Abstract
Biotin is an essential cofactor required for carboxylation and decarboxylation reactions in all domains of life. While biotin biosynthesis in most Bacteria and Eukarya is well studied, the complete pathway for this vitamer in Archaea is still not known. Detailed genome searches indicated the presence of possible bio gene clusters only in Methanococcales and Thaumarchaeota. Therefore, we analysed the functionality of the predicted genes bioA, bioB, bioD and bioF in the Thaumarchaeon Nitrososphaera gargensis Ga2.9 which are essential for the later steps of biotin synthesis. In complementation tests, the gene cluster-encoded N. gargensis bioABD genes except bioF restored growth of corresponding E. coli Rosetta-gami 2 (DE3) deletion mutants. To find out how biotin biosynthesis is initiated, we searched the genome for a possible bioH analogue encoding a pimeloyl-ACP-methylester carboxylesterase. The respective amino acid sequence of the ORF estN1 showed weak conserved domain similarity to this class of enzymes (e-value 3.70e−42). Remarkably, EstN1 is a promiscuous carboxylesterase that complements E. coli ΔbioH and Mesorhizobium loti ΔbioZ mutants for growth on biotin-free minimal medium. Additional 3D-structural models support the hypothesis that EstN1 is a BioH analogue. Thus, this is the first report providing experimental evidence that Archaea carry functional bio genes.
Similar content being viewed by others
Introduction
Biotin-dependent carboxylases, decarboxylases and transcarboxylases are key enzymes of different ubiquitous reactions and are e.g. involved in CO2 fixation, fatty acid synthesis and amino acid catabolism1,2. With the exception of RuBisCO (ribulose-1,5-bisphosphate carboxylase/oxygenase), carboxylases from all domains of life require biotin as cofactor. Biotin can either be taken up from the environment or it can be synthesised as it has been shown for many bacteria, plants and fungi. Surprisingly, biosynthesis of biotin in Archaea has not been well studied.
Within Bacteria like E. coli, biotin is being synthesised with a bioABCDF operon and bioH. The first step of biotin synthesis, in which a biotin precursor is formed, is catalysed by BioC, a SAM-dependent methyl transferase (Fig. 1)3. The precursor molecule malonyl-ACP (acyl-carrier-protein) methyl ester is processed to pimeloyl-ACP methyl ester within the fatty acid synthesis cycle. This forms the substrate of BioH, a methyl ester carboxylesterase, which diverts pimeloyl-ACP methyl ester from the fatty acid cycle to the biotin synthesis pathway. BioH is thus responsible for a crucial step and has a “gatekeeper” function4. After hydrolysis and release of one molecule of methanol, pimeloyl-ACP undergoes an enzyme cascade finally leading to the biotin molecule5. In this process, a 7-keto-8-amino pelargonic acid (KAPA) synthase (BioF), a 7,8-diaminopelargonic acid (DAPA) aminotransferase (BioA), a dethiobiotin synthetase (BioD) and a biotin synthase [BioB, radical S-adenosylmethionin (SAM) superfamily] are involved. Regulation of this synthesis pathway is carried out by BirA, a biotin-protein ligase acting at the same time as transcriptional regulator of the bio gene cluster. Enzymes that carry out the same reaction like E. coli BioH and are thus also responsible for the cleavage of pimeloyl-ACP-methyl esters have been discovered for instance in Haemophilus influenza [BioG6], Helicobacter pylori [BioV7], Francisella species [BioJ8] and in cyanobacteria [BioK9]. The amino acid sequences of these enzymes differ clearly from each other and can be considered as evolutionary distinct although they have the same catalytic triad typical for α/β-hydrolases (usually formed by Ser, Asp/Glu and His) and similar structural properties7,10. In Mesorhizobium loti, BioZ shows amino acid sequence similarity to a β-ketoacyl-ACP synthase III, also known as FabH, but it was able to complement E. coli ΔbioH mutants11. While bioH in E. coli and bioV in H. influenzae are not integrated into the bio cluster, bioH in P. aeruginosa is and thus can be co-transcribed. The expression level of the operon-encoded bioH in P. aeruginosa is higher12. While the operon encoded genes bioABDF are highly conserved in Bacteria, bioC and bioH can be exchanged. In certain Bacillus species, the BioI (cytochrome P450) and BioW (pimeloyl-CoA synthetase) pathway has been discovered. Unlike the carbon-carbon bond cleaving BioI, BioW is essential for biotin synthesis and converts pimelic acid from the fatty acid synthesis pathway into pimeloyl-CoA13.
Synthesis pathway of biotin as it is known for Bacteria. In E. coli, BioC and BioH provide the pimeloyl moiety. In Bacillus spp., BioW produces the precursor molecule for biotin. The other genes bioFADB synthesizing the two fused rings and finally biotin are highly conserved5. E. coli BioF can convert both pimeloyl-CoA and -ACP while Bacillus subtilis BioF can only accept pimeloyl-CoA42.
Compared to this variety of different enzymes involved in bacterial biotin synthesis, biotin synthesis in Archaea is far less investigated. By looking into genome datasets, it can be concluded that solely the euryarchaeal Methanococcales seem to have a complete set of biotin synthesis genes with bioW, bioF, bioA, bioD and bioB. Archaea from many other different clades like Lokiarchaeota, Korarchaeota, Sulfolobales and Thermococcales seem to contain only bioB genes. This leads to the assumption that these Archaea might synthesise biotin either in an unknown pathway or their biotin synthesis enzymes do not show enough similarity to their bacterial counterparts to be recognised as such. Other members of Thermoproteales, Methanosarcinales or Archaeoglobi might not be able to synthesise biotin at all as not even bioB can be found with database searches. Thaumarchaeota possess an incomplete set of biotin synthesis genes. Considering the impact and influential role they play for biogeochemical processes and for which they require biotin, their essential bio genes to fulfil these tasks have been investigated in this study more closely.
Thaumarchaeota represent the third largest archaeal phylum behind Euryarchaeota and Crenarchaeota14. Members of the thaumarchaeal lineage are highly abundant in marine, freshwater and soil ecosystems. Most of them are mesophilic ammonia-oxidisers that are capable of autotrophic CO2 fixation15. Additionally, some members are able to shift to a mixotrophic lifestyle by utilizing organic compounds16. The significant fixation rates of Thaumarchaeota for global carbon- and nitrogen-cycles have long been underestimated17,18,19. Still, thaumarchaeal genomes possess high numbers of genes with general function prediction only and even genes encoding hypothetical proteins with no known conserved domains at all. Within six thaumarchaeal genomes (Nitrososphaera spp. and Nitrosocosmicus spp.) deposited in the IMG/MER database20, between 46 and 51% of the predicted proteins are without function prediction, and therefore many pathways still remain incomplete.
In this study, we investigated the soil group I.1b Thaumarchaeon Nitrososphaera gargensis which was originally isolated from hot spring microbial mats and its hypothetical proteins involved in biosynthesis of the essential cofactor biotin. Genome analyses of the mesothermophilic ammonia-oxidiser N. gargensis revealed potential enzymatic pathways for the citric acid cycle, gluconeogenesis and the oxidative pentose-phosphate-pathway21,22,23. Like other ammonia-oxidizing Archaea (AOA), N. gargensis couples ammonia-oxidation with carbon fixation and assimilates inorganic carbon via an energy-efficient autotrophic 3-hydroxypropionate/4-hydroxybutyrate (HP/HB) cycle. This carbon-fixation pathway is even more efficient in Thaumarchaeota than in Crenarchaeota which use a modified HP/HB-pathway17. As the carboxylation reactions involved are biotin-dependent, they can be completely inhibited in presence of the specific inhibitor avidin. The N. gargensis genome is composed of 2.8 Mb of DNA encoding 3,562 genes of which so far only 52.4% have a function prediction (source: IMG). N. gargensis contains a predicted biotin synthesis gene cluster with bioF, bioA, bioD and bioB. Their function has not been demonstrated until now. The early stage synthesis genes bioH, bioC or bioW/I were not annotated or present as such. Our aim was to find out how this Archaeon produces biotin and therefore, the putative bioFADB genes and different α/β-hydrolases from N. gargensis that could have methyl ester carboxylesterase activity similar to BioH were studied for their functionality.
Materials and Methods
Cultivation of N. gargensis and genomic DNA extraction
N. gargensis was grown for at least eight weeks at 46 °C in a 300 ml Erlenmeyer flask containing 100 ml mineral salts medium24. For larger cell quantities, 5 l flasks containing 1.5 l of medium were incubated at 46 °C under slow stirring. The biotin-free medium was supplemented with trace element solution25 and contained 10 mM NaCl and 5 mM NH4Cl. The pH was controlled with cresol red indicator and was steadily adjusted with a solution of 5% (w/v) NaHCO3 to a pH of approx. 7.0. DNA was extracted from the cell pellet of a 1.5 l N. gargensis culture using a phenol-chloroform extraction method with RNase (1 mg/ml) containing lysis buffer and a proteinase K (10 mg/ml) incubation step for 1 h at 37 °C.
Cloning of putative bio- and α/β-hydrolase-genes from N. gargensis into E. coli and activity screening on tributyrin (TBT)
From the genome sequence of N. gargensis (NCBI GenBank accession no. CP002408), the putative bioF, bioA, bioD and bioB genes were chosen for cloning (Table S1). Six different ORFs that were annotated as putative α/β-hydrolase fold proteins and could therefore serve as counterpart of the bacterial BioH were also selected as well as four possible candidates that could serve as bioC analogues (bioC1-4). The ORFs Ngar_c21820, Ngar_c24650, Ngar_c14400 (i. e. estN1), Ngar_c30910 (i. e. estN2), Ngar_c32780 and Ngar_c35080, which could possibly encode a BioH analogue, were amplified with specific primers, cloned into pDrive (PCR Cloning Kit, Qiagen, Hilden, Germany) and transformed into competent E. coli DH5α cells. As they encode carboxylesterases, the different clones were first tested on TBT agar plates at 37 °C for general activity. The plates were incubated for up to 6 days at 37 °C in order to see halos around active E. coli colonies expressing active carboxylesterases from N. gargensis. The genes estN1 and estN2 from the active clones and the bioFADBC genes were further separately cloned into the multiple-cloning site of pET21a(+) expression vectors (Novagen/Merck, Darmstadt, Germany) encoding a C-terminal six-fold histidine tag (His6-tag).
Construction of biotin auxotrophic E. coli mutants
To test the functionality of the different N. gargensis bio-genes recombinantly, E. coli Rosetta-gami 2 (DE3) was chosen as a strain for creating insertion mutants of various biotin synthesis genes (bioA, bioB, bioC, bioD, bioF and bioH). Proteins from N. gargensis were much better expressed in this strain compared to e.g. BL21. Using the Gene Bridges “E. coli Gene Deletion Kit”, the manufacturer’s protocols were followed with prolonged incubation times as adjustment to the slower growth of Rosetta-gami 2 compared to other E. coli expression strains (approx. 1.5- to 2-times longer). As recombination plasmid, pRedET transferring ampicillin resistance was used. For each target ORF, a forward and reverse oligo nucleotide was designed that contained 50 bp of the target sequence and a homology arm region for the FRT-PGK-gb2-neo-FRT cassette that provides resistance to kanamycin (Table S1). Finally, the insertion mutants contained a 1,737 bp kan-cassette instead of their original gene. On medium without biotin, the strains were unable to grow.
Functional complementation studies in E. coli on M9 medium
The E. coli Rosetta-gami 2 insertion mutants ΔbioA, ΔbioB, ΔbioC, ΔbioD, ΔbioF and ΔbioH were electroporated with the corresponding E. coli genes ligated into pET21a serving as controls to restore wild type activities and the corresponding genes from N. gargensis to check for their ability to complement growth (Table S1). Empty pET21a vector served as negative control. Syntheses of genes from N. viennensis EN76 (NCBI acc. no. AIC14436) and N. evergladensis SR1 (NTE_01571) with similarity to estN1 were performed at Eurofins (Ebersberg, Germany). The corresponding genes were also cloned into pET21a and transformed into the ΔbioH mutants. All clones were tested for growth on M9 minimal medium26 with and without 4 nM D-biotin supplemented. The medium contained 0.8% (w/v) glucose, 100 µg/ml thiamin and 0.1% (w/v) casamino acids suitable for vitamin assays. As selection pressure and in order to induce expression of the plasmid-encoded genes under the T7 promoter, the agar plates contained 25 µg/ml chloramphenicol, 50 µg/ml ampicillin, 15 µg/ml kanamycin and 10 µg/ml IPTG. This IPTG amount showed the same result only with a faster colony growth compared to 100 µg/ml IPTG. After one to two days of incubation at 37 °C, the plates were checked for visible growth of colonies.
Complementation of Mesorhizobium loti ΔbioZ on Rhizobial Defined Medium (RDM)
The genes estN1 and estN2 from N. gargensis as well as bioH from E. coli were cloned from pET21a including His-tag coding sequence into the broad-host-range vector pBBR1MCS-5 via the ApaI and PstI restriction sites. M. loti ΔbioZ11 was transformed with the constructs and an empty vector control by triparental conjugation using E. coli DH5α as donor and E. coli K12 H101 with the helper plasmid pRK2013. M. loti ΔbioZ colonies were grown at 28 °C and selected on TY agar containing 200 µg/ml neomycin and 50 µg/ml gentamicin. In order to select for growth on biotin-free medium, RDM was prepared27 containing glucose (10 mM), trace elements25 and either none or 20 nM biotin.
Overexpression of EstN1 in E. coli Rosetta-gami 2 (DE3) and enzyme purification
As mentioned above, the gene estN1 (Ngar_c14400) encoding an active esterase was cloned into the pET21a expression vector (Table S1). The plasmid was transformed into E. coli BL21 [DE3; F− ompT hsdSg(rg−mg−) gal dcm] and Rosetta-gami 2 [DE3; Δara-leu7 679 ΔlacX74 ΔphoA PvuII phoR araD139 ahpC galE galK rpsL F−(lac lacI pro) gor522::Tn10 trxB pRARE2] for comparative expression studies (both purchased from Novagen/Merck, Darmstadt, Germany). While the BL21 overnight culture containing estN1::pET21a was grown on medium with 100 µg/ml ampicillin, cultures of E. coli Rosetta-gami 2 strains were grown on LB medium supplemented with 34 µg/ml chloramphenicol, 3 µg/ml tetracycline and 100 µg/ml ampicillin. One litre of fresh LB medium containing the above mentioned antibiotics was inoculated in a 3 litre flask with 1% (v/v) of the overnight culture and incubated at 37 °C under stirring until an OD600 of 0.8 was reached. Therefore, the BL21 cultures were incubated for approx. 4 hours at 37 °C. The Rosetta-gami 2 cultures were slower in growth and reached an OD600 of 0.8 after approx. 7 hours. Overexpression of the enzyme was then induced by addition of IPTG to the cultures to a final concentration of 1 mM. The BL21 culture was incubated and stirred for further 4 hours at 37 °C and the Rosetta-gami 2 culture for further 16 to 18 hours at 28 °C. Cells containing recombinantly produced EstN1 were harvested by centrifugation, resuspended in lysis buffer and crude cell extracts were prepared by two passages through a French pressure cell press. Cell debris was collected by centrifugation. The target protein which contained a C-terminal His6-tag was purified from the clear supernatant via immobilised nickel ion affinity chromatography using Protino®Ni-NTA agarose (Macherey-Nagel, Düren, Germany) and eluted with a buffer containing 250 mM imidazole. The enzyme was dialysed and concentrated by centrifugation in a Vivaspin®6 ultrafiltration unit (MWCO 10,000; Sartorius, Goettingen, Germany) by washing twice with 0.1 M potassium phosphate buffer (pH 7.0). The protein was analysed by SDS-PAGE with gels containing 12% (separating gel) and 7% (stacking gel) acrylamide.
Characterization of EstN1’s activity
Substrate spectrum on para-nitrophenyl esters (pNP)
Nine different pNP-ester compounds with a distinct residue chain length between 2 and 18 carbon atoms have been tested as substrates in a final concentration of 0.5 to 1 mM in 0.1 M potassium-phosphate buffer at pH 7.0. Unless otherwise indicated, this phosphate buffer was also used in the following assays. The reactions took place at 40 °C. 15 µg of purified EstN1 were added to 200 µl of the reaction mixture and incubated for 20 minutes before the reaction product p-nitrophenol was quantified spectrophotometrically at 405 nm.
Determination of esterase activity against p-nitrophenyl propionate (pNPP)
This substrate was tested in a final concentration of 1 mM in 0.1 M potassium-phosphate buffer at pH 7.0. The reaction was initiated by the addition of a stock solution (100 mM in acetonitrile) of pNPP to 195 µl of 0.1 M potassium-phosphate buffer containing 0.1 mg purified protein in 96-well microtiter plates. Reaction was followed at 405 nm by UV-VIS spectrophotometry in a Synergy HT Multi-Mode Microplate Reader. Initial rates were determined from linear fits of the absorbance versus time (<10% conversion) and corrected for the rate of un-catalysed hydrolysis. One unit (U) of enzyme activity was defined as the number of enzyme required to transform 1 µmol substrate in 1 min under the assay conditions using the reported extinction coefficient (∑pNPP at 405 nm = 15200 M−1 cm−1). Reactions (performed in triplicate) were maintained at 40 °C.
Activity on mono- and dimethyl esters
Methyl acetate, methyl propionate, methyl butyrate, methyl hexanoate, methyl octanoate and dimethyl pimelate were tested as substrates. All substrates were diluted to a concentration of 100 mg/ml in dimethylsulfoxide (DMSO) and enzymatic hydrolysis was monitored in triplicates over 15 hours in a microtiter plate-based pH-shift assay using phenol red as indicator in a Synergy HT Microplate Reader at room temperature (25 °C). During hydrolysis, the increasingly acidic pH within the reactions was measured spectrophotometrically at 550 nm with an extinction coefficient of 8450 M−1 cm−1 28.
Determination of acyl-CoA thioesterase activity
Acyl-CoA thioesterase activity was measured spectrophotometrically at 412 nm with 5,5′-dithiobis(2-nitrobenzoic acid) [DTNB29]. The medium contained 0.1 M potassium-phosphate buffer, 0.05 mM DTNB, 10 μM of acyl-CoAs (acetyl-CoA, n-propionyl-CoA, methylmalonyl-CoA, malonyl-CoA, succinyl-CoA or palmitoyl-CoA) and 0.1 mg purified protein. Reactions were followed at 412 nm by UV-VIS spectrophotometry in a Synergy HT Multi-Mode Microplate Reader. Initial rates were determined from linear fits of the absorbance versus time (<10% conversion) corrected for the rate of un-catalyzed hydrolysis. In all cases, one unit (U) of enzyme activity was defined as the number of enzyme required to transform 1 µmol substrate in 1 min under the assay conditions. An E412 = 13600 m−1 cm−1 was used to calculate the activity. Reactions were performed like in all other assays in triplicate and maintained at 40 °C.
Temperature optimum and temperature stability
Due to its higher stability against auto-hydrolysis, 0.5 mM pNP-C6 (hexanoate) was used for measuring the enzyme activity at different temperatures. To find the optimal temperature, the enzyme was pre-incubated between 10 °C and 90 °C for 5 minutes before the substrate was added to 0.5 mM concentration and the reactions were incubated for further 30 minutes. The temperature stability was tested by incubating the enzyme at 40 °C and 60 °C for up to 3.5 hours. The residual activity was then measured after addition of pNP-C6 and incubation at 40 °C for 30 minutes.
pH optimum
Enzyme reactions were carried out at 40 °C for 30 minutes with pNP-C6 using different citrate (pH 4.0 to 5.6), phosphate (pH 5.6 to 8.0), Tris-HCl (8.0 to 9.0) and glycine-NaOH (pH 9.0 to 10.6) buffers to find the optimal pH.
Influence of cofactors, solvent stability and stability against inhibitors and detergents
Ca2+, Co2+, Cu2+, Fe3+, Mg2+, Mn2+, Rb2+ and Zn2+ in 1 mM and 10 mM concentration were added to the standard enzyme reaction (30 min incubation at pH 7.0 and 40 °C with pNP-C6) in order to see if the enzyme activity depends on cofactors. As solvents, DMSO, isopropanol, methanol, dimethyl formamide (DMF), acetone, acetonitrile or ethanol were added to the standard reaction in 10% and 30% (v/v) concentration. As detergents, sodium dodecyl sulfate (SDS), Triton-X-100 and Tween80 [1 and 5% (v/v)] and as inhibitors in 1 and 10 mM concentration EDTA, dithiothreitol (DTT) or phenylmethylsulfonyl fluoride (PMSF) were tested to assay the residual activity after treatment with these substances.
Database searches, structural prediction and Hidden-Markov-Model (HMM) search for similar enzymes
The genome sequence of N. gargensis21 was retrieved from the NCBI GenBank database [NCBI Resource Coordinators30] under accession no. CP002408. Together with other archaeal genomes deposited in the KEGG database (www.kegg.jp) and the Integrated Microbial Genomes (IMG) database of the Joint Genome Institute [https://img.jgi.doe.gov]31, the genomes were searched for genes related to the biotin biosynthetic pathway. Analysis of biotin synthesis enzymes from all archaeal phyla was accomplished using similarity analyses, KEGG pathway analyses, GenBank and IMG and included BioW, BioC, BioH, BioF, BioA, BioD and BioB. Conserved domain searches were performed using the NCBI web tool [https://www.ncbi.nlm.nih.gov/cdd]32 using amino acid sequences of the proteins of interest.
Amino acid sequences of two enzymes originating from closely related Thaumarchaeota that show relatively high similarity to EstN1 were retrieved from the NCBI database after performing BlastX and BlastP searches [NCBI Resource Coordinators]30. The sequences were aligned with T-coffee33 and a profile HMM was built with the alignment using the hmmbuild function of the HMMER package (http://hmmer.org). The Uniprot database34 was searched with this HMM to find similar sequences. These sequences were aligned and a new HMM was built by going through the same procedure as mentioned above. Highly conserved amino acid residues were identified after visual inspection of a HMM logo created with the webtool Skylign35. The relationship between the amino acid sequences of the different proteins was visualised in a neighbour-joining phylogenetic tree constructed with MEGA636.
A homology model of the possible structure of EstN1 was calculated using the Robetta Full-chain Protein Structure Prediction Server [http://robetta.bakerlab.org]37 with a Ginzu prediction method. PDB files were created or retrieved from the PDB database (https://www.rcsb.org/) that was processed and visualised with the program Chimera [https://www.cgl.ucsf.edu/chimera]38. Pairwise comparison of protein structures was accomplished using the Dali server [http://ekhidna.biocenter.helsinki.fi/dali_lite]39.
Results and Discussion
Within the KEGG database, 140 available archaeal genome sequences from 22 different phyla were searched for biotin synthesis genes (Table 1). In all but two phyla, the synthesis pathways are fragmentary although 119 of the 140 genomes contain a predicted bioB gene (Fig. 2). In Thaumarchaeota, genes for the synthesis starting from pimeloyl-CoA to biotin have been predicted. However no genes involved in the early synthesis steps were identified. The Methanococcales appear to be the only group within the archaeal domain that seems to have a complete biotin synthesis gene set and in which pimeloyl-CoA could be provided by the 6-carboxyhexanoate-CoA-ligase BioW. Remarkably, none of these genes has been verified for functionality.
Within this framework, we decided to investigate the functionality of the predicted bio genes in the phylum Thaumarchaeota and especially in N. gargensis as the way it obtains pimeloyl-ACP or -CoA biotin precursor molecules is not known.
Complementation of E. coli Rosetta-gami 2 (DE3) ΔbioF, ΔbioA, ΔbioD, ΔbioB and ΔbioC mutants
Because of the large phylogenetic distance of the N. gargensis genes to E. coli and the known problems concerning expression of archaeal proteins, E. coli Rosetta-gami 2 was chosen as host for complementation assays. It carries genes for seven additional tRNAs that help to solve the rare codon usage problem and increase protein yield40 as well as trxB/gor mutations for enhanced disulfide bond formation41. To investigate the functionality and compatibility of N. gargensis genes annotated as enzymes involved in biotin synthesis, five different auxotrophic E. coli Rosetta-gami 2 mutant strains were created. Each of the mutants ΔbioF, ΔbioA, ΔbioD, ΔbioB and ΔbioC carries a kanamycin resistance gene instead of the respective gene. The mutants were verified by PCR and failed to grow on M9 media not supplemented with nanomolar amounts of biotin. Furthermore, the mutant strains transformed with an empty pET21a vector as negative control also failed to grow without biotin (Fig. 3). In our experiments, the mutant strains were complemented with the corresponding E. coli gene and with the putative N. gargensis counterpart. On biotin-free medium, E. coli ΔbioA, ΔbioB and ΔbioD showed sufficient growth when complemented with both the E. coli and the N. gargensis version of the corresponding genes. E. coli ΔbioA and ΔbioD could be restored to approximately half of the full growth by N. gargensis’ bioA and bioD. The E. coli ΔbioF mutant, however, could only be complemented with E. coli’s bioF and not with the N. gargensis equivalent (Fig. 3). Since bioDABF are located in a gene cluster (NCBI acc. no. Ngar_c17540-Ngar_c17580), we speculate that bioF is active, but maybe only on pimeloyl-CoA and not on pimeloyl-ACP provided by E. coli. This is also the case for Bacillus subtilis BioF42 and would make sense as no ACPs are known for Archaea43.
Auxotrophic insertion mutants of E. coli Rosetta-gami 2 (DE3) bioA, bioB, bioD and bioF were complemented with the corresponding bioA/bioB/bioD/bioF-genes from E. coli and N. gargensis and an empty vector pET21a served as control. The clones were assayed for growth on M9 medium without biotin and under supplementation of 4 nM biotin. All complementation assays were carried out in triplicate in at least three independent experiments.
An O-methyltransferase and bioC-analogue was not found in the genome. Concerning bioC, only four out of 15 possible bioC candidates annotated as methyltransferases from N. gargensis were tested and failed to complement the E. coli ΔbioC mutant. Further research is necessary in this case.
We conclude that N. gargensis possesses at least the functional biotin synthesis genes bioA, bioD and bioB.
Conserved domains of the six putative α/β-hydrolases from N. gargensis
Six different α/β-hydrolase genes without further specified function prediction were chosen from the N. gargensis genome in order to be tested for functionality and for possible BioH-activities. According to a conserved domain search, three of the candidate enzymes (Ngar_c21820, Ngar_c24650 and Ngar_c35080) showed domains for being members of the α/β-hydrolase superfamily (e-values between 1.9e−44 and 2.1e−17), while three others even showed possible pimeloyl-ACP methyl ester carboxylesterase domains [Ngar_c14400 (i.e. EstN1), Ngar_c30910 (i.e. EstN2) and Ngar_c32780; e-values between 4.2e−41 and 2.0e−20].
Hydrolytic activity of carboxylesterases from N. gargensis after expression in E. coli
The six different putative α/β-hydrolase genes Ngar_c21820, Ngar_c24650, Ngar_c14400 (estN1), Ngar_c30910 (estN2), Ngar_c32780 and Ngar_c35080 were cloned into E. coli DH5α. As it is known that BioH enzymes are slightly promiscuous and able to hydrolyse the triglyceride tributyrin (TBT44), the respective clones were screened for activity on LB agar plates containing 1% of TBT. This step was performed to ensure that only active enzymes were further pursued. After incubation of one day at 37 °C and one day at 46 °C, three of the clones showed reproducible activity and can therefore be further specified as carboxylesterases (EC 3.1.1). The clones contained the genes estN1 (786 bp), estN2 (834 bp) and Ngar_c35080 (789 bp). Due to its very low activity, the clone containing Ngar_c35080 was not further investigated.
Sequence analysis of the N. gargensis carboxylesterases EstN1 and EstN2
The amino acid (aa) sequences of the two enzymes EstN1 (261 aa, estimated size of 29.6 kDa) and EstN2 (277 aa, estimated size of 31.1 kDa) show similarities to enzymes from other Thaumarchaeota. EstN1 (GenBank AFU58376) has similarity to other not further specified putative α/β-hydrolases from a soil-metagenome-derived Thaumarchaeon (89% identity, e-value 1e−175), Candidatus Nitrososphaera evergladensis SR1 (72%, 8e−132) and two Nitrososphaera viennensis strains (72%, 8e−132 and 1e−131, respectively) according to a BlastP search. It has an identity of less than 52% to other thaumarchaeal hydrolases. In case of EstN2 (GenBank and PDB acc. no. 5A62_A45), only one enzyme from Nitrososphaera evergladensis SR1 annotated as putative hydrolase or acyltransferase of the α/β-superfamily shows a fairly higher similarity with 70% identity and an e-value of 1e−130. The next best hits belonged to Sulfolobus islandicus strains (30–31% identity, e-values 4e−25 to 2e−24), but also to Bacteria like Chloroflexi (29%, 5e−28) or Rhodobacter sp. (30%, 2e−26).
Complementation studies in E. coli Rosetta-gami 2 ΔbioH and Mesorhizobium loti ΔbioZ mutants
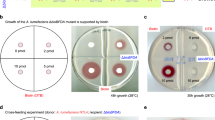
BioH is an essential enzyme for biotin biosynthesis46 and ΔbioH mutants are dependent on biotin supplementation. An E. coli Rosetta-gami 2 ΔbioH mutant was created and transformed with pET21a as an empty vector control, with bioH::pET21a from E. coli to restore wild-type growth, and with estN1 and estN2 ligated into pET21a. After one to two days of incubation at 37 °C, all strains were grown on M9 medium with 4 nM biotin supplemented. On biotin-free medium, only the mutant strains containing bioH::pET21a and estN1::pET21a were able to grow (Fig. 4A). Thus (unlike EstN2) EstN1 catalyses the same hydrolysis reaction on pimeloyl-ACP-methyl ester like BioH.
Complementation of E. coli ΔbioH (A) and M. loti ΔbioZ (B) auxotrophic mutants on M9 and RDM medium. The mutants were complemented with empty vector and the plasmid encoded genes from E. coli and N. gargensis. The N. gargensis gene estN1 was able to restore growth in both bacterial strains in contrast to the other esterase estN2.
Within the α-proteobacterium M. loti, the 3-oxoacyl-ACP-synthase BioZ bypasses BioH and leads to pimeloyl-ACP production in another way. BioZ is a FabH (β-ketoacyl-ACP synthase) homologue and not an α/β-hydrolase. It has a catalytic Cys/His/Asn triad47,48. Thus, it is not able to cleave the methyl ester like BioH with its catalytic Ser. However, E. coli’s BioH complemented an auxotrophic M. loti ΔbioZ mutant11 and therefore, M. loti ΔbioZ was transformed with an empty broad-host-range vector pBBR1MCS-5 as well as with bioH, estN1 and estN2 in pBBR1MCS-5. All strains were able to grow at 28 °C on RDM medium supplemented with 20 nM biotin after four days. Without the addition of biotin, growth was significantly impaired for cells containing the empty vector and estN2, while E. coli’s bioH was able to restore wild-type growth which is consistent with previous findings11. Growth of M. loti ΔbioZ was even stronger with estN1 that could apparently fully complement bioZ (Fig. 4B).
Recombinant production and biochemical characterization of EstN1
EstN1 was overexpressed with a C-terminal His6-tag in E. coli BL21 and Rosetta-gami 2 to study its enzymatic properties. As previously mentioned, Rosetta-gami 2 contains an extra plasmid providing tRNAs for rare codons and imparts improved disulfide bond formation and protein folding. The enzyme was expressed in this strain with much higher yields compared to BL21. The enzyme was purified with Ni-NTA-agarose (Fig. S1) and subjected to buffer change and concentration before its activity was characterised. EstN1 was mostly active on short-chain pNP-esters as model substrates. On pNP-esters, the highest esterase activity was measured with 10.1 ± 0.19 U/mg on pNPP (-C3). It was significantly higher than the activity observed for other shorter or longer pNP esters, namely, pNP-acetate (-C2) with 0.2 U/mg followed by 0.1 U/mg on pNP- butyrate (- C4) and 0.07 U/mg on pNP-hexanoate (-C6). This substrate spectrum on pNP esters and the preference for short acyl chains are very similar to the ones of E. coli BioH49.
The carboxylesterase was mostly active (>60% relative activity) in a moderately thermophilic range between 30 and 55 °C in agreement with the optimal growth temperature of N. gargensis (∼46 °C). The highest activity was measured at 40 °C. Beneath 20 °C and above 60 °C, only low activity could be detected (<35% rel. activity) (Fig. S2, left). EstN1 is most active in a range between pH 5 and 8 with an optimum at pH 6 in phosphate buffer (Fig. S2, middle). This differs to the rather narrow pH optimum of BioH from E. coli (pH 8.0–8.5). EstN1 has a half-life-time (50% residual activity) of approx. 135 minutes at 40 °C and 15 minutes at 60 °C (Fig. S2, right).
EstN1 showed activity on methyl esters, but the activity was comparably low with 0.06 U/mg on methyl hexanoate. The activity was lower on shorter and longer residues. A higher activity was observed for dimethyl pimelate with 0.35 U/mg. Dimethyl pimelate resembles the proposed natural substrate pimeloyl-ACP-methyl ester; still, it is a much smaller molecule. Concerning the specific activity, it is noteworthy that the monomethyl and dimethyl ester assays could not be carried out at 40 °C due to restraints in experimental setup, but at room temperature (25 °C) and that the values are expected to be higher under optimal conditions.
EstN1 was finally tested for thioesterase activity against diverse acyl-CoAs to learn more about its promiscuity and possible activity on substrates similar to pimeloyl-CoA. It showed its overall highest activity at 40 °C towards n-propionyl-CoA (14.6 ± 0.3 U/mg) and to a much lower extent for acetyl-CoA (0.26 ± 0.03 U/mg), but no considerable activity on any of the other derivatives tested with more complex residues. Given the fact that thirteen out of 22 structurally different substrates were hydrolysed, EstN1 can be considered as enzyme with slightly promiscuous activity.
EstN1 is a cofactor-independent carboxylesterase as none of the tested ions Ca2+, Co2+, Cu2+, Fe3+, Mg2+, Mn2+, Rb2+ and Zn2+ enhanced its activity significantly. It was susceptible to enzyme inhibitors and retained between 30% (10 mM PMSF) and 66% (10 mM EDTA) residual activity after treatment for 15 minutes. DTT diminished its activity completely. While Triton X-100 and Tween 80 did not significantly affect the enzyme, SDS inhibited EstN1 completely. When treated with the solvents DMSO, isopropanol, methanol, DMF, acetone, acetonitrile and ethanol, EstN1 generally kept its activity between 81% (isopropanol) or even enhanced it to 105% (DMF) when 10% of the solvents were applied. In higher concentrations of 30%, the solvents decreased the enzymatic activity to values between 43% (acetone and isopropanol) and 90% (acetonitrile). DMSO and DMF enhanced the activity in 30% concentration to 123 and 108%, respectively.
Homology model of EstN1
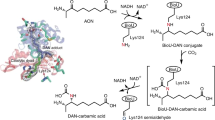
A model of EstN1’s protein structure has been calculated with the prediction server Robetta and the Ginzu PDB template identification and domain prediction protocol. Regions of EstN1’s protein chains were aligned to PDB templates. Altogether, the structural model has been calculated out of 15 different X-ray structures of α/β-hydrolases from different Bacteria like Burkholderia xenovorans (PDB code 2XUA), Pseudomonas fluorescens (2D0D and 1IUN) and a metagenomic α/β-hydrolase (4Q3L). The identities were rather low with a maximum of 29.45% to a meta-cleavage product hydrolase CumD from P. fluorescens (2D0D). The comparative model shows a central β-sheet composed of eight parallel β-strands, one of them in antiparallel orientation (Fig. 5A). The catalytic Ser93, His238 and Asp211 are positioned in the centre and can be accessed by small molecules through a hydrophobic tunnel (Fig. S3A). Intriguingly, the protein structure of BioH from E. coli (1M33) was not considered in this automated modelling process, because its amino acid sequence similarity to EstN1 is too low (22% on the total sequence). As both enzymes catalyse the same reaction, the structure of BioH was retrieved from the PDB database and compared to EstN1 manually (Fig. 5B). For BioH, a proof that pimeloyl-ACP-methyl ester is the natural substrate has been made by co-crystallization with this substrate4. The structures of EstN1 and BioH show many similarities regarding their tertiary structure, surface hydrophobicity and architecture of their catalytic domain (Fig. S3). The structural similarity between the two enzymes is given with a Z-score of 28.8 and a RMSD (root-mean-square deviation) of 2.3 based on a pairwise structural alignment using the DALI tool. It is visualised by superimposition of the two structures (Fig. S3C). Next to its highly similar tertiary structure, BioH has a catalytic triad composed of a Ser82, His235, and Asp20749, but the catalytic serine is surrounded by a GWSLGG motif in comparison to the GSSFGG motif of the three enzymes from the Nitrososphaera species which have been taken into account in a sequence alignment (Fig. S4). It is known that esterases like BioG from Haemophilus influenza, BioK from cyanobacteria, BioJ from Francisella spp. and BioV from Helicobacter spp. have the same function like BioH7. The enzymes can be structurally different from each other, but, based on homology models, all have similar components including eight β-sheets, six α-helices and the catalytic residues Ser, His and Asp situated on loops.
Structural comparison between the homology modelled enzyme structure of N. gargensis EstN1 (A) and the crystal structure of E. coli BioH (B); (PDB 1M33). The catalytic serine residue of both structures as well as the N- and C-termini are indicated. The EstN1 structure was calculated with Robetta using the Ginzu prediction protocol.
Comparison between EstN1 and similar esterases from N. evergladensis and N. viennensis
Next to an enzyme from an uncultured Thaumarchaeon (OLC87659), the two esterases from the closely related N. evergladensis SR1 (AIF83634) and N. viennensis EN76 (WP_084790873) show the highest similarity to EstN1 on amino acid level with an identity of 72% and an E-value of 8e−132. The two esterases were synthesised, cloned into E. coli Rosetta-gami 2 ΔbioH and tested for their ability to perform the same BioH activity as EstN1. However, the enzymes were not able to complement for growth on M9 agar plates without biotin despite their relatively high similarity to EstN1 (Fig. S5). Within the amino acid alignment of the three sequences together with BioH, some residues become apparent that only EstN1 and BioH have in common and might explain this difference (Fig. S4). Only few amino acid substitutions can shift the activity of a carboxylesterase towards BioH activity50. It was supposed in previous studies that an esterase which is actually involved in another pathway could have shifted its activity by successive point-mutations and gene duplications to a higher specificity for pimeloyl-ACP-methyl esters7,51. This protein evolution could have happened in N. gargensis and could also explain EstN1’s slightly promiscuous activity on other substrates than pimeloyl-ACP or -CoA methyl ester. The biotin synthesis operon of N. evergladensis and N. viennensis are arranged differently to the operon from N. gargensis which is also a sign that the ability to synthesise biotin evolved separately within the species (Fig. 6). It is possible that within these two Nitrososphaera species another esterase has taken over the pimeloyl-ACP or -CoA methyl esterase function or perhaps only the CoA-form can be hydrolysed. Nevertheless, it cannot be fully excluded that EstN1 rather coincidentally cleaves pimeloyl-ACP methyl ester and randomly fulfils the BioH activity with its promiscuous activity on methyl esters, an activity that the other two enzymes might not have in the same way.
Biotin synthesis operons of three Nitrososphaera species. ORF context of estN1 and genes from N. evergladensis and N. viennensis with the highest similarity to estN1. Function prediction for N. gargensis and N. evergladensis retrieved from NCBI, for N. viennensis from IMG (coding regions named according to the NCBI database). The amino acid sequence identities of the N. evergladensis and N. viennensis biotin synthesis proteins to the amino acid sequences from N. gargensis are indicated as percentages beneath the corresponding genes. *Functional proof is missing.
Phylogenetic analysis of EstN1 homologues based on amino acid sequence similarities
We took amino acid sequences of the α/β-hydrolases from N. evergladensis SR1 and N. viennensis EN76 that showed the highest similarity to EstN1 in order to construct a Hidden-Markov-model (HMM) that points out conserved residues and motifs. Application of this HMM in Uniprot database searches with a significance E-value of 0.01 set as cut-off resulted in two other Thaumarchaeal α/β-hydrolase hits from a metagenomic Thaumarchaeon 13_1_40cm (GenBank OLC87659) and Candidatus Nitrosotenuis chungbukensis (WP_042686824). A refined HMM was calculated out of these five sequences and applied for a new database search that resulted in 49 hits. All hits belonged to members of the Thaumarchaeota and a large fraction originates from uncultivated marine Thaumarchaeota (Fig. 7). Other hits group to members of the genus Nitrosopumilus, Nitrosotalea or Nitrososphaera. The HMM logo (Fig. S6) leading to these sequences shows the conserved catalytic triad with a Ser (position 96), Asp (pos. 214) and His (pos. 242). Highly conserved amino acids are furthermore tryptophan (W, pos. 38, 210), proline (P, pos. 43, 65,142, 176, 218, 244, 249) and glycine (G, pos. 30, 31, 58, 60, 123, 211). Three other glycine residues (pos. 94, 98, 99) surround the serine within the catalytic triad and form a GSSXGG motif.
Phylogenetic reconstruction of protein sequences found with our HMM-based search within the Uniprot database using the Maximum Likelihood Method. Search for similar amino acid sequences was carried out using a HMM model constructed out of five different sequences with high similarity to EstN1 and resulted in 49 different thaumarchaeal sequences. Number of Bootstrap Replications: 100. Uniprot accession numbers are indicated in brackets. The tree can be grouped into at least four different branches. Orange: Nitrososphaera lineage; purple: Nitrosotalea lineage; light blue: Nitrosopumilus lineage; petrol: marine uncultured lineage.
Synthesis of biotin in Archaea
Many metabolic processes are strictly dependent on biotin. Within 14 out of 21 different phyla, at least a few representatives contain a biotin synthase bioB gene and therefore should be able to synthesise the cofactor (Fig. 2; Table 1). Considering the extreme environmental conditions in many archaeal habitats, biotin uptake into the cells of extremophiles is often impossible. Biotin is generally sensitive to both acids and bases, but relatively stable against light, heat or oxidizing and reducing agents (source: https://toxnet.nlm.nih.gov/). Mostly mesophilic and anaerobic Methanosarcinales, however, possess biotin-dependent enzymes like pyruvate carboxylases but do not have biotin synthesis genes. They have a BioY biotin uptake transporter instead.
Three different classes of enzymes require biotin as cofactor to translocate carbon dioxide, namely carboxylases, decarboxylases and transcarboxylases. Carboxylases catalyse CO2-fixation and enzymes like pyruvate carboxylase (gluconeogenesis), acetyl-CoA carboxylase (fatty acid synthesis), propionyl-CoA carboxylase (amino acid and fatty acid oxidation), beta-methylcrotonyl-CoA carboxylase (amino acid catabolism), geranyl-CoA carboxylase (isoprenoid catabolism) and urea amidolyase (nitrogen metabolism) belong to this group2. Acetyl-CoA carboxylase and propionyl-CoA carboxylase are the CO2-fixing enzymes in the 3-hydroxypropionate bicycle and the 3-hydroxypropionate-4-hydroxybutyrate cycle and are therefore mandatory for Archaea with these autotrophic carbon fixation pathways52. Depending on their metabolism, most members of the Archaea are able to perform at least one pathway that requires biotin. Exceptions are for instance the obligate symbiotic Nanoarchaeum equitans (Nanoarchaeota) or Thermoplasmatales that do not seem to have biotin dependent carboxylases. One possible BioB that occurs in the KEGG database from a Thermoplasmatales archaeon might as well be a radical SAM protein according to a BlastX search. How biotin synthesis e. g. in Sulfolobales and Thermococcales takes place remains to be elucidated as no enzymes with similarity to the other biotin synthesis genes could be found (Table 1). While Methanococcales possess the genetic equipment for a Bacillus-like bioW pathway, this study shows that Thaumarchaeota like N. gargensis could be able to synthesise biotin following the E. coli-like bioH pathway.
Still, many open questions remain about be origin of the pimeloyl-CoA methyl ester required for biotin synthesis as until now, no fatty acid synthesis pathway for Archaea is known although several Archaea have been reported to be capable of performing it53. A possible fatty acid biosynthesis pathway that is currently being discussed for Archaea could function like the reversed β-oxidation of fatty acids53. It could similarly work with CoA instead of ACP. The enzymatic equipment proposed for this pathway is present in N. gargensis as it possesses an acetyl-CoA C-acetyltransferase from the mevalonate pathway (NCBI-acc. no. Ngar_c00180) that catalyses the transfer of C2 fragments on acetyl-CoA. The proposed fatty acid synthesis could continue with a 3-hydroxyacyl-CoA dehydrogenase (Ngar_c14000), which is also annotated as enoyl-CoA hydratase, producing the intermediate acyl-2-enoyl-CoA. Finally, an acyl-CoA dehydrogenase (Ngar_c13350) and a possible ketoacyl-CoA thiolase (Ngar_c17490) could complete this possible biosynthesis pathway. N. gargensis has two putative adjacent acetyl-CoA/propionyl-CoA carboxylases (Ngar_c01870 and Ngar_c01880) that could convert acetyl-CoA into malonyl-CoA for the initiation of biotin synthesis. According to KEGG, N. gargensis might even encode a solitary 3-ketoacyl-ACP-reductase FabG (Ngar_c11430). Experimental verification of this hypothesis is needed.
Conclusions
Our study shows that bioA, bioB and bioD from N. gargensis are functional and able to complement E. coli deletion mutants. This is a first function-based proof of biotin synthesis in Thaumarchaeota and in the Archaea in general. Furthermore, the investigated and characterised carboxylesterase EstN1 has a full BioH activity and striking structural similarities to its bacterial counterpart although no significant similarity can be found on sequence-level. BioH is the key-enzyme that takes precursor molecules from the fatty acid cycle in order to feed them into the biotin-forming enzyme cascade. The significance of this study lies in the fact that essential global carbon cycles Archaea are involved in and metabolic pathways they possess strictly depend on biotin as cofactor.
References
Satiaputra, J., Shearwin, K. E., Booker, G. W. & Polyak, S. W. Mechanisms of biotin-regulated gene expression in microbes. Synthetic and systems biotechnology 1, 17–24, https://doi.org/10.1016/j.synbio.2016.01.005 (2016).
Waldrop, G. L., Holden, H. M. & St Maurice, M. The enzymes of biotin dependent CO2 metabolism: what structures reveal about their reaction mechanisms. Protein science: a publication of the Protein Society 21, 1597–1619, https://doi.org/10.1002/pro.2156 (2012).
Lin, S. & Cronan, J. E. The BioC O-methyltransferase catalyzes methyl esterification of malonyl-acyl carrier protein, an essential step in biotin synthesis. The Journal of biological chemistry 287, 37010–37020, https://doi.org/10.1074/jbc.M112.410290 (2012).
Agarwal, V., Lin, S., Lukk, T., Nair, S. K. & Cronan, J. E. Structure of the enzyme-acyl carrier protein (ACP) substrate gatekeeper complex required for biotin synthesis. Proceedings of the National Academy of Sciences of the United States of America 109, 17406–17411, https://doi.org/10.1073/pnas.1207028109 (2012).
Cronan, J. E. Biotin and Lipoic Acid: Synthesis, Attachment, and Regulation. EcoSal Plus 6, https://doi.org/10.1128/ecosalplus.ESP-0001-2012 (2014).
Shi, J., Cao, X., Chen, Y., Cronan, J. E. & Guo, Z. An Atypical alpha/beta-Hydrolase Fold Revealed in the Crystal Structure of Pimeloyl-Acyl Carrier Protein Methyl Esterase BioG from Haemophilus influenzae. Biochemistry 55, 6705–6717, https://doi.org/10.1021/acs.biochem.6b00818 (2016).
Bi, H., Zhu, L., Jia, J. & Cronan, J. E. A Biotin Biosynthesis Gene Restricted to Helicobacter. Scientific reports 6, 21162, https://doi.org/10.1038/srep21162 (2016).
Feng, Y. et al. The Atypical Occurrence of Two Biotin Protein Ligases in Francisella novicida Is Due to Distinct Roles in Virulence and Biotin Metabolism. mBio 6, e00591, https://doi.org/10.1128/mBio.00591-15 (2015).
Rodionov, D. A., Mironov, A. A. & Gelfand, M. S. Conservation of the biotin regulon and the BirA regulatory signal in Eubacteria and Archaea. Genome research 12, 1507–1516, https://doi.org/10.1101/gr.314502 (2002).
Shapiro, M. M., Chakravartty, V. & Cronan, J. E. Remarkable diversity in the enzymes catalyzing the last step in synthesis of the pimelate moiety of biotin. PloS one 7, e49440, https://doi.org/10.1371/journal.pone.0049440 (2012).
Sullivan, J. T., Brown, S. D., Yocum, R. R. & Ronson, C. W. The bio operon on the acquired symbiosis island of Mesorhizobium sp. strain R7A includes a novel gene involved in pimeloyl-CoA synthesis. Microbiology 147, 1315–1322, https://doi.org/10.1099/00221287-147-5-1315 (2001).
Cao, X., Zhu, L., Hu, Z. & Cronan, J. E. Expression and Activity of the BioH Esterase of Biotin Synthesis is Independent of Genome Context. Scientific reports 7, 2141, https://doi.org/10.1038/s41598-017-01490-0 (2017).
Manandhar, M. & Cronan, J. E. Pimelic acid, the first precursor of the Bacillus subtilis biotin synthesis pathway, exists as the free acid and is assembled by fatty acid synthesis. Molecular microbiology 104, 595–607, https://doi.org/10.1111/mmi.13648 (2017).
Spang, A. et al. Distinct gene set in two different lineages of ammonia-oxidizing archaea supports the phylum Thaumarchaeota. Trends in microbiology 18, 331–340, https://doi.org/10.1016/j.tim.2010.06.003 (2010).
Stieglmeier, M., Alves, R. J. E. & Schleper, C. In The Prokaryotes – Other Major Lineages of Bacteria and the Archaea Vol. Fourth Edition (eds Rosenberg, E. et al.) Ch. 26, 347–358 (Springer, 2014).
Mussmann, M. et al. Thaumarchaeotes abundant in refinery nitrifying sludges express amoA but are not obligate autotrophic ammonia oxidizers. Proceedings of the National Academy of Sciences of the United States of America 108, 16771–16776, https://doi.org/10.1073/pnas.1106427108 (2011).
Konneke, M. et al. Ammonia-oxidizing archaea use the most energy-efficient aerobic pathway for CO2 fixation. Proceedings of the National Academy of Sciences of the United States of America 111, 8239–8244, https://doi.org/10.1073/pnas.1402028111 (2014).
Berg, C., Vandieken, V., Thamdrup, B. & Jurgens, K. Significance of archaeal nitrification in hypoxic waters of the Baltic Sea. The ISME journal 9, 1319–1332, https://doi.org/10.1038/ismej.2014.218 (2015).
Zhang, L. M., Hu, H. W., Shen, J. P. & He, J. Z. Ammonia-oxidizing archaea have more important role than ammonia-oxidizing bacteria in ammonia oxidation of strongly acidic soils. The ISME journal 6, 1032–1045, https://doi.org/10.1038/ismej.2011.168 (2012).
Markowitz, V. M. et al. IMG: the integrated microbial genomes database and comparative analysis system. Nucleic acids research 40, D115–D122 (2012).
Spang, A. et al. The genome of the ammonia-oxidizing Candidatus Nitrososphaera gargensis: insights into metabolic versatility and environmental adaptations. Environ Microbiol 14, 3122–3145, https://doi.org/10.1111/j.1462-2920.2012.02893.x (2012).
Lebedeva, E. V. et al. Moderately thermophilic nitrifying bacteria from a hot spring of the Baikal rift zone. FEMS microbiology ecology 54, 297–306, https://doi.org/10.1016/j.femsec.2005.04.010 (2005).
Hatzenpichler, R. et al. A moderately thermophilic ammonia-oxidizing crenarchaeote from a hot spring. Proceedings of the National Academy of Sciences of the United States of America 105, 2134–2139, https://doi.org/10.1073/pnas.0708857105 (2008).
Krümmel, A. & Harms, H. Effect of organic matter on growth and cell yield of ammonia-oxidizing bacteria. Archives of Microbiology 133, 50–54, https://doi.org/10.1007/BF00943769 (1982).
Widdel, F. In The Prokaryotes - A Handbook on the Biology of Bacteria: Ecophysiology, Isolation, Identification, Applications. (eds Balows, A. et al.) 3352–3378 (Springer-Verlag New York, 1992).
Sambrook, J. & Russel, D. W. (Cold Spring Harbor Laboratory Press, New York, USA, 2001).
Ronson, C. W. & Primrose, S. B. Carbohydrate Metabolism in Rhizobium trifolii: Identification and Symbiotic Properties of Mutants. Microbiology 112, 77–88, https://doi.org/10.1099/00221287-112-1-77 (1979).
Martinez-Martinez, M. et al. Determinants and Prediction of Esterase Substrate Promiscuity Patterns. ACS chemical biology 13, 225–234, https://doi.org/10.1021/acschembio.7b00996 (2018).
Berge, R. K. & Farstad, M. Long-chain fatty acyl-CoA hydrolase from rat liver mitochondria. Methods in enzymology 71(Pt C), 234–242 (1981).
Database Resources of the National Center for Biotechnology Information. Nucleic acids research 45, D12–D17, doi:10.1093/nar/gkw1071 (2017).
Chen, I. A. et al. IMG/M: integrated genome and metagenome comparative data analysis system. Nucleic acids research 45, D507–D516, https://doi.org/10.1093/nar/gkw929 (2017).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic acids research 45, D200–D203, https://doi.org/10.1093/nar/gkw1129 (2017).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. Journal of molecular biology 302, 205–217, https://doi.org/10.1006/jmbi.2000.4042 (2000).
Bateman, A. et al. UniProt: the universal protein knowledgebase. Nucleic acids research 45, D158–D169 (2017).
Wheeler, T. J., Clements, J. & Finn, R. D. Skylign: a tool for creating informative, interactive logos representing sequence alignments and profile hidden Markov models. Bmc Bioinformatics 15 (2014).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Molecular biology and evolution 30, 2725–2729, https://doi.org/10.1093/molbev/mst197 (2013).
Song, Y. et al. High-resolution comparative modeling with RosettaCM. Structure 21, 1735–1742, https://doi.org/10.1016/j.str.2013.08.005 (2013).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. Journal of computational chemistry 25, 1605–1612, https://doi.org/10.1002/jcc.20084 (2004).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic acids research 38, W545–549, https://doi.org/10.1093/nar/gkq366 (2010).
Kim, R. et al. Overexpression of archaeal proteins in Escherichia coli. Biotechnology letters 20, 207–210 (1998).
Prinz, W. A., Aslund, F., Holmgren, A. & Beckwith, J. The role of the thioredoxin and glutaredoxin pathways in reducing protein disulfide bonds in the Escherichia coli cytoplasm. The Journal of biological chemistry 272, 15661–15667 (1997).
Manandhar, M. & Cronan, J. E. A Canonical Biotin Synthesis Enzyme, 8-Amino-7-Oxononanoate Synthase (BioF), Utilizes Different Acyl Chain Donors in Bacillus subtilis and Escherichia coli. Applied and environmental microbiology 84, https://doi.org/10.1128/AEM.02084-17 (2018).
Lombard, J., Lopez-Garcia, P. & Moreira, D. An ACP-independent fatty acid synthesis pathway in archaea: implications for the origin of phospholipids. Molecular biology and evolution 29, 3261–3265, https://doi.org/10.1093/molbev/mss160 (2012).
Shi, Y. et al. Molecular cloning of a novel bioH gene from an environmental metagenome encoding a carboxylesterase with exceptional tolerance to organic solvents. BMC biotechnology 13, 13, https://doi.org/10.1186/1472-6750-13-13 (2013).
Kaljunen, H., Chow, J., Streit, W. R. & Mueller-Dieckmann, J. Cloning, expression, purification and preliminary X-ray analysis of EstN2, a novel archaeal alpha/beta-hydrolase from Candidatus Nitrososphaera gargensis. Acta crystallographica. Section F, Structural biology communications 70, 1394–1397, https://doi.org/10.1107/S2053230X14018482 (2014).
Lin, S., Hanson, R. E. & Cronan, J. E. Biotin synthesis begins by hijacking the fatty acid synthetic pathway. Nature chemical biology 6, 682–688, https://doi.org/10.1038/nchembio.420 (2010).
Qiu, X. et al. Crystal structure of beta-ketoacyl-acyl carrier protein synthase III. A key condensing enzyme in bacterial fatty acid biosynthesis. The Journal of biological chemistry 274, 36465–36471 (1999).
McKinney, D. C. et al. Antibacterial FabH Inhibitors with Mode of Action Validated in Haemophilus influenzae by in vitro Resistance Mutation Mapping. ACS infectious diseases 2, 456–464, https://doi.org/10.1021/acsinfecdis.6b00053 (2016).
Sanishvili, R. et al. Integrating structure, bioinformatics, and enzymology to discover function: BioH, a new carboxylesterase from Escherichia coli. The Journal of biological chemistry 278, 26039–26045, https://doi.org/10.1074/jbc.M303867200 (2003).
Flores, H., Lin, S., Contreras-Ferrat, G., Cronan, J. E. & Morett, E. Evolution of a new function in an esterase: simple amino acid substitutions enable the activity present in the larger paralog, BioH. Protein engineering, design & selection: PEDS 25, 387–395, https://doi.org/10.1093/protein/gzs035 (2012).
Espinosa-Cantu, A., Ascencio, D., Barona-Gomez, F. & DeLuna, A. Gene duplication and the evolution of moonlighting proteins. Frontiers in genetics 6, 227, https://doi.org/10.3389/fgene.2015.00227 (2015).
Berg, I. A. et al. Autotrophic carbon fixation in archaea. Nature reviews. Microbiology 8, 447–460, https://doi.org/10.1038/nrmicro2365 (2010).
Dibrova, D. V., Galperin, M. Y. & Mulkidjanian, A. Y. Phylogenomic reconstruction of archaeal fatty acid metabolism. Environ Microbiol 16, 907–918, https://doi.org/10.1111/1462-2920.12359 (2014).
Acknowledgements
We would like to thank John T. Sullivan and Clive Ronson (University of Otago, Dunedin, New Zealand) for providing the ΔbioZ insertion mutant of Mesorhizobium loti. The authors gratefully acknowledge Cristina Coscolín (CSIC, Madrid, Spain) for her contribution to EstN1 protein expression and purification used for the pNPP and acyl-CoA ester assays. Funding by the German Ministry of Education and Research (BMBF) within the frameworks of “Biokatalyse 2021” project P44 (“Biokatalyse in hochviskosen Medien”) and iVDT-V2 is also gratefully acknowledged. This project received funding from the European Union’s Horizon 2020 research and innovation program [Blue Growth: Unlocking the potential of Seas and Oceans] under grant agreement no. [634486] (project acronym INMARE). This research was also supported by the grants PCIN-2014-107 (within ERA NET IB2 grant nr. ERA-IB-14-030 MetaCat), PCIN-2017-078 (within the ERA-MarineBiotech grant ProBone), BIO2014-54494-R, and BIO2017-85522-R from the Spanish Ministry of Economy, Industry and Competitiveness. The authors gratefully acknowledge financial support provided by the European Regional Development Fund (ERDF).
Author information
Authors and Affiliations
Contributions
J.C. and W.R.S. conceived the idea. J.C. wrote the manuscript, prepared the figures and carried out the experiments concerning cultivation, cloning, purification, biochemical characterization, database searches and complementation assays. D.D. carried out HMM model building and HMM database search. M.F. had the idea for pNPP and acyl-CoA thioesterase activity assays and carried them out. All authors discussed the results.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chow, J., Danso, D., Ferrer, M. et al. The Thaumarchaeon N. gargensis carries functional bioABD genes and has a promiscuous E. coli ΔbioH-complementing esterase EstN1. Sci Rep 8, 13823 (2018). https://doi.org/10.1038/s41598-018-32059-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-32059-0
Keywords
This article is cited by
-
Advances and prospects in microbial production of biotin
Microbial Cell Factories (2024)
-
Phylogenomics suggests oxygen availability as a driving force in Thaumarchaeota evolution
The ISME Journal (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.