264-α-P and Ser-271-α-P. In this case, considering the debate about the strength of its influence on the PDC-E1 catalytic activity, we can hypothesize that this modification does not lead to enzyme inhibition or the inhibition is obtained by a different mechanism.
In this work, because of the lack of an experimentally solved atomistic structure of E1 in complex with the other subunits, as well as, the too large computational resources required to perform long enough atomistic simulations of the entire PDC, we chose to focus our investigation on a ‘minimal model’ able to recapitulate the most important features of the complex. Consequently, once more complete structural information and more potent computational resources become available, additional investigations should be carried out to gain a deeper understanding the effects of phosphorylation in the context of the entire system.
Finally, from a pharmacological point of view, the results of this work suggest that an allosteric modulation of the PDC-E1 is possible, and it could be a viable strategy to design new molecular tools aimed to modulate the enzymatic activity.
In particular, small organic molecules or peptides could be designed to mimic the effects of Ser-264-α-P. Alternatively, in the case of PDHA1 deficiency, the inhibitory effects of the two posttranslational modifications could be selectively blocked, hindering the interaction of PDK(s) and the PDC-E1 subunit near Ser-264-α for which the most potent inhibition was reported15,16,21.
Computational Methods
MD simulations
Molecular dynamics simulations were carried out starting from the structure of the E1-PDC human protein (PDB ID: 1NI4)46. The coordinates of PYR, not present in the 1NI4 crystal structure, were obtained by structural alignment with the structure of the E1-PDC protein from Geobacillus stearothermophilus (PDB ID: 3DV0)11. Furthermore, selenomethionine residues were mutated to methionine.
All molecular complexes were fully solvated in a TIP3P water box47 with a minimum distance of 10 Å from the protein surface. The charge neutrality of the systems was ensured by adding a proper number of counterions (Na+ ions).
The systems were described by Amber Force Field (ff14SB48 for the protein, phosaa1049,50 for the phosphorylated residues, GAFF51 for pyruvate and cofactors; the parameters for Na+ and Mg2+ were taken from references52 and53, respectively). Atomic charges for ThDP and PYR were computed with the AM1-bcc approach implemented in the ANTECHAMBER module of Amber16. Finally, MD simulations reported in this study were carried out with the GPU accelerated PMEMD code available in Amber1654.
To avoid distortions of the complexes, the systems were first minimized for 10.000 steps or until an energy gradient of 0.2 kcal mol−1Å−1 was reached. First, only water and counterions were left free to move, while the backbone atoms were restrained with a harmonic potential of 20 kcal mol−1Å−1. Then, the restraints were removed and the minimization procedure repeated. Finally, the temperature of the systems was increased to 298.5 K in 400 ps, and the pressure increased to 1 atm. During the MD simulations, the temperature and pressure were kept constant using the Langevin thermostat55 and Monte Carlo barostat56, respectively. Furthermore, the distance between the activated ThDP carbon and C2@PYR was restricted in the range between 3.3 Å and 3.7 Å by applying a flat-bottom harmonic (k = 80 kcal/mol/Å2) distance restraint.
Analysis of the MD simulations
MD trajectories were analyzed using specific tools implemented in the programs VMD57, cpptraj58 and ALMOST59. The Lindemann coefficient38 was computed considering backbone atoms by the PCA suite program (http://mmb.pcb.ub.es/software/pcasuite/pcasuite.html). The Lindemann coefficient was first proposed to determine the solid or liquid character of an infinite system60,61 and subsequently applied to proteins by Karplus and coworkers38.
Protein tunnel analysis was done with the Caver 3.0 software43 on 1250 equally spaced snapshots (1 every 800 ps) extracted from the 1 μs MD simulations. All the water molecules and the pyruvate were removed from the structures. Furthermore, the origin of the tunnels was set to be the geometrical center of the following four atoms: (1) the nucleophile carbon of ThDP (Cn in Fig. 1), (2) Cα@Phe61α, (3) Cα@Tyr89α and (4) Cα@Phe86β (residue numbering refers to the human E1-PDC structure, PDB ID: 1NI446). Finally, considering the size of PYR, the search of tunnels was performed using a probe with a radius of 1.5 Å.
In Caver analysis, each tunnel is described as an ensemble of spheres whose radius can fit the tunnel diameter. All identified tunnels were clustered by hierarchical average link using the pairwise distances between the centers of the spheres. The predicted tunnels were ranked according to the Carver score, designed to give priority to wide paths with a reasonable length43.
The affinity between the proteins and the substrate (PYR) was estimated using the MM-GBSA approach62.
MM-GBSA calculations were run on 2500 frames extracted (one every 400 ps) from the entire trajectories according to the following procedure: (1) water molecules and counterions were removed, (2) the same parameters used during MD simulations were assigned to both protein and ligand, and (3) the polar contribution to solvation energy was computed by the Onufriev, Bashford and Case model setting the dielectric constant to 1 for the solute and 80 for the solvent63. Per-residue decomposition (PRD) was used to estimate the contribution of single protein residues to the binding energy. All the calculations were run using the MMPBSA.py62 module available in AmberTools16.
The maximum information spanning tree (MIST) approach of Tidor and coworkers64,65 is a well-established method to estimate entropy from MD simulations66,67,68. The influence of the protein phosphorylations on conformational entropy were assessed using the implementation of the method in the recently released PDB2ENTROPY program67. To address this aim, 2500 snapshots were extracted from the MD simulations and processed using the following command line options ‘-mi -nt 16 -ne 40’.
The entropic contribution to the free energy caused by phosphorylation was defined as:
How phosphorylation influences the hydrophobicity/hydrophilicity of PDC-E1 was assessed by computing the molecular hydrophobicity potential (MHP) using the PLATINUM webserver69. The difference between the sum of the MHP values over the protein solvent-accessible surface calculated for the phosphorylated and unphosphorylated protein (ΔMHP) was considered as measure of the effect of phosphorylation.
References
Patel, M. S., Nemeria, N. S., Furey, W. & Jordan, F. The pyruvate dehydrogenase complexes: structure-based function and regulation. J Biol Chem 289, 16615–16623 (2014).
Solaini, G., Sgarbi, G. & Baracca, A. Oxidative phosphorylation in cancer cells. Biochimica et Biophysica Acta (BBA) - Bioenergetics 1807, 534–542 (2011).
Ahlqvist, K. J., Suomalainen, A. & Hämäläinen, R. H. Stem cells, mitochondria and aging. Biochimica et Biophysica Acta (BBA) - Bioenergetics 1847, 1380–1386 (2015).
He, H., Xia, H., Xia, Q., Ren, Y. & He, H. Design and optimization of N-acylhydrazone pyrimidine derivatives as E. coli PDHc E1 inhibitors: Structure-activity relationship analysis, biological evaluation and molecular docking study. Bioorg. Med. Chem. 25, 5652- (2017).
Zhou, Y. et al. Design, Synthesis, and Potency of Pyruvate Dehydrogenase Complex E1 Inhibitors against Cyanobacteria. Biochemistry 56, 6491–6502 (2017).
Saunier, E., Benelli, C. & Bortoli, S. The pyruvate dehydrogenase complex in cancer: An old metabolic gatekeeper regulated by new pathways and pharmacological agents. Int. J. Cancer 138, 809–817 (2016).
Chen, J. et al. Compartmentalized activities of the pyruvate dehydrogenase complex sustain lipogenesis in prostate cancer. Nat. Genet. 50, 219–228 (2018).
Bunik, V. I. et al. Specific inhibition by synthetic analogs of pyruvate reveals that the pyruvate dehydrogenase reaction is essential for metabolism and viability of glioblastoma cells. Oncotarget 6, 40036–40052 (2015).
Deshmukh, A., Deshpande, K., Arfuso, F., Newsholme, P. & Dharmarajan, A. Cancer stem cell metabolism: a potential target for cancer therapy. Molecular Cancer 15, 69, x (2016).
Byron, O. & Lindsay, J. G. The Pyruvate Dehydrogenase Complex and Related Assemblies in Health and Disease. Subcell. Biochem. 83, 523–550 (2017).
Pei, X. Y., Titman, C. M., Frank, R. A. W., Leeper, F. J. & Luisi, B. F. Snapshots of Catalysis in the E1 Subunit of the Pyruvate Dehydrogenase Multienzyme Complex. 16, 1860–1872, (2008).
Korotchkina, L. G. & Patel, M. S. Site specificity of four pyruvate dehydrogenase kinase isoenzymes toward the three phosphorylation sites of human pyruvate dehydrogenase. J. Biol. Chem. 276, 37223–37229 (2001).
Kolobova, E., Tuganova, A., Boulatnikov, I. & Popov, K. M. Regulation of pyruvate dehydrogenase activity through phosphorylation at multiple sites. Biochem. J. 358, 69–77 (2001).
Korotchkina, L. G. & Patel, M. S. Probing the mechanism of inactivation of human pyruvate dehydrogenase by phosphorylation of three sites. J. Biol. Chem. 276, 5731–5738 (2001).
Sale, G. J. & Randle, P. J. Analysis of site occupancies in [32P]phosphorylated pyruvate dehydrogenase complexes by aspartyl-prolyl cleavage of tryptic phosphopeptides. Eur. J. Biochem. 120, 535–540 (1981).
Sale, G. J. & Randle, P. J. Role of individual phosphorylation sites in inactivation of pyruvate dehydrogenase complex in rat heart mitochondria. Biochem. J 203, 99–108 (1982).
Stacpoole, P. W. The pyruvate dehydrogenase complex as a therapeutic target for age-related diseases. Aging cell 11, 371–377 (2012).
Stacpoole, P. W. Therapeutic Targeting of the Pyruvate Dehydrogenase Complex/Pyruvate Dehydrogenase Kinase (PDC/PDK) Axis in Cancer. J. Natl. Cancer. Inst. 109, (2017).
Thibodeau, A., Geng, X., Previch, L. & Ding, Y. Pyruvate dehydrogenase complex in cerebral ischemia-reperfusion injury. Brain Circulation 2, 61–66 (2016).
Lee, I.-K. The Role of Pyruvate Dehydrogenase Kinase in Diabetes and Obesity. Diabetes Metab. Journal 38, 181–186 (2014).
Kato, M. et al. Structural basis for inactivation of the human pyruvate dehydrogenase complex by phosphorylation: role of disordered phosphorylation loops. Structure 16, 1849–1859 (2008).
Seifert, F. et al. Phosphorylation of serine 264 impedes active site accessibility in the E1 component of the human pyruvate dehydrogenase multienzyme complex. Biochemistry 46, 6277–6287 (2007).
Magistrato, A., Sgrignani, J., Krause, R. & Cavalli, A. Single or Multiple Access Channels to the CYP450s Active Site? An Answer from Free Energy Simulations of the Human Aromatase Enzyme. J. Phys. Chem. Lett. 8, 2036–2042 (2017).
Culka, M., Gisdon, F. J. & Ullmann, G. M. In Advances in Protein Chemistry and Structural Biology Vol. 109 (ed Tatyana Karabencheva-Christova) 77–112 (Academic Press, 2017).
Sgrignani, J. & Magistrato, A. Influence of the membrane lipophilic environment on the structure and on the substrate access/egress routes of the human aromatase enzyme. A computational study. J. Chem. Inf. Model. 52, 1595–1606 (2012).
Chiappori, F., Mattiazzi, L., Milanesi, L. & Merelli, I. A novel molecular dynamics approach to evaluate the effect of phosphorylation on multimeric protein interface: the alphaB-Crystallin case study. BMC Bioinformatics 17(Suppl 4), 57 (2016).
Zhu, J., Lv, Y., Han, X., Xu, D. & Han, W. Understanding the differences of the ligand binding/unbinding pathways between phosphorylated and non-phosphorylated ARH1 using molecular dynamics simulations. Sci. Rep. 7, 12439 (2017).
Montenegro, M., Masgrau, L., Gonzalez-Lafont, A., Lluch, J. M. & Garcia-Viloca, M. Influence of the enzyme phosphorylation state and the substrate on PKA enzyme dynamics. Biophys. Chem. 161, 17–28 (2012).
Missimer, J. H., Steinmetz, M. O., van Gunsteren, W. F. & Dolenc, J. Influence of63Ser phosphorylation and dephosphorylation on the structure of the stathmin helical nucleation sequence: a molecular dynamics study. Biochemistry 51, 8455–8463 (2012).
Lehman, W. et al. Phosphorylation of Ser283 Enhances the Stiffness of the Tropomyosin Head-to-Tail Overlap Domain. Arch. Biochem. Biophys. 571, 10–15 (2015).
Bueren-Calabuig, J. A. & Michel, J. Impact of Ser17 Phosphorylation on the Conformational Dynamics of the Oncoprotein MDM2. Biochemistry 55, 2500–2509 (2016).
Gone, S., Alfonso-Prieto, M., Paudyal, S. & Nicholson, A. W. Mechanism of Ribonuclease III Catalytic Regulation by Serine Phosphorylation. Sci. Rep. 6, 25448 (2016).
Groban, E. S., Narayanan, A. & Jacobson, M. P. Conformational Changes in Protein Loops and Helices Induced by Post-Translational Phosphorylation. PLOS Comput. Biol. 2, e32 (2006).
Espinoza-Fonseca, L. M., Kast, D. & Thomas, D. D. Thermodynamic and Structural Basis of Phosphorylation-Induced Disorder-to-Order Transition in the Regulatory Light Chain of Smooth Muscle Myosin. J. Am. Chem. Soc. 130, 12208–12209, https://doi.org/10.1021/ja803143g (2008).
Hezaveh, S., Zeng, A. P. & Jandt, U. Full Enzyme Complex Simulation: Interactions in Human Pyruvate Dehydrogenase Complex. J. Chem. Inf. Mod. 58, 362–369 (2018).
Hezaveh, S., Zeng, A.-P. & Jandt, U. Investigation of Core Structure and Stability of Human Pyruvate Dehydrogenase Complex: A Coarse-Grained Approach. ACS Omega 2, 1134–1145 (2017).
Hezaveh, S., Zeng, A. P. & Jandt, U. Human Pyruvate Dehydrogenase Complex E2 and E3BP Core Subunits: New Models and Insights from Molecular Dynamics Simulations. J. Phys. Chem. B 120, 4399–4409 (2016).
Zhou, Y., Vitkup, D. & Karplus, M. Native proteins are surface-molten solids: application of the lindemann criterion for the solid versus liquid state11Edited by A. R. Fersht. J. Mol. Biol. 285, (1371–1375 (1999).
Sheng, X. & Liu, Y. Theoretical study of the catalytic mechanism of E1 subunit of pyruvate dehydrogenase multienzyme complex from Bacillus stearothermophilus. Biochemistry 52, 8079–8093 (2013).
Arjunan, P. et al. Structure of the pyruvate dehydrogenase multienzyme complex E1 component from Escherichia coli at 1.85 A resolution. Biochemistry 41, 5213–5221 (2002).
Polyansky, A. A. & Zagrovic, B. Protein Electrostatic Properties Predefining the Level of Surface Hydrophobicity Change upon Phosphorylation. J. Phys. Chem. Lett. 3, 973–976 (2012).
Nishi, H., Shaytan, A. & Panchenko, A. R. Physicochemical mechanisms of protein regulation by phosphorylation. Frontiers in Genetics 5, (2014).
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLoS Comput Biol 8, e1002708 (2012).
Musiani, F. et al. Protein Tunnels: The Case of Urease Accessory Proteins. J. Chem. Theory Comput. 13, 2322–2331 (2017).
Ebert, M. C. C. J. C., Dürr, S. L., Houle, A. A., Lamoureux, G. & Pelletier, J. N. Evolution of P450 Monooxygenases toward Formation of Transient Channels and Exclusion of Nonproductive Gases. ACS Catalysis 6, 7426–7437 (2016).
Ciszak, E. M., Korotchkina, L. G., Dominiak, P. M., Sidhu, S. & Patel, M. S. Structural basis for flip-flop action of thiamin pyrophosphate-dependent enzymes revealed by human pyruvate dehydrogenase. J. Biol. Chem. 278, 21240–21246 (2003).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, L. M. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Maier, J. A. et al. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Homeyer, N., Horn, A. H., Lanig, H. & Sticht, H. AMBER force-field parameters for phosphorylated amino acids in different protonation states: phosphoserine, phosphothreonine, phosphotyrosine, and phosphohistidine. J. Mol. Mod. 12, 281–289 (2006).
Steinbrecher, T., Latzer, J. & Case, D. A. Revised AMBER parameters for bioorganic phosphates. J. Chem. Theory Comput. 8, 4405–4412 (2012).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. J. Comp. Chem. 25, 1157–1174 (2004).
Joung, I. S. & Cheatham, T. E. Determination of alkali and halide monovalent ion parameters for use in explicitly solvated biomolecular simulations. J. Phys. Chem. B 112, 9020–9041 (2008).
Li, P., Song, L. F. & Merz, K. M. Jr. Parameterization of highly charged metal ions using the 12-6-4 LJ-type nonbonded model in explicit water. J. Phys. Chem. B 119, 883–895 (2015).
Le Grand, S., Götz, A. W. & Walker, R. C. SPFP: Speed without compromise—A mixed precision model for GPU accelerated molecular dynamics simulations. Comput. Phys. Commun. 184, 374–380 (2013).
Loncharich, R. J., Brooks, B. R. & Pastor, R. W. Langevin dynamics of peptides: the frictional dependence of isomerization rates of N-acetylalanyl-N’-methylamide. Biopolymers 32, 523–535 (1992).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the Cartesian equations of motion of a system with constraints; molecular dynamics of n-alkanes. J. Comp. Phys. 23, 327–341 (1977).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Roe, D. R. & Cheatham, T. E. PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Fu, B. et al. ALMOST: an all atom molecular simulation toolkit for protein structure determination. J. Comput. Chem. 35, 1101–1105 (2014).
Bilgram, J. H. Dynamics at the solid-liquid transition: Experiments at the freezing point. Phys. Rep. 153, 1–89 (1987).
Löwen, H. Melting, freezing and colloidal suspensions. Phys. Rep. 237, 249–324 (1994).
Miller, B. R. et al. MMPBSA.py: An Efficient Program for End-State Free Energy Calculations. J. Chem. Theory Comput. 8, 3314–3321 (2012).
Onufriev, A., Bashford, D. & Case, D. A. Exploring protein native states and large-scale conformational changes with a modified generalized born model. Proteins 55, 383–394 (2004).
King, B. M. & Tidor, B. MIST: Maximum Information Spanning Trees for dimension reduction of biological data sets. Bioinformatics 25, 1165–1172, btp109 (2009).
King, B. M., Silver, N. W. & Tidor, B. Efficient Calculation of Molecular Configurational Entropies Using an Information Theoretic Approximation. J. Phys Chem. B 116, 2891–2904 (2012).
Fleck, M., Polyansky, A. A. & Zagrovic, B. PARENT: A Parallel Software Suite for the Calculation of Configurational Entropy in Biomolecular Systems. J. Chem. Theory Comput. 12, (2055–2065 (2016).
Fogolari, F., Maloku, O., Dongmo Foumthuim, C. J., Corazza, A. & Esposito, G. PDB2ENTROPY and PDB2TRENT: Conformational and Translational–Rotational Entropy from Molecular Ensembles. J. Chem. Inf. Mod. 58, 1319–1324 (2018).
Suárez, E., Díaz, N., Méndez, J. & Suárez, D. CENCALC: A computational tool for conformational entropy calculations from molecular simulations. J. Comp. Chem. 34, 2041–2054 (2013).
Pyrkov, T. V., Chugunov, A. O., Krylov, N. A., Nolde, D. E. & Efremov, R. G. PLATINUM: a web tool for analysis of hydrophobic/hydrophilic organization of biomolecular complexes. Bioinformatics 25, 1201–1202 (2009).
Acknowledgements
AC acknowledge the Swiss National Supercomputing Center (CSCS) for the availability of high-performance computing resources. This study was supported by grants from Krebsliga Schweiz (KLS-3839-02-2016-R) and the Swiss National Science Foundation (31003A-166472) to A.C. We thank Dr. Dmitry E. Nolde for help with PLATINUM calculations and Prof. F. Fologari for support in the use of the pdb2entropy program.
Author information
Authors and Affiliations
Contributions
J.S. and A.C. designed the research, J.S. performed the simulations, J.S. and A.C. analyzed the simulations, J.S., A.C., J.C. and A.A. discussed and analyzed the results, J.S. and A.C. wrote the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sgrignani, J., Chen, J., Alimonti, A. et al. How phosphorylation influences E1 subunit pyruvate dehydrogenase: A computational study. Sci Rep 8, 14683 (2018). https://doi.org/10.1038/s41598-018-33048-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-33048-z
Keywords
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.