Abstract
RNA-induced silencing complex (RISC) plays a critical role in small interfering RNA (siRNA) and microRNAs (miRNA) pathways. Accumulating evidence has demonstrated that the major RISC members (AGO, DICER, TRBP, PACT and GW182) represent expression discrepancies or multiple orthologues/paralogues in different species. To elucidate their evolutionary characteristics, an integrated evolutionary analysis was performed. Here, animal and plant AGOs were divided into three classes (multifunctional AGOs, siRNA-associated AGOs and piRNA-associated AGOs for animal AGOs and multifunctional AGOs, siRNA-associated AGOs and complementary functioning AGOs for plant AGOs). Animal and plant DICERs were grouped into one class (multifunctional DICERs) and two classes (multifunctional DICERs and siRNA-associated DICERs), respectively. Protista/fungi AGOs or DICERs were specifically associated with the siRNA pathway. Additionally, TRBP/PACT/GW182 were identified only in animals, and all of them functioned in the miRNA pathway. Mammalian AGOs, animal DICERs and chordate TRBP/PACT were found to be monophyletic. A large number of gene duplications were identified in AGO and DICER groups. Taken together, we provide a comprehensive evolutionary analysis, describe a phylogenetic tree-based classification of the major RISC members and quantify their gene duplication events. These findings are potentially useful for classifying RISCs, optimizing species-specific RISCs and developing research model organisms.
Similar content being viewed by others
Introduction
RNA-induced silencing complex (RISC) plays an important role in small interfering RNA (siRNA)- and microRNA (miRNA)-mediated gene regulation1,2,3,4,5,6. The siRNA pathway regulates target gene expression, provides antiviral responses, and restricts transposons. The miRNA pathway represses target gene expression at the post-transcriptional level and participates in physiological and pathophysiological processes4,5,6. The siRNA/miRNA have found widespread applications as research and clinical techniques, allowing simple yet effective knockdown of target genes of interest and disease therapy7. Further, miRNAs are important biomarkers for disease diagnosis and prognosis8.
The major members of RISC have been identified and characterized, including AGO (argonaute), DICER, TRBP (TARBP2), PACT (PRKRA) and GW182 (GAWKY). The crystal structure of the AGO protein from Pyrococcus furiosus at 2.25 Å resolution has been resolved9. Dicer, a member of the RNase III family, processes miRNA precursors (pre-miRNAs) to mature miRNAs. Recent studies have demonstrated that AGO/DICER are not required for guide hairpin RNA function10,11. Within the RISC loading complex, DICER/TRBP assists in processing pre-miRNAs to mature miRNAs and then load them onto AGO2. AGO2 bound to the mature miRNA constitutes the minimal RISC and may subsequently dissociate from DICER and TRBP. TRBP/PACT found in RISC binds to double-stranded RNAs (dsRNAs) and ensures regular miRNA biogenesis12,13. GW182 directly interacts with AGO2, and GW182 proteins are essential for miRNA-guided gene silencing in various organisms. The N-terminal region of GW182 contains multiple GW repeats, which directly associate with an AGO214.
Previously, few phylogenetic analyses of RISC members have been performed. Several conclusions are summarized as follows: (I) vertebrates lack siRNA-class AGO proteins and vertebrate AGOs display low rates of molecular evolution15; (II) Dicers might have duplicated and diversified independently and have evolved for various functions in invertebrates16; (III) Loquacious identified in insects may be ancestral to both TRBP and PACT; and (IV) significant acceleration in the accumulation of amino acid changes of GW182-binding regions indicates its early origin and adaptive evolution17. However, a systematic, integrative evolutionary analysis is still lacking. In our study, we applied improved and integrated bioinformatic softwares/algorithms for investigating the evolution of the major RISC members, and these results provide a novel insight into RISC evolution and RISC-mediated gene regulation.
Results
Evolution of RISC-related AGO members
The AGO protein family plays a central role in RISC-mediated gene regulation. AGOs mainly involve four characteristic domains: an N-terminal, PAZ (which is responsible for small RNA binding), Mid and a C-terminal PIWI (which confers catalytic activities) domain (Supplementary Figs S1 and S2)18,19. A number of noncoding RNAs are their substrates, including miRNAs, siRNAs and Piwi-interacting RNAs (piRNAs)20. Small RNAs guide AGOs to their specific targets through sequence complementarity, which typically leads to silencing of the target mostly by post-transcriptional inhibition or mRNA degradation.
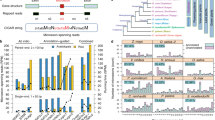
AGO2 and its homologues have been identified in Chordata, Arthropoda, Nematoda and Platyhelminthes, and a phylogenetic tree of AGOs was constructed (Fig. 1A and Supplementary Fig. S3). According to their RNA-binding characteristics or functions, these AGOs were divided into three classes: (I) multifunctional AGOs; (II) siRNA-associated AGOs; and (III) piRNA-associated AGOs. Class I AGOs contain chordate AGO1–4, in which all human AGOs associate with both siRNAs and miRNAs. As shown in Fig. 1A, chordate AGO1-4 contain two subclasses: AGO2 and AGO1/3/4. Only AGO2 protein functions as an endonuclease, cleaving mRNA within regions that base pair with perfectly complementary siRNAs or miRNAs. AGO1/3/4 are slicing-incompetent AGOs21,22 and remove passenger strands via the bypass mechanism. Class II AGOs specifically bind to siRNAs, which contain Arthropoda (Drosophila AGO2) and Nematoda AGOs (TAG-76/ERGO1/WAGO1/YQ53/NRDE3). AGOs in Arthropoda and Nematoda have evolutionary complexities compared with the monophyletic group of mammalian AGOs (Fig. 1A and Supplementary Fig. S3). Drosophila AGO2, as a part of the RISC complex, is required for the unwinding of siRNA duplex and subsequent assembly of siRNA into RISC in Drosophila embryos. Drosophila AGO2 shared a close evolutionary relationship with plant/Chordata AGOs (Supplementary Fig. S3). AGO1 is dispensable for efficient RNAi in Drosophila embryos23, but it is unreviewed in UniProt and unannotated in Fig. 1A. The phylogenetic tree and Pfam/SMART-based domain structures show that TAG-76 (Caenorhabditis elegans, C. elegans) has the modest similarities with mammalian AGOs (Supplementary Fig. S2). Unfortunately, its biological function is still unclear, but other Nematoda proteins participate in the RNAi pathway. ERGO-1 serves as an AGO and functions in the endogenous RNAi pathway24. WAGO1 is a worm-specific AGO and silences certain genes, transposons, pseudogenes, and cryptic loci25. Uncharacterized YQ53 with PAZ and PIWI domains shown in Supplementary Fig. S2 has a possible function of endogenous and exogenous RNAi18. Another nuclear AGO protein, NRDE3, binds to siRNAs and is required for nuclear RNAi, and thus transports specific classes of small regulatory RNAs to distinct cellular compartments to regulate gene expression26. Class III AGOs are composed of chordate PIWIL1-4, Arthropoda AGO3/AUB/ PIWI/SIWI and Platyhelminthes PIWIL/PIWI1/PIWI2. PIWIL1-4, the members of the PIWI family, bind to piRNAs and are exclusively expressed in germ-line cells, but other AGOs are ubiquitously expressed in most tissues27. In addition, Supplementary Fig. S3 shows that other Arthropoda AGO2-related proteins are close to the subclade of chordate PIWIL1-4, and they are directed by piRNAs to cleave transposon transcripts and instruct Piwi to suppress transposon transcription to protect the germline genome in Drosophila ovarian germ cells. The members of the PIWI protein family (PIWIL, PIWI1 and PIWI2) have also been identified in Platyhelminthes (Dugesia japonica and Schmidtea mediterranea). They have domains similar to those of animal AGOs and may be required for stem cell function and piRNA biogenesis (Supplementary Figs S2 and S3)28,29.
Previous studies have determined three clades of AGOs in plants30. For example, Arabidopsis thaliana (A. thaliana) exhibits an equal distribution of its ten members within three clades: (I) AGO1, AGO5, AGO10; (II) AGO2, AGO3, AGO7; and (III) AGO4, AGO6, AGO8, AGO9. Consistent with these results, our analyses showed three major clades, and these plant AGOs were artificially grouped into three classes based on their functions: (I) multifunctional AGOs; (II) siRNA-associated AGOs; and (III) complementary functioning AGOs. However, protein sequence identity-based BLAST scores do not provide the more precise classifications (Fig. 1A and Supplementary Fig. S3). Class I AGOs are AGO1/AGO5/AGO10 (A. thaliana) and AGO1A/AGO1B/AGO1C/AGO1D/AGO11/AGO12/AGO13/AGO14/AGO17/AGO18/PNH1/MEL1 (Oryza sativa, O. sativa). AGO1 acts in the miRNA and siRNA pathways, and AGO10 possibly participates in these pathways, at least in some tissues; AGO5, localized in both the nucleus and the cytoplasm, binds to small RNAs and regulates RNA-mediated post-transcriptional gene silencing31. PNH1 (O. sativa) probably influences the formation of the shoot apical meristem and leaf adaxial cell specification and the RNA silencing pathway32,33. MEL1 probably mediates small RNA-related gene silencing and its function may be performed by another AGO in Arabidopsis33. Class II AGOs contain AGO2/AGO3/AGO7 (A. thaliana) and AGO2/AGO3/AGO7(O. sativa). AGO7 acts in the RDR6/SGS3/DCL4/AGO7 trans-acting siRNA pathway involved in leaf developmental timing. AGO2/3 lack the DDH motif in the PIWI domain and possibly have similar activities; and AGO2/AGO3 mutants show no developmental defects. Class III AGOs contain AGO4/AGO6/AGO8/AGO9 (A. thaliana) and AGO4A/AGO4B/AGO15/AGO16 (O. sativa). These AGOs have complementary biological functions. AGO4 is possibly shared in different nuclear complexes and in a distinct pathway34. AGO6 is required for heterochromatin siRNA and transcriptional gene silencing pathways, and AGO6 activity is partially redundant with the activity of AGO4. AGO8/9 (A. thaliana) mRNAs have a different tissue distribution, and their mutants have no effect on plant phenotypes. Based on sequence identities, there is a clear group of Class I members (orange), but there is an overlapping region between Class II (green) and Class III members (dark green) (Fig. 1A). AtAGOs have been studied for decades, but many questions still remain. OsAGOs appear to have higher diversities and gene duplications because many isoforms were identified as shown in Fig. 1A and Supplementary Fig. S3.
A single AGO containing PIWI domains is identified in the Protista Giardia intestinalis (Giardia lamblia) genome and regulates variant-specific surface protein (VSP) expression on the surface of each parasite via an RNAi pathway (Supplementary Figs S2 and S3)35. Surprisingly, a single AGO1 with PAZ and Piwi domains from yeast was found in fungi and closely located at the third clade of plants (Supplementary Figs S2 and S3). It may possess sequence specific DNA binding activity, and its detailed role in the RISC could be explored in depth.
Evolution of RISC-related Dicer members
The RNAi process arises from an interaction between RNA molecules and RISC. Dicer anchors a dsRNA molecule and cuts it to produce short dsRNAs as a primary RNA recognition and processing enzyme in the RNAi process. Dicer is a member of the RNase III family and highly conserved in the evolution. Sequence analyses show that Dicer of each species has similar domains. In most species, the N-terminus of Dicer is an RNA helicase domain followed by a PAZ domain (Supplementary Fig. S1). The C-terminus of Dicer has two RNase III domains and a dsRNA binding domain36.
Dicers widely exist in eukaryotes. The current phylogenetic tree of the DICER family shows its independent diversification in animals, plants and fungi (Fig. 1B and Supplementary Fig. S4). Distinct with the phylogenetic tree of animal AGOs, a monophyletic group of animal DICER was found to include Chordata, Arthropoda and Nematoda and one class was functionally generated: multifunctional DICERs. These DICERs possess the dual function of recognizing a hairpin or dsRNA and processing them into mature miRNA-miRNA*/siRNA duplexes. It is noteworthy that Arthropoda DICER (Drosophila DCR1) is required for miRNA biogenesis, and the unreviewed Drosophila DICER paralogue DCR2 in UniProt may be for generating siRNAs. The helicase motifs of Nematoda DICER (C. elegans DCR1) are required for siRNA, but not miRNA, processing37,38,39,40. Dicer required for piRNA processing remains unidentified. The additional constructed chordate subclade includes DDX58, DHX58, IFIH1, FANCM and RNC (Fig. 1B and Supplementary Fig. S4). They all have helicase activities, but DDX58 and DHX58 bind to DNA, and IFIH1 and FANCM have RNA binding affinities. Functionally, they could participate in RISC-like complexes and respond to exogenous stress. RNC encodes dsRNA-specific RNase III.
There are four DICERs (DCL1-4) in plants38. Supplementary Fig. S4 shows a monophyletic group of plant DICERs containing four subclades: DCL1, DCL2, DCL3 and DCL4. Based on the maturation types of plant small RNAs, these DICERs were grouped into two classes: (I) multifunctional DICERs (DCL1); and (II) siRNA-associated DICERs (DCL2-4). DCL1 participates in RISC formation to process miRNA/siRNA precursors. AtDCL2–4 generate siRNAs and are implicated in virus defense and production of siRNAs from natural cis-acting antisense transcripts, chromatin modification guidance or vegetative phase change regulation39,40,41. RTL3 (O. sativa) belongs to the DCL1 subclade and the RNase III family, suggesting that it possibly involves the miRNA/siRNA pathway. Furthermore, more DICER isoforms were found in O. sativa, indicating that a gene duplication event might have occurred during rice DICER evolution. A plant clade locates at outgroup, including RTL3 (A. thaliana) and RTL2 (A. thaliana and O. sativa), which are ribonucleases cleaving dsRNA and producing small RNAs.
Fungi (Ascomycota) Dicers show two subclades supported by a bootstrap value of 64: (I) DCL1, DCR1(Schizosaccharomyces pombe, S. pombe); and (II) DCL2 (Fig. 1B and Supplementary Fig. S4). In vegetative cells, DCL2 is a major Dicer enzyme in the process of siRNA biogenesis, but DCL1 has a redundant role42. However, only DCL1 is specifically expressed and required for meiotic silencing during meiosis43. In the siRNA maturation process, dsRNA precursors are processed by DCR1 with similar domains of canonical DICER in animals. DCR1 (S. pombe) is a Dicer homologue in fission yeast. MPH1 is an ATP-dependent DNA helicase associated with DNA damage response to maintain genome integrity44. Yeast MFH2 has DNA binding and DNA helicase activities. Other clades are mainly from bacterial RNCs (Actinobacteria, Aquificae, Firmicutes, Proteobacteria, Tenericutes and Thermotogae) which encode RNase and cleave RNAs (Fig. 1B and Supplementary Fig. S4).
Evolution of RISC-related TRBP and PACT members
TRBP is implicated in HIV-1 gene expression45, possibly linking miRNAs and the response of the IFN-PKR pathway to HIV-1 infection46. In vertebrates, TRBP is a paralogue to the protein kinase R (PKR)-activating protein or PACT46,47. They regulate PKR as an inhibitor (TRBP) or activator (PACT). RISC-related biological functions of TRBP and/or PACT with three DRSM domains (which bind to dsRNAs and mediate protein-protein interaction), shown in Supplementary Fig. S2, are the following: recruiting substrates to Dicer, facilitating Dicer-mediated processing of immature miRNAs, removing the Dicer product and controlling which type of dsRNA is loaded onto AGOs12,13.
In our phylogenetic trees, TRBPs/PACTs are shown only in Chordata (Fig. 1C,D and Supplementary Figs S5 and S6). Otherwise, Drosophila R2D2 and C. elegans RDE-4, both known participants in RNAi processing, are distantly related to TRBP/PACT and do not emerge in chordate clades of TRBP/PACT. Loquacious has been identified in insects, but they are not considered because of low sequence identity and unreviewed entry in UniProt (Fig. 1C,D). Other evolutionarily related RNA/DNA binding genes are DSRAD (Chordata), STAU1/2/STAUH (Chordata, Mollusca and Arthropoda), ILF3 (Chordata), STRBP (Chordata), RED1/2 (Chordata) and RNC (Chloroflexi and Proteobacteria). DSRADs are involved in RNA editing and facilitate loading of miRNA onto RISC48. Chordate STAU1/2/STAUH seems to function in mRNA transport or distribution49,50, and similar to miRNA precursors, Exportin-5 transports Staufen-dsRNA complexes out of the nucleus51. Chordate ILF3 binds to RNA and functions in the biogenesis of circular RNAs (circRNAs). Chordate STRBP regulates spermatogenesis and sperm function through binding dsDNA/RNA. Chordate RED1 catalyzes A-to-I RNA editing to affect gene expression and function. Chordate RED2 with adenosine deaminase activity and dsRNA/ssRNA binding affinity prevents the binding of other ADAR enzymes to targets and reduces the efficiency of these enzymes. RNCs in Chloroflexi and Proteobacteria belong to RNase III family. Unexpectedly, DRB2 (O. sativa) and IIV6-340R (Invertebrate iridescent virus 6) were located at the clades of chordate DSRAD and bacterial RNCs in our phylogenetic tree of PACT, respectively (Fig. 1D and Supplementary Fig. S6). They possibly bind to and cleave RNAs in plants or their host, but their RISC-related functions are still unknown.
Evolution of RISC-related GW182 members
GW182 identified in animals is a key component of RISC, interacts PIWI domain of AGO1 via its N-terminal region for miRNA-mediated gene regulation (Supplementary Fig. S1) and exhibits high diversities in sequence length, conservation and composition. In this study, the full-length sequences of GW182 and its orthologues were used for a reliable phylogenetic reconstruction, which was consistent with previous results (Supplementary Fig. S7)17. The results indicate that mammalian TNRC6C is the founding member of the chordate gene family representing and diverging from the orthologue of nonchordate GW182 genes.
In our study, a limited number of GW182 were identified and characterized, including human, mouse and fly (Fig. 1E and Supplementary Fig. S7). A further database BLAST search analysis of GW182 mRNA/protein showed no additional homologues with highly convinced sequence similarities and biological functions, even in Nematoda. Instead, two functional analogues, AIN-1 and AIN-2, are encoded in the genome of C. elegans. The observation is not surprising because more than half of the genes encoded in Nematoda are unique52.
Gene duplication analysis of the major RISC members
Gene duplication is a crucial driving force of phenotype diversity, the cause of human diseases, and evolution. In our study, we quantified gene duplication events of the major RISC members. There were 51 gene duplications identified in the tree of AGOs (Supplementary Fig. S8). Class I, II and III of animal AGOs contained 6, 4 and 14 gene duplications, respectively. Class I, II and III of plant AGOs included 13, 3 and 5 gene duplications, respectively. In the tree of DICERs, 61 gene duplications were determined (Supplementary Fig. S9). Of this class of animal DICERs, 4 gene duplications arose from Chordata. In the two classes of plant DICERs, only class II had 6 gene duplications. For TRBP/PACT/GW182, there were 68/27/10 gene duplications, in which the numbers of their closely associated gene duplications were 1, 3 and 2, respectively (Supplementary Figs 10–12). These results suggest that plenty of gene duplications in AGOs and DICERs may be the main contributor to evolutionary diversity.
Discussion
RISC-mediated gene regulation is vital for growth, development and metabolic disorders such as mitochondrial uncoupling proteins-mediated obesity and diabetes53,54,55. RISC is a large protein complex, in which its composition and protein copy numbers apparently affect its function. Previous investigations have demonstrated that RISCs originating from different species have distinct compositions; this phenomenon may mainly result from evolutionary selection pressures and the complexity of RISCs. To explore RISC evolution, phylogenetic trees and gene duplications of the major RISC members (AGOs, DICERs, TRBPs, PACTs and GW182s) were analyzed. These RISC components have various distributions in animals, plants, protista and fungi. TRBP/PACT/GW182 were observed only in animals. The monophyletic groups in the phylogenetic trees were as follows: mammalian AGOs, animal DICERs, chordate TRBP/PACT. RISC compositions in Arthropoda and Nematoda showed the evolutionary complexities, possibly derived from their unique functions.
Recently, Niels Wynant et al. have investigated the evolution of animal AGOs and reported three conserved AGO functional lineages: siRNA-class AGO, miRNA-class AGO and PIWI AGOs15. Here, we provide a refined classification of animal AGOs: multifunctional AGOs, siRNA-associated AGOs and piRNA-associated AGOs. Also, Fig. 1A points out a group of species- and sequence identity-based classes: Chordata (3 subclasses: AGO2, AGO1/3/4 and PIWIL1–4), Arthropoda (2 subclasses: AGO2, AGO3/AUB/PIWI/SIWI), Nematoda (2 subclasses: TAG-76, ERGO1/WAGO1/YQ53/NRDE3) and Platyhelminthes. Homologous prokaryotic AGO proteins were discovered, but their functions are elusive because Archaea and Bacteria are deficiency of RNAi pathway56.
MiRNAs are identified in Giardia intestinalis (G. intestinalis), and they all target the open reading frame, but not the classic 3′-UTRs because their 3′-UTRs of genes are too short57. Therefore, G. intestinalis may generate a special RISC to interact with the miRNAs.
DICERs belong to the RNase III family which recognizes dsRNAs and cleaves them at specific targeted locations. In this study, many RNase IIIs were identified and investigated from eukaryotes to prokaryotes. Based on the structural differences of RNase IIIs, they were grouped into four classes: class 1 (bacteria and bacteriophage), class 2 (fungi), class 3 and class 4. DICER is classified as class 4 RNase III because it possesses both helicase and PAZ domains (Supplementary Fig. S2). A PAZ domain and two RNase III domains from G. intestinalis have been discovered by X-ray crystallography58.
Like function-dependent animal AGOs, plant AGO/DICER families have diversified extensively and appeared to possess plant-specific AGOs/DICERs. The identification of the 5′ terminal nucleotide of plant small RNA and miRNA-duplex structure strongly affect the loading of small RNAs onto specific AGOs. DCL1 processes miRNA, DCL1–4 process hairpin-derived siRNAs, and natural antisense siRNAs are processed by DCL1, DCL2 or DCL3. The precursors of secondary siRNA transcribed by Pol II are trimmed by DCL2/DCL4 to be siRNA. Heterochromatic siRNAs maturation is mediated by DCL3. It is likely that the diversification and specialization of RISC compositions in plants probably represent their roles in adaptation to a sessile lifestyle.
Yeast is miRNA pathway-deficient, but in fission yeast, AGO, DICER and RNA-dependent RNA polymerase factors are identified and used for the RNAi pathway, which are valuable for heterochromatin formation at the centromeres and mating type region. Therefore, fission yeast could be a model for studying the RNAi pathway. Moreover, the synthesis of dsRNAs is mediated by RDRs. The three major clades of eukaryotic RDRs are RDRα, RDRβ and RDRγ. In the fission yeast S. pombe, RDRγ is implicated in transcriptional gene silencing. These results indicate that yeast AGO/DICER/RDRs evolution may be integrated.
The related genes of chordate TRBP/PACT shown in Fig. 1C,D are mostly able to bind to RNA/DNA and have a variety of functions, such as RNA editing, mRNA transport, circRNAs biogenesis and RNA cleavage. These findings indicate that TRBP/PACT possibly perform the biological functions of their evolutionarily related genes in RISC besides the substrates recruitment and loading.
Gene duplication acts as an evolutionary engine so that it constitute a necessary force for evolutionary innovation and provide new genetic materials for new genes. A series of gene duplication events were found in RISC members, including AGOs and DICERs, which mostly existed in higher organisms. Gene duplications may be produced by unequal crossingover, retrotransposition, duplicated DNA transposition and polyploidization, but the molecular mechanisms of gene duplication in the major RISC members still remain ambiguous. Sometimes, gene duplications result in functional redundancies, such as plant complementary functioning AGOs as classified in our study. To minimize the genome sizes, CRISPR system has been widely used for gene editing59,60. The current results may provide assistance in improving or minimizing RISC and the genomes for achieving better efficiency and accuracy.
In summary, we systematically analyzed the evolution of the major RISC members, including AGO, DICER, TRBP, PACT and GW182. The findings provide potential support of species-specific RNAi/miRNA related RISC system optimization and model organism development.
Methods
Protein (amino acid) sequences retrievals
Key protein sequences of the major RISC members (AGO2 Homo sapiens-Q9UKV8, DICER Homo sapiens- Q9UPY3, TRBP2 Homo sapiens- Q15633, PRKRA Homo sapiens- O75569, GAWKY Drosophila melanogaster- Q8SY33) as the queries are used for searching all RISC-related members via BLAST algorithm, and subsequently, all reviewed protein sequences were retrieved from UniProt61,62. Homology cut-offs were E-values ≤0.0001 with the consideration of BLAST scores62. All detailed protein/gene information was listed in Supplementary Tables S1–5.
Multiple sequence alignments
There are 93 (AGOs), 205 (DICERs), 124 (TRBPs) 57 (PACTs) and 15 (GW182s) protein entries used for multiple sequence alignments. Initial multiple sequence alignments were performed using the program MUSCLE (MUltiple Sequence Comparison by Log- Expectation) for reaching better average accuracy and speed63, The default settings (Gap Open:−2.9, Hydrophobicity Multiplier: 1.2, Clustering Methods: UPGMB, Min Diag Length: 24.) in MUSCLE were chosen for best accuracy and speed.
Phylogenetic analysis
Several methods in MEGA7 are capable of constructing phylogenetic trees. In the present study, similar results were obtained from these methods, so the phylogenetic tree constructing by Neighbor-joining method was shown64,65. The percentage of replicate trees evaluated by booststrap test (1,000 replicates) were labeled next to the branches66. Possion correction method computed their evolutionary distances which were in the units of the number of amino acid substitutions per site.
Domain architecture analysis
SMART (a Simple Modular Architecture Research Tool) was applied for identifying, annotating and analyzing domain architecture of the major RISC members67. Protein sequences obtained from UniProt were input into SMART server which searched Pfam domains of all proteins of interest.
Finding gene duplications
Gene duplications were analyzed by “Find gene duplications” module in MEGA764,68. Gene duplications were identified by searching for all branching points in the topology with at least one species present in both subtrees of the branching point. An unrooted gene tree was used for the analysis such that the search for duplication events was performed by finding the placement of the root on a branch or branches that produced the minimum number of duplication events.
References
Castanotto, D. & Rossi, J. J. The promises and pitfalls of RNA-interference-based therapeutics. Nature 457, 426–433 (2009).
Pratt, A. J. & MacRae, I. J. The RNA-induced silencing complex: a versatile gene-silencing machine. J. Biol. Chem. 284, 17897–17901 (2009).
Ambros, V. microRNAs: tiny regulators with great potential. Cell 107, 823–826 (2001).
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Yu, Y., Jia, T. & Chen, X. The ‘how’ and ‘where’ of plant microRNAs. New Phytol. 216, 1002–1017 (2017).
Soifer, H. S., Rossi, J. J. & Saetrom, P. MicroRNAs in disease and potential therapeutic applications. Mol. Ther. 15, 2070–2079 (2007).
Hayes, J., Peruzzi, P. P. & Lawler, S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol. Med. 20, 460–469 (2014).
Song, J. J., Smith, S. K., Hannon, G. J. & Joshua-Tor, L. Crystal structure of Argonaute and its implications for RISC slicer activity. Science 305, 1434–1437 (2004).
Ohno, S. et al. Development of novel small hairpin RNAs that do not require processing by Dicer or AGO2. Mol. Ther. 24, 1278–1289 (2016).
Eckenfelder, A. et al. Argonaute proteins regulate HIV-1 multiply spliced RNA and viral production in a Dicer independent manner. Nucleic Acids Res. 45, 4158–4173 (2017).
Wilson, R. C. et al. Dicer-TRBP complex formation ensures accurate mammalian microRNA biogenesis. Mol. Cell 57, 397–407 (2015).
Heyam, A., Lagos, D. & Plevin, M. Dissecting the roles of TRBP and PACT in double-stranded RNA recognition and processing of noncoding RNAs. RNA 6, 271–289 (2015).
Pfaff, J. et al. Structural features of Argonaute-GW182 protein interactions. Proc. Natl. Acad. Sci. USA 110, E3770–3779 (2013).
Wynant, N., Santos, D. & Vanden Broeck, J. The evolution of animal Argonautes: evidence for the absence of antiviral AGO Argonautes in vertebrates. Sci. Rep. 7, 9230 (2017).
Gao, Z., Wang, M., Blair, D., Zheng, Y. & Dou, Y. Phylogenetic analysis of the endoribonuclease Dicer family. PloS One 9, e95350 (2014).
Zielezinski, A. & Karlowski, W. M. Early origin and adaptive evolution of the GW182 protein family, the key component of RNA silencing in animals. RNA Biol. 12, 761–770 (2015).
Hutvagner, G. & Simard, M. J. Argonaute proteins: key players in RNA silencing. Nat. Rev. Mol. Cell Biol. 9, 22–32 (2008).
Frank, F., Sonenberg, N. & Nagar, B. Structural basis for 5’-nucleotide base-specific recognition of guide RNA by human AGO2. Nature 465, 818–822 (2010).
Hansen, T. B. et al. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 30, 4414–4422 (2011).
Liu, J. et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441 (2004).
Meister, G. et al. Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 15, 185–197 (2004).
Williams, R. W. & Rubin, G. M. ARGONAUTE1 is required for efficient RNA interference in Drosophila embryos. Proc. Natl. Acad. Sci. USA 99, 6889–6894 (2002).
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Gu, W. et al. Distinct argonaute-mediated 22G-RNA pathways direct genome surveillance in the C. elegans germline. Mol. Cell 36, 231–244 (2009).
Guang, S. et al. An Argonaute transports siRNAs from the cytoplasm to the nucleus. Science 321, 537–541 (2008).
Han, B. W. & Zamore, P. D. piRNAs. Curr. Biol. 24, R730–733 (2014).
Palakodeti, D., Smielewska, M., Lu, Y. C., Yeo, G. W. & Graveley, B. R. The PIWI proteins SMEDWI-2 and SMEDWI-3 are required for stem cell function and piRNA expression in planarians. RNA 14, 1174–1186 (2008).
Reddien, P. W., Oviedo, N. J., Jennings, J. R., Jenkin, J. C. & Alvarado, A. S. SMEDWI-2 is a PIWI-like protein that regulates planarian stem cells. Science 310, 1327–1330 (2005).
Zhang, H., Xia, R., Meyers, B. C. & Walbot, V. Evolution, functions, and mysteries of plant ARGONAUTE proteins. Curr. Opin. Plant Biol. 27, 84–90 (2015).
Vaucheret H. Plant ARGONAUTES. Trends Plant Sci. 13, 350–358 (2008).
Nishimura, A., Ito, M., Kamiya, N., Sato, Y. & Matsuoka, M. OsPNH1 regulates leaf development and maintenance of the shoot apical meristem in rice. Plant J. 30, 189–201 (2002).
Nonomura, K. I. et al. A germ cell-specific gene of the ARGONAUTE family is essential for the progression of premeiotic mitosis and meiosis during sporogenesis in rice. Plant Cell 19, 2583–2594 (2007).
Agorio, A. & Vera, P. ARGONAUTE4 is required for resistance to Pseudomonas syringae In Arabidopsis. Plant Cell 19, 3778–3790 (2007).
Li, W., Saraiya, A. A. & Wang, C. C. The profile of snoRNA-derived microRNAs that regulate expression of variant surface proteins in Giardia lamblia. Cell. Microbiol. 14, 1455–1473 (2012).
Mukherjee, K., Campos, H. & Kolaczkowski, B. Evolution of animal and plant dicers: early parallel duplications and recurrent adaptation of antiviral RNA binding in plants. Mol. Biol. Evol. 30, 627–641 (2013).
Welker, N. C. et al. Dicer’s helicase domain is required for accumulation of some, but not all, C. elegans endogenous siRNAs. RNA 16, 893–903 (2010).
Deleris, A. et al. Hierarchical action and inhibition of plant Dicer-like proteins in antiviral defense. Science 313, 68–71 (2006).
Ma, X., Shao, C., Jin, Y., Wang, H. & Meng, Y. Long non-coding RNAs: A novel endogenous source for the generation of Dicer-like 1-dependent small RNAs in Arabidopsis thaliana. RNA Biol. 11 (2014).
MacKay, C. R., Wang, J. P. & Kurt-Jones, E. A. Dicer’s role as an antiviral: still an enigma. Curr. Opin. Immunol. 26, 49–55 (2014).
Margis, R. et al. The evolution and diversification of Dicers in plants. FEBS Lett. 580, 2442–2450 (2006).
Catalanotto, C. et al. Redundancy of the two dicer genes in transgene-induced posttranscriptional gene silencing in Neurospora crassa. Mol. Cell. Biol. 24, 2536–2545 (2004).
Alexander, W. G. et al. DCL-1 colocalizes with other components of the MSUD machinery and is required for silencing. Fungal Genet. Biol. 45, 719–727 (2008).
Silva, S. et al. Mte1 interacts with Mph1 and promotes crossover recombination and telomere maintenance. Genes Dev. 30, 700–717 (2016).
Dorin, D. et al. TheTAR RNA-binding protein, TRBP, stimulates the expression of TAR-containing RNAs in vitro and in vivo independently of its ability to inhibit the dsRNA-dependent kinase PKR. J. Biol. Chem. 278, 4440–4448 (2003).
Rossi, J. J. Mammalian Dicer finds a partner. EMBO Rep. 6, 927–929 (2005).
Haase, A. D. et al. TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep. 6, 961–967 (2005).
Ota, H. et al. ADAR1 forms a complex with Dicer to promote microRNA processing and RNA-induced gene silencing. Cell 153, 575–589 (2013).
Duchaine, T. F. et al. Staufen2 isoforms localize to the somatodendritic domain of neurons and interact with different organelles. J. Cell Sci. 115, 3285–3295 (2002).
Liu, J., Hu, J. Y., Wu, F., Schwartz, J. H. & Schacher, S. Two mRNA-binding proteins regulate the distribution of syntaxin mRNA in Aplysia sensory neurons. J. Neurosci. 26, 5204–5214 (2006).
Macchi, P. et al. The brain-specific double-stranded RNA-binding protein Staufen2: nucleolar accumulation and isoform-specific exportin-5-dependent export. J. Biol. Chem. 279, 31440–31444 (2004).
Parkinson, J. et al. A transcriptomic analysis of the phylum Nematoda. Nat. Genet. 36, 1259–1267 (2004).
Nakanishi, K. Anatomy of RISC: how do small RNAs and chaperones activate Argonaute proteins? WIREs RNA 7, 637–660 (2016).
Kobayashi, H. & Tomari, Y. RISC assembly: Coordination between small RNAs and Argonaute proteins. Biochim. Biophys. Acta 1859, 71–81 (2016).
Cuellar, T. L. & McManus, M. T. MicroRNAs and endocrine biology. J. Endocrinol. 187, 327–332 (2005).
Hegge, J. W., Swarts, D. C. & van der Oost, J. Prokaryotic Argonaute proteins: novel genome-editing tools? Nat. Rev. Microbiol. 16, 5–11 (2018).
Saraiya, A. A., Li, W., Wu, J., Chang, C. H. & Wang, C. C. The microRNAs in an ancient protist repress the variant-specific surface protein expression by targeting the entire coding sequence. PLoS Pathog. 10, e1003791 (2014).
Lau, P. W., Potter, C. S., Carragher, B. & MacRae, I. J. Structure of the human Dicer-TRBP complex by electron microscopy. Structure 17, 1326–1332 (2009).
Koonin, E. V., Makarova, K. S. & Zhang, F. Diversity, classification and evolution of CRISPR-Cas systems. Curr. Opin. Microbiol. 37, 67–78 (2017).
Jiang, F. & Doudna, J. A. CRISPR-Cas9 Structures and Mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
The UniProt, C. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC bioinformatics 5, 113 (2004).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Felsenstein, J. Confidence-Limits on Phylogenies - an Approach Using the Bootstrap. Evolution 39, 783–791 (1985).
Letunic, I. & Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 46, D493–D496 (2018).
Zmasek, C. M. & Eddy, S. R. A simple algorithm to infer gene duplication and speciation events on a gene tree. Bioinformatics 17, 821–828 (2001).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31470716, 31000323 and 31070672) and the Natural Science Foundation of Jiangsu Province (BK20131272).
Author information
Authors and Affiliations
Contributions
R.Z., Y.J., H.Z., Y.N., C.L., J.W., K.Z. and C.-Y. Z. performed the experiment and analyzed data. D.L. designed and performed the experiments, analyzed data, and drafted the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, R., Jing, Y., Zhang, H. et al. Comprehensive Evolutionary Analysis of the Major RNA-Induced Silencing Complex Members. Sci Rep 8, 14189 (2018). https://doi.org/10.1038/s41598-018-32635-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32635-4
Keywords
This article is cited by
-
miRNome profiling reveals differential miRNAs associated with embryogenic potential in the somatic embryogenesis of Araucaria angustifolia
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
-
Comparative Analyses of Human Exosome Proteomes
The Protein Journal (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.