Abstract
Expression of the chromatin-associated protein HMGA2 correlates with progression, metastasis and therapy resistance in pancreatic ductal adenocarcinoma (PDAC). Hmga2 has also been identified as a marker of a transient subpopulation of PDAC cells that has increased metastatic ability. Here, we characterize the requirement for Hmga2 during growth, dissemination, and metastasis of PDAC in vivo using conditional inactivation of Hmga2 in well-established autochthonous mouse models of PDAC. Overall survival, primary tumour burden, presence of disseminated tumour cells in the peritoneal cavity or circulating tumour cells in the blood, and presence and number of metastases were not significantly different between mice with Hmga2-wildtype or Hmga2-deficient tumours. Treatment of mice with Hmga2-wildtype and Hmga2-deficient tumours with gemcitabine did not uncover a significant impact of Hmga2-deficiency on gemcitabine sensitivity. Hmga1 and Hmga2 overlap in their expression in both human and murine PDAC, however knockdown of Hmga1 in Hmga2-deficient cancer cells also did not decrease metastatic ability. Thus, Hmga2 remains a prognostic marker which identifies a metastatic cancer cell state in primary PDAC, however Hmga2 has limited if any direct functional impact on PDAC progression and therapy resistance.
Similar content being viewed by others
Introduction
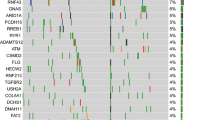
Pancreatic ductal adenocarcinoma (PDAC) is among the most fatal of all malignancies1. Due to its high rate of metastatic spread and poor response to therapy the five-year survival rate for PDAC patients remains below ten percent and PDAC is projected to be the second most common cause of cancer deaths by 20302. While PDAC is well characterized genomically,3,4 the mechanisms that enable cancer cells to leave their primary site and form distant metastases remain largely unknown5,6,7,8,9,10. The most common driver mutations in human PDAC are oncogenic mutations in KRAS and loss of the tumour suppressors CDKN2A, SMAD4, and TP53. These alterations are present in both primary tumours and metastases and additional genomic alterations specific to metastases have not been identified6,11. Several recent studies have suggested that PDAC metastasis is driven by epigenetic alterations, metabolic changes, and microenvironment-induced gene expression alterations10,12,13,14,15. Due to its highly metastatic nature, most PDAC patients are diagnosed at an advanced disease stage with local or distant metastasis and are therefore not candidates for potentially curative surgery. Thus, the majority of patients are treated with chemotherapy. However, chemotherapeutic regimens and targeted therapies are largely ineffective, and tumours that do respond rapidly develop resistance12,15,16,17.
Genetically engineered mouse models of PDAC recapitulate many aspects of the human disease. Pancreas-specific expression of oncogenic KrasG12D and Cre/loxP-based inactivation of the tumour suppressors that are frequently inactivated in human PDAC results in the development of murine PDAC that is histologically and molecularly similar to the human disease18,19. These models recapitulate the entire course of the disease, from the development of preneoplastic lesions to invasive carcinomas and the development of widespread metastatic disease19,20,21,22. Importantly, these models also allow the impact of additional genetic alterations to be investigated during the initiation and progression of pancreatic cancer, as well as response to therapy in vivo23,24,25.
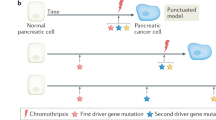
HMGA2 (high mobility group A2) is a chromatin-associated DNA-binding protein that is expressed in embryonic stem cells and during development, but is absent from most adult somatic cells26,27. Interestingly, HMGA2 is re-expressed in many human malignancies, including PDAC, where high expression correlates with lymph node metastasis, increased tumour grade, and reduced patient survival28,29,30,31,32,33,34. HMGA2 has been shown to interact with histones and alter DNA topology through binding to the minor groove of DNA26. Due to its high expression in multiple cancer types at advanced stages as well as in metastases, it has been suggested that HMGA2 functions to drive metastatic ability27,28,30,34,35,36,37,38,39. Recently, using genetically engineered mouse models, we defined a transient, Hmga2-positive subpopulation of PDAC cells that has increased metastatic ability13. However, it remains unclear whether Hmga2 is a functionally important driver of the pro-metastatic properties of these cells or only a marker of this state.
In this study, we directly investigated the importance of Hmga2 during PDAC development, metastasis, and therapy response in vivo. We conditionally inactivated Hmga2 in well-established autochthonous PDAC models and found no effect of Hmga2 inactivation on tumour initiation, progression, or metastasis. The absence of Hmga2 also did not dramatically alter the response of autochthonous PDAC to gemcitabine treatment.
Results
Hmga2 deficiency does not impact the survival of PDAC bearing mice
To investigate the role of Hmga2 in PDAC, we first confirmed Hmga2 expression in the carcinoma stage of pancreatic cancer tissue isolated from the well-established autochthonous KrasLSL-G12D/+;p53LSL-R172H/+;Rosa26LSL-tdTomato/+;Pdx1-Cre (KP172CT) mouse model of PDAC (Fig. 1a, Supplementary Fig. S1a). Recently, we uncovered Hmga2 as a marker of a transient subpopulation of pancreatic cancer cells with increased metastatic ability in KP172CT mice13. To mark Hmga2positive cells within autochthonous PDAC, we utilized a conditional Hmga2 allele that converts from the unrecombined Hmga2CK conformation to a null-allele/GFP-knock-in reporter (Hmga2GFP) upon Cre-mediated inversion of a splice-acceptor-GFP-splice-donor cassette13,40.
Absence of Hmga2 does not alter overall survival in autochthonous mouse models of PDAC. (a) Alleles in the KP172CT (KrasLSL-G12D/+;p53LSL-R172H/+;R26LSL-Tom/+;Pdx1-Cre) and the KPHetCT (KrasLSL-G12D/+;p53flox/+;R26LSL-Tom/+;Pdx1-Cre) models before and after allele recombination. (b) Hmga2 expression in pancreatic tumors in KP172CT;Hmga2CK/+ mice overlaps with GFP in Tomato positive cancer cells. Hmga2 is absent in KP172CT;Hmga2CK/CK tumours. Representative of >10 tumors. Scale bars = 50 µm. (c,d) Kaplan-Meier survival curves of KP172CT (c) and KPHetCT (d) mice shows no significant difference with (Hmga2CK/CK) or without (Hmga2+/+) Hmga2 inactivation.
To initially investigate whether pancreas-specific inactivation of Hmga2 would grossly alter pancreatic development and size, we generated Hmga2CK/CK;Pdx1-Cre mice. These mice were born at expected Mendelian ratios and developed normally. Pancreata from 4-week and 11-week-old mice showed no signs of malformation of the exocrine or endocrine compartments, as assessed both by organ weight and histology (Supplementary Fig. S1b,c).
To investigate the role of Hmga2 during pancreatic tumourigenesis, we incorporated the Hmga2CK allele and a Rosa26LSL-tdTomato/+ reporter allele into the KrasLSL-G12D/+;p53LSL-R172H/+;Pdx1-Cre (KP172C) background (Fig. 1a). We generated KP172CT;Hmga2CK/+ and KP172CT;Hmga2CK/CK tumour-bearing mice. Immunohistochemistry for Hmga2, tdTomato, and the GFP fusion protein confirmed that Hmga2 expression in neoplastic cells in KP172CT;Hmga2CK/+ mice overlaps with GFP expression and that Hmga2 protein is absent from all tumours in KP172CT;Hmga2CK/CK mice (Fig. 1b). To analyse whether Hmga2 deficiency impacts the development and progression of PDAC, we generated cohorts of KP172CT;Hmga2CK/CK and KP172CT;Hmga2+/+ mice. Gross cancer-associated phenotypes were comparable, with mice showing similar cachexic symptoms as well as the development of ascites and jaundice in some mice (data not shown). Kaplan-Meier analysis showed no significant survival difference between KP172CT;Hmga2CK/CK and KP172CT;Hmga2+/+ mice. There was even a trend towards slightly shorter survival of KP172CT;Hmga2CK/CK mice (Fig. 1c).
To further investigate the impact of Hmga2 deficiency on PDAC growth, we employed a second mouse model of PDAC, where instead of expression of a dominant negative p53 allele (p53LSL-R172H) we incorporated a p53floxed allele to enable conditional heterozygous deletion of p53 (p53flox/+, Fig. 1a right panel). We generated KrasLSL-G12D/+;p53flox/+;Rosa26LSL-tdTomato/+;Pdx1-Cre;Hmga2+/+ (KPHetCT;Hmga2+/+) and KPHetCT;Hmga2CK/CK mice and assessed overall survival. Consistent with our data from the KP172CT model, Hmga2-deficiency did not significantly alter overall survival and gross disease-associated phenotypes in both groups were comparable (Fig. 1d and data not shown). Collectively, these data suggest that Hmga2 inactivation has limited, if any, effect on overall pancreatic tumour growth.
Hmga2 deficiency does not change PDAC phenotypes
To investigate the impact of Hmga2 deficiency on pancreatic cancer phenotypes in KP172CT and KPHetCT mice, we assessed tumour histology. At the time of analysis, most KP172CT;Hmga2+/+ and KP172CT;Hmga2CK/CK animals had developed pancreatic adenocarcinoma (Fig. 2a,c). In these mice, tumours were of similar size and were characterized by the presence of mPanINs and glandular structures accompanied by an extensive desmoplastic reaction (data not shown). Regardless of genotype, tumours had both well- and poorly-differentiated areas which is typical for tumours in this model18. Detailed histological analyses did not uncover any differences in the histological features of mice with Hmga2-deficient or Hmga2-wildtype tumours. Similarly, KPHetCT;Hmga2+/+ and KPHetCT;Hmga2CK/CK mice developed pancreatic cancer with full penetrance and comparable histology (Fig. 2b,c). Thus, Hmga2-deficiency does not affect the histological phenotype of mouse pancreatic cancer.
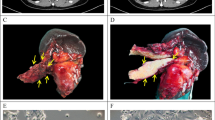
Hmga2 deficiency does not alter the phenotype of pancreatic cancer in multiple autochthonous mouse models. (a,b) Representative histology of primary PDAC tumours and metastases from KP172CT (a) and KPHetCT (b) mice with (Hmga2+/+) or without (Hmga2CK/CK) Hmga2. Scale bars = 100 µm. (c) Incidence of primary PDAC and metastases in KP172CT and KPHetCT mice with (Hmga2+/+) or without (Hmga2CK/CK) Hmga2. All comparisons showed no significant differences. (d,e) Incidence of circulating tumour cells (CTC) and disseminated tumour cells (DTC) is comparable in KP172CT;Hmga2+/+ and KP172CT;Hmga2CK/CK mice (e) as detected by FACS (d) for tdTomatopos, lineageneg, DAPIneg cancer cells in blood (CTC) and pleural fluid (DTC). No significant differences in CTC number were observed.
To assess the effect of Hmga2-deficiency on cancer cell dissemination and metastasis, we quantified the development of -metastases by fluorescence-stereomicroscopy. Distant metastases were present in the liver, peritoneal cavity, and lung in about 30–50% of mice. Hmga2-deficient and Hmga2-wildtype metastases in the KP172CT and KPHetCT models were histologically indistinguishable (Fig. 2a–c).
We also utilized the bright fluorescent marking conferred by the Rosa26tdTomato/+ allele to assess the presence of disseminated tumour cells (DTCs) in the peritoneal fluid and circulating tumour cells (CTCs) in the blood by flow cytometry for tdTomato+ cancer cells (Fig. 2d). DTCs were present in the peritoneal cavity of almost all KP172CT;Hmga2+/+ and KP172CT;Hmga2CK/CK mice and CTCs were present in five out of eight KP172CT;Hmga2+/+ and four out of eleven KP172CT;Hmga2CK/CK mice. The number of CTCs detectable in mice was variable, but was not significantly different between KP172CT;Hmga2+/+ and KP172CT;Hmga2CK/CK mice (Fig. 2e and Supplemental Fig. S2). Collectively, these data suggest that Hmga2 is not required for dissemination and metastasis of pancreatic cancer.
Hmga2 deficiency does not alter PDAC sensitivity to gemcitabine
The nucleoside analogue gemcitabine is one of the standard agents of care for PDAC patients12. In addition to its association with more advanced and metastatic human PDAC, HMGA2 has also been suggested to promote gemcitabine resistance of PDAC41,42. To investigate whether Hmga2 deficiency would increase PDAC sensitivity to gemcitabine in vivo, we assessed the impact of Hmga2 deficiency on overall survival with and without gemcitabine treatment. In the KrasLSL-G12D/+;p53flox/flox;Pdx1-Cre (KPKOC) model, pancreatic cancer develops rapidly and advanced disease is already present 4 weeks after birth. KPKOC mice typically succumb to their very aggressive disease between 50 and 60 days of age43. To determine whether Hmga2 deficiency leads to increased sensitivity to gemcitabine, we generated KPKOC;Hmga2+/+ and KPKOC;Hmga2CK/CK mice (Fig. 3a). These mice were either treated with four intraperitoneal injections of gemcitabine (120 mg/kg) or vehicle control on days 28, 32, 35 and 39 of age (Fig. 3b). We monitored overall survival as a metric of tumour response. Consistent with published reports, gemcitabine provided a small survival benefit to KPKOC;Hmga2+/+ mice (median survival of 61 days in control treated mice versus 77 days in gemcitabine treated mice)24,44. However, the increase in survival was equally small in KPKOC;Hmga2CK/CK mice (16-day increase in survival in KPKOC;Hmga2+/+ mice treated with gemcitabine versus a 14-day increase in survival in KPKOC;Hmga2CK/CK mice treated with gemcitabine) and no significant difference in survival was detectable between gemcitabine treated KPKOC;Hmga2+/+ and gemcitabine treated KPKOC;Hmga2CK/CK mice (p = 0.515). This suggests that the absence of Hmga2 does not increase PDAC sensitivity to this standard chemotherapeutic agent. Furthermore, histological analysis of the tumours from gemcitabine treated KPKOC;Hmga2+/+ and KPKOC;Hmga2CK/CK mice uncovered comparable histology, with complete destruction of normal pancreatic architecture and development of undifferentiated and anaplastic PDAC (Fig. 3c,d). Thus, Hmga2 deficiency did not lead to gemcitabine sensitivity in vivo.
Hmga2 deficiency does not dramatically alter PDAC sensitivity to gemcitabine. (a–c) KPKOC;Hmga2+/+ and KPKOC;Hmga2CK/CK mice (a) were treated with four intraperitoneal gemcitabine injections at 28, 32, 35, and 39 days of age (b) and compared to their respective vehicle-treated control littermates. Gemcitabine treatment slightly increased median survival in both genotypes (65.5 days in KPKOC;Hmga2CK/CK and 77 days in KPKOC;Hmga2+/+ mice) compared to untreated controls (51.5 days in KPKOC;Hmga2CK/CK and 61 days in KPKOC;Hmga2+/+ mice), but no significant survival difference was observed between gemcitabine -treated KPKOC;Hmga2CK/CK and KPKOC;Hmga2+/+ mice (p = 0.47) (c). (d) Representative histology of tumours in KPKOC;Hmga2+/+ and KPKOC;Hmga2CK/CK mice treated with gemcitabine. Across all mice, no consistent changes in histology were observed. Scale bars = 50 µm.
Hmga1 expression overlaps with Hmga2 expression, but Hmga1 does not compensate for Hmga2 deficiency
HMGA2 belongs to the family of HMGA proteins that consists of two family members, HMGA1 and HMGA226. Both proteins are considered architectural transcription factors that regulate transcription indirectly by altering DNA conformation. In normal adult cells, both family members are almost undetectable though they are abundant during embryonic development. Both family members have been shown to regulate cell signalling, proliferation, and differentiation with at least partially overlapping functions26,27,31,45. The expression of both HMGA1 and HMGA2 has been shown to correlate with advanced tumour stages and metastasis in PDAC patients32. To investigate whether Hmga1 expression overlaps with Hmga2 expression, we performed immunohistochemistry and western blot analysis on murine and human PDAC tissue samples. In tumours from KP172CT;Hmga2CK/+ mice, Hmga1 and Hmga2 were expressed in primary tumour areas as well as in metastases in liver and lung (Supplemental Fig. 3a). In PDAC from KPHetCT;Hmga2+/+ mice the expression of Hmga1 and Hmga2 generally overlapped, with Hmga1 also being highly expressed in advanced primary tumours as well as in metastases (Fig. 4a). In PDAC from KPHetCT;Hmga2CK/CK mice, Hmga1 was still expressed at similar levels. Western blot analysis for Hmga1 and Hmga2 on sorted (lineagenegative, DAPInegative, Tomatopositive) cancer cells isolated from pancreatic tumours from KPHetCT;Hmga2+/+ and KPHetCT;Hmga2CK/CK mice confirmed the absence of Hmga2, the presence of the GFP fusion protein in Hmga2-deficient tumours, and unaltered Hmga1 expression (Fig. 4b). Similarly, HMGA1 and HMGA2 proteins are often co-expressed in human PDAC (Fig. 4c). In TCGA PDAC dataset, HMGA1 and HMGA2 expression was also moderately correlated (0.414 Pearson correlation coefficient or 0.581 Spearman correlation coefficient, respectively). Correlation of overall survival with HMGA1 and HMGA2 protein expression showed that PDAC patients with tumours that were both HMGA1negative and HMGA2negative had better overall survival than patients whose tumours expressed HMGA1 and/or HMGA2 (Fig. 4d,e; p = 0.0022).
Hmga1 and Hmga2 are co-expressed in mouse and human pancreatic cancer. (a) Immunohistochemistry for Hmga1 and Hmga2 in wild type (WT), KrasLSL-G12D/+;Pdx1-Cre (KC) and KrasLSL-G12D/+;p53flox/+;R26LSL-Tom/+;Pdx1-Cre (KPHetCT) mice shows co-expression of Hmga1 and Hmga2 in PDAC and metastases but not in preneoplastic lesions or healthy pancreata. Scale bars = 50 µm. (b) Western Blot on sorted cancer cells from KPHetCT;Hmga2+/+ and KPHetCT;Hmga2CK/CK mice documents the expression of the GFP-fusion reporter gene and the absence of Hmga2 in cells from KPHetCT;Hmga2CK/CK mice. Hmga1 expression is not affected. Corresponding full length images and details regarding probing of the membranes are available in Supplemental Fig. S4c. (c) Tissue microarrays from human pancreatic cancer patients were stained for HMGA1 and HMGA2. Scale bars = 50 µm. (d) Kaplan-Meier survival analysis for patients with HMGA1 and HMGA2 positive or negative tumours. (e) Median survival in months of PDAC patients with expression of HMGA1 and HMGA2 shows that patients with HMGA1negative/HMGA2negative tumours have significantly (p = 0.0022) better overall survival than patients who have tumors that express of either HMGA1 or HMGA2 or both. (f–i) Knockdown of Hmga1 in a cell line (2648LN) isolated from a lymph node metastasis from a KP172CT;Hmga2CK/CK mouse has no significant impact on their metastatic potential upon intravenous injection into recipient mice. (f) qPCR was used to assess knockdown; mean +/− SEM of triplicate wells is shown. (g) Representative brightfield and fluorescent images of one lung lobe and corresponding histology. Scale bars in the upper and middle panels = 4 mm; scale bars in the lower panels = 50 µm. (h,i) Quantification of %-Tomatopositive area (h) and number of tdTomatopositive tumours in the lung (i), n = 3 mice per group; the red line indicates the mean.
To test whether Hmga1 compensates for Hmga2 in PDAC metastasis, we generated PDAC cell lines from tumours from KP172CT;Hmga2+/+ and KP172CT;Hmga2CK/CK mice (n = 4 each). We performed qPCR analysis for Hmga1 and Hmga2 expression. Hmga1 expression was not affected by Hmga2-deficiency (Supplemental Fig. S3c,d). To investigate if Hmga1 could functionally compensate for Hmga2, we stably knocked down Hmga1 in a cancer cell line generated from a KP172CT;Hmga2CK/CK lymph node metastasis. Knockdown at the RNA level was confirmed by qPCR (Fig. 4f). However, no significant difference in metastatic ability between control and Hmga1 knockdown cells was observed upon intravenous injection into immunocompetent recipient mice (Fig. 4g,h,i). Additionally, we stably knocked down Hmga1 or Hmga2 in a cell line generated from a liver metastasis in a KP172CT;Hmga2+/+ mouse (Supplemental Fig. S4a). Intravenous transplantation of Hmga1 knockdown, Hmga2 knockdown, or control cells into immunocompetent recipient mice to seed metastases into the lungs showed no difference in the metastatic ability of control cells and cells deficient for either Hmga1 or Hmga2 (Supplemental Fig. S4b).
Discussion
To investigate the function of the chromatin-associated protein Hmga2 during pancreatic cancer growth, metastasis, and treatment response, we took a classical genetic approach of specifically inactivating Hmga2 protein expression in the well-established KP172CT, KPHetCT, and KPKOC mouse models of PDAC. To our surprise, Hmga2-deficient tumours formed in the pancreata of KP172CT;Hmga2CK/CK, KPHetCT;Hmga2CK/CK, and KPKOC;Hmga2CK/CK mice with similar rate and morphology and within a comparable time course as in control KP172CT, KPHetCT and KPKOC mice (Fig. 2). By all metrics, from overall survival to histology of primary tumours and metastases to the presence and number of CTCs and DTCs, Hmga2-deficiency did not perturb PDAC development in any significant way.
Numerous in vitro studies across several different human cancer types have suggested a role for HMGA2 in controlling phenotypes associated with malignant transformation and metastasis32,36,42,46,47. However, very few studies have investigated Hmga2 function in autochthonous cancer models in vivo. In a mouse model of Wnt1-driven colorectal cancer, tumour formation was reduced in Hmga2−/− mice, suggesting a role for Hmga2 in cancer development48. However, Hmga2−/− mice have severe dwarfism49, therefore the direct importance of Hmga2 in colorectal cancer cells in vivo remains unclear. Furthermore, in squamous skin cancer mouse models, Hmga2 was found to be dispensable for tumour initiation and progression50. While we recently showed that Hmga2 is a marker of a transient metastatic subpopulation in PDAC in vivo13, our current study shows that neither deletion of Hmga2 in autochthonous mouse models of PDAC, nor knockdown of Hmga2 in an aggressive pancreatic cancer cell line impacted pancreatic tumour initiation, progression, or metastatic ability. In the genetically engineered mouse models that we used, Hmga2 deletion occurs concomitantly with expression of KrasG12D and inactivation of p53 during tumour initiation, and therefore tumours develop in the complete absence of Hmga2. This could lead to an alternative, Hmga2-independent mechanism for PDAC growth and progression. Second hit mouse models of PDAC could provide additional information on whether acute Hmga2 inactivation in PDAC impacts growth, progression, or therapy response.
In contrast to human non-small cell lung cancer cell lines35, cell lines derived from the KrasG12D-driven p53-deficient mouse lung adenocarcinoma model38, and the mouse breast cancer cell line 4T148, Hmga2 knockdown in murine PDAC cells did not reduce metastatic ability in vivo. This is particularly interesting as PDAC is among the most genetically stable cancer entities and its progression is based on epigenetic changes rather than acquisition of additional DNA mutations11,14, suggesting an even greater importance of epigenetic reprogramming of malignant cells39,51.
HMGA2 has been previously implicated in resistance to gemcitabine in human pancreatic cancer cell lines in vitro, where HMGA2 knockdown sensitized PDAC cells to gemcitabine treatment42. HMGA2 mediated gemcitabine resistance in three-dimensional collagen cultures of PDAC cell lines via increased histone acetylation41. To test whether Hmga2 mediates gemcitabine resistance in vivo, we utilized a well-established clinically relevant treatment model44,52,53. Neither overall survival nor histology of the tumours in mice treated with gemcitabine were significantly affected by Hmga2-deficiency. While these models have limited resolution to detect modest effects, our results suggest that Hmga2 has limited if any effect on gemcitabine resistance in vivo. Despite Hmga2 having no effect in gemcitabine-treated mouse models of pancreatic cancer, it remains possible that HMGA2 may still be a determinant of therapy sensitivity in human PDAC or of response to other chemotherapies.
We also investigated whether HMGA1, a HMGA2 family member with related functions, compensates for HMGA2 during PDAC metastasis. We and others have shown that HMGA1 and HMGA2 overlap in their expression in human PDAC and in KP172CT and KPHetCT mice (Fig. 4, ref.32). Additionally, the expression of HMGA1 and HMGA2 correlates with poor survival of PDAC patients32. Notably, PDAC patients with HMGA1negative HMGA2negative tumours have significantly better survival than all other patients (Fig. 4d). Cancer cells from KPHetCT;Hmga2+/+ and KPHetCT;Hmga2CK/CK mice express Hmga1 at similar levels, therefore Hmga1 could compensate for Hmga2 deficiency to enable malignant transformation and metastasis. Knockdown of Hmga1 in a PDAC cancer cell line generated from a KP172CT;Hmga2CK/CK mouse had no impact on metastatic ability after intravenous transplantation. Hmga1 knockdown in our system might not have been sufficient to entirely remove Hmga1 function, with remaining Hmga1 levels still being adequate to enable metastasis. Alternatively, a pro-metastatic effect of Hmga1/Hmga2 could be stable and not depend on continued Hmga1 expression. In vivo models that incorporate genetic deletion of both Hmga1 and Hmga2 will allow this question to be addressed definitively.
Taken together, we provide evidence that Hmga2 is functionally dispensable for the malignant transformation, progression, and metastatic ability of pancreatic cancer in vivo. Our results support Hmga2 being solely of prognostic value, as has been shown for multiple cancer types29,30,31,34. Hmga2 remains a prognostic marker which identifies an advanced cancer cell state in primary pancreatic tumours and marks a metastasis-driving subpopulation of cancer cells.
Methods
Mice
KrasLSL-G12D, p53LSL-R172H, p53flox, Pdx1-Cre, Rosa26LSL-tdTomato, and Hmga2CK mice have been described40,54,55,56,57,58. For gemcitabine treatment, 28 day old KrasLSL-G12D/+;p53flox/flox;R26LSL-Tom/+;Pdx1-Cre and KrasLSL-G12D/+;p53flox/flox;Hmga2CK/CK;R26LSL-Tom/+;Pdx1-Cre were randomly separated into two groups each (mixed gender) and treated with four intraperitoneal injections of gemcitabine (120 mg/kg) or saline on day 28, 32, 35, and 39 of age. For transplantation of knockdown cell lines, 2 × 104 cells were injected into the lateral tail vein of male 129/Bl6 F1 mice (Jackson Laboratories, Stock number 101043). Mice were analysed 3 weeks after injection. No statistical method was used to predetermine sample size. All experimental protocols were approved by the Stanford University Animal Care and Use Committee and performed in accordance with their guidelines.
Histology and immunohistochemistry on mouse PDAC
Tissues were fixed in 4% formalin in PBS overnight and transferred to 70% ethanol prior to paraffin embedding. After de-parafinization, re-hydration, and antigen retrieval, IHC was performed on 4 μm sections with the ABC Vectastain kit (PK-4001, Vector Laboratories Inc.) with antibodies to Hmga2 (59170AP, BioCheck Inc.), Tomato (600-401-379, Rockland Inc.), GFP (ab6673, Abcam), CK19 (TROMA-III, Developmental Studies Hybridoma Bank), Insulin (A0564, Dako) and Hmga1 (sc-8982, Santa Cruz). Sections were developed with DAB and counterstained with haematoxylin. Haematoxylin and Eosin (H&E) staining was performed using standard methods.
Tumour dissociation and cancer cell flow cytometric analysis
Pancreatic tumours and metastasis were dissociated into a single cell suspension prior to FACS analysis. Briefly, tumours were minced with sharp scissors and incubated for 20 minutes at 37oC in HBSS medium (Corning, 21-022-CV) with collagenase IV (Worthington, 1 mg/ml, LS004188) and dispase (Corning, 354235) followed by quenching with ice-cold L-15 medium (Invitrogen, 21083-027) containing 10% FBS and DNase (20 μg/mL). Cells were stained with Allophycocyanin (APC)-conjugated antibodies to CD45 (30-F11), CD31 (390), F4/80 (BM8), and Ter119 (TER-119, all from BioLegend) to exclude hematopoietic and endothelial cells. DAPI was used to exclude dead cells. Cells within the peritoneal cavity were collected immediately after euthanasia by making a small incision in the peritoneum followed by introduction of 1 ml PBS. For circulating tumour cell analysis blood was collected immediately after euthanasia by cardiac puncture. Red blood cell lysis was performed by incubation with ACK lysis buffer (155 mM NH4Cl, 10 mM KHCO3, 0.1 mM EDTA). BD LSR II analysers and FACSAria sorters (BD Biosciences) were used and FlowJo software was utilized for analysis.
Immunohistochemistry for HMGA1 and HMGA2 on human PDAC
Expression of HMGA1 and HMGA2 was assessed using IHC on 4 μm sections of paraffin-embedded tissue from each TMA. Sections were heated at 60 °C for 60 minutes, de-paraffinized in xylene and rehydrated through a graded alcohol series. Three percent H2O2 block was used to quench endogenous peroxidase activity. Antigen retrieval was performed in a de-cloaking chamber using a Borg Decloaker RTU antigen retrieval solution (Biocare Medical). Sections were then blocked in normal horse serum (Vector Laboratories) and incubated in a humidified chamber overnight at 4 °C with a 1:1000 dilution of primary anti-HMGA1 antibody (sc-8982, Santa Cruz) or anti-HMGA2 antibody (59170AP, BioCheck). Slides were washed in PBS and incubated with biotinylated anti-rabbit secondary antibody (Vector Laboratories) for 30 minutes at room temperature. Staining was visualized using DAB (Vector Laboratories) at room temperature. Slides were then counterstained with haematoxylin, rinsed in water and dehydrated through a graded alcohol series and xylene, and mounted with VectaMount (Vector Laboratories). HMGA1 and HMGA2 staining of individual samples was evaluated manually and an arbitrary cutoff of staining intensity was used to determine if a sample was considered negative or positive. Overall survival (OS) was defined as the time of surgery to the date of death from any cause. OS was analysed using Kaplan–Meier and log-rank tests. Significance was determined as a p-value < 0.05.
This study was approved by the Stanford University Institutional Review Board and performed in accordance with their guidelines, with a waiver of informed consent to use patient tissue and specimens.
Generation and selection of murine PDAC cell lines
We generated polyclonal cell lines from primary tumours and metastases that formed in the autochthonous KrasLSL-G12D/+;Trp53LSL-R172H/+;RosaLSL-tdTomato;Pdx1-Cre pancreatic cancer mouse model with (Hmga2+/+) and without (Hmga2CK/CK) Hmga2 expression22,54. To establish the cell lines, a piece of the tumour or macro-metastasis was dissociated into a single cell suspension, washed twice with cold PBS, minced with a scalpel and transferred to a tissue culture dish containing DMEM media (high glucose with 10% FBS and antibiotics). Cells were allowed to attach and grown for one week with two media changes. Then cells were passaged at least 3 times to select away from fibroblast contamination. Purity was confirmed by FACS for tdTomato and MycoAlert Mycoplasma detection kit (Lonza) was used to verify the lack of Mycoplasma contamination.
Lentiviral knockdown and qRT-PCR analysis
Hmga1 and Hmga2 were knocked down using pLKO lentiviral vectors; mouse shHmga11 (TRCN0000182651) and mouse shHmga2 (TRCN0000126044). The control vector was pLKO-shLuciferase. Virus production, cell infection, and selection were performed as previously described59. Knockdown was confirmed by qPCR and western blotting. qRT-PCR for mouse Hmga1 (Hmga1_Fw GGGGCAGACCCAAGAAAC and Hmga1_Rv GGCACTGCGAGTGGTGAT) and Hmga2 (Hmga2_Fw GGATCCTGGCAGAAACTTCC and Hmga2_Rv AACGGGACAGAGATAGAGACTGA) was performed using standard SYBR green qPCR protocols and normalized to mouse Gapdh (Gapdh_Fw TTTGATGTTAGTGGGGTCTCG and Gapdh-Rv AGCTTGTCATCAACGGGAAG).
Western blot analyses
Cells were pelleted before lysis with standard RIPA buffer supplemented with 1/100 volume of protease inhibitor cocktail (Sigma-Aldrich, P8340). Lysates were denatured in Laemmli buffer with β-mercaptoethanol before loading onto a precast 4–12% Bis-Tris PAGE gel (Invitrogen, NP0321BOX). Separated samples were then transferred onto a PVDF membrane (Bio-Rad, 162-0177) before staining with the following antibodies overnight: anti-Hmga2 (Biocheck, 59170AP, or Thermo Scientific, P52926), anti-Hsp90 (BD Biosciences, 610418), and anti-Hmga1 (Santa Cruz, sc-8982 or sc-393213). For the secondary antibodies, goat anti-rabbit IgG-HRP (Santa Cruz Biotech, sc-2004), goat anti-mouse IgG-HRP (Santa Cruz Biotech, sc-2005), and donkey anti-goat IgG-HRP (Santa Cruz Biotech, sc-2020) were used. For enhanced chemiluminescence, Thermo Scientific Pierce ECL 2 Western Blotting Substrate (PI80196) was used.
Statistical analysis
Graphs and statistics were generated using GraphPad Prism software. Significance, where indicated, was calculated using the Wilcoxon test for non-normally distributed data. Survival was compared using Kaplan–Meier plots and log-rank tests. Significance was determined as a p-value < 0.05. No statistical method was used to predetermine sample size.
Data Availability
No datasets were generated during the current study.
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2018. CA Cancer J Clin 68, 7–30 (2018).
Rahib, L. et al. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res 74, 2913–2921 (2014).
Ying, H. et al. Genetics and biology of pancreatic ductal adenocarcinoma. Genes Dev 30, 355–385 (2016).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015).
Yachida, S. & Iacobuzio-Donahue, C. A. The pathology and genetics of metastatic pancreatic cancer. Archives of pathology & laboratory medicine 133, 413–422 (2009).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Moffitt, R. A. et al. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nature genetics 47, 1168–1178 (2015).
Jones, S. et al. Core signaling pathways in human pancreatic cancers revealed by global genomic analyses. Science 321, 1801–1806 (2008).
Knudsen, E. S., O’Reilly, E. M., Brody, J. R. & Witkiewicz, A. K. Genetic Diversity of Pancreatic Ductal Adenocarcinoma and Opportunities for Precision Medicine. Gastroenterology 150, 48–63 (2016).
Whittle, M. C. et al. RUNX3 Controls a Metastatic Switch in Pancreatic Ductal Adenocarcinoma. Cell 161, 1345–1360 (2015).
Makohon-Moore, A. P. et al. Limited heterogeneity of known driver gene mutations among the metastases of individual patients with pancreatic cancer. Nature genetics 49, 358–366 (2017).
Kleeff, J. et al. Pancreatic cancer. Nature reviews. Disease primers 2, 16022 (2016).
Chiou, S. H. et al. BLIMP1 Induces Transient Metastatic Heterogeneity in Pancreatic Cancer. Cancer discovery 7, 1184–1199 (2017).
McDonald, O. G. et al. Epigenomic reprogramming during pancreatic cancer progression links anabolic glucose metabolism to distant metastasis. Nature genetics 49, 367–376 (2017).
Neureiter, D., Jager, T., Ocker, M. & Kiesslich, T. Epigenetics and pancreatic cancer: pathophysiology and novel treatment aspects. World J Gastroenterol 20, 7830–7848 (2014).
Herreros-Villanueva, M., Hijona, E., Cosme, A. & Bujanda, L. Adjuvant and neoadjuvant treatment in pancreatic cancer. World J Gastroenterol 18, 1565–1572 (2012).
Binenbaum, Y., Na’ara, S. & Gil, Z. Gemcitabine resistance in pancreatic ductal adenocarcinoma. Drug resistance updates: reviews and commentaries in antimicrobial and anticancer chemotherapy 23, 55–68 (2015).
Hruban, R. H. et al. Pathology of genetically engineered mouse models of pancreatic exocrine cancer: consensus report and recommendations. Cancer Res 66, 95–106 (2006).
Mazur, P. K. & Siveke, J. T. Genetically engineered mouse models of pancreatic cancer: unravelling tumour biology and progressing translational oncology. Gut (2011).
Aguirre, A. J. et al. Activated Kras and Ink4a/Arf deficiency cooperate to produce metastatic pancreatic ductal adenocarcinoma. Genes Dev 17, 3112–3126 (2003).
Bardeesy, N. et al. Smad4 is dispensable for normal pancreas development yet critical in progression and tumor biology of pancreas cancer. Genes Dev 20, 3130–3146 (2006).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Algul, H. et al. Pancreas-specific RelA/p65 truncation increases susceptibility of acini to inflammation-associated cell death following cerulein pancreatitis. J Clin Invest 117, 1490–1501 (2007).
Ardito Gruner et al. EGF Receptor Is Required for KRAS-Induced Pancreatic Tumorigenesis. Cancer Cell 22, 304–317 (2012).
Mazur, P. K. et al. Notch2 is required for progression of pancreatic intraepithelial neoplasia and development of pancreatic ductal adenocarcinoma. Proc Natl Acad Sci USA 107, 13438–13443 (2010).
Cleynen, I. & Van de Ven, W. J. The HMGA proteins: a myriad of functions (Review). International journal of oncology 32, 289–305 (2008).
Pfannkuche, K., Summer, H., Li, O., Hescheler, J. & Droge, P. The high mobility group protein HMGA2: a co-regulator of chromatin structure and pluripotency in stem cells? Stem Cell Rev 5, 224–230 (2009).
Abe, N. et al. An increased high-mobility group A2 expression level is associated with malignant phenotype in pancreatic exocrine tissue. Br J Cancer 89, 2104–2109 (2003).
Meyer, B. et al. HMGA2 overexpression in non-small cell lung cancer. Molecular carcinogenesis 46, 503–511 (2007).
Motoyama, K. et al. Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clin Cancer Res 14, 2334–2340 (2008).
Pallante, P., Sepe, R., Puca, F. & Fusco, A. High mobility group a proteins as tumor markers. Frontiers in medicine 2, 15 (2015).
Piscuoglio, S. et al. HMGA1 and HMGA2 protein expression correlates with advanced tumour grade and lymph node metastasis in pancreatic adenocarcinoma. Histopathology 60, 397–404 (2012).
Sun, M. et al. HMGA2/TET1/HOXA9 signaling pathway regulates breast cancer growth and metastasis. Proc Natl Acad Sci USA 110, 9920–9925 (2013).
Wang, X. et al. Overexpression of HMGA2 promotes metastasis and impacts survival of colorectal cancers. Clin Cancer Res 17, 2570–2580 (2011).
Di Cello, F. et al. HMGA2 participates in transformation in human lung cancer. Molecular cancer research: MCR 6, 743–750 (2008).
Hristov, A. C. et al. HMGA2 protein expression correlates with lymph node metastasis and increased tumor grade in pancreatic ductal adenocarcinoma. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc 22, 43–49 (2009).
Raskin, L. et al. Transcriptome profiling identifies HMGA2 as a biomarker of melanoma progression and prognosis. The Journal of investigative dermatology 133, 2585–2592 (2013).
Winslow, M. M. et al. Suppression of lung adenocarcinoma progression by Nkx2-1. Nature 473, 101–104 (2011).
Fusco, A. & Fedele, M. Roles of HMGA proteins in cancer. Nature reviews. Cancer 7, 899–910 (2007).
Chiou, S. H. et al. A conditional system to specifically link disruption of protein-coding function with reporter expression in mice. Cell reports 7, 2078–2086 (2014).
Dangi-Garimella, S., Sahai, V., Ebine, K., Kumar, K. & Munshi, H. G. Three-dimensional collagen I promotes gemcitabine resistance in vitro in pancreatic cancer cells through HMGA2-dependent histone acetyltransferase expression. PLoS One 8, e64566 (2013).
Xiao, G., Wang, X. & Yu, Y. CXCR4/Let-7a Axis Regulates Metastasis and Chemoresistance of Pancreatic Cancer Cells Through Targeting HMGA2. Cellular physiology and biochemistry: international journal of experimental cellular physiology, biochemistry, and pharmacology 43, 840–851 (2017).
Gruner, B. M. et al. Modeling Therapy Response and Spatial Tissue Distribution of Erlotinib in Pancreatic Cancer. Mol Cancer Ther 15, 1145–1152 (2016).
Mazur, P. K. et al. Combined inhibition of BET family proteins and histone deacetylases as a potential epigenetics-based therapy for pancreatic ductal adenocarcinoma. Nat Med 21, 1163–1171 (2015).
Sgarra, R. et al. High Mobility Group A (HMGA) proteins: Molecular instigators of breast cancer onset and progression. Biochim Biophys Acta 1869, 216–229 (2018).
Watanabe, S. et al. HMGA2 maintains oncogenic RAS-induced epithelial-mesenchymal transition in human pancreatic cancer cells. Am J Pathol 174, 854–868 (2009).
Jiang, W. et al. miRNA-101 Suppresses Epithelial-to-Mesenchymal Transition by Targeting HMGA2 in Pancreatic Cancer Cells. Anti-cancer agents in medicinal chemistry 16, 432–439 (2016).
Morishita, A. et al. HMGA2 is a driver of tumor metastasis. Cancer Res 73, 4289–4299 (2013).
Zhou, X., Benson, K. F., Ashar, H. R. & Chada, K. Mutation responsible for the mouse pygmy phenotype in the developmentally regulated factor HMGI-C. Nature 376, 771–774 (1995).
White, A., Flores, A., Ong, J. & Lowry, W. E. Hmga2 is dispensable for cutaneous squamous cell carcinoma. Experimental dermatology 25, 409–412 (2016).
Sgarra, R. et al. Nuclear phosphoproteins HMGA and their relationship with chromatin structure and cancer. FEBS Lett 574, 1–8 (2004).
Gruner, B. M. et al. Modeling therapy response and spatial tissue distribution of erlotinib in pancreatic cancer. Mol Cancer Ther (2016).
Olive, K. P. et al. Inhibition of Hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science 324, 1457–1461 (2009).
Hingorani, S. R. et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell 4, 437–450 (2003).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev 15, 3243–3248 (2001).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nature neuroscience 13, 133–140 (2010).
Olive, K. P. et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell 119, 847–860 (2004).
Jackson, E. L. et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res 65, 10280–10288 (2005).
Gruner, B. M. et al. An in vivo multiplexed small-molecule screening platform. Nat Methods 13, 883–889 (2016).
Acknowledgements
We thank the Stanford Shared FACS Facility for expert assistance; Pauline Chu for histology services; Teri Longacre for the analysis of human PDAC IHC samples; Grace Kim for the analysis of mouse PDAC histology samples; Jennifer Brady, Pawel Mazur, the Winslow and the Grüner laboratories for helpful comments; and Sean Dolan and Alexandra Orantes for administrative support. This work was supported by a Pancreatic Cancer Action Network-AACR Award in memory of Skip Viraugh 13-20-25-WINS (to MMW), and in part by the Stanford Cancer Institute support grant (P30-CA124435) from the National Cancer Institute. B.M.G. was supported by the Pancreatic Cancer Action Network – AACR Fellowship in memory of Samuel Stroum (14-40-25-GRUE), was a Hope Funds for Cancer Research Fellow supported by the Hope Funds for Cancer Research (HFCR-15-06-07), and is a recipient of an Emmy Noether Award from the German Research Foundation (DFG).
Author information
Authors and Affiliations
Contributions
S.-H.C., B.M.G. and M.M.W. designed the project. B.M.G., S.-H.C., M.D. and E.K. performed experiments. S.N. genotyped mice. M.K. performed human TMA-IHC analysis. A.C.K. advised on the project. B.M.G. and M.M.W. wrote the manuscript with comments from all authors.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chiou, SH., Dorsch, M., Kusch, E. et al. Hmga2 is dispensable for pancreatic cancer development, metastasis, and therapy resistance. Sci Rep 8, 14008 (2018). https://doi.org/10.1038/s41598-018-32159-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32159-x
Keywords
This article is cited by
-
Decoding high mobility group A2 protein expression regulation and implications in human cancers
Discover Oncology (2024)
-
Epigenetic reprogramming-induced guanidinoacetic acid synthesis promotes pancreatic cancer metastasis and transcription-activating histone modifications
Journal of Experimental & Clinical Cancer Research (2023)
-
LINC00963 affects the development of colorectal cancer via MiR-532-3p/HMGA2 axis
Cancer Cell International (2021)
-
Role of non-coding RNAs and RNA modifiers in cancer therapy resistance
Molecular Cancer (2020)
-
An updated review of the H19 lncRNA in human cancer: molecular mechanism and diagnostic and therapeutic importance
Molecular Biology Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.