Abstract
The water soluble fullerene C60 daughter product - fullerols C60(OH)24 (FNP) possesses a great potential of modifying secondary metabolites biosynthesis. In order to clarify the extent of interaction, the impact of FNP (10, 100 and 1000 ng mL−1) on aflatoxin production and the available precursors of biosynthesis pathway from Aspergillus flavus NRRL 3251 was determined, in both the mycelia and yeast extract sucrose (YES) medium, during a 168-hour growth period at 29 °C in the dark. The FNP of 8 nm in diameter, and with a zeta potential of −33 mV affected mycelial growth at 1000 ng mL−1 while conidia production was slightly affected at 10 ng mL−1. The FNP effect on aflatoxin and it biosynthetic precursors was concentration dependent and alteration of the sterigmatocystin (ST) export from the cell was observed. Most of the monitored aflatoxin precursors, except norsolorinic acid, were detected in both mycelia and YES medium. However, observed precursor concentrations were much higher in mycelia, with exception of ST. The study shows the loss of FNP antioxidative effect after 120 hours of growth, and strong concentration dependent aflatoxigenic effect after that time. Thus, this data is relevant to guide future considerations on FNP-fungal interactions in the environments and on risk assessment.
Similar content being viewed by others
Introduction
Fullerols C60(OH)24 (FNP) are water soluble fullerene C60 daughter products with poorly defined environmental reactivity, distinctive material properties, and biological activities, which have gained great attention today. There are several available toxicity studies of FNP1,2,3,4,5; however, these studies were not carried out on fungal species. For example, Vávrová et al.1 stated that fullerols are low-toxic substances and Çavaş et al.2 reported that co-exposure with fullerol significantly reduced the cytotoxicity and genotoxicity of acetamiprid in IMR-90 cells. In addition, Milic Torres et al.3 reported the antiproliferative properties and protective effects of fullerol against doxorubicin cytotoxicity mediated by antioxidative and hydroxyl radical scavenging activity. Furthermore, Grebowski et al.4 reported that fullerol C60(OH)36 protects human erythrocyte membrane against high-energy electrons due to their reactive oxidative species (ROS) scavenging ability. Kazmierska-Grebowska et al.5, however, reported that fullerol C60(OH)36 exhibited dose-dependent effect by impairing hippocampal theta oscillations (in vivo and in vitro) and triggering epilepsy (in vitro) at relatively high concentrations (60 μM and 80 μM in vitro; 0.2 μg μL−1 in vivo) but exerting no apparent effects at lower concentrations (20 μM, 40 μM in vitro; 0.05 μg μL−1, 0.1 μg μL−1, 0.15 μg μL−1 in vivo).
FNP can be used as biomedical agents in chemotherapy, neurodegenerative diseases and radiobiology due to their antioxidant properties, which endows them as intensive research and promising application agents6,7,8. Increased use, environmental release, mineralization time up to 16 weeks, and limited literature about FNP – fungi interaction due to inherent properties of both mycotoxigenic fungi and FNP, are considerations that make this topic of prime importance. Fullerene C60 production is elevated by several tonnes per year while mycotoxigenic fungi migration is provoked by raising the environmental temperature in the face of climate change9,10,11. FNPs are mineralised in soils where the most important mycotoxin-producing saprophytic soil fungus, Aspergillus flavus, resides. These fungi are involved in pre-harvest and post-harvest crop colonization and contamination with aflatoxins. The mechanism of biological activity of FNP is related to their antioxidative properties, while A. flavus is sensitive to oxidative status perturbations. Literature on impact of FNP on A. flavus secondary metabolism, as a representative of mycotoxigenic fungi metabolism, is severely limited. In a bid to resolve the cause and consequences of such interaction, the impact of FNP on aflatoxin biosynthesis precursors, in both the mycelia and growth medium were determined.
Results and Discussion
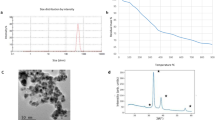
The particle size distribution by number of FNP in solution (10 µg mL−1) indicated that most of the particles had a hydrodynamic radius of 8 nm (Fig. 1a). The mean value of the zeta potential of the FNP aqueous solution was −33 mV (Fig. 1b). The data of the particle size distribution by number are in a very good agreement with the results of the particle size distribution by volume. FNP particle sizes ranged from 5 nm to 25 nm, with mean size of 8 nm as above mentioned. The DLS and zeta potential measurements of the FNP solution are consistent with previous reports12,13,14,15. Only particles with diameter smaller than 100 nm were present in the prepared FNP solution, this is in accordance with the EC Recommendation for the definition of the term “nanomaterial”16. In addition, the particle size and zeta potential data (Fig. 1), as well as theoretically calculated fullerenol C60(OH)24 molecule radius (1.1 nm), and the absence of non-agglomerated (single) molecules detection in this study, indicate the high tendency of FNPs to form agglomerates. The antimicrobial effect of FNPs is nanoparticle size-dependent17. Therefore, the 8 nm FNP agglomerates effect on A. flavus NRRL 3251 growth and secondary metabolism may have been due to interactions with and/or adsorption onto the fungal cell wall as proposed by Chen et al.18 and Quiao et al.19, as well as due to cell oxidative status modulation as established by Kovač et al.13.
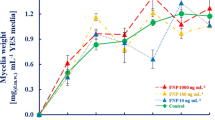
The effect of FNP on A. flavus mycelial growth is presented in Table 1. For measurable quantity of mycelia, both in FNP-treated and non-treated samples, at least 48 hours of incubation period was needed. Altogether, there was no statistically significant effect on mycelial growth (p > 0.05). However, it was found that A. flavus growth rate was stimulated in presence of the 1000 ng mL−1 of FNP. Unlike other FNP concentrations, the 10 ng mL−1 treated samples revealed mycelial mass reduction between 72 and 144 hours, indicating growth suppression during the logarithmic growth phase. The results of A. flavus mycelial growth stimulation under the influence of 1000 ng mL−1 FNP in this study are in disagreement with the reports of Aoshima et al.20 who reported the inhibition of Malasesia furfur and Candida albicans with different types of FNP at 60 to 100 times higher concentrations than those here applied. However, our report is partly in accordance with that of Gao et al.21 who reported stimulating effect of FNP on A. niger growth at concentrations higher than 10 µg mL−1. The disparity observed between the reports of Gao et al.21 and the present study, where the 100 ng mL−1 of FNP only stimulated growth at 72 and 120 hours, may have been due to the reproductive structures examined in both studies – conidia for Gao et al.21 and mycelia for the present study. The inhibition of mycelial growth at 10 ng mL−1 in the present study also agrees with the reduction of biomass yield of A. flavus, A. parasiticus and A. ochraceus by single low dose of FNP (5.2 ng mL−1) during a 120 h growth study22.
The effect of FNP on A. flavus conidia production ability is presented in Fig. 2. Collectively, FNP does not affect the conidia production ability of the fungus at statistically significant rate until 96 hours of growth. This is the first available data on conidia production under FNP influence. Furthermore, there is limited data and a knowledge gap regarding the FNP and mycotoxigenic fungi interaction, with the exception of reports of Unković et al.22 and Kovač et al.13,23. The relation between oxidative and/or drought stress, conidia production and aflatoxin biosynthesis are well documented24,25,26,27, and a change in the oxidative status of aflatoxigenic fungi due to FNP aflatoxin B1 (AFB1) production modulation was reported by Kovač et al.9,13. The results presented in Fig. 2 therefore probably reflects the impact of FNP on A. flavus cell oxidative status (up to the 96th hour of growth) as reported by Kovač et al.13, which brings the potential for reflection on the aflatoxin production.
Therefore, if the antiaflatoxigenic effect is expected, according to Holmes et al.28 there is obvious assumption that FNP could alter signalling inputs perceived by fungus cell, interfere with signal transduction and gene expression regulatory networks upstream of AFB1 biosynthesis or block a biosynthetic enzyme activity. Due to such assumption, the available aflatoxin precursors of the biosynthetic pathway (Figs 3–5) of A. flavus NRRL 3251 secondary metabolism in FNP presence were determined, in both the mycelia and growth medium. In addition, listing of aflatoxin precursors in the group of “emerging toxins”29 is additional reason to determine their concentrations in the presence of FNP. Such data are necessary for risk assessment, i.e. potential future legislation regarding the FNP production and release.
The aflatoxin biosynthetic pathway is proposed to originate in peroxisomes (Fig. 3), continues with multiple fusion events in vesicles and ends with formation of aflatoxisomes – the large bodies containing aflatoxins and their precursors (Figs 4 and 5)27,30,31.
The initial reactions of this energy expensive biosynthetic pathway (1 AC-CoA, 9 MA-CoA, <10 NADPH, 2 SAM, 9 ATP) begins in peroxisomes by norsorolinic acid (Nor-AC) synthesis31,32,33. The effect of FNP on Nor-AC concentration is shown in Fig. 3. Nor-AC was detected only in mycelial samples; both in FNP treated and control samples. The most probable reason for lack of Nor-AC in growth medium is its rapid conversion in further steps of the pathway. Such results for Nor-AC are in accordance with the model for compartmentalization, translocation, and aflatoxisome biogenesis during biosynthesis of aflatoxin in Aspergilli as proposed by Rose et al.34. However, FNP at all tested concentrations caused an average decrease of 67% of Nor-AC in mycelia until 72 hours of growth in comparison to the control growth. The Nor-AC concentration increased after 72 hours of growth under FNP influence while growth concentration remained constant in control samples. After 144 hours of growth, FNP caused increase of Nor-AC biosynthesis up to 42% at 100 ng mL−1 while at 10 and 1000 ng mL−1 FNP increase of Nor-AC production was at 25% rate (Fig. 3). This implies possible involvement of FNP in early stages of aflatoxin biosynthesis.
The next two steps of the aflatoxin biosynthetic pathway occur in the cytosol (Figs 4 and 5). The pathway enzymes and precursors are translocated into the cytosol via vesicles that originate from budding of peroxisomes and mitochondria, and they fuse with trafficking and secretory vesicles, altogether forming aflatoxisomes - large bodies containing substantial aflatoxin concentrations, pathway enzymes and aflatoxin precursors31,34,35.
At first, Nor-AC is converted to averantin (AVN) then to averufin (AVR), with averufanin (AFN) as intermediate. After multiple reactions, AVR is converted to versicolorins B (VB), A (VA) and C (VC)36. The effects of FNP on concentrations of the hydroxyantraquinones AVN, AFN, AVR, VA and VC are shown in Fig. 4. VB is synthesised and converted to aflatoxin by multiple enzyme reactions (Fig. 5). At first, sterigmatocystin (ST) and O-methylsterigmatocystin (OMST) are formed, and later the aflatoxins B1 and B2 at the end of the biosynthetic pathway. All of mentioned aflatoxin precursors (Figs 4 and 5) were biosynthesised during the entire incubation period, and were present in mycelia and also released into the growth medium. The concentrations of AVN, AVR, AFN, VA and VC in the mycelia were on average 2.3–3.6, 7.9–14.4, 105–255, 2.6–6.0 and 61.2–80.1 times higher than in the medium, respectively. The sum of AVN concentrations (mycelia and medium) was not significantly affected by FNP, except at 100 and 1000 ng mL−1 FNP for 72 hours of growth where an increase of 95 and 72%, respectively, were recorded. In the case of AFN, an increase in the sum of mycelia and medium concentrations was recorded after 144 hours of growth for 10 ng mL−1 (32%) and 100 ng mL−1 (up to 53%) FNP. The sum of AVR concentrations in mycelia and medium, however, showed about 23% decrease until 72 hours of growth, while an increase in AVR concentration was observed for all FNP concentrations after 144 hours, especially at 100 ng mL−1 (up to 80%). Furthermore, nidirufin (NID), C-2′hydroxyl derivative of AVN37, was observed only in mycelial samples. Its concentrations decreased by 51% under FNP influence until 72 hours and increased by as much as 58% after 144 hours of growth (Fig. 5). NID is one of the by-products in the transformation process of AVN to VA36. VA is the result of multiple enzyme transformation of AVR to VB, which can be further transformed to VC. The sum of VA concentrations decreased under FNP presence at 72 hours of growth. VC is a “side shunt metabolite” of the aflatoxin biosynthesis pathway and is regarded as non-toxic36. The sum of its concentrations in mycelia and medium decreased under FNP presence from 48 to 144 hours of growth (Fig. 4).
The most toxic of the AFB1 precursors, which has been assigned to group 2 (possibly carcinogenic to humans) by the International Agency for Research on Cancer (IARC 2012), is ST38. ST is one of the few metabolites found in higher concentrations in medium than in mycelia, indicating possible active transport out of the fungal cell. However, the concentrations in the sum of ST (mycelia and medium) showed 50% increase until 72 hours, and the 95% decrease after 96 hours of growth, all in the presence of FNP. This implies an early export of ST from the cell until 72 hours of growth due to FNP. The results recorded for ST are also in accordance with the observed sum of concentrations of OMST. The sum of AFB1 concentrations in mycelia and medium decreased under FNP presence until 120 hours of growth at average rate of 58%, but increased afterwards in a concentration dependent manner at an average rate of 22% until 168 hours of growth, where the highest FNP concentration applied showed the strongest aflatoxigenic effect. However, a higher FNP effect occurred in the case of AFB2, where a concentration dependent aflatoxigenic effect leading to a 50% average concentration increase at highest applied FNP concentration after 120 hours of growth was observed. In addition, AFM1 was detected in both mycelia and medium, and the sum of the concentrations revealed the strongest aflatoxigenic effect at an average rate of 93% until 144 hours of growth at the lowest FNP concentration (Fig. 5). AFM1 is primarily found in animal tissues and fluids (milk and urine) as a metabolic product of AFB139 but is also found as a product of secondary metabolism of Aspergillus species40,41,42,43. There was no observed statistically significant difference between the ratio of aflatoxins B1 and B2 for the applied FNP concentrations. In general the ratio between AFB1 to AFB2 was in accordance with literature, on average 93 ± 4% of AFB1, and 7 ± 4% of AFB2 was produced44. Furthermore, the correlation between conidia production and AFB1 production was statistically significant for control samples (r = 0.85838; p = 0.0159) and FNP treated samples at 10 ng mL−1 (r = 0.86229; p = 0.0152), 100 ng mL−1 (r = 0.90389; p = 0.0189) and 1000 ng mL−1 (r = 0.22522; p = 0.00011).
This work presents updated information on the most detailed and available study on FNP effect on growth and aflatoxin producing ability of A. flavus13,23. The novelty of this study is in the determination of every available aflatoxin precursor in the biosynthetic pathway of A. flavus NRRL 3251 secondary metabolism. Previously Kovač et al. (2017)13 demonstrated the antiaflatoxigenic effect of FNP in YES medium and the published data did not include mycelial AFB1 content; thus, the hypothesis that mycelial mycotoxin content could be different was formulated. The main reason for this assumption is that the antioxidative potential of FNP is limited with time such that the proven ability of FNP to modulate oxidative stress within the fungal cell, even at relatively low doses, disappear after some time (Fig. 5). Sequel to the initial antioxidative activity of FNP, the AFB1 production increases due to oxidative stress modulation (i.e. higher oxidative stress levels), and the increased oxidative stress is one that is known to trigger AFB1 biosynthesis45. The data in the present study expatiates the previous work by Kovač et al.13, which was on a preliminary investigation into the modulation of mycotoxin production by FNP. Furthermore, this present study addresses issues of study design, i.e. importance of metabolite determination in both mycelial and media, FNP alteration ability into aflatoxin biosynthetic pathway precursors export, as well as AFB1 accumulation within the cells.
Conclusion
This study has shown that FNP modulates aflatoxin production by affecting the biosynthetic pathway of metabolites reported herein. When considering environmental risk estimation, the sum of mycotoxin concentrations from mycelia and growth medium should be considered to determine FNP impact on toxigenic fungi. Nanoparticles as an environmental threat are concentration-dependent, and trends of FNP concentration increase are expected in the future due to increased production, usage, release, and time-lag in the mineralisation processes in the environment. Thus, the environmentally occurring FNP levels are capable of posing threats to the toxigenic potentials of mycotoxigenic moulds (e.g. A. flavus), especially in the face of climate change. In this study, FNP exerted a concentration-dependent effect on the aflatoxin biosynthesis pathway and an alteration in ST export from the cell occurred. Prior to 120 hours of growth of A. flavus, FNP exerted antioxidative potential which vanished afterwards to cause strong and concentration-dependent rise in biosynthesis of aflatoxins. Consequently, such negative prospects of FNP application evoke thoughts on investigating photosensitised FNP action on A. flavus and other mycotoxigenic fungal species as an option of controlling aflatoxigenicity in the future.
Material and Methods
Chemicals
Yeast extract, potato dextrose agar and sucrose were purchased from Biolife (Italy) while aflatoxin standard mix (B1, G1, B2, G2) was purchased from Biopure (Austria). Acetonitrile and methanol (both HPLC grade) were obtained from Merck (Germany). Ammonium acetate and glacial acetic acid (p.a.) were purchased from Sigma Aldrich (Vienna, Austria). A Purelab Ultra system (ELGA LabWater, Celle, Germany) was used for ultrapure water preparation. Standards of A. flavus metabolites were purchased from Various research groups or from the following commercial sources: Romer Labs®Inc.(Tulln, Austria), Sigma–Aldrich (Vienna, Austria), Iris Biotech GmbH (Marktredwitz, Germany), Axxora Europe (Lausanne, Switzerland) And LGC Promochem GmbH (Wesel, Germany) and prepared as described by Malachova et al.46.
Fullerol C60(OH)24 synthesis, preparation and characterisation of nanoparticles solution
Fullerol C60(OH)24 was synthesised and nanoparticle solution prepared in ultrapure water as previously described by Mirkov et al.47. Dynamic light scattering (DLS) technique was used for the determination of hydrodynamic size, and electrophoretic light scattering (ELS) for the measurements of the surface charge (zeta potential (ζ)) of analysed samples. The measurements were conducted on a Zetasizer Nano ZS instrument (Malvern Instruments Inc., UK). All DLS analyses were performed in triplicates at 633 nm wavelength and a measurement angle of 173o (backscatter detection), in an aqueous solution at room temperature. Zeta potential (ζ) measurements were performed in duplicate.
Aspergillus flavus growth and spore count determination
The preparation of conidia suspension of A. flavus NRRL 3251, inoculation, as well as mycelial growth in aflatoxin-inducing YES medium were conducted in the dark at 29 °C which favours aflatoxin production as previously described (Kovač et al.)13. The inoculated flasks were incubated on a rotary shaker (KS 260 basic, IKA, Germany) at 200 rpm for 168 hours in the presence of environmentally plausible C60(OH)24 concentrations (0, 10, 100 and 1000 ng mL−1). Samples of medium and mycelia were collected from the flasks every 24 hours from the 48th to 168th hour of incubation. Mycelia were separated from the media by filtration, mycelia precooled and stored in 2 mL tubes at −80 °C for at least 24 hours until lyophilisation (Christ, Alpha 1–4 LD, Germany). Drying conditions were as follows: freezing temperature −55 °C; temperature of sublimation −35 to 0 °C; vacuum level 0.220 mbar. The temperature of isothermal desorption varied from 0 to 22 °C under the vacuum of 0.060 mbar. Freeze-drying lasted until the constant mass of mycelia was obtained, which was about 5 h. In addition, a part of mycelia prior to −80 °C storage and lyophilisation was dried until constant mass (24 hr at 105 °C) in order to determine dry mycelial weight.
The conidia count in YES medium after separation was performed as previously described13.
Estimation of A. flavus metabolites in mycelia and culture media
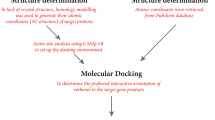
For the estimation of A. flavus metabolites in mycelia and culture medium, the multi-analyte “dilute and shoot” LC-MS/MS method was used as previously described46.
The 125 mg of the lyophilised mycelia were mixed with 1 mL of extraction solvent (acetonitrile/water/acetic acid 79:20:1, v/v/v) and extracted for 90 min at GLF 3017 rotary shaker (GLF, Germany). Extracts were transferred into glass vials and two-fold diluted with dilution solvent (acetonitrile/water/acetic acid 20:79:1, v/v/v). Vial contents were vigorously mixed and 5 µL was injected directly into LC-MS/MS system. The metabolites in YES medium were estimated by following a ten-fold dilution with dilution solvent in glass vials without any pre-treatment, as well. The screening and detection of metabolites was the same as described by Malachová et al.46, in brief the QTrap 5500 MS/MS detector (Applied Biosystems, Foster City, CA) equipped with TurboV electrospray ionization (ESI) source, and Agilent 1290 binary UHPLC system (Agilent Technologies, Waldbronn, Germany) was used. For the separation of the metabolites the Gemini® C18 column (150 × 4.6 mm i.d., 5 µm particle size) was combined with the C18 security guard pre-column (4 × 3 mm i.d.) (Phenomenex, Torrance, CA, US). The eluents were freshly prepared, the gradient and the flow rate were followed exactly as described by Malachová et al.46. The Scheduled selected reaction monitoring (sSRM) mode was used, and two runs per sample were used (each for one mode). The detection window was set to ±27 s in positive and ±42 s in negative mode due to number of monitored metabolites. The ESI source parameters were followed exactly as described by Malachová et al.46. Two sSRM transitions were monitored per metabolite (quantifier, and qualifier), and according to the validation guidelines, the ratio between two transitions were used as additional identity conformation point.
Statistical analysis
Data presented in this paper are expressed as the mean value ± SEM from three separate experiments. The pooled datasets were checked for distribution normality by Shapiro-Wilk test and compared by nonparametric statistics methods (Friedman ANOVA and Kendall coefficient of concordance; Kruskal-Wallis ANOVA). The programme package Statistica 12.0 (Dell, 2015) was used and differences were considered significant when the p value was < 0.05.
References
Vávrová, J., Řezáčová, M. & Pejchal, J. Fullerene nanoparticles and their anti-oxidative effects: a comparison to other radioprotective agents. J. Appl. Biomed. 10, 1–8 (2012).
Çavaş, T., Çinkiliç, N., Vatan, O. & Yilmaz, D. Effects of fullerenol nanoparticles on acetamiprid induced cytoxicity and genotoxicity in cultured human lung fibroblasts. Pestic. Biochem. Physiol. 114, 1–7 (2014).
Torres, V. M. et al. Fullerenol C60(OH)24prevents doxorubicin-induced acute cardiotoxicity in rats. Pharmacol. Reports 62, 707–718 (2010).
Grebowski, J. et al. Fullerenol C60(OH)36protects human erythrocyte membrane against high-energy electrons. Biochim. Biophys. Acta - Biomembr. 1860, 1528–1536 (2018).
Kazmierska-Grebowska, P. et al. Fullerenol C60(OH)36at relatively high concentrations impairs hippocampal theta oscillations (in vivo and in vitro) and triggers epilepsy (in vitro) – A dose response study. Exp. Mol. Pathol. 105, 98–109 (2018).
Injac, R. et al. Protective effects of fullerenol C60(OH)24against doxorubicin-induced cardiotoxicity and hepatotoxicity in rats with colorectal cancer. Biomaterials 30, 1184–1196 (2009).
Gitsov, I. et al. Polymer-assisted biocatalysis: Unprecedented enzymatic oxidation of fullerene in aqueous medium. J. Polym. Sci. Part A Polym. Chem. 50, 119–126 (2012).
Sachkova, A. S. et al. On mechanism of antioxidant effect of fullerenols. Biochem. Biophys. Reports 9, 1–8 (2017).
Medina, Á., Rodríguez, A., Sultan, Y. & Magan, N. Climate change factors and Aspergillus flavus: effects on gene expression, growth and aflatoxin production. World Mycotoxin J. 8, 171–179 (2015).
Medina, A. et al. Interactions between water activity and temperature on the Aspergillus flavus transcriptome and aflatoxin B 1 production. Int. J. Food Microbiol. 256, 36–44 (2017).
Battilani, P. et al. Aflatoxin B1 contamination in maize in Europe increases due to climate change. Sci. Rep. 6, 24328 (2016).
Letenko, D. G. et al. Study of aqueous solutions of fullerenol-d by the dynamic light scattering method. Russ. J. Appl. Chem. 84, 50–53 (2011).
Kovač, T. et al. Fullerol C60(OH)24 nanoparticles and mycotoxigenic fungi: a preliminary investigation into modulation of mycotoxin production. Env. Sci Pollut Res 60, 1–9 (2017).
Borišev, M. et al. Drought impact is alleviated in sugar beets (Beta vulgaris L.) by foliar application of fullerenol nanoparticles. PLoS One 11, 1–20 (2016).
Vraneš, M. et al. Self-assembling, reactivity and molecular dynamics of fullerenol nanoparticles. Phys. Chem. Chem. Phys. 19, 135–144 (2017).
Rauscher, H. et al. Towards a review of the EC Recommendation for a definition of the term ‘nanomaterial’, Part 3: Scientific-technical evaluation of options to clarify the definition and to facilitate its implementation, https://doi.org/10.2788/678452 (2015).
Lyon, D. Y., Adams, L. K., Falkner, J. C. & Alvarez, P. J. J. Antibacterial activity of fullerene water suspensions: Effects of preparation method and particle size. Environ. Sci. Technol. 40, 4360–4366 (2006).
Chen, R. et al. Differential uptake of carbon nanoparticles by plant and mammalian cells. Small 6, 612–617 (2010).
Qiao, R., Roberts, A. P., Mount, A. S. & Klaine, S. J. Translocation of C 60 and Its Derivatives Across a Lipid Bilayer. Nanoletters 7, 614–619 (2007).
Aoshima, H. et al. Antimicrobial of fellerenes and their OH derivatives.pdf. Biocontrol Sci. 14, 69–72 (2009).
Gao, J. et al. Polyhydroxy fullerenes (fullerols or fullerenols): Beneficial effects on growth and lifespan in diverse biological models. PLoS One 6, 1–8 (2011).
Unković, N. et al. Aspergilli response to benzalkonium chloride and novel-synthesized fullerenol/benzalkonium chloride nanocomposite. Sci. World J. 2015, (2015).
Kovač, T. et al. Antiaflatoxigenic effect of fullerene C60 nanoparticles at environmentaly plausible concentrations. AMB Express 8, 14 (2018).
Jayashree, T. & Subramanyan, C. Oxidative stress as a prerequisite for aflatoxin production by Aspergillus parasiticus. Free Radic. Biol. Med. 29, 981–985 (2000).
Narasaiah, K. V., Sashidhar, R. B. & Subramanyam, C. Biochemical analysis of oxidative stress in the production of aflatoxin and its precursor intermediates. Mycopathologia 162, 179–189 (2006).
Linz, J. E., Hong, S. Y. & Roze, L. V. Oxidative stress-related transcription factors in the regulation of secondary metabolism. Toxins (Basel). 5, (683–702 (2013).
Fountain, J. C. et al. Responses of Aspergillus flavus to oxidative stress are related to fungal development regulator, antioxidant enzyme, and secondary metabolite biosynthetic gene expression. Front. Microbiol. 7, (2016).
Holmes, R. A., Boston, R. S. & Payne, G. A. Diverse inhibitors of aflatoxin biosynthesis. Appl Microbiol Biotechnol 78, 559–572 (2008).
Kovalsky, P. et al. Co-Occurrence of Regulated, Masked and Emerging Mycotoxins and Secondary Metabolites in Finished Feed and Maize—An Extensive Survey. Toxins (Basel). 8, (1–29 (2016).
Chanda, A. et al. A key role for vesicles in fungal secondary metabolism. Proc. Natl. Acad. Sci. USA 106, 19533–19538 (2009).
Keller, N. P. Translating biosynthetic gene clusters into fungal armor and weaponry. Nat. Chem. Biol. 11, 671–677 (2015).
Yabe, K. & Nakajima, H. Enzyme reactions and genes in aflatoxin biosynthesis. Appl. Microbiol. Biotechnol. 64, 745–755 (2004).
Reverberi, M. et al. How peroxisomes affect aflatoxin biosynthesis in Aspergillus flavus. PLoS One 7, e48097 (2012).
Roze, L. V., Chanda, A. & Linz, J. E. Compartmentalization and molecular traffic in secondary metabolism: A new understanding of established cellular processes. Fungal Genet. Biol. 48, 35–48 (2011).
Kistler, H. C. & Broz, K. Cellular compartmentalization of secondary metabolism. Front. Microbiol. 6, 1–11 (2015).
Ezekiel, C. N., Sulyok, M., Warth, B., Odebode, A. C. & Krska, R. Natural occurrence of mycotoxins in peanut cake from Nigeria. Food Control 27, 338–342 (2012).
Murphy, R. A. & Cava, M. P. Stereochemistry of Nidurufin: Synthesis of 6,8-Dideoxynidurufin and 6,8-Dideoxyepinidurufin. J. Am. Chem. Soc. 106, 7630–7632 (1984).
Varga, J., Baranyi, N., Chandrasekaran, M., Vágvölgyi, C. & Kocsubé, S. Mycotoxin producers in the Aspergillus genus: An update. Acta Biol. Szeged. 59, 151–167 (2015).
Varga, J., Frisvad, J. C. & Samson, R. A. A reappraisal of fungi producing aflatoxins. World Mycotoxin J. 2, 263–277 (2009).
Yabe, K. et al. Production of M-/GM-group aflatoxins catalyzed by the OrdA enzyme in aflatoxin biosynthesis. Fungal Genet. Biol. 49, 744–754 (2012).
Chala, A. et al. Multimycotoxin analysis of sorghum (Sorghum bicolor L. Moench) and finger millet (Eleusine coracana L. Garten) from Ethiopia. Food Control 45, 29–35 (2014).
Sulyok, M. et al. Quantitation of multiple mycotoxins and cyanogenic glucosides in cassava samples from Tanzania and Rwanda by an LC-MS/MS-based multi-toxin method. Food Addit. Contam. - Part A Chem. Anal. Control. Expo. Risk Assess. 32, 488–502 (2015).
Abdallah, M. F., Girgin, G., Baydar, T., Krska, R. & Sulyok, M. Occurrence of multiple mycotoxins and other fungal metabolites in animal feed and maize samples from Egypt using LC-MS/MS. J. Sci. Food Agric. 97, 4419–4428 (2017).
Kensler, T. W., Roebuck, B. D., Wogan, G. N. & Groopman, J. D. Aflatoxin: A 50-year Odyssey of mechanistic and translational toxicology. Toxicol. Sci. 120, 28–48 (2011).
Reverberi, M. et al. Antioxidant enzymes stimulation in Aspergillus parasiticus by Lentinula edodes inhibits aflatoxin production. Appl. Microbiol. Biotechnol. 69, 207–215 (2005).
Malachová, A., Sulyok, M., Beltrán, E., Berthiller, F. & Krska, R. Optimization and validation of a quantitative liquid chromatography-tandem mass spectrometric method covering 295 bacterial and fungal metabolites including all regulated mycotoxins in four model food matrices. J. Chromatogr. A 1362, 145–156 (2014).
Mirkov, S. M. et al. Nitric oxide-scavenging activity of polyhydroxylated fullerenol, C 60(OH) 24. Nitric Oxide - Biol. Chem. 11, 201–207 (2004).
Acknowledgements
This work was funded by Josip Juraj Strossmayer University of Osijek (Young Scientist Grant) while nanoparticles characterization was supported by the Ministry of Education, Science and Technological Development of the Republic of Serbia, Project number III 45005.
Author information
Authors and Affiliations
Contributions
T. Kovač and B. Šarkanj conceived and designed experiments. T. Kovač and I. Borišev carried out FNP suspension preparation and characterization. T. Kovač and B. Crevar carried out fungal growth and prepared mycelial samples while T. Kovač, B. Crevar and M. Kovač prepared growth media samples for LC-MS/MS analysis. M. Sulyok, B. Šarkanj, T. Kovač and R. Krska performed LC-MS/MS analysis and carried out qualification and quantification of metabolites. T. Kovač and B. Šarkanj prepared data for statistical analysis; statistical analysis and figure preparation was carried out by T. Kovač and F. Čačić Kenjerić. T. Kovač, B. Šarkanj, M. Sulyok, R. Krska, I. Strelec, and C.N. Ezekiel analysed the data and interpreted results of statistical analysis. T. Kovač wrote the manuscript. B. Šarkanj, M. Kovač, I. Borišev, C.N. Ezekiel, I. Strelec, M. Sulyok and R. Krska reviewed and revised manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kovač, T., Borišev, I., Crevar, B. et al. Fullerol C60(OH)24 nanoparticles modulate aflatoxin B1 biosynthesis in Aspergillus flavus. Sci Rep 8, 12855 (2018). https://doi.org/10.1038/s41598-018-31305-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31305-9
This article is cited by
-
Impact of fullerol C60(OH)24 nanoparticles on the production of emerging toxins by Aspergillus flavus
Scientific Reports (2020)
-
Thiosemicarbazone nano-formulation for the control of Aspergillus flavus
Environmental Science and Pollution Research (2020)
-
Restricting mycotoxins without killing the producers: a new paradigm in nano-fungal interactions
Applied Microbiology and Biotechnology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.