Abstract
Gravitational forces can impose physical stresses on the human body as it functions to maintain homeostasis. It has been reported that astronauts exposed to microgravity experience altered biological functions and many subsequent studies on the effects of microgravity have therefore been conducted. However, the anticancer mechanisms of simulated microgravity remain unclear. We previously showed that the proliferation of human Hodgkin’s lymphoma (HL) cells was inhibited when these cells were cultured in time-averaged simulated microgravity (taSMG). In the present study, we investigated whether taSMG produced an anticancer effect. Exposure of human HL cells to taSMG for 2 days increased their reactive oxygen species (ROS) production and NADPH oxidase family gene expression, while mitochondrial mass, ATPase, ATP synthase, and intracellular ATP levels were decreased. Furthermore, human HL cells exposed to taSMG underwent autophagy via AMPK/Akt/mTOR and MAPK pathway modulation; such autophagy was inhibited by the ROS scavenger N-acetylcysteine (NAC). These results suggest an innovative therapeutic approach to HL that is markedly different from conventional chemotherapy and radiotherapy.
Similar content being viewed by others
Introduction
The advent of human space exploration has produced novel research in to the effects of space travel on human health and diseases. In space, microgravity and cosmic radiation are considered the most consequential environmental factors1,2. Several studies have found that astronauts and experimental animals in space experience physiological changes owing to microgravity during and after space flights; the effects of microgravity include muscle atrophy, bone loss, immune dysregulation, and abnormal cellular functions3,4,5,6,7,8. Microgravity also affects major cellular functions such as cell growth, cell cycle, self-renewal and differentiation7,9,10,11,12. As such, it was hypothesized that microgravity has anticancer potential through cell growth inhibition, and many studies were conducted to investigate this notion2,13,14,15. Additionally, microgravity has been reported to cause cellular oxidative stress that leads to the production of reactive oxygen species (ROS), as well as endoplasmic reticulum stress12,16,17,18. However the mechanism by which microgravity elicits these cellular responses remain poorly understood.
Autophagy is a catabolic process that helps maintain cellular homeostasis through the degradation of bulk cytoplasm, long-lived proteins, and organelles in response to stresses such as nutrient deprivation, viral infection, and genotoxicity19,20,21. Recent evidence suggests that autophagy is an important mediator of pathological response and of the cell’s response to oxidative stress caused by ROS and reactive nitrogen species22,23,24,25. Autophagy involves the formation of double-membrane-bound structures called autophagosomes as initiated by the phosphoinositide 3-kinase (PI3K) type III-Atg6/Beclin-1 cascade26,27. The classic PI3K/Akt/mammalian target of rapamycin (mTOR) signaling pathway is an important negative regulator of autophagosome formation. Recent studies have found that activation of adenosine monophosphate-activated protein kinase (AMPK) results in autophagy via the negative regulation of mTOR and direct phosphorylation of Unc-51 like autophagy activating kinase 1 (ULK1)28,29,30. The role of autophagy induction in cells is depend on the nature of the stimulus as well as the cell type. In cancer therapy, autophagy increases cell migration, invasion, and chemoresistance, paradoxically autophagy can also induce cell death in response to certain stimuli and causes dysregulated cell energy metabolism31,32,33,34,35,36. Therefore, it is necessary to fully understand the function and mechanism of autophagy in cancer therapy.
Hodgkin’s lymphoma (HL) is a malignant tumor originating from B cells, while its precise cause is unknown, approximately 9,000 new patients are diagnosed annually in the United States37. HL is more common in males than in females, and most commonly occurs in individuals aged 15–40 or over 50 years but rarely in those under 10 years. If diagnosed found at an early stage, it can be treated with chemotherapy or radiotherapy, whereupon the 5-year survival rate is as high as 86%. Classical therapies have greatly improved the chance of cure, although their side effects are often severe38,39,40. Furthermore, recurrence after successful treatment is common41,42,43,44,45. Therefore, novel anticancer therapies that avoid the severe side effects and recurrence rates of existing modalities are needed.
In this study, we simulated a time-averaged microgravity environment to investigate if such a milieu produces an anticancer effect against human HL cells. The time-averaged simulated microgravity (taSMG) environment was produced using a clinostat as validated in a previous study46. The clinostat rotates in a manner that produces a constantly varying gravity vector in a non-repeating pattern, thereby producing a vector-free gravity environment by continuously averaging the vector. Using a clinostat can provide gravitational stress and microgravity-like effects in cells. In our previous studies, we confirmed that the proliferation of human HL cells is inhibited in a taSMG environment46. In this study, we hypothesized that taSMG produces cellular stress in human HL cells. We found that taSMG induces autophagy through mitochondrial dysregulation via the AMPK/Akt/mTOR and MAPK pathways.
Results
taSMG induces mitochondrial dysfunction in human HL cells
Microgravity has been reported to affect cell proliferation47,48. Our previous studies also showed that proliferation of human HL cells (L-540 and HDLM-2) is inhibited under taSMG conditions46. Since cell proliferation is mediated by mitochondrial regulation49,50,51, we investigated whether taSMG induces mitochondrial stress in human HL cells. We found that intracellular ROS levels were increased under taSMG compared to normal gravity, 1 G (Fig. 1a). As ROS is generated by nicotinamide adenine dinucleotide phosphate (NADPH) oxidase52,53,54, we also found that the expression levels of NADPH oxidase family genes (gp91-, p22-, p47-, and p67-phox) are higher under taSMG than under 1 G conditions (Fig. 1b). To observe changes in mitochondrial biogenesis, we measured the mitochondrial mass using MitoTracker labeling and found it to be significantly decreased under taSMG (Fig. 1c). Furthermore, the mRNA expression levels of ATPase (ATP1A1) and ATP synthase (ATP5A1) were notably lower than 1 G levels, as determined by RT-PCR and qRT-PCR (Fig. 1d,e). These findings confirmed that intracellular ATP levels were significantly lower under taSMG than 1 G (Fig. 1f). These results suggest that taSMG causes ROS generation and mitochondrial dysfunction in human HL cells.
Time-averaged simulated microgravity induces mitochondrial dysfunction. Human HL cell lines (L-540 and HDLM-2) incubated in 1 G or taSMG conditions for 2 days were harvested. (a) ROS generation was determined by 2′,7′–dichlorofluorescin diacetate (DCFDA) staining. (b) RT-PCR analysis of NADPH oxidase family genes (gp91-, p22, p47, and p67-phox). (c) Mitochondrial mass was analyzed by MitoTrackerTM Green FM. (d) RT-PCR and (e) qRT-PCR analysis of ATPase (ATP1A1) and ATP synthase (ATP5A1) mRNA levels. (f) Analysis of intracellular ATP levels in L-540 and HDLM-2 cells. Data represent the mean ± SEM, n = 4. **p < 0.01 and ***p < 0.001 vs. the 1 G group. +++p < 0.0001 vs. the 1 G group of L-540 cells. #p < 0.05, ##p < 0.01 and ###p < 0.001 vs. the 1 G group of HDLM-2 cells. The grouping of gels cropped from different gels. 1 G: normal gravity conditions; taSMG: time-averaged simulated microgravity.
Mitochondrial dysregulation under taSMG leads to human HL cell autophagy
Mitochondrial dysfunction, as evidenced by increased ROS generation and reduced ATP levels, triggers autophagy20,22,55. Therefore, we measured the expression levels of the autophagy-related genes (ULK1, ATG14, BECN1 and LC3); all were found to be upregulated under taSMG (Fig. 2a,b). Levels of phosphorylated ULK1, ATF4, Beclin-1, and microtubule-associated protein 1 light chain 3 (LC3) were increased, while expression of the Bcl-2 family proteins Bcl-2 and Mcl-1, which inhibit autophagy by directly binding to the BH3 domain of Beclin-1/Atg6, were decreased under taSMG conditions (Fig. 2c). Additionally, we found that the LC3-II/I ratio56,57,58, an indicator of autophagy, increased under taSMG (Fig. 2d). These results showed that taSMG contributes to the induction of autophagy in human HL cells.
Time-averaged simulated microgravity contributes to induce autophagy. (a) RT-PCR and (b) qRT-PCR analysis of mRNA levels of autophagy-related genes. (c) Western blot analysis of autophagy-related proteins. GAPDH was used as the loading control. Densitometric analysis of protein expression was determined after normalization to GAPDH. The red dotted line indicates the base line. (d) Densitometric analysis of the LC3-II/I ratio was determined after normalization to GAPDH. Data represent the mean ± SEM, n = 4. +++p < 0.0001 vs. the 1 G group of L-540 cells. #p < 0.05, ##p < 0.01 and ###p < 0.001 vs. the 1 G group of HDLM-2 cells. **p < 0.01 and ***p < 0.001 vs. the 1 G group. The grouping of gels and blots cropped from different gels. 1 G: normal gravity conditions; taSMG: time-averaged simulated microgravity.
AMPK/Akt/mTOR and MAPK signaling are differentially regulated under taSMG
To further understand the mechanism of taSMG-induced autophagy, we investigated various cell signaling pathways by Western blotting. Since the liver kinase B1 (LKB1)/AMPK pathway is known to be activated under conditions of low intracellular ATP to inhibit cell growth and induce autophagy59, we have confirmed whether activation of these pathway under taSMG. We found that LKB1 and activated AMPK increased under taSMG (Fig. 3a). We also confirmed that activated AMPK inhibits the Akt/mTOR/S6K pathway (a negative regulator of autophagy) and activates the Akt suppressor protein PTEN (Fig. 3a). Previous studies have shown that ROS activates MAPK signaling, and that this elicits a variety of downstream signaling events60,61,62. To that end, we found that taSMG activates RAS, ERK, and JNK (Fig. 3b). Additionally, ROS-induced JNK and ERK activation are known to induce both autophagy and apoptosis63; however, no taSMG-induced apoptosis was observed (Fig. 3c). These results suggest that only autophagy is induced by modulating the AMPK/Akt/mTOR and MAPK signaling pathways under taSMG.
The regulation of AMPK/Akt/mTOR and MAPK pathways under time-averaged simulated microgravity. (a) Representative blots showing the phosphorylation levels of AMPK, PTEN, Akt, mTOR and S6K and total levels of LKB1, AMPK, PTEN, Akt, mTOR, and S6K in L-540 and HDLM-2 cells were determined by Western blotting. (b) Representative blots showing the activation levels of the MAPK pathway components RAS, ERK and JNK. (c) Representative blots showing apoptosis markers (PARP, caspase-3 and caspase-9). The dashed line is cleaved size that indicates apoptosis activation. All experiments were performed at least in quadruplicate. The grouping of gels cropped from different blots. 1 G: normal gravity conditions; taSMG: time-averaged simulated microgravity.
ROS scavenging attenuates the mitochondrial dysfunction induced by taSMG
We used the ROS scavenger N-acetylcysteine (NAC) to determine whether the abovementioned phenomena under taSMG can be inhibited. We treated the cells with NAC during clinostat operation and observed no increase in the mRNA levels of NADPH oxidase family genes (Fig. 4a). Using RT-PCR (Fig. 4b) and qRT-PCR (Fig. 4c), we also found that mRNA levels of ATPase and ATP synthase were not reduced under taSMG in the presence of NAC.
N-acetylcysteine protects mitochondrial dysfunction by time-averaged simulated microgravity. (a) RT-PCR data shows the mRNA levels of NADPH oxidase family genes. (b) RT-PCR and (c) qRT-PCR analysis of mRNA levels of ATPase and ATP synthase. Data represent the mean ± SEM, n = 4. *p < 0.05, **p < 0.01 and ***p < 0.001 vs. the 1 G group or taSMG. The grouping of gels cropped from different blots. 1 G: normal gravity conditions; taSMG: time-averaged simulated microgravity. Treatment of NAC proceeded under the same conditions.
In addition, we confirmed that taSMG-induced autophagy was inhibited by NAC treatment. In the presence of NAC, we found that there was no changes in autophagy-related mRNA (Fig. 5a,b) and protein levels, including the LC3-II/I ratio (Fig. 5c).
N-acetylcysteine prevents autophagy by time-averaged simulated microgravity. (a) RT-PCR and (b) qRT-PCR shows mRNA levels of autophagy-related genes. (c) Western blot analysis of protein levels of autophagy-related genes. (d) Densitometric analysis of LC3-II/I ratio was determined after normalization to GAPDH. Data represent the mean ± SEM, n = 4. *p < 0.05, **p < 0.01 and ***p < 0.001 vs. the 1 G group or taSMG. The grouping of gels cropped from different blots. 1 G: normal gravity conditions; taSMG: time-averaged simulated microgravity. Treatment of NAC proceeded under the same conditions.
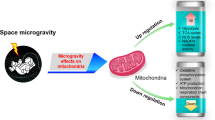
Taken together, these results indicate that autophagy was induced by mitochondrial dysfunction due to ROS generation under taSMG, suggesting that inhibition of ROS generation would prevent these phenomena (Fig. 6).
Schematic representation of the time-averaged simulated microgravity (taSMG)-induced autophagy mechanism in human Hodgkin’s lymphoma cells. taSMG induces the activation of AMPK, PTEN, and MAPK, and as well as the suppression of Akt, leading to the inhibition of mTOR activity. The downstream regulator S6K, which inhibits autophagy, is consequently suppressed, resulting in autophagy in human HL cells.
Discussion
Primary chemoradiotherapy results in cure rates of 90% and 80% in HL patients with early- and advanced-stage disease, respectively64. However, these treatments lead to a significant risk of short- and long-term toxicities, which can also cause secondary malignancies65,66. Therefore, new therapies based on suppressing disease progression mechanisms, including molecule-specific inhibition are needed for successful treatment. We sought to investigate the use of microgravity as a potential new treatment for HL, given our previous findings that it inhibits cell proliferation46.
As taSMG can be achieved artificially, studies using this technique have advanced rapidly. Based on symptoms experienced by astronauts in the space such as muscle atrophy, bone loss, and immunodeficiency, microgravity has been investigated in the treatment of various diseases2,4,6,7,10,17. Our research team designed a clinostat that uses a specific algorithm aimed at randomizing the gravitational vector pattern and achieving a nullified time-averaged vector46.
Microgravity in space has been reported to cause oxidative stress such as ROS production67,68,69. However, the study of microgravity induced oxidative stress is not yet fully understood. In this study, we validated our hypothesis that taSMG induces oxidative cellular stress in human HL cells, as demonstrated by elevated ROS and impaired mitochondrial function.
NADPH oxidase-dependent ROS production is implicated in many physiologic and pathophysiologic processes. NADPH oxidase mediated ROS can alter parameters of signal transduction, mitochondrial damage, cell proliferation, cell death and autophagy22,23,54,69. We also found that taSMG upregulates NADPH oxidase family genes while decreasing the mitochondrial mass, and lowering ATPase, and ATP synthase levels, resulting in reduced intracellular ATP levels.
Autophagy mediates the bulk degradation of intracellular through lysosomal-dependent mechanisms and is necessary for the maintenance of cellular homeostasis19,20,22. Autophagy is also induced in response to oxidative stress caused by ROS and RNS22,24,25. We hypothesized that autophagy would occur due to increased ROS, and interestingly, autophagy was induced under taSMG.
Recently, some research reported that autophagy is induced in simulated microgravity and inhibits cancer proliferation and metastasis18,21,70. However, its molecular mechanisms are unclear yet. As an intracellular energy sensor, AMPK signaling serves as a mitochondrial function regulator to maintain energy homeostasis29. And recent studies have found that activation of AMPK results in autophagy through negative regulation of mTOR and phosphorylation of ULK127,28,29,30. Interestingly, we showed that the mitochondrial dysfunction produced by taSMG promotes autophagy via modulation of the AMPK/Akt/mTOR and MAPK signaling pathways. Overall, the effect of taSMG was more prominent in L-540 cells than in HDLM-2 cells; L-540 cells have a higher proliferation rate, they are likely more sensitive to taSMG than HDLM-2 cells.
In fact, the role of autophagy in cancer remains controversial given that it varies according to the type of cancer. Autophagy may contribute to cancer progression by increasing cell migration and invasion; conversely, it can have an anticancer effect by decreasing cell proliferation, and promoting cancer cell death28,31,34,35,36.
Taken together, our research findings suggest that taSMG-induced autophagy following the induction of oxidative stress in human HL cells has an anticancer effect. For the scavenger of ROS generation, we confirmed that phenomena of oxidative stress and autophagy were prevented under taSMG.
We can’t know whether the effect of taSMG occurs in normal cells. Previous studies confirmed that proliferation of human dermal fibroblast (HDF) cells did not change under taSMG46. However, as we know that it is necessary to compare the effects of taSMG on normal lymphocytes and HL cells, we will proceed further study.
Our data ought to provide important insight concerning the effect of taSMG on cancer cells, as well as our understanding of human HL cell mechanisms. These findings could lead the way to promote new treatment methods for cancer patients.
Materials and Methods
Control of 3D Clinostat
3D clinostat, which consists of two perpendicular rotating axes, were developed and validated in our previous research41. The 3D clinostat provides taSMG, which is time-averaged acceleration (including gravitational, centrifugal, and tangential acceleration) less than 10−3G after 24 h operation. Coriolis, radial, and frictional forces were not considered in the simulation because angular velocities of two axes were set to sufficiently small values (inner frame: 0.683 rpm, outer frame: 0.913 rpm). Operation of 3D clinostat with designed angular velocity was proven to avoid repeated pattern of gravitational vector. For more detailed information, previously published paper can be helpful.
Cell culture
Human Hodgkin’s lymphoma (HL) cell lines L-540 and HDLM-2 were obtained from the German Collection of Microorganisms and Cell Cultures (DSMZ, Braunschweig, Germany)71. The cell lines were maintained in RPMI 1640 (Life Technologies, USA) supplemented with 10% fetal bovine serum (FBS, Life Technologies, USA) and 1% penicillin/streptomycin solution (Life Technologies, USA) at 37 °C in 5% CO2. For cellular ROS inhibition, the cells were added with N-acetylcysteine (NAC, Sigma Aldrich, USA) at 10 mM in complete medium for operating clinostat. Simulation of microgravity using clinostat were previously described46. All experiments were performed after human HL cells were cultured under 1 G or taSMG condition for 2 days.
ROS detection assay
For detection of cellular ROS, we used DCFDA cellular ROS detection assay kit (Abcam, UK). The collected cells were stained with 20 μM DCFDA in 1X buffer for 30 min at 37 °C. Without the washing procedure, stained cells were immediately carried out by flow cytometry (FACS) using FACS LSRFortessa (BD Bioscience, USA) at Ex488 nm/Em535 nm beam. Each determination was based on the mean fluorescence intensity of 10,000 cells.
Mitochondrial mass analysis
Mitochondrial mass per cell was measured by flow cytometry using MitoTracker Green FM (Thermo Fisher Scientific, USA). Cells were collected, resuspended in 0.5 ml of PBS, and stained with 40 nM MitoTracker green (MTG) for 15 min at 37 °C in the dark. Cells were then washed with PBS, resuspended in FACS buffer (eBioscience, USA), and added 200 ng/ml of DAPI (Sigma Aldrich, USA). Stained cells were analyzed using FACS LSRFortessa at Ex488 nm/Em535 nm beam. Each determination was based on the mean fluorescence intensity of 25,000 cells.
Reverse transcription (RT)-PCR and quantitative real-time RT-PCR (qRT-PCR)
Total RNA were extracted using a RNAiso Plus reagent (Takara, Japan) and cDNA was synthesized using ReverTra Ace qPCR RT Master Mix kit (TOYOBO, Japan). Quantitative real time-PCR was performed using the SYBR Green PCR mix (Applied Biological Material, Canada) with an Applied Biosystems 7300 Real-time PCR system (Life Technologies, USA) and the raw data were analyzed using comparative Ct quantification. All primers were purchased from Qiagen (Qiagen, USA). The as basic PCR amplification conditions were 58 °C annealing temperature and 35 cycles.
ATP assay
Levels of intracellular ATP was measured using an ATP Bioluminescence assay kit (Roche, Switzerland) according to manufacturer’s instruction. Cells were collected and heated tubes to at least 95 °C for 7 min. To remove pellet cell debris, spin down at 14000 RPM for 3 min. And then each sample were transferred to a black 96 well plate and quickly added luciferase reagent to each well. Luminescence was measured at 0.1 sec/well using the luminescence program.
Western blot
Cells were lysed in a lysis buffer (50 mM Tris-HCl, pH 7.4, 350 mM NaCl, 0.5% NonidetP-40, 10% glycerol, 0.1% SDS, and 1% Triton X-100). The lysates were centrifuged at 13200 rpm for 10 min at 4 °C and protein amounts were quantified using a Bio-Rad protein assay (Bio-Rad, USA). Proteins were separated by SDS-poly acrylamide gel electrophoresis (SDS-PAGE) and transferred onto nitrocellulose membranes (Whatman, Atlanta, USA). The membranes were blocked in blocking buffer (5% skim milk in 150 mM NaCl, 25 mM Tris-HCl, pH 7.4, and 0.1% Tween 20) and subsequently incubated with specific primary antibodies for the target molecules. The membranes were then washed with Tris-buffered saline containing 0.1% Tween 20 (TBS-T) and further incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1 h at room temperature. After washing with TBS-T, the signals were visualized using the ECL Plus Western blotting substrate (Thermo Fisher Scientific, USA).
Statistical analysis
All experiments were performed by more than four times. The results are represented as means with standard error of the mean (SEM). Statistical significance was determined based on a two-tailed Student’s t-test and analyzed using Graph Pad Prism 6 (Graph Pad Software, INC., USA).
References
Ran, F., An, L., Fan, Y., Hang, H. & Wang, S. Simulated microgravity potentiates generation of reactive oxygen species in cells. Biophysics Reports 2, 100–105, https://doi.org/10.1007/s41048-016-0029-0 (2016).
Kurisu, K., Takeda, M., Okazaki, T., Kawahara, Y. & Yuge, L. Effects of Simulated Microgravity on Proliferation and Chemosensitivity in Malignant Glioma Cells. Neuro-Oncology 16, iii36–iii36, https://doi.org/10.1093/neuonc/nou208.50 (2014).
Fitts, R. H. et al. Prolonged space flight-induced alterations in the structure and function of human skeletal muscle fibres. The Journal of Physiology 588, 3567–3592, https://doi.org/10.1113/jphysiol.2010.188508 (2010).
Trappe, S. W. et al. Comparison of a space shuttle flight (STS-78) and bed rest on human muscle function. Journal of Applied Physiology 91, 57–64, https://doi.org/10.1152/jappl.2001.91.1.57 (2001).
Williams, D., Kuipers, A., Mukai, C. & Thirsk, R. Acclimation during space flight: effects on human physiology. CMAJ: Canadian Medical Association Journal 180, 1317–1323, https://doi.org/10.1503/cmaj.090628 (2009).
Foster, J. S., Khodadad, C. L. M., Ahrendt, S. R. & Parrish, M. L. Impact of simulated microgravity on the normal developmental time line of an animal-bacteria symbiosis. Scientific Reports 3, 1340, https://doi.org/10.1038/srep01340 (2013).
Wei, L. et al. Effect of Change in Spindle Structure on Proliferation Inhibition of Osteosarcoma Cells and Osteoblast under Simulated Microgravity during Incubation in Rotating Bioreactor. PLoS ONE 8, e76710, https://doi.org/10.1371/journal.pone.0076710 (2013).
Grimm, D. et al. The impact of microgravity on bone in humans. Bone 87, 44–56, https://doi.org/10.1016/j.bone.2015.12.057 (2016).
Maier, J. A. M., Cialdai, F., Monici, M. & Morbidelli, L. The Impact of Microgravity and Hypergravity on Endothelial Cells. Bio Med Research International 2015, 434803, https://doi.org/10.1155/2015/434803 (2015).
Lin, S.-C. et al. Simulated Microgravity Disrupts Cytoskeleton Organization and Increases Apoptosis of Rat Neural Crest Stem Cells Via Upregulating CXCR4 Expression and RhoA-ROCK1-p38 MAPK-p53 Signaling. Stem Cells and Development 25, 1172–1193, https://doi.org/10.1089/scd.2016.0040 (2016).
Blaber, E. A. et al. Microgravity Reduces the Differentiation and Regenerative Potential of Embryonic Stem Cells. Stem Cells and Development 24, 2605–2621, https://doi.org/10.1089/scd.2015.0218 (2015).
Feger, B. J. et al. Microgravity induces proteomics changes involved in endoplasmic reticulum stress and mitochondrial protection. Scientific Reports 6, 34091, https://doi.org/10.1038/srep34091 (2016).
Arun, R. P., Sivanesan, D., Vidyasekar, P. & Verma, R. S. PTEN/FOXO3/AKT pathway regulates cell death and mediates morphogenetic differentiation of Colorectal Cancer Cells under Simulated Microgravity. Scientific Reports 7, 5952, https://doi.org/10.1038/s41598-017-06416-4 (2017).
Riwaldt, S. et al. Pathways Regulating Spheroid Formation of Human Follicular Thyroid Cancer Cells under Simulated Microgravity Conditions: A Genetic Approach. International Journal of Molecular Sciences 17, 528, https://doi.org/10.3390/ijms17040528 (2016).
Kopp, S. et al. Identifications of novel mechanisms in breast cancer cells involving duct-like multicellular spheroid formation after exposure to the Random Positioning Machine. Scientific Reports 6, 26887, https://doi.org/10.1038/srep26887 (2016).
Adrian, A. et al. The oxidative burst reaction in mammalian cells depends on gravity. Cell Communication and Signaling: CCS 11, 98–98, https://doi.org/10.1186/1478-811X-11-98 (2013).
Verhaar, A. P. et al. Dichotomal effect of space flight-associated microgravity on stress-activated protein kinases in innate immunity. Scientific Reports 4, 5468, https://doi.org/10.1038/srep05468 (2014).
Morabito, C. et al. Transient increases in intracellular calcium and reactive oxygen species levels in TCam-2 cells exposed to microgravity. Scientific Reports 7, 15648, https://doi.org/10.1038/s41598-017-15935-z (2017).
Poillet-Perez, L., Despouy, G., Delage-Mourroux, R. & Boyer-Guittaut, M. Interplay between ROS and autophagy in cancer cells, from tumor initiation to cancer therapy. Redox Biology 4, 184–192, https://doi.org/10.1016/j.redox.2014.12.003 (2015).
Filomeni, G., De Zio, D. & Cecconi, F. Oxidative stress and autophagy: the clash between damage and metabolic needs. Cell Death and Differentiation 22, 377, https://doi.org/10.1038/cdd.2014.150 (2014).
Sambandam, Y. et al. Microgravity control of autophagy modulates osteoclastogenesis. Bone 61, 125–131, https://doi.org/10.1016/j.bone.2014.01.004 (2014).
Lee, J., Giordano, S. & Zhang, J. Autophagy, mitochondria and oxidative stress: cross-talk and redox signalling. Biochemical Journal 441, 523–540, https://doi.org/10.1042/BJ20111451 (2012).
Pal, R. et al. Src-dependent impairment of autophagy by oxidative stress in a mouse model of Duchenne muscular dystrophy. Nature communications 5, 4425–4425, https://doi.org/10.1038/ncomms5425 (2014).
Wang, Q., Liang, B., Shirwany, N. A. & Zou, M.-H. 2-Deoxy-D-Glucose Treatment of Endothelial Cells Induces Autophagy by Reactive Oxygen Species-Mediated Activation of the AMP-Activated Protein Kinase. PLoS ONE 6, e17234, https://doi.org/10.1371/journal.pone.0017234 (2011).
Sureshbabu, A., Ryter, S. W. & Choi, M. E. Oxidative stress and autophagy: Crucial modulators of kidney injury. Redox Biology 4, 208–214, https://doi.org/10.1016/j.redox.2015.01.001 (2015).
Xie, Z. & Klionsky, D. J. Autophagosome formation: core machinery and adaptations. Nature Cell Biology 9, 1102, https://doi.org/10.1038/ncb1007-1102 (2007).
Yu, Y. et al. Akt/AMPK/mTOR pathway was involved in the autophagy induced by vitamin E succinate in human gastric cancer SGC-7901 cells. Molecular and Cellular Biochemistry 424, 173–183, https://doi.org/10.1007/s11010-016-2853-4 (2017).
He, C. & Klionsky, D. J. Regulation Mechanisms and Signaling Pathways of Autophagy. Annual review of genetics 43, 67–93, https://doi.org/10.1146/annurev-genet-102808-114910 (2009).
Kim, J., Kundu, M., Viollet, B. & Guan, K.-L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nature cell biology 13, 132–141, https://doi.org/10.1038/ncb2152 (2011).
Shaw, R. J. LKB1 and AMPK control of mTOR signalling and growth. Acta physiologica (Oxford, England) 196, 65–80, https://doi.org/10.1111/j.1748-1716.2009.01972.x (2009).
Grandér, D. & Panaretakis, T. Autophagy: cancer therapy’s friend or foe? Future Medicinal Chemistry 2, 285–297, https://doi.org/10.4155/fmc.09.155 (2010).
Laane, E. et al. Cell death induced by dexamethasone in lymphoid leukemia is mediated through initiation of autophagy. Cell Death and Differentiation 16, 1018, https://doi.org/10.1038/cdd.2009.46 (2009).
Buentke, E. et al. Glucocorticoid-induced cell death is mediated through reduced glucose metabolism in lymphoid leukemia cells. Blood Cancer Journal 1, e31, https://doi.org/10.1038/bcj.2011.27 (2011).
Thorburn, A., Thamm, D. H. & Gustafson, D. L. Autophagy and Cancer Therapy. Molecular Pharmacology 85, 830–838, https://doi.org/10.1124/mol.114.091850 (2014).
Yang, Z. J., Chee, C. E., Huang, S. & Sinicrope, F. A. The Role of Autophagy in Cancer: Therapeutic Implications. Molecular cancer therapeutics 10, 1533–1541, https://doi.org/10.1158/1535-7163.MCT-11-0047 (2011).
Sui, X. et al. Autophagy and chemotherapy resistance: a promising therapeutic target for cancer treatment. Cell Death & Amp; Disease 4, e838, https://doi.org/10.1038/cddis.2013.350 (2013).
Surveillance, Epidemiology, and End Results Program. Cancer Strat Facts: Hodgkin Lymphoma. NIH. National Cancer Institute (2018).
Eichenauer, D. A. et al. Therapy-related acute myeloid leukemia and myelodysplastic syndromes in patients with Hodgkin lymphoma: a report from the German Hodgkin Study Group. Blood 123, 1658 (2014).
Rutenberg, M. S., Flampouri, S. & Hoppe, B. S. Proton therapy for Hodgkin lymphoma. Current Hematologic Malignancy Reports 9, 203–211, https://doi.org/10.1007/s11899-014-0212-7 (2014).
Franklin, J., Eichenauer, D. A., Becker, I., Monsef, I. & Engert, A. Optimisation of chemotherapy and radiotherapy for untreated Hodgkin lymphoma patients with respect to second malignant neoplasms, overall and progression‐free survival: individual participant data analysis. Cochrane Database of Systematic Reviews. https://doi.org/10.1002/14651858.CD008814.pub2 (2017).
Canellos, G. P. et al. Chemotherapy of Advanced Hodgkin’s Disease with MOPP, ABVD, or MOPP Alternating with ABVD. New England Journal of Medicine 327, 1478–1484, https://doi.org/10.1056/NEJM199211193272102 (1992).
Duggan, D. B. et al. Randomized Comparison of ABVD and MOPP/ABV Hybrid for the Treatment of Advanced Hodgkin’s Disease: Report of an Intergroup Trial. Journal of Clinical Oncology 21, 607–614, https://doi.org/10.1200/JCO.2003.12.086 (2003).
Chisesi, T. et al. Long-Term Follow-Up Analysis of HD9601 Trial Comparing ABVD Versus Stanford V Versus MOPP/EBV/CAD in Patients With Newly Diagnosed Advanced-Stage Hodgkin’s Lymphoma: A Study From the Intergruppo Italiano Linfomi. Journal of Clinical Oncology 29, 4227–4233, https://doi.org/10.1200/JCO.2010.30.9799 (2011).
Engert, A. et al. Reduced-intensity chemotherapy and PET-guided radiotherapy in patients with advanced stage Hodgkin’s lymphoma (HD15 trial): a randomised, open-label, phase 3 non-inferiority trial. The Lancet 379, 1791–1799, https://doi.org/10.1016/S0140-6736(11)61940-5 (2012).
Song, E. J. et al. Consolidation Radiation Therapy for Patients With Advanced Hodgkin Lymphoma in Complete Metabolic Response According to PET-CT or Gallium Imaging. Clinical Lymphoma Myeloma and Leukemia, https://doi.org/10.1016/j.clml.2017.12.007 (2017).
Kim, Y. J. et al. Time-averaged simulated microgravity (taSMG) inhibits proliferation of lymphoma cells, L-540 and HDLM-2, using a 3D clinostat. BioMedical Engineering OnLine 16, 48, https://doi.org/10.1186/s12938-017-0337-8 (2017).
Dhwani, V. J., Raosaheb, K. K. & Rana, P. S. Microgravity Alters Cancer Growth and Progression. Current Cancer Drug Targets 14, 394–406, https://doi.org/10.2174/1568009614666140407113633 (2014).
Yan, M. et al. The effects and mechanisms of clinorotation on proliferation and differentiation in bone marrow mesenchymal stem cells. Biochemical and Biophysical Research Communications 460, 327–332, https://doi.org/10.1016/j.bbrc.2015.03.034 (2015).
Antico Arciuch, V. G., Elguero, M. E., Poderoso, J. J. & Carreras, M. C. Mitochondrial Regulation of Cell Cycle and Proliferation. Antioxidants & Redox Signaling 16, 1150–1180, https://doi.org/10.1089/ars.2011.4085 (2012).
Yan, X.-J. et al. Mitochondria play an important role in the cell proliferation suppressing activity of berberine. Scientific Reports 7 41712, https://doi.org/10.1038/srep41712 https://www.nature.com/articles/srep41712#supplementary-information (2017).
Holmuhamedov, E. et al. Suppression of human tumor cell proliferation through mitochondrial targeting. The FASEB Journal 16, 1010–1016, https://doi.org/10.1096/fj.01-0996com (2002).
Meitzler, J. L. et al. NADPH Oxidases: A Perspective on Reactive Oxygen Species Production in Tumor Biology. Antioxidants & Redox Signaling 20, 2873–2889, https://doi.org/10.1089/ars.2013.5603 (2014).
Chen, S., Meng, X.-F. & Zhang, C. Role of NADPH Oxidase-Mediated Reactive Oxygen Species in Podocyte Injury. BioMed Research International 2013, 7, https://doi.org/10.1155/2013/839761 (2013).
Bedard, K. & Krause, K.-H. The NOX Family of ROS-Generating NADPH Oxidases: Physiology and Pathophysiology. Physiological Reviews 87, 245–313, https://doi.org/10.1152/physrev.00044.2005 (2007).
Apostolova, N. & Victor, V. M. Molecular Strategies for Targeting Antioxidants to Mitochondria: Therapeutic Implications. Antioxidants & Redox Signaling 22, 686–729, https://doi.org/10.1089/ars.2014.5952 (2015).
Aparicio, I. M. et al. Autophagy-related proteins are functionally active in human spermatozoa and may be involved in the regulation of cell survival and motility. Scientific Reports 6, 33647, https://doi.org/10.1038/srep33647 (2016).
Kadowaki, M. & Karim, M. R. In Methods in Enzymology Vol. 452 199–213 (Academic Press, 2009).
Wang, Y. et al. Loss of Macroautophagy Promotes or Prevents Fibroblast Apoptosis Depending on the Death Stimulus. The Journal of biological chemistry 283, 4766–4777, https://doi.org/10.1074/jbc.M706666200 (2008).
Shackelford, D. B. & Shaw, R. J. The LKB1-AMPK pathway: metabolism and growth control in tumor suppression. Nature reviews. Cancer 9, 563–575, https://doi.org/10.1038/nrc2676 (2009).
Son, Y. et al. Mitogen-Activated Protein Kinases and Reactive Oxygen Species: How Can ROS Activate MAPK Pathways? Journal of Signal Transduction 2011, 6, https://doi.org/10.1155/2011/792639 (2011).
Torres, M. & Forman, H. J. Redox signaling and the MAP kinase pathways. BioFactors 17, 287–296, https://doi.org/10.1002/biof.5520170128 (2003).
Valko, M. et al. Free radicals and antioxidants in normal physiological functions and human disease. The International Journal of Biochemistry & Cell Biology 39, 44–84, https://doi.org/10.1016/j.biocel.2006.07.001 (2007).
Zhou, Y.-Y., Li, Y., Jiang, W.-Q. & Zhou, L.-F. MAPK/JNK signalling: a potential autophagy regulation pathway. Bioscience Reports 35, e00199, https://doi.org/10.1042/BSR20140141 (2015).
Lowry, L., Hoskin, P. & Linch, D. Developments in the management of Hodgkin’s lymphoma. The Lancet 375, 786–788, https://doi.org/10.1016/S0140-6736(09)61878-X (2010).
Stein, H. et al. Down-regulation of BOB.1/OBF.1 and Oct2 in classical Hodgkin disease but not in lymphocyte predominant Hodgkin disease correlates with immunoglobulin transcription. Blood 97, 496 (2001).
Berglund, M. et al. Molecular cytogenetic characterization of four commonly used cell lines derived from Hodgkin lymphoma. Cancer Genetics and Cytogenetics 141, 43–48, https://doi.org/10.1016/S0165-4608(02)00656-8 (2003).
Tian, Y. et al. The Impact of Oxidative Stress on the Bone System in Response to the Space Special Environment. International Journal of Molecular Sciences 18, 2132, https://doi.org/10.3390/ijms18102132 (2017).
Stein, T. P. & Leskiw, M. J. Oxidant damage during and after spaceflight. American Journal of Physiology-Endocrinology and Metabolism 278, E375–E382, https://doi.org/10.1152/ajpendo.2000.278.3.E375 (2000).
Mao, X. W. et al. Role of NADPH Oxidase as a Mediator of Oxidative Damage in Low-Dose Irradiated and Hindlimb-Unloaded Mice. Radiation Research 188, 392–399, https://doi.org/10.1667/RR14754.1 (2017).
Yoo, Y.-M., Han, T.-Y. & Kim, H. S. Melatonin Suppresses Autophagy Induced by Clinostat in Preosteoblast MC3T3-E1 Cells. International Journal of Molecular Sciences 17, 526, https://doi.org/10.3390/ijms17040526 (2016).
Kim, B. H. et al. A small molecule compound identified through a cell-based screening inhibits JAK/STAT pathway signaling in human cancer cells. Molecular cancer therapeutics 7, 2672–2680, https://doi.org/10.1158/1535-7163.MCT-08-0309 (2008).
Acknowledgements
This work was supported by the Brain Korea 21 PLUS program. The National Research Foundation of Korea (NRF) funded by the Korea government (MISP; grant No. 2014R1A2A1A11053203 and 2017R1A2B2006839 to S.K. Ye and 2018M1A3A3A02065779 to S. Kim).
Author information
Authors and Affiliations
Contributions
A.J.J. performed whole biological experiments and wrote the manuscript. Y.J.K. manufactured clinostat hardware and development the algorithm. M.H.L. participated in clinostat algorithm development. H.L., K.N. and B.H.K. conducted parts of the experiments. J.W.C. and C.H.C. provided materials. S.K.Y. and S.K. planned and supervised experimental design.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jeong, A.J., Kim, Y.J., Lim, M.H. et al. Microgravity induces autophagy via mitochondrial dysfunction in human Hodgkin’s lymphoma cells. Sci Rep 8, 14646 (2018). https://doi.org/10.1038/s41598-018-32965-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32965-3
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.