Abstract
Beside its unique nutritional content breast milk also contains live cells from the mother. Fate of these cells in the offspring has not been adequately described. In this study, we aimed to detect and identify maternal cells in the suckling’s blood and the brain. Green fluorescent protein expressing transgenic female mice (GFP+) were used as foster mothers to breastfeed wildtype newborn pups. One week and two months after the birth, blood samples and brains of the sucklings were analyzed to detect presence of GFP+ cells by fluorescence activated cell sorting, polymerase chain reaction and immunohistochemistry on the brain sections and optically cleared brains. The tests confirmed that maternal cells were detectable in the blood and the brain of the pups and that they differentiated into both neuronal and glial cell types in the brain. This phenomenon represents breastfeeding – induced microchimerism in the brain with functional implications remain to be understood.
Similar content being viewed by others
Introduction
The breast milk is a unique secretion that contains different bioactive substances such as hormones, enzymes, immunoglobulins, growth factors, cytokines, anti-inflammatory agents and anti-microbial factors beside the nutritional content composed of proteins, carbohydrates, fats and vitamins1. It also harbors live cells of different types that vary in distribution significantly. While leukocytes constitute 13–70% of all breast milk cells (BMCs) under normal conditions, this rate may rise up to 94% in an infection2. Epithelial cells from the ducts of mammary glands are always among the normal cellular population3. Another group of cells in the breast milk is mammary gland stem cells (BSc) that provide the formation of new mammary tissue during lactogenesis4. While BSc are found in few numbers or inactive in a normal mammary gland, they actively regenerate the mammary gland with pregnancy and breastfeeding. They can differentiate into alveolar, ductal and myoepithelial cells of mammary tissue5. Indeed, an entirely new breast formation was achieved in BSc transplanted mice6. Rather curiously, beside BSc, the breast milk contains other types of stem cells that express embryonic markers like nestin, cytokeratin, OCT4, SOX2, NANOG, SSEA4 and TRA-14,7,8. These cells were successfully differentiated into neurons, hepatocytes, pancreatic beta cells, osteoblasts, and adipocytes under in vitro conditions4.
The breast milk – born cells have been shown to survive the challenging conditions of gastrointestinal tract of an infant and pass to the intestinal wall9 and blood circulation that carries them to the liver10 and the spleen11. However, exact distribution of these cells in the body and their fate are largely unknown. Possibility of breast milk stem cells to differentiate and to get integrated into different tissues has been speculated but not conclusively proved12.
A first attempt to decipher transfer and potential integration of breastmilk-derived stem cells, along with immune cells, to the offspring was recently carried out by Hassiotou (now Kakulas) et al. with positive results showing integration and differentiation in various organs of the nursed offspring in a murine model13. Consistent with these earlier reports, in this study, we have shown that, breast milk stem cells pass to the pups, reach to the brain, settle there and differentiate into neuron and glial cells in mice.
Results
Detection of GFP+ Cells in the Bloodstream and Brain of Pups by Flow Cytometry
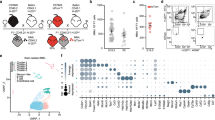
To detect the transfer of milk cells, we made GFP+ female mice breastfeed WT pups postpartum. Background noise and threshold of the GFP and anti-GFP signal was determined using positive (GFP+) and negative (WT) control cell suspensions prepared from freshly dissected brain tissues with isotype controls. The cells that had both GFP signal and anti-GFP staining were considered to be of breast milk origin. As expected, we found >99% of GFP+ cells in the positive control group while <0.1% in the negative control group (Fig. 1). We found GFP+ cells in the bloods of pups which were nursed by GFP+ dams for 1 week (5.18 ± 2.1%) and for 2 months (4.7 ± 1.6%). We also detected GFP+ cells in the brain tissue of pups which were nursed by GFP+ dams for 1 week (0.15 ± 0.1%) and for 2 months (0.21 ± 1%). There was no statistically significant difference in GFP+ cell numbers between 1 week and 2 months nursing periods (Supplementary Table S1). Cells identified as GFP+ by flow cytometry were sorted and examined by confocal microscope and GFP and anti-GFP signals were verified (Fig. 1).
Experimental design and flow cytometric analyses. (A) Breeding and nursing diagram of WT and GFP+ mice. WT newborn pups were immediately delivered to GFP+ foster mothers to be breastfed. Flow cytometry plots present the confirmation of expected GFP expression in the brains of GFP+ and WT mice. (B) Brain samples of WT and GFP+ mice were analyzed with GFP/FSC plot to determine the threshold of GFP signal for further flow cytometry analysis. (C) GFP+ cells were detected in both blood and brain of pups by the end of first week (n = 6) and two months (n = 6) of nursing. Error bars in the percentage of GFP+ cells in total cells graphs are S.D. (D) GFP+ cells detected in the brain by flow cytometry were sorted to verify GFP signal microscopically (scale bars: 10 µm).
Characterization of GFP+ Cells in Brain by Immunohistochemistry
Flow cytometry analysis showed that maternal cells can be transferred to the blood and brain of the pups via breast milk. To verify presence of these cells and identify their phenotype in the pups’ brains tissue clearing and immunocytochemistry were performed. For this, brain sections were double stained with primary antibodies (anti-GFP/anti-GFAP and anti-GFP/anti-NeuN). Triple positive cells (GFP+/anti-GFP+/anti-NeuN+ and GFP+/anti-GFP+/anti-GFAP+) were examined in whole coronal brain sections from Bregma +2 mm to −3,5 mm. These cells were widely distributed from the cerebral cortex to the different midbrain regions. We detected them in the deep cortical plate, dispersed among fiber tracts such as corpus callosum, in and around the hippocampal region, striatum, thalamus and hypothalamus. Throughout the brain we could not identify a specific brain region where GFP+ maternal cells were preferentially concentrated. The GFP signal was co-localized both with neuronal marker NeuN and glial protein GFAP immunoreactivity (Figs 2 and 3, see also Supplementary Figs S1 and S2 and Movies S1–S3).
Immunohistochemically stained brain sections of sucklings nursed by GFP+ mothers. Breast milk borne maternal cells are randomly distributed in the pup’s brain. (A,B) Representative zoomed images of GFP+ cells in the hippocampal region of the brain. (C–F) Glial cell marker (anti-GFAP) and neuronal marker (anti-NeuN) were used to label different cell types, counterstained with nuclear dye TO-PRO3. GFP+ cells differentiated to neuronal and glial cells were detected after one week and two months of nursing (scale bars: 5 µm).
Tissue clearing method and immunohistochemical staining of sucklings brain. iDISCO method was used to determine the specific localization and 3-dimensional morphology of the GFP+ cells in the pup’s brain. GFP+ cells showed a random distribution like in sections. However, the morphology and localization of these cells were much better elucidated in cleared tissue. (A–D) GFAP+ and GFP+ cells can be seen in cleared brain section of pups (250 µm z-optical section of 2 mm thick section). (E) Close proximity of a GFP+ maternal cell to a blood vessel is shown. As it also expresses glial marker GFAP, it is likely to be an astrocyte associated with the blood vessel to contribute the blood brain barrier. (F) Orthogonal section of GFP+ cell detected at the top of the blood vessel (scale bars: 15 µm) (See also Movie S1).
Detection of EGFP Copy Number in Brain of Pups by qPCR
CT values for each sample were taken as average of each replicate, log copy numbers were calculated using the standard curve equation. After comparison from log copy number to copy number, presence of breast milk originating GFP+ cell in brain tissues of 1-week and 2-months old pups were determined quantitatively as shown in Table 1.
Discussion
Breast milk has always attracted research, traditionally as an invaluable nutritional source and recently as a medium containing various biologically active compounds and living cells14,15,16. The relationship of the cells in breast milk with the baby was first demonstrated in 198317 and subsequent studies on the subject often focused on their immune interactions9,10,18,19. By the demonstration of immunoreactivity for multipotent stem cell marker nestin in the BMCs8 possibility of more diverse functions became the subject of new investigations. It was shown that embryonic stem cell - like cells isolated from the breast milk can differentiate into cells of various tissues from 3 different germ layers3,4. Hassiotou et al. performed a detailed analysis of BMCs’ fate in wildtype pups breastfed by TdTomato expressing foster mothers and reported their presence in the blood, stomach, thymus, liver pancreas, spleen and brain in stem cell or differentiated forms13. In the current study, we confirm and extend some of these findings by demonstrating that stem cells in the breast milk not only pass to the suckling’s blood circulation but also home in on the brain tissue where they differentiate into both neurons and glial cells.
The first challenge for the BMCs is to survive the harsh conditions in the gastrointestinal tract and then to enter the blood circulation. However, the neonatal organisms have some compromises for both. First, digestive enzymes and gastric acid in the digestive system are weaker in newborns20,21. It was reported that BMCs collected from stomach of 2-week old mice following suckling had a viability of 80%22. Second, the permeability of the intestines to macromolecules and cells in the neonatal period is higher than that of a fully developed intestine23. Experiments with different animal models, including mouse, baboon and lamb, demonstrated that milk leukocytes can penetrate the intestinal wall by diapedesis and pass into the blood circulation10,17,24,25,26,27. It has also been reported that the permeability of the Peyer’s patches in the intestines is greater than in other intestinal regions. Indeed, maternal T lymphocytes from the breast milk were detected in this tissue9. Once they pass to portal circulation, the first destination of BMCs would be the liver10,28,29. Consistent with other reports13, we found quite a large number of maternal cells in the suckling’s blood even after two months, a finding emphasizing the significance of the phenomenon. Another standing question is how BMC’s cross the blood brain barrier (BBB). Activated leukocytes may pass through the vascular endothelium to the brain parenchyma by a process consisting of cell adhesion, diapedesis and cell migration30. Intravascularly delivered neural and mesenchymal stem cells have been reported to cross BBB and migrate into the brain tissue by various mechanisms31,32,33. Interestingly, according to a recent report, stem cells may use a peculiar type of extravasation, referred to as angiopellosis, in which the stem cell is relatively passive while vascular wall undergoes remodeling to extrude it34. Moreover, new born BBB may be more permeable35,36, though some oppose this idea37. Though this study does not directly address the issue, existing literature suggests that like others, stem cells in the breast milk may also have strategies to cross BBB.
The presence of cells expressing pluripotency stemness markers such as Oct4, SOX2, NANOG, SSEA4 and TRA-1-60 in the breast milk has been shown and they were successfully differentiated into various cell types including neurons in culture4,12. In this study, we detected BMCs in the suckling’s brains that had differentiated both into neuron and glial cell types expressing NeuN and GFAP, respectively. It is noteworthy that these cells were detectable in the brain as early as P7. This is in line with others’ finding that BMCs can pass to the suckling from the beginning of the breast-feeding period before the functional development of the digestive tract is completed3,17. The fact that these cells are still present in the brain 2 months after birth, as Hassiotou et al. also demonstrated this in multiple tissue13, implies that this is not a simple early-life epiphenomenon, though its significance is not clear. Harboring foreign cells from another individual is called microchimerism, which occurs during pregnancy through the placenta, and postnatally by blood transfusion or stem cell, bone marrow, organ and tissue transplantation38,39,40. What we report in this study can be defined as breastfeeding-induced maternal microchimerism and probably has commonality with trans-placental version. The latter has been known for long and shown to involve many organ and tissues from the bone marrow to the heart28,29,41 though no report on the brain involvement has been published so far, which adds to the novelty of our findings. Fetal cells can also pass through the placenta and lead to fetal microchimerism42, which is out of scope of this paper.
Maternal microchimerism is quite common in human and may occur more than one fifth of the population, persists for decades and involves multiple organs and tissues43,44. This study supports the idea that breastfeeding may be partially responsible for this high incidence3. Based on the experimental data and theoretical calculations, daily viable BMCs transfer to a breastfed human baby was estimated to be 4 × 106 to 17 × 109, among which there are substantial number of stem cells3. Biological significance of maternal microchimerism is rather complicated29. Breastfeeding-induced maternal microchimerism is thought to induce immune system maturation and tolerance to non-inherited maternal antigens, which may also explain how these foreign cells persist in the pup tissues without inducing immune response28,45. Indeed, transplanted organ recepients who were breastfed as infants have higher tolerance to tissues from their mothers46,47,48. It is reported that maternal cells may engraft into child’s organs and contribute to the host tissue function as in the case of maternal cells detected in child’s heart as differentiated cardiomyocytes28,49. What significant roles maternal cells in the newborn’s brain may play is an enigmatic question. It has been proposed that they may contribute to tissue maturation and support the host cells by secreting growth factors12. Some authors suggest that breastmilk may cause epigenetic changes in the infant through, for example, exosomal microRNA content and this may even lead to a type of kinship even among non-siblings breastfed by the same female50,51,52,53,54. Engrafted maternal cells may act as a permanent source of exosome – mediated secretions and keep affecting the host tissue epigenetically and/or chemically, even after the weaning.
Maternal microchimerism has been often associated with pathological states especially with autoimmune diseases, notably systemic sclerosis, systemic lupus erythematosus and neonatal lupus syndrome55,56. Elevated levels of circulating maternal cells in the offspring are associated with type 1 diabetes57,58. Potential involvement of chimeric brain cells with diseases, especially of autoimmune nature should be considered and investigated. On the other hand, some authors emphasize the potential of breastmilk to easily and non-invasively harvest stem cells for emerging cell replacement therapies for various genetic and acquired diseases and injuries3,4,12.
In conclusion, this study has firmly established that BMCs may lead to microchimerism in the brain of mouse sucklings, while spectrum of implications remains to be elucidated.
Materials and Methods
Experimental Groups
All animal experiments were reviewed and approved by the Animal Research Ethics Committee of Istanbul Medipol University. All procedures were conducted in conformity with institutional guidelines that are in compliance with European Economic Community Council Directive 86/609. C57BL/6-Tg(CAG-EGFP)1Osb/J mice were obtained from the Jackson Laboratory (ME, USA, #003291). This mouse line, with an enhanced GFP (EGFP) cDNA under the control of a chicken beta-actin promoter and cytomegalovirus enhancer, have widespread EGFP fluorescence, with the exception of erythrocytes and hair, referred to as GFP+. Animals were housed in temperature, water, and humidity-controlled cages that alternated between 12 h light and dark cycles.
The basic experimental paradigm of the study was to let GFP+ mice breastfeed wildtype C57BL/6 (WT) pups and to track GFP+ cells in the brain tissue of pups at different time points (Fig. 1A). For this purpose, two main groups were formed: (1) Newborn WT pups were nursed by GFP+ dams (n = 24, all WT pups were nursed by 6 GFP+ mice), (2) WT pups were nursed by WT dams as a negative control (n = 12). Brain and blood samples from fostered pups were collected at 1-week (n = 12) and 2-months (n = 12) of nursing for experimental assays. 6 mice in each group were used for flow cytometric and real time PCR analysis while 6 mice were used for immunohistochemical staining and microscopic evaluations. Flow cytometry and real time PCR analysis were performed immediately after the animals were sacrificed. For microscopic evaluations, mice were transcardially perfused by 10 ml PBS followed by 10 ml 4% paraformaldehyde (PFA). After perfusion, brain samples were dissected and split into two parts by coronally cutting into 2 mm pieces; one part was processed for tissue clearing method as described below, the other part was incubated in 20% sucrose-PBS until samples sank at the bottom of the tube, embedded in OCT compound (Tissue-Tek®) and stored at −80 °C until sectioning.
Isolation of Cells from Bloodstream and Brain
Cells were isolated from blood and brain samples for flow cytometric analysis to identify GFP+ cells in the tissues.
Blood samples were lysed to remove erythrocytes. Briefly, 2 ml of BD Pharm Lyse ™ (BD Biosciences, 555899) was added to 200 μl of blood sample and incubated for 15 min at RT. Samples centrifuged at 200 g for 5 min and supernatant was discarded. then 2 ml PBS was added onto the pellet and centrifuged at 200 g for 5 min. After the supernatant was discarded, 500 μl PBS was added onto the pellet to obtain cells.
Brain tissues were divided into pieces with a thickness of 1 mm with a razor blade and taken into a Hibernate-A (ThermoFisher Scientific, A1247501) solution. For enzymatic digestion, they were incubated with 4 mg/ml papain (Sigma-Aldrich, P3125) for 1 h at 30 °C and then triturated. After centrifugation for 5 min at 500 G, the supernatant was discarded and cells were obtained in pellet. Obtained cells were fixed with BD Cytofix/Cytoperm (BD Biosciences, 554722).
Flow Cytometry
Isolated cells were first treated with blocking solution (3% bovine serum albumin, 0.3% sodium azide, 1% goat serum, 0.1% Triton X-100) to block Fc receptors. They were then labeled with Alexa Fluor conjugated anti-GFP antibodies to confirm and enhance GFP signals. Subsequently, each sample was stained with DAPI or TO-PRO3 and then analyzed by flow cytometry. Flow cytometry was performed with BD Influx™ Cell Sorter (BD Biosciences). Analysis and absolute cell count were performed after acquiring at least 50,000 events. Doublet and multiplet discrimination were performed by using side scatter (SSC)-W vs. SSC-H and forward scatter (FSC)-W vs. FSC-H plots. Recorded data were analyzed using FlowJo (Tree Star) software.
Gating Strategy
The first orientating gate was selected using FSC vs. SSC plot to exclude particles and debris smaller than the cells were excluded from the analysis. Subsequently, DAPI or TO-PRO3 vs. FSC plots were used to select intact cells with nuclei. After that, respectively, GFP and anti-GFP vs. FSC plots were used to identify the cells which was transferred through breast milk of GFP+ foster mothers to WT pups. Identified cells were sorted in separate tubes from the samples for microscopic evaluation.
DNA Extraction from Brain Tissue
Brain tissues of 1-week and 2-months old pups were collected and total genomic DNA were isolated using Qiagen DNeasy Tissue Kit (Qiagen, Valencia, CA, USA) by applying protocol provided by the manufacturer. DNA concentrations were determined using Qubit dsDNA HS Assay Kit (Thermo Scientific, Q32851).
Real-Time PCR
Real-time PCR was performed to determine the copy number of GFP+ cells in DNA lysates isolated from brain tissues of the pups. To determine the exact copy number of GFP+ cells, standard curve changing between 2 to 2 × 105 copy number/µl was constructed with serially diluted pEGFP-N3 plasmids which is a kind gift from Dr. Fahri Akbaş. For ng/µl to copy number/µl conversion of pEGFP-N3 plasmid, formula below is used as Joshi et al. reported59.
Equation used for ng/µl to copy number/µl conversion (n: DNA length, bp: base pair, m: mass, average molecular weight of 1 mole dsDNA molecule: 660 g/mole).
For achieving high sensitivity to detect single copy number of genomic DNA, probe based real-time PCR was performed. Probe and primer sequences specific to C57BL/6-Tg(CAG-EGFP)1Osb/J mice strain were obtained from Jackson Laboratory and synthesized in Ella Biotech. Probe, forward and reverse primer sequences were 5′-Fam-TTC AAG TCC GCC ATG CCC GAA-Tamra-3′, 5′-AGT GCT TCA GCC GCT ACC-3′, 5′-GAA GAT GGT GCG CTC CTG-3′ respectively. Each PCR reaction was conducted as triplicate in 20 µl volume using Fast Probe Master Mix (Biotium, 31005) with Bio-Rad CFX Connect™ (Bio Rad, CA, USA). Non-template, negative and positive controls were included in all experiments. Transgene copy numbers in each DNA lysate were calculated from the measured CT values using the copy number-CT equation depicted from the standard curve.
Immunohistochemistry
Brains were cut using a cryostat (Leica Biosystems, CM 1850) (coronal sections of 20 µm thickness) and mounted on positively charged glass slides (ThermoFisher Scientific, 10143352). Sections were fixed in 4% paraformaldehyde (PFA)/0.1 mol/L phosphate buffered saline (PBS) for 30 mins, washed and immersed for 1 h in blocking solution (0.1 mol/L PBS containing 0.3% Triton X-100 (PBS-T)/10% normal goat serum). Sections were incubated overnight (o/n) at 4 °C with Alexa Fluor 555-conjugated monoclonal mouse anti-NeuN (Merck Millipore, MAB377A5, 1:500), DyLight 550-conjugated monoclonal Mouse anti-GFAP (Novus Biologicals, NBP2-33184R, 1:1000) and Alexa Fluor 405-conjugated rabbit polyclonal anti-GFP (Novus Biologicals, NB600-310AF405, 1:100). After that, sections were washed with PBS, counterstained with TO-PRO®3 (ThermoFisher Scientific, T3605) and analyzed using a laser scanning confocal microscope (LSM 780, Carl Zeiss, Jena, Germany).
Whole Tissue Immunohistochemical Staining and Clearing
Tissue clearing and immunohistochemically staining method were used for brain samples as described60. Briefly, dissected brains were coronally cut into 2 mm pieces. Brain slices were fixed in 4% PFA at 4 °C o/n with shaking and washed in PBS for 30 mins at room temperature (RT). Before the immunohistochemical staining, sections were pretreated with methanol/H2O series: 20%, 40%, 60%, 80%, 100%; 1 h each. After that, sections were incubated in 66% dichloromethane (Sigma Aldrich, 270997, DCM)/33% methanol at RT. Then, the samples were washed twice in 100% methanol at RT and chilled at 4 °C. Sections were bleached in chilled fresh 5% H2O2 in methanol o/n at 4 °C and rehydrated with methanol/H2O series: 80%, 60%, 40%, 20%, PBS; 1 h each at RT. After pretreatment, sections were incubated at 37 °C and permeabilized in PBS-T for one day and blocked in blocking solution for another day. Subsequently, they were incubated with Alexa Fluor 555 or DyLight 550 conjugated primary antibodies (anti-NeuN and anti-GFAP) for 2 days at 37 °C. Washed in PBS-T for 4–5 times until the next day and counter stained with TO-PRO®3.
After immunohistochemical staining, sections were cleared for further microscopic investigations. Sections were dehydrated in methanol/H2O series: 20%, 40%, 60%, 80%, 100%; 1 h each and incubated in 66% DCM/33% methanol for 3 h at RT. Then they were incubated in 100% DCM for 15 mins twice to wash methanol. Finally, sections were incubated in DiBenzyl Ether (Sigma Aldrich, 108014, DBE) until the sections were optically transparent. After the tissue clearing, brain samples directly imaged in laser scanning confocal microscope in a chamber filled with DBE.
Statistics
The Graph Pad Prism 6 (GraphPad Software Inc., CA, USA) was used for statistical analysis. Student’s t test, one-way ANOVA and Pearson’s correlation analyses were used where appropriate.
References
Walker, A. Breast milk as the gold standard for protective nutrients. The Journal of pediatrics 156, S3–7 (2010).
Hassiotou, F. et al. Maternal and infant infections stimulate a rapid leukocyte response in breastmilk. Clinical & translational immunology 2, e3 (2013).
Hassiotou, F., Geddes, D. T. & Hartmann, P. E. Cells in human milk: state of the science. Journal of human lactation: official journal of International Lactation Consultant Association 29, 171–182 (2013).
Hassiotou, F. et al. Breastmilk is a novel source of stem cells with multilineage differentiation potential. Stem cells (Dayton, Ohio) 30, 2164–2174 (2012).
Visvader, J. E. Keeping abreast of the mammary epithelial hierarchy and breast tumorigenesis. Genes & development 23, 2563–2577 (2009).
Kordon, E. C. & Smith, G. H. An entire functional mammary gland may comprise the progeny from a single cell. Development 125, 1921–1930 (1998).
Fan, Y., Chong, Y. S., Choolani, M. A., Cregan, M. D. & Chan, J. K. Unravelling the mystery of stem/progenitor cells in human breast milk. PloS one 5, e14421 (2010).
Cregan, M. D. et al. Identification of nestin-positive putative mammary stem cells in human breastmilk. Cell and tissue research 329, 129–136 (2007).
Cabinian, A. et al. Transfer of Maternal Immune Cells by Breastfeeding: Maternal Cytotoxic T Lymphocytes Present in Breast Milk Localize in the Peyer’s Patches of the Nursed Infant. PloS one 11, e0156762 (2016).
Zhou, L. et al. Two independent pathways of maternal cell transmission to offspring: through placenta during pregnancy and by breast-feeding after birth. Immunology 101, 570–580 (2000).
Arvola, M. et al. Immunoglobulin-secreting cells of maternal origin can be detected in B cell-deficient mice. Biology of reproduction 63, 1817–1824 (2000).
Twigger, A. J., Hodgetts, S., Filgueira, L., Hartmann, P. E. & Hassiotou, F. From breast milk to brains: the potential of stem cells in human milk. Journal of human lactation: official journal of International Lactation Consultant Association 29, 136–139 (2013).
Hassiotou, F., Mobley, A., Geddes, D., Hartmann, P. & Wilkie, T. Breastmilk Imparts the Mother’s Stem Cells to the Infant. The FASEB Journal 29(876), 878 (2015).
Ghosh, M. K., Nguyen, V., Muller, H. K. & Walker, A. M. Maternal Milk T Cells Drive Development of Transgenerational Th1 Immunity in Offspring Thymus. Journal of immunology (Baltimore, Md.: 1950) 197, 2290–2296 (2016).
Kulinich, A. & Liu, L. Human milk oligosaccharides: The role in the fine-tuning of innate immune responses. Carbohydrate research 432, 62–70 (2016).
Yesildal, F., Koc, E., Tas, A. & Ozgurtas, T. Angiopoietins in Human Breast Milk. Breastfeeding medicine the official journal of the Academy of Breastfeeding Medicine 11, 366–369 (2016).
Weiler, I. J., Hickler, W. & Sprenger, R. Demonstration that milk cells invade the suckling neonatal mouse. American journal of reproductive immunology AJRI: official journal of the American Society for the Immunology of Reproduction and the International Coordination Committee for Immunology of Reproduction 4, 95–98 (1983).
Dixon, D. L. & Forsyth, K. D. Leukocytes in expressed breast milk of asthmatic mothers. Allergologia et immunopathologia (2016).
Jansen, M. A. et al. Decreased memory B cells and increased CD8 memory T cells in blood of breastfed children: the generation R study. PloS one 10, e0126019 (2015).
Jakaitis, B. M. & Denning, P. W. Human Breast Milk and the Gastrointestinal Innate Immune System. Clinics in Perinatology 41, 423–435 (2014).
Tatematsu, M., Takahashi, M., Tsuda, H., Hirose, M. & Furihata, C. Precocious differentiation of immature chief cells in fundic mucosa of infant rats induced by hydrocortisone. Cell differentiation 4, 285–294 (1975).
Ma, L. J., Walter, B., Deguzman, A., Muller, H. K. & Walker, A. M. Trans-epithelial immune cell transfer during suckling modulates delayed-type hypersensitivity in recipients as a function of gender. PloS one 3, e3562 (2008).
Cummins, A. G., Steele, T. W., LaBrooy, J. T. & Shearman, D. J. Maturation of the rat small intestine at weaning: changes in epithelial cell kinetics, bacterial flora, and mucosal immune activity. Gut 29, 1672–1679 (1988).
Jain, L. et al. In vivo distribution of human milk leucocytes after ingestion by newborn baboons. Archives of disease in childhood 64, 930–933 (1989).
Michie, C. A. The long term effects of breastfeeding: a role for the cells in breast milk? J Trop Pediatr 44, 2–3 (1998).
Tuboly, S., Bernath, S., Glavits, R., Kovacs, A. & Megyeri, Z. Intestinal absorption of colostral lymphocytes in newborn lambs and their role in the development of immune status. Acta Vet Hung 43, 105–115 (1995).
Schnorr, K. L. & Pearson, L. D. Intestinal absorption of maternal leucocytes by newborn lambs. Journal of reproductive immunology 6, 329–337 (1984).
Dutta, P. & Burlingham, W. J. Stem cell microchimerism and tolerance to non-inherited maternal antigens. Chimerism 1, 2–10 (2010).
Stevens, A. M. Maternal microchimerism in health and disease. Best Pract Res Clin Obstet Gynaecol 31, 121–130 (2016).
Wilson, E. H., Weninger, W. & Hunter, C. A. Trafficking of immune cells in the central nervous system. The Journal of clinical investigation 120, 1368–1379 (2010).
Andres, R. H. et al. The CCR2/CCL2 interaction mediates the transendothelial recruitment of intravascularly delivered neural stem cells to the ischemic brain. Stroke 42, 2923–2931 (2011).
Diaz-Coranguez, M. et al. Transmigration of neural stem cells across the blood brain barrier induced by glioma cells. PloS one 8, e60655 (2013).
Matsushita, T. et al. Mesenchymal stem cells transmigrate across brain microvascular endothelial cell monolayers through transiently formed inter-endothelial gaps. Neurosci Lett 502, 41–45 (2011).
Allen, T. A. et al. Angiopellosis as an Alternative Mechanism of Cell Extravasation. Stem cells (Dayton, Ohio) 35, 170–180 (2017).
Kniesel, U., Risau, W. & Wolburg, H. Development of blood-brain barrier tight junctions in the rat cortex. Brain Res Dev Brain Res 96, 229–240 (1996).
Risau, W. & Wolburg, H. Development of the blood-brain barrier. Trends in neurosciences 13, 174–178 (1990).
Ek, C. J., Dziegielewska, K. M., Habgood, M. D. & Saunders, N. R. Barriers in the developing brain and Neurotoxicology. Neurotoxicology 33, 586–604 (2012).
Gammill, H. S. & Nelson, J. L. Naturally acquired microchimerism. The International journal of developmental biology 54, 531–543 (2010).
Stikvoort, A. et al. Long-Term Stable Mixed Chimerism after Hematopoietic Stem Cell Transplantation in Patients with Non-Malignant Disease, Shall We Be Tolerant? PloS one 11, e0154737 (2016).
Konuma, T. et al. Early phase mixed chimerism in bone marrow does not affect long-term outcomes of myeloablative single-unit cord blood transplantation for adult patients with hematological malignancies. Leukemia & lymphoma 1–7 (2016).
Tuffrey, M., Bishun, N. P. & Barnes, R. D. Porosity of the mouse placenta to maternal cells. Nature 221, 1029–1030 (1969).
Tan, K. H., Zeng, X. X., Sasajala, P., Yeo, A. & Udolph, G. Fetomaternal microchimerism: Some answers and many new questions. Chimerism 2, 16–18 (2011).
Maloney, S. et al. Microchimerism of maternal origin persists into adult life. The Journal of clinical investigation 104, 41–47 (1999).
Lo, Y. M., Lau, T. K., Chan, L. Y., Leung, T. N. & Chang, A. M. Quantitative analysis of the bidirectional fetomaternal transfer of nucleated cells and plasma DNA. Clinical chemistry 46, 1301–1309 (2000).
Moles, J. P. et al. Breastmilk cell trafficking induces microchimerism-mediated immune system maturation in the infant. Pediatr Allergy Immunol 29, 133–143 (2018).
Zhang, L., van Bree, S., van Rood, J. J. & Claas, F. H. Influence of breast feeding on the cytotoxic T cell allorepertoire in man. Transplantation 52, 914–916 (1991).
Hanson, L. A. The mother-offspring dyad and the immune system. Acta paediatrica (Oslo, Norway: 1992) 89, 252–258 (2000).
Dutta, P. & Burlingham, W. J. Microchimerism: tolerance vs. sensitization. Current opinion in organ transplantation 16, 359–365 (2011).
Stevens, A. M., Hermes, H. M., Rutledge, J. C., Buyon, J. P. & Nelson, J. L. Myocardial-tissue-specific phenotype of maternal microchimerism in neonatal lupus congenital heart block. Lancet 362, 1617–1623 (2003).
Ozkan, H., Tuzun, F., Kumral, A. & Duman, N. Milk kinship hypothesis in light of epigenetic knowledge. Clin Epigenetics 4, 14 (2012).
Melnik, B. C. et al. Milk miRNAs: simple nutrients or systemic functional regulators? Nutr Metab (Lond) 13, 42 (2016).
Melnik, B. C., John, S. M., Carrera-Bastos, P. & Schmitz, G. Milk: a postnatal imprinting system stabilizing FoxP3 expression and regulatory T cell differentiation. Clin Transl Allergy 6, 18 (2016).
Melnik, B. C., John, S. M. & Schmitz, G. Milk is not just food but most likely a genetic transfection system activating mTORC1 signaling for postnatal growth. Nutr J 12, 103 (2013).
Melnik, B.C. & Schmitz, G. Milk’s Role as an Epigenetic Regulator in Health and Disease. Diseases 5 (2017).
Adams, K. M. & Nelson, J. L. Microchimerism: an investigative frontier in autoimmunity and transplantation. Jama 291, 1127–1131 (2004).
Nelson, J. L. et al. Microchimerism and HLA-compatible relationships of pregnancy in scleroderma. Lancet 351, 559–562 (1998).
Ye, J., Vives-Pi, M. & Gillespie, K. M. Maternal Microchimerism: Increased in the Insulin Positive Compartment of Type 1 Diabetes Pancreas but Not in Infiltrating Immune Cells or Replicating Islet Cells. PloS one 9, e86985 (2014).
Nelson, J. L. et al. Maternal microchimerism in peripheral blood in type 1 diabetes and pancreatic islet beta cell microchimerism. Proceedings of the National Academy of Sciences of the United States of America 104, 1637–1642 (2007).
Joshi, M., Keith Pittman, H., Haisch, C. & Verbanac, K. Real-time PCR to determine transgene copy number and to quantitate the biolocalization of adoptively transferred cells from EGFP-transgenic mice. BioTechniques 45, 247–258 (2008).
Erturk, A., Lafkas, D. & Chalouni, C. Imaging cleared intact biological systems at a cellular level by 3DISCO. Journal of visualized experiments: JoVE (2014).
Acknowledgements
This project was supported by “The Scientific and Technological Research Council of Turkey” (TUBITAK) with a grant number 114R078.
Author information
Authors and Affiliations
Contributions
M.S.A., and G.O. designed experiments and wrote the manuscript. M.S.A., E.E. and G.O. analyzed data, interpreted results. M.SA., and E.N.Y., performed the experiment model, real time PCR analysis and microscopic imaging. E.V. performed flow cytometric analysis.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aydın, M.Ş., Yiğit, E.N., Vatandaşlar, E. et al. Transfer and Integration of Breast Milk Stem Cells to the Brain of Suckling Pups. Sci Rep 8, 14289 (2018). https://doi.org/10.1038/s41598-018-32715-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32715-5
Keywords
This article is cited by
-
Human breast tissue engineering in health and disease
EMBO Molecular Medicine (2024)
-
Feasibility of intranasal human milk as stem cell therapy in preterm infants with intraventricular hemorrhage
Journal of Perinatology (2024)
-
The cellular and immunological dynamics of early and transitional human milk
Communications Biology (2023)
-
Bovine milk-derived cells express transcriptome markers of pluripotency and secrete bioactive factors with regenerative and antimicrobial activity
Scientific Reports (2023)
-
Regenerative Potential of Human Breast Milk: A Natural Reservoir of Nutrients, Bioactive Components and Stem cells
Stem Cell Reviews and Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.