Abstract
The fibrotic reaction is a characteristic feature of human pancreatic ductal adenocarcinoma (PDAC) tumors. It is associated with activation and proliferation of pancreatic stellate cells (PSCs), which are key regulators of fibrosis in vivo. While there is increasing interest in the regulation of PD-L1 expression in cancer and immune cells, the expression and regulation of PD-L1 in other stromal cells, such as PSCs, has not been fully evaluated. Here we show that PSCs in vitro express higher PD-L1 mRNA and protein levels compared to the levels present in PDAC cells. We show that inhibitors targeting bromodomain and extra-terminal (BET) proteins and BRD4 knockdown decrease interferon-γ (IFN-γ)-induced PD-L1 expression in PSCs. We also show that c-MYC, one of the well-established targets of BET inhibitors, does not mediate IFN-γ-regulated PD-L1 expression in PSCs. Instead we show that interferon regulatory factor 1 (IRF1) mediates IFN-γ-induced PD-L1 expression in PSCs. Finally, while we show that BET inhibitors do not regulate IFN-γ-induced IRF1 expression in PSCs, BET inhibitors decrease binding of IRF1 and BRD4 to the PD-L1 promoter. Together, these results demonstrate the interplay between IRF1 and BRD4 in the regulation of PD-L1 in PSCs.
Similar content being viewed by others
Introduction
PD-L1 expression is seen in a number of human cancers, with increased expression of PD-L1 by cancer cells in some tumor types can be associated with responses to antibodies targeting PD-1/PD-L11. For example, increased expression of PD-L1 in non-small cell lung cancers has predictive value for response to antibodies targeting PD-1/PD-L12,3. However, the predictive value for response to immune checkpoint inhibitors in other tumor types, e.g., bladder cancer4, correlates with PD-L1 expression by infiltrating immune cells and not by cancer cells1. While there is increasing interest in identifying mechanisms regulating PD-L1 expression in cancer and immune cells, the expression and regulation of PD-L1 in other stromal cells, such as fibroblasts, has not been fully evaluated. Importantly, the stroma in some tumors can account for a significant portion of the tumor mass. For example, the stroma in human pancreatic ductal adenocarcinoma (PDAC) tumors, which can account for as much as 80–90% of the tumor mass, is associated with proliferation of pancreatic stellate cells (PSCs)5,6. The PSCs are the predominant fibroblasts present in human PDAC tumors and the key regulators of fibrosis in vivo5.
Recently, we showed that inhibitors targeting bromodomain (BRD) and extra-terminal (BET) proteins induce primary PSCs isolated from human PDAC tumors to become quiescent7. The BET proteins, which include BRD2, BRD3, BRD4, and the testis-specific BRDT, regulate transcription of genes involved in several human diseases8,9. These proteins bind to acetylation motifs present in histones and enable recruitment of transcription factors and other chromatin regulators during RNA transcription8,9. Importantly, a number of selective and potent small-molecule BET inhibitors are currently being evaluated in clinical trials for solid tumors8,9. In addition, inhibitors that promote degradation of BET proteins through proteolysis-targeting chimera (PROTAC) are also being developed10,11,12,13. While BET inhibitors were recently shown to repress PD-L1 expression in lymphoma and ovarian cancer cells14,15, the contribution of BET proteins to the regulation of PD-L1 expression in PSCs has yet to be evaluated.
In this report, we show that PSCs express higher PD-L1 mRNA and protein levels compared to the levels present in PDAC cells. We show that BET inhibitors, BET PROTAC and BRD4 knockdown decrease interferon-γ (IFN-γ)-induced PD-L1 expression in PSCs. We also show that c-MYC, one of the well-established targets of BET inhibitors16,17,18, does not mediate IFN-γ-mediated PD-L1 expression in PSCs. Instead we show that interferon regulatory factor 1 (IRF1) mediates IFN-γ-induced PD-L1 expression in PSCs. Finally, while we show that BET inhibitors do not regulate IFN-γ-induced IRF1 expression in PSCs, BET inhibitors decrease binding of IRF1 and BRD4 to the PD-L1 promoter. Together, these results demonstrate the interplay between IRF1 and BRD4 in the regulation of PD-L1 in PSCs.
Results
PSCs express PD-L1
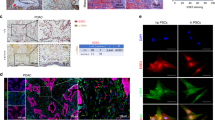
Since activated PSCs are the predominant fibroblasts present in the pancreatic stromal reaction5, and the immune cells are very likely to encounter these cells in vivo, we evaluated expression of PD-L1 in an immortalized PSC cell line and in primary PSCs isolated from human PDAC tumors using the outgrowth assay19. Human PSCs express PD-L1 mRNA, which in most cases exceeded the PD-L1 mRNA levels present in human PDAC cell lines (Fig. 1A). In agreement with the mRNA findings, both the immortalized PSC cell line and the primary PSCs demonstrated higher PD-L1 protein levels relative to the levels present in the CD18 PDAC cell line (Fig. 1A). However, treatment with IFN-γ resulted in robust induction of PD-L1 in CD18 cells to levels comparable to basal PD-L1 levels present in PSCs (Fig. 1B).
Pancreatic stellate cells (PSCs) express PD-L1. (A) Human PD-L1 mRNA expression was analyzed in a panel of human PDAC cell lines, in an immortalized human pancreatic stellate cell line (Im PSCs), and in primary pancreatic stellate cells (Pri PSCs) isolated from human PDAC tumors. The relative expression was normalized to PD-L1 mRNA levels present in the Panc1 cells. PD-L1 protein expression was determined in CD18 PDAC cells, in Im PSCs and in Pri PSC #2 using HSP90 as loading control. PD-L1 and HSP90 bands from three independent experiments were quantified by densitometry and data expressed as relative protein ratios. (B) CD18 PDAC cells, Im PSCs, and Pri PSC #2 were treated with IFN-γ (0.2 μg/mL) for 24 hours. The effect on PD-L1 mRNA expression was determined by qRT-PCR and the effect on PD-L1 protein expression was determined by Western blotting. PD-L1 and HSP90 bands from three independent experiments were quantified by densitometry and data expressed as relative protein ratios. (C) Mouse PD-L1 mRNA was analyzed in a panel of mouse PDAC cell lines established from tumors arising in the KPC (Kras/p53) mouse model, in mouse Pan02 cells and in two immortalized mouse pancreatic stellate cell lines (Im mPSCs). The relative expression was normalized to mouse PD-L1 mRNA levels present in the KPC1199 cells. KPC1245 PDAC cells and Im mPSC #3 were treated with mouse IFN-γ (0.05 μg/mL) for 4 hours. The effect on mouse PD-L1 mRNA expression was determined by qRT-PCR. The gene expression results are representative of three independent experiments. Bar graphs represent means +/−S.D. *p < 0.05; **p < 0.01; ***p < 0.001 relative to control samples.
We also evaluated the expression of PD-L1 mRNA expression in a panel of mouse PDAC cell lines and in 2 immortalized mouse PSC cell lines. Similar to our findings with human PDAC and PSC cells (Fig. 1A), mouse PSC cells show relatively increased PD-L1 mRNA levels compared to levels present in mouse Pan02 PDAC cell line and in mouse PDAC cell lines generated from tumors arising in the KPC transgenic mouse model of pancreatic cancer (Fig. 1C). Similarly, treatment of mouse PDAC cells with mouse IFN-γ resulted in induction of mouse PD-L1 mRNA to levels comparable to PD-L1 mRNA levels present in mouse PSC cell lines (Fig. 1C).
BET inhibitors decrease PD-L1 expression in PDAC cells and PSCs
Recently, we showed that BET inhibitors can modulate stellate cell function by inducing primary PSCs to become quiescent, with decreased α-SMA and collagen I expression7. Thus, we evaluated the effect of BET inhibitors JQ1 and I-BET151 on PD-L1 expression in PSCs and in PDAC cells. JQ1 and I-BET151 decreased IFN-γ-induced PD-L1 expression at the mRNA and protein levels in CD18 and AsPC1 PDAC cells (Fig. 2A and D, and Supplemental Fig. 1A). BET inhibitors also decreased IFN-γ-induced PD-L1 mRNA and protein expression in primary PSCs isolated from human PDAC tumors (Fig. 2B and E, and Supplemental Fig. 1B) and in the stellate cell line (Fig. 2C and F). Significantly, IFN-γ rapidly induced PD-L1 expression in PSCs and in PDAC cells, which was inhibited by pre-treatment with JQ1 (Supplemental Fig. 1C).
BET inhibitors decrease IFN-γ-induced PD-L1 expression in PDAC cells and PSCs. CD18 PDAC cells, primary PSC #1, and the pancreatic stellate cell line were pre-treated with the BET inhibitors JQ1 (1 μM) or I-BET151 (I-BET; 1 μM) for 30 minutes and treated with IFN-γ (0.2 μg/mL) for 24 hours. (A–C) The effect on PD-L1 mRNA expression was determined by qRT-PCR. (D–F) The effect on PD-L1 protein expression was determined by Western blotting and by FACS analysis. PD-L1 and HSP90 bands from three independent experiments were quantified by densitometry and data expressed as relative protein ratios. The gene expression results are representative of five independent experiments. Bar graphs represent means +/−S.D. *p < 0.05; **p < 0.01; ***p < 0.001 relative to control samples.
BET PROTAC decreases PD-L1 expression in PSCs
We also evaluated the effect of BET PROTAC, which can promote degradation of BET proteins10,11,12,13, on PD-L1 expression in PSCs. Treatment with the BET PROTAC ARV-825 resulted in loss of BRD3 and BRD4 proteins without decreasing BRD2 protein levels. Significantly, ARV-825 decreased IFN-γ-induced PD-L1 mRNA and protein expression in primary PSCs and in the stellate cell line (Fig. 3A–D).
BET PROTAC ARV-825 decreases IFN-γ-induced PD-L1 expression in PSCs. Primary PSC #1 and the pancreatic stellate cell line were pre-treated with the BET PROTAC ARV-825 (0.25 μM) for 30 minutes and treated with IFN-γ (0.2 μg/mL) for 24 hours. (A,B) The effect on BRD2, BRD3, BRD4 and PD-L1 protein expression was determined by Western blotting. PD-L1 and HSP90 protein bands from three independent experiments were quantified by densitometry and data expressed as relative protein ratios. (C,D) The effect on PD-L1 mRNA expression was determined by qRT-PCR. The gene expression results are representative of three independent experiments. Bar graphs represent means +/−S.D. ns, not significant; *p < 0.05; **p < 0.01 relative to control samples.
BRD4 regulates PD-L1 expression in PSCs
We next evaluated the relative contribution of the BET proteins BRD2, BRD3 and BRD4 to the regulation of IFN-γ-induced PD-L1 expression in primary PSCs and in the stellate cell line. Knocking down BRD2 or BRD3 had minimal effects on IFN-γ-induced PD-L1 mRNA in both primary PSCs (Fig. 4A,C and E, and Supplemental Fig. 2) and the stellate cell line (Fig. 4B,D and F). In contrast, BRD4 knockdown decreased IFN-γ-induced PD-L1 mRNA expression in primary PSCs and in the stellate cell line (Fig. 4 and Supplemental Fig. 2).
BRD4 knockdown decreases IFN-γ-induced PD-L1 expression in PSCs. (A,B) Primary PSC #1 and the pancreatic stellate cell line were transfected with control siRNA or with siRNAs against BRD2, BRD3 or BRD4 for 48 hours. The cells were then treated with IFN-γ (0.2 μg/mL) for 24 hours. (A,B) The effect on the individual BRD proteins was determined by Western blotting. Bands from three independent experiments were quantified by densitometry and data expressed as relative ratios. (C,D) The effect on the individual BRD mRNAs was determined by qRT-PCR. (E,F) The effect on PD-L1 mRNAs was determined by qRT-PCR. The gene expression results are representative of four independent experiments. Bar graphs represent means +/− S.D. ns, not significant; *p < 0.05; **p < 0.01; ***p < 0.001 relative to control samples.
c-MYC knockdown does not suppress IFN-γ-induced PD-L1 expression in PSCs and PDAC cells
Recently, c-MYC was shown to regulate PD-L1 expression in mouse and human tumor cells20,21, with suppression of c-MYC decreasing PD-L1 mRNA and protein levels. Since we have published that BET inhibitors can repress c-MYC levels both in the stellate cell line and in PDAC cells7,22,23, we evaluated the role of c-MYC in IFN-γ-induced PD-L1 mRNA expression. Suppressing c-MYC in primary PSCs and in the stellate cell line did not decrease IFN-γ-induced PD-L1 expression (Fig. 5A and B). Similarly, c-MYC knockdown in CD18 and AsPC1 PDAC cells did not decrease IFN-γ-induced PD-L1 expression (Supplemental Fig. 3A). Significantly, c-MYC knockdown in fact increased PD-L1 expression in some of these cells (Fig. 5A and Supplemental Fig. 3A). These results suggest that c-MYC does not mediate the suppressive effects of BET inhibitors on IFN-γ-induced PD-L1 in PSCs and PDAC cells.
c-MYC knockdown does not decrease IFN-γ-induced PD-L1 expression. (A,B) Primary PSCs and the pancreatic stellate cell line were transfected with control siRNA or with siRNA against c-MYC for 48 hours. The cells were then treated with IFN-γ (0.2 μg/mL) for 24 hours. The effect on c-MYC was determined by qRT-PCR and by Western blotting, and the effect on PD-L1 mRNA was determined by qRT-PCR. c-MYC and GAPDH bands from three independent experiments were quantified by densitometry and data expressed as relative ratios. The gene expression results are representative of three independent experiments. Bar graphs represent means +/− S.D. ns, not significant; *p < 0.05; **p < 0.01; ***p < 0.001 relative to control samples.
IRF1 knockdown decreases IFN-γ-induced PD-L1 expression
Previously, it was shown that interferon regulatory factor 1 (IRF1) regulated IFN-γ-induced PD-L1 expression in lung, liver and colon cancer cells24. Thus, we evaluated the extent to which BET inhibitors regulated IRF1 in PSCs and PDAC cells. Initially, we evaluated the role of IRF1 on IFN-γ-induced PD-L1 expression in PSCs. Knockdown of IRF1 decreased IFN-γ-induced PD-L1 expression in primary PSCs (Fig. 6A) and in the stellate cell line (Fig. 6B).
IRF1 knockdown decreases IFN-γ-induced PD-L1 expression. (A,B) Primary PSCs and the pancreatic stellate cell line were transfected with control siRNA or with siRNA against IRF1 for 48 hours. The cells were then treated with IFN-γ (0.2 μg/mL) for 24 hours. The effect on IRF1 was determined by qRT-PCR and by Western blotting, and the effect on PD-L1 mRNA was determined by qRT-PCR. IRF1 and GAPDH bands from three independent experiments were quantified by densitometry and data expressed as relative ratios. The gene expression results are representative of three independent experiments. Bar graphs represent means +/−S.D. ns, not significant; *p < 0.05; **p < 0.01 relative to control samples.
BET inhibitors do not suppress IFN-γ-induced IRF1 protein expression
We next evaluated the effects of JQ1 and I-BET151 on IFN-γ-induced IRF1 expression. While IFN-γ increased IRF1 expression, pre-treatment with JQ1 or I-BET151 had minimal effects on IFN-γ-induced IRF1 expression in primary PSCs and in the stellate cell line (Fig. 7A). BET inhibitors also had minimal effects on IFN-γ-induced IRF1 expression in the PDAC cell lines CD18 and AsPC1 (Supplemental Fig. 3B). Moreover, pre-treatment with the BET PROTAC ARV-825 (Supplemental Fig. 3C), or knockdown of BRD4 (Fig. 7B), also had minimal effects on IFN-γ-induced IRF1 expression in primary PSCs and the stellate cell line.
BET inhibitors do not suppress IFN-γ-induced IRF1 protein expression. (A) Primary PSCs and the pancreatic stellate cell line were pre-treated with the BET inhibitors JQ1 (1 μM) or I-BET151 (I-BET; 1 μM) for 30 minutes and then treated with IFN-γ (0.2 μg/mL) for 4 hours. The effect on IRF1 mRNA and protein expression was determined by qRT-PCR and by Western blotting. (B) Primary PSCs and the pancreatic stellate cell line were transfected with control siRNA or with siRNA against BRD4 for 48 hours. The cells were then treated with IFN-γ (0.2 μg/mL) for 4 hours. The effect on BRD4 protein was determined by qRT-PCR and by Western blotting, and the effect on IRF1 mRNA was determined by qRT-PCR. Bands from three independent experiments were quantified by densitometry and data expressed as relative ratios. The gene expression results are representative of three independent experiments. Bar graphs represent means +/− S.D. ns, not significant; *p < 0.05; **p < 0.01; ***p < 0.001 relative to control samples.
BET inhibitors decrease binding of IRF1 to PD-L1 promoter
Since BRD4 can bind to the PD-L1 promoter in cancer cells14,15, we initially evaluated the effect of BET inhibitors on the recruitment of BRD4 to the PD-L1 promoter in the stellate cell line. Treatment with IFN-γ increased binding of BRD4 to the PD-L1 promoter in the stellate cell line, which was inhibited in the presence of JQ1 (Fig. 8A). We next evaluated the effect of BET inhibitors on binding of IRF1 to the PD-L1 promoter in PSCs. As previously shown15, treatment with IFN-γ increased binding of IRF-1 to the PD-L1 promoter (Fig. 8B). Significantly, while JQ1 did not decrease IFN-γ-induced IRF1 protein expression (Fig. 7A), JQ1 decreased IRF1 binding to the PD-L1 promoter (Fig. 8B).
BET inhibitors decrease binding of IRF1 to PD-L1 promoter. (A, B) The pancreatic stellate cell line was pre-treated with the BET inhibitor JQ1 (1 μM) for 30 minutes and then treated with IFN-γ (0.2 μg/mL) for 4 hours. The cells were subjected to ChIP analysis using anti-BRD4 antibody (A) or anti-IRF1 antibody (B). An isotype-matched IgG was used as a negative control. The association with the PD-L1 gene promoter was quantified by qPCR. The results are representative of three independent experiments. Bar graphs represent means +/−S.D. **p < 0.01; ***p < 0.001 relative to control samples.
Discussion
Since PD-L1 levels can correlate with response to immune checkpoint therapy in certain tumors2,3, there is increasing interest in understanding PD-L1 regulation1. However, while PD-L1 regulation has been extensively studied in cancer cells1, the regulation of PD-L1 in stromal cells is less well understood. Also, since the stroma in PDAC tumors predominates5,6, it is very likely that cytolytic T cells in PDAC tumors will encounter PSCs. In this report we show that PSCs, which contribute to the pronounced stroma that is seen in human PDAC tumors5,6, express PD-L1 and that BET inhibitors can repress PD-L1 in PSCs. Our data are in agreement with recent studies demonstrating that BET inhibitors can decrease PD-L1 expression in lymphoma and ovarian cancer cells14,15. In this report we also show that BET PROTAC and knockdown of BRD4, but not knockdown of BRD2 or BRD3, decrease PD-L1 expression in PSCs.
We have previously shown that BET inhibitors can target c-MYC in a human stellate cell line and in PDAC cells7,22,23. However, we have found that targeting c-MYC does not decrease PD-L1 levels in PSCs or in PDAC cell lines, indicating that the effect of BET inhibitors in these cells is not indirect through c-MYC repression. In fact, c-MYC knockdown in PDAC cells increased IFN-γ-induced PD-L1 expression, in agreement with a recent study showing that c-MYC knockdown in hepatocellular carcinoma cells results in increased IFN-γ-induced PD-L1 expression25. While our findings contrast with a study demonstrating that targeting c-MYC by shRNA or JQ1 treatment decreases PD-L1 in melanoma and lung cancer cells20,21, our findings are in agreement with another study showing that JQ1 regulation of PD-L1 in lymphoma cells is not mediated by c-MYC15. Instead, BET inhibitors directly decrease binding of BRD4 to the PD-L1 promoter in lymphoma cells15. We have also found that BET inhibitors directly decrease binding of BRD4 to the PD-L1 promoter in PSCs.
We also demonstrate that BET inhibitors decrease binding of IRF1 to the PD-L1 promoter in PSCs. IRF1 is well known to regulate PD-L1 in a number of different cancer types24,26. IRF1 rapidly binds to the PD-L1 promoter following IFN-γ treatment and down-regulation of IRF1 decreases PD-L1 expression24. We also show that IRF1 regulates PD-L1 expression in PSCs. Significantly, we have found that BET inhibitors, BET PROTAC and BRD4 siRNA do not decrease IRF1 protein expression in PSCs. Instead, we show that BET inhibitors decrease binding of IRF1 protein to the PD-L1 promoter. Our results are in contrast to the data in lymphoma cells in which BET inhibitors did not affect binding of IRF1 to the PD-L1 promoter15. It is possible that the effect of BET inhibitors on IRF1 binding to the PD-L1 promoter may vary depending on the cell type.
While the role of fibrosis in PDAC progression is controversial, and whether the pancreatic stroma is pro- or anti-tumorigenic continues to be debated27,28,29, recent reports have shown that normalizing the stroma30, instead of eliminating the stroma28,29, can enhance drug delivery and improve survival. We have recently also shown that BET inhibitors can normalize the stroma by inducing stellate cells to become quiescent as reflected by re-accumulation of lipid droplets, decreased collagen production and decreased α-SMA expression7. As we now show that BET inhibitors decrease PD-L1 expression, and since BET inhibitors can enhance response to anti-PD-1 antibody in MYC-expressing lymphoma tumors15, it is possible that BET inhibitors may also enhance response to immune checkpoint inhibitors in pancreatic cancer. However, since BET inhibitors decrease PD-L1, and thereby decrease the ‘target’ for the PD-1/PD-L1 checkpoint inhibitors, in future studies we will evaluate the relative efficacy of combining BET inhibitors with anti-PD-1/PD-L1 antibodies or with other immune checkpoint inhibitors, such as anti-CTLA4 antibodies31, in transgenic mouse models of pancreatic cancer.
In summary, we show that PSCs, the predominant fibroblasts in human PDAC tumors5,6, express high levels of PD-L1 and that the interplay between IRF1 and BRD4 regulates PD-L1 expression in PSCs. These results, together with our earlier studies evaluating the effect of BET inhibitors on pancreatic cancer cells and PSCs7,22, provide impetus for the evaluation of BET inhibitors in patients with pancreatic cancer.
Methods
Chemicals/reagents
General tissue culture materials were obtained from VWR International. Antibodies against BRD2 (ab139690), BRD3 (ab50818), BRD4 (ab128874) and IRF1 (ab26109) antibody for chromatin immunoprecipitation (ChIP) were obtained from Abcam. Antibodies against c-MYC (5605), human PD-L1 (E1L3N clone; 13684), IRF1 (8478) and control IgG (2729) antibody for ChIP were purchased from Cell Signaling Technology. Anti-BRD4 (A301–985A) antibody for ChIP was obtained from Bethyl Laboratories. PE-conjugated human PD-L1 (MIH1 clone; 12-5983-42) antibody was from Thermo Fisher. The GAPDH (MAB374) antibody was from Millipore, and α-tubulin (sc-8035) antibody and HSP90 (sc-7940) antibody were obtained from Santa Cruz Biotechnology. Secondary anti-mouse IgG (A4416) and anti-rabbit IgG (A6667) antibodies were purchased from Sigma. The BET inhibitors JQ1 and I-BET151 were obtained from Tocris Bioscience, the BET PROTAC ARV-825 was obtained from MedChem Express, and human and mouse IFN-γ was purchased from Thermo Fisher Scientific. The siRNAs against BRD2, BRD3, BRD4, c-MYC, and IRF1 were purchased from Thermo Fisher Scientific.
Cell culture
Human PDAC cell lines AsPC1, BxPC3, CD18/HPAF-II, MiaPaca2, Panc1 and Panc02–03 were obtained from ATCC. Chemo-resistant CD18 (CD18-CR) cells were generated by treating parental CD18 cells with increasing concentration of 5-fluorouracil (5-FU) over a period of 3 months32. KPC22, KPC105, KPC163 and KPC344 – all derived from PDAC tumor developing in the KPC (Kras/p53) mouse model in B6 background – were obtained from Sam Grimaldo (University of Illinois, Chicago, US). KPC1245 and KPC1199 cells, both derived from KPC tumors developing in B6 background, were provided by David Tuveson (Cold Spring Harbor Laboratories, Cold Spring Harbor, New York, US)33,34. Mouse Pan02 cells were purchased from the NCI.
A human pancreatic stellate cell line, which was immortalized with telomerase and SV40 large T antigen35,36,37, was obtained from Dr. Rosa F. Hwang (MD Anderson Cancer Center, Houston, Texas, US). The immortalized mouse pancreatic stellate cell lines mPSC#2 and mPSC#3, which were immortalized with SV40 large T antigen38, were obtained from Dr. Raul Urrutia (Mayo, Rochester, Minnesota, US). Primary PSCs were generated from de-identified fresh human PDAC specimens using primary outgrowth cultures, which have been shown to be a reliable source of human pancreatic stellate cells19. The primary PSCs were used within 5–6 passages from isolation.
Transfection
The human stellate cell line and the primary PSCs were transfected with siRNA against BRD2, BRD3, BRD4, c-MYC, IRF1 or control siRNA (20 nM) using RNAimax (Thermo Fisher Scientific) according to the manufacturer’s instructions23,32.
qRT-PCR analysis
Quantitative gene expression was performed with gene-specific TaqMan probes, TaqMan Universal PCR Master Mix, and the 7500 Fast Real-time PCR System from Applied Biosystems. The data were then quantified with the comparative CT method for relative gene expression23,32.
ChIP
The human stellate cell line was pre-treated with JQ1 for 30 minutes, treated with IFN-γ for 4 hours, and then treated with formaldehyde to create DNA-protein cross-links. Chromatin fragments were prepared using EZ-Zyme Chromatin Prep kit (17–375, Millipore) and ChIP performed using the EZ-Magna ChIP A/G Chromatin Immunoprecipitation kit (17–10086, Millipore) and anti-BRD4 antibody (A301-985A, Bethyl Laboratories), anti-IRF1 antibody (ab26109, Abcam) or control IgG antibody (2729, Cell Signaling). Purified DNA was then analyzed by PCR using KiCqStart SYBR Green qPCR ReadyMix (KCQS02, Sigma) and primers specific for the PD-L1 locus: forward 5′-AAGCCATATGGGTCTGCTC-3′ and reverse 5′-TTATCAGAAAGGCGTCCCCC-3′15.
Immunoblotting
Immunoblotting for BRD2, BRD3, BRD4, c-MYC, IRF1, PD-L1, α-tubulin, GAPDH and HSP90 was done as described previously23,32.
Flow Cytometric Analysis
PDAC cells and PSCs in suspension were incubated with PE-conjugated anti-PD-L1 antibody and co-stained with DAPI prior to analysis with BD LSR Fortessa 6-Laser flow cytometer7,32.
Study Approval
Pancreatic tissue was obtained from patients with PDAC undergoing resection on a protocol approved by the Institutional Review Board of Northwestern University. Informed consent was obtained from patients prior to resection. The resected specimens were deidentified and subsequently processed for isolation of primary PSCs. All experiments were performed in accordance with relevant guidelines and regulations.
Statistical analysis
All statistical analyses were done using Microsoft Excel using a two-tailed t-test analysis. Error bars represent standard deviation.
Data Availability
All data supporting the findings of this study are available with the article or from corresponding authors upon reasonable request.
References
Sun, C., Mezzadra, R. & Schumacher, T. N. Regulation and Function of the PD-L1 Checkpoint. Immunity 48, 434–452 (2018).
Borghaei, H. et al. Nivolumab versus Docetaxel in Advanced Nonsquamous Non-Small-Cell Lung Cancer. N Engl J Med 373, 1627–1639 (2015).
Fehrenbacher, L. et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 387, 1837–1846 (2016).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Apte, M. V., Wilson, J. S., Lugea, A. & Pandol, S. J. A starring role for stellate cells in the pancreatic cancer microenvironment. Gastroenterology 144, 1210–1219 (2013).
Shields, M. A., Dangi-Garimella, S., Redig, A. J. & Munshi, H. G. Biochemical role of the collagen-rich tumor microenvironment in pancreatic cancer progression. Biochem J 441, 541–552 (2012).
Kumar, K. et al. BET inhibitors block pancreatic stellate cell collagen I production and attenuate fibrosis in vivo. JCI Insight 2, e88032 (2017).
Abedin, S. M., Boddy, C. S. & Munshi, H. G. BET inhibitors in the treatment of hematologic malignancies: current insights and future prospects. Onco Targets Ther 9, 5943–5953 (2016).
Sahai, V., Redig, A. J., Collier, K. A., Eckerdt, F. D. & Munshi, H. G. Targeting bet bromodomain proteins in solid tumors. Oncotarget 7, 53997–54009 (2016).
Lu, J. et al. Hijacking the E3 Ubiquitin Ligase Cereblon to Efficiently Target BRD4. Chem Biol 22, 755–763 (2015).
Raina, K. et al. PROTAC-induced BET protein degradation as a therapy for castration-resistant prostate cancer. Proc Natl Acad Sci USA 113, 7124–7129 (2016).
Saenz, D. T. et al. Novel BET protein proteolysis-targeting chimera exerts superior lethal activity than bromodomain inhibitor (BETi) against post-myeloproliferative neoplasm secondary (s) AML cells. Leukemia 31, 1951–1961 (2017).
Sun, B. et al. BET protein proteolysis targeting chimera (PROTAC) exerts potent lethal activity against mantle cell lymphoma cells. Leukemia 32, 343–352 (2017).
Zhu, H. et al. BET Bromodomain Inhibition Promotes Anti-tumor Immunity by Suppressing PD-L1 Expression. Cell Rep 16, 2829–2837 (2016).
Hogg, S. J. et al. BET-Bromodomain Inhibitors Engage the Host Immune System and Regulate Expression of the Immune Checkpoint Ligand PD-L1. Cell Rep 18, 2162–2174 (2017).
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917 (2011).
Filippakopoulos, P. et al. Selective inhibition of BET bromodomains. Nature 468, 1067–1073 (2010).
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011).
Han, S. et al. Primary outgrowth cultures are a reliable source of human pancreatic stellate cells. Lab Invest 95, 1331–1340 (2015).
Casey, S. C. et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 352, 227–231 (2016).
Casey, S. C., Baylot, V. & Felsher, D. W. MYC: Master Regulator of Immune Privilege. Trends Immunol 38, 298–305 (2017).
Sahai, V. et al. BET bromodomain inhibitors block growth of pancreatic cancer cells in three-dimensional collagen. Mol Cancer Ther 13, 1907–1917 (2014).
Kumar, K. et al. GLI2-dependent c-MYC upregulation mediates resistance of pancreatic cancer cells to the BET bromodomain inhibitor JQ1. Sci Rep 5, 9489 (2015).
Lee, S. J. et al. Interferon regulatory factor-1 is prerequisite to the constitutive expression and IFN-gamma-induced upregulation of B7-H1 (CD274). FEBS Lett 580, 755–762 (2006).
Zou, J. et al. MYC inhibition increases PD-L1 expression induced by IFN-gamma in hepatocellular carcinoma cells. Mol Immunol 101 (2018).
Garcia-Diaz, A. et al. Interferon Receptor Signaling Pathways Regulating PD-L1 and PD-L2 Expression. Cell Rep 19, 1189–1201 (2017).
Gore, J. & Korc, M. Pancreatic cancer stroma: friend or foe? Cancer Cell 25, 711–712 (2014).
Rhim, A. D. et al. Stromal elements act to restrain, rather than support, pancreatic ductal adenocarcinoma. Cancer Cell 25, 735–747 (2014).
Ozdemir, B. C. et al. Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell 25, 719–734 (2014).
Sherman, M. H. et al. Vitamin D receptor-mediated stromal reprogramming suppresses pancreatitis and enhances pancreatic cancer therapy. Cell 159, 80–93 (2014).
Mellman, I., Coukos, G. & Dranoff, G. Cancer immunotherapy comes of age. Nature 480, 480–489 (2011).
Kumar, K. et al. Differential Regulation of ZEB1 and EMT by MAPK-Interacting Protein Kinases (MNK) and eIF4E in Pancreatic Cancer. Mol Cancer Res 14, 216–227 (2016).
Sherman, M. H. et al. Stromal cues regulate the pancreatic cancer epigenome and metabolome. Proc Natl Acad Sci USA 114, 1129–1134 (2017).
Nielsen, S. R. et al. Macrophage-secreted granulin supports pancreatic cancer metastasis by inducing liver fibrosis. Nat Cell Biol 18, 549–560 (2016).
Hwang, R. F. et al. Cancer-associated stromal fibroblasts promote pancreatic tumor progression. Cancer Res 68, 918–926 (2008).
Krantz, S. B. et al. MT1-MMP cooperates with Kras(G12D) to promote pancreatic fibrosis through increased TGF-beta signaling. Mol Cancer Res 9, 1294–1304 (2011).
Shields, M. A. et al. Snail Cooperates with KrasG12D to Promote Pancreatic Fibrosis. Mol Cancer Res 11, 1078–1087 (2013).
Mathison, A. et al. Pancreatic stellate cell models for transcriptional studies of desmoplasia-associated genes. Pancreatology 10, 505–516 (2010).
Acknowledgements
We thank David Tuveson (Cold Spring Harbor Laboratories, Cold Spring Harbor, New York, US) for providing KPC1199 and KPC1245 mouse cell lines. This work was supported by grants R01CA186885 (to H.G. Munshi) and R21CA220625 (to K. Kumar) from the NCI, and a Merit award I01BX002922 (to H.G. Munshi) from the Department of Veterans Affairs.
Author information
Authors and Affiliations
Contributions
K.E. and K.K. (co-first authors) designed the studies, performed the experiments, analyzed the data, and wrote the manuscript. B.T.D., T.N.P., K.A.C., M.S., and D.R.P. performed the experiments and analyzed the data. M.A.S. and P.J.G. helped with the design of in vivo studies and edited the manuscript. S.G. provided K.P.C. cell lines, while R.F.H and R.U. provided immortalized pancreatic stellate cell lines. D.J.B. helped with procuring of the human PDAC specimens. H.G.M. designed the studies, analyzed the data, wrote and edited the manuscript, and secured funding.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ebine, K., Kumar, K., Pham, T.N. et al. Interplay between interferon regulatory factor 1 and BRD4 in the regulation of PD-L1 in pancreatic stellate cells. Sci Rep 8, 13225 (2018). https://doi.org/10.1038/s41598-018-31658-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31658-1
This article is cited by
-
High Expression Level of BRD4 Is Associated with a Poor Prognosis and Immune Infiltration in Esophageal Squamous Cell Carcinoma
Digestive Diseases and Sciences (2023)
-
Statins abrogate gemcitabine-induced PD-L1 expression in pancreatic cancer-associated fibroblasts and cancer cells with improved therapeutic outcome
Cancer Immunology, Immunotherapy (2023)
-
New insights into epigenetic regulation of resistance to PD-1/PD-L1 blockade cancer immunotherapy: mechanisms and therapeutic opportunities
Experimental Hematology & Oncology (2022)
-
Epigenetic regulation of pancreatic adenocarcinoma in the era of cancer immunotherapy
Journal of Gastroenterology (2022)
-
The BET family in immunity and disease
Signal Transduction and Targeted Therapy (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.