Abstract
Biological routes to the production of fuels from renewable feedstocks hold significant promise in our efforts towards a sustainable future. The fatty acid decarboxylase enzyme (OleTJE) is a cytochrome P450 enzyme that converts long and medium chain fatty acids to terminal alkenes and shares significant similarities in terms of structure, substrate scope and mechanism with the hydroxylase cytochrome P450 (P450BSβ). Recent reports have demonstrated that catalytic pathways in these enzymes bifurcate when the heme is in its iron-hydroxo (compound II) state. In spite of significant similarities, the fundamental underpinnings of their different characteristic wild-type reactivities remain ambiguous. Here, we develop point charges, modified parameters and report molecular simulations of this crucial intermediate step. Water occupancies and substrate mobility at the active site are observed to be vital differentiating aspects between the two enzymes in the compound II state and corroborate recent experimental hypotheses. Apart from increased substrate mobility in the hydroxylase, which could have implications for enabling the rebound mechanism for hydroxylation, OleTJE is characterized by much stronger binding of the substrate carboxylate group to the active site arginine, implicating it as an important enabling actor for decarboxylation.
Similar content being viewed by others
Introduction
Cytochrome P450 (P450s) superfamily of enzymes have been found in all life forms and have evolved to catalyze a wide variety of important chemical transformations ranging from hydroxylations, oxidations, epoxidations, dehydrogenation, deamination, dehalogenations and decarboxylation. P450s achieve these transformations with the help of a cysteine coordinated iron porphyrin group or the heme1. One of the recently characterized transformations involves conversion of fatty acids to terminal alkenes2 which enables the enzymatic production of hydrocarbons from common biological metabolites, and thus offers a promising route for the sustainable production of fuels that are amenable with current energy infrastructure and can replace conventional petroleum derived fuels3. In 2011, Rude et al. demonstrated that the Jeotgallicoccus bacterial species produces a P450 enzyme (OleTJE) that catalyzes the decarboxylation of saturated fatty acids (eg: myristic acid-C14, palmitic acid-C16, stearic acid-C18 and arachidic acid-C20) to terminal alkenes2. In 2014, the crystal structure of OleTJE was experimentally determined and provided molecular level insight into the structural basis for this decarboxylase enzyme4. The structure revealed significant similarity in its secondary and tertiary protein structure with other P450 enzymes especially P450SPα and P450BSβ peroxygenases which hydroxylate fatty acids5,6. OleTJE and the hydroxylase P450s not only share structural similarities but also operate on the same substrates, which has made understanding the basis for their specific reactivity particularly intriguing.
Since 2014, there have been a number of experimental and computational studies aimed at probing various aspects of OleTJE ranging from the mechanism, substrate range and protein engineering strategies to improve enzyme efficiency. Grant et al. in 2015 observed for the first time, the presence of a ferryl-oxo radical species (compound I) in OleTJE suggesting that the 1st step of the reaction mechanism involves the abstraction of a hydrogen from the substrate by the oxygen on compound I7. A subsequent study published in 2016 from the same group, confirmed the presence of a 2nd intermediate as the ferryl-hydroxo species (compound II), which was observed to be stable over the picosecond time scale8.
One of the most interesting aspects of OleTJE is its unique decarboxylating activity in spite of its close similarity in terms of structure and substrate with the hydroxylating enzymes P450SPα and P450BSβ. The reaction mechanism (Fig. 1) in these enzymes has been explored with DFT and QM/MM studies and portions of the mechanism have been corroborated in more recent experimental studies8,9,10 It has been established that the first step of the mechanism involves the abstraction of a hydrogen atom from the substrate fatty acid (from the α carbon for P450SPα and the β carbon for the P450BSβ and OleTJE) by compound I7,10. This generates the intermediate state of the system constituting a radical on the substrate and compound II8,10. The next step of the mechanism is crucial in determining whether the resulting product is a hydroxylated fatty acid or an alkene and a CO2 molecule. The current hypothesis involves the hydroxylase enzyme enabling the rebound mechanism wherein the substrate radical is hydroxylated when it approaches compound II, while the decarboxylase enzyme enables decarboxylation by avoiding this rebound8,10. Hence, despite the similarity in their sequence and overall structures, OleTJE and P450BSβ employ unique enabling factors to catalyze their respective reactions.
(Top) The reaction mechanism: The first step of H-abstraction by compound I is common to both hydroxylases and decarboxylases followed by catalytic bifurcation at the intermediate state with the substrate radical and compound II. (Bottom: Left) Canonical P450 structural fold in OleTJE with α-helices colored as follows A,A’-blue; B,B’-red; C,D-orange; E,F-yellow; G,H-tan; I,J-gray; J’,K-green. The β-sheets are colored in white and the loops in pink. (Bottom: Right) Structural comparison between OleTJE (silver) and P450BSβ (white). The differences in the F-G loop are highlighted with the longer OleTJE loop shown in Blue and the shorter loop in P450BSβ shown in red.
There have been numerous experimental studies focused on enzyme engineering strategies to widen the substrate range and improve the efficiency of OleTJE. For example, there has been success in enabling OleTJE to decarboxylate short aromatic and linear carboxylic acids and also in recent evaluation of the ability of other redox partners in activating OleTJE11,12,13,14. More recently, the role of key OleTJE active site residues (Phe79, His85 and Arg245) in substrate binding and catalytic activity has been investigated15, revealing the crucial catalytic role of the arginine residue. The His85 has been proposed to play a role in decarboxylation, since it is replaced by a glutamine in the hydroxylase enzyme. While the activity of both the H85Q and F79A OleTJE mutants were reduced, it was still observed to have similar product slates compared to the wild-type15.
The energetics for the competing decarboxylation and hydroxylation mechanisms have also been explored using QM/MM calculations providing insight into the importance of the Fe spin state and its impact on reaction barriers10. Upon the initial H-abstraction step, it was suggested that the H-bonding networks and polarity of the active site arginine-carboxylate interaction destabilize the rebound processes that enable hydroxylation10. In a more recent study, Du et al. employ classical molecular dynamics to study active site tunnels, binding energies and identified a key interaction aiding substrate binding for short (C12), medium and (C16) and long (C20) substrates in OleTJE16. This study mainly focused on the impact of substrate chain length on OleTJE activity considering the enzyme in its compound I state and the substrate in its reactant form. Computational studies on the catalytic pathway bifurcation step of the enzyme have been hindered by the lack of parameters to adequately describe the compound II state. The fundamental enabling actors and the mechanistic underpinnings of this catalytic bifurcation step are thus of compelling interest.
While most studies have focused solely on OleTJE to address the above question, we look to compare and contrast the wild-type forms of OleTJE and P450BSβ enzymes structures and their dynamics during all stages of the reaction mechanism to glean differences and understand the basis for their respective catalytic activities. The parameters for compound II, developed in this study, thereby enable molecular dynamics simulations of the crucial state wherein catalytic pathway bifurcation is hypothesized to occur. We start with a comparison of the crystal structures and highlight important differences between the two enzymes. We then simulate the dynamics of the apo and substrate bound enzyme in its reactant (compound I, substrate) and intermediate (compound II, substrate radical) states and monitor protein fluctuations, characterize key substrate enzyme interactions and substrate access, egress tunnels. Here, we choose to conduct substrate-bound simulations in both enzymes with myristate (C14) in order to eliminate substrate-associated differences. Myristate is chosen since it has been demonstrated to have the most decarboxylase activity in OleTJE and has also been known to be hydroxylated by P450BSβ17,18. The simulations reveal contrasting substrate binding and dynamics in OleTJE and P450BSβ active sites implicating the increased binding to active site arginine, sustained active site water occupancies in the former, and increased substrate mobility in the latter as key players in establishing characteristic catalytic activities.
Methods
Structural Comparisons
The structures for P450BSβ PDB ID 1IZO and OleTJE PDB ID 4L40 were obtained from the protein data bank4,18. Chain A of both structures was used for analyses as well as for conducting molecular dynamics simulations. The structures were aligned using the Matchmaker tool in Chimera molecular visualization and analysis package19. The aligned sequences and structures were analyzed for positional differences in amino acids.
Molecular Dynamics Simulations
The crystal structure of the OleTJE enzyme from Jeotgalicoccus sp. 8456 (PDB ID: 4L40) and the P450BSβ hydroxylase from Bacillus subtilis (PDB ID:1IZO) were used for setting up MD simulations, which were run using the Amber16 package20. The geometric parameters for the heme including the cysteine binding residue were obtained from Shahroukh et al.21 Both OleTJE and P450BSβ were simulated at various stages of the reaction mechanism namely apo, reactant (substrate bound compound I) and intermediate (substrate in its radical form with compound II) states. The point charges for compound I and the compound II were calculated based on quantum mechanical (QM) calculations of a reduced system. The QM calculations were performed at the B3LYP22 level of theory and LANL2DZ23 basis set for Fe and 6–31 G(d) for C, H, O, N, and S atoms with point charges derived via Restricted Electrostatic Potential methodology22 using the R.E.D server24. The frcmod files and the prep files for heme are included as part of the supplemental information. The ff14SB force field25 was used to describe the protein, GAFF26 for the fatty acid substrate and intermediate, and TIP3P27 for water molecules. In order to calculate the point charges on the β radical substrate, the structure was initially optimized at the HF/6–31 G level using the Gaussian package followed by the derivation of its RESP point charges using the antechamber package28.
The online pKa prediction server H++ was used to estimate the protonation states for the amino acid residues of the protein29. The protein systems were solvated with a water buffer of 10 Å and then minimized for 2000 steps using the steepest gradient methodology and for 5000 steps in the conjugate gradient method. Subsequently, the system was equilibrated in the isobaric isothermal ensemble (NPT) at 300 K with a non-bonded interaction cutoff of 10 Å for 1 ns. The MD runs employed a 2 fs time step with lengths of bonds to hydrogen atoms in the system restrained using the SHAKE algorithm30. Post equilibration, 150 ns trajectories were simulated for production runs and data analysis.
Analyses
Substrate access to and from the active site has been one of the most well characterized features in P450 enzyme systems31. The CAVER 3.0 software package was used to characterize tunnels and channels connecting the active site of the protein to its external surface in OleTJE and P450BSβ systems in their apo and substrate-bound states32. A total of 1500 frames from 150 ns were analyzed considering a probe radius of 1 Å and a clustering threshold of 3.5, as employed by Du et al. for their CAVER results on OleTJE16. The CAVER analyses for substrate-bound states were done in the presence of the substrate.
The MMPBSA.py utility in AMBER was utilized for the calculation of overall and residue-based decomposition binding energies from MD trajectories33. A total of 7500 frames (1 every 20 ps from the 150 ns trajectory) were used for the MMGBSA binding energy calculations. An in-house VMD script was utilized to identify residue-based substrate – enzyme contact lists, which were used for performing decomposition analysis of the MMGBSA binding energies. This has previously been successfully used to identify amino acid residues important for substrate binding34,35. The trajectory analysis tool ptraj in concert with pytraj was used for RMSF, RMSD and dynamic cross-correlation analysis36. The trajectory alignment protocol used the coordinates of the α-helices and β-sheets of the first frame of each trajectory as the reference. The residue-based RMSFs for the enzymes were calculated considering the Cα atoms. VMD was used for calculating volumetric occupancy maps (volmaps) and hydrogen bond analyses37. Volmaps were calculated for water occupancies within 10 Å of heme from aligned trajectories on a grid with a resolution of 0.5 Å. The hydrogen bond occupancies were calculated over the 150 ns trajectory considering a 3 Å distance and 1200 angle cutoff.
Although simulations of both enzymes were conducted in the apo, reactant (compound I and substrate) and intermediate (compound II and substrate radical) states, the results of the reactant state are only discussed in the supplemental information. The intermediate state, which is more relevant for the interest of this study as it is the crucial catalytic bifurcation step, is discussed along with the apo state.
Results and Discussion
Sequence and structural comparisons between OleTJE and P450BSβ
OleTJE and P450BSβ share the canonical P450 structural domain and overall fold characterized by 14 α-helices and 6 β-sheets (Fig. 1 Left). The established naming convention for the secondary structure elements was adopted and the helices color coded for the interpretation of RMSF plots in subsequent sections38. The crystal structures share ~41% sequence identity but the overall 3-D structural alignment (Fig. 1 Right) of the two crystal structures yields a backbone RMSD of 1.34 Å across 410 atom pairs. A closer analysis of the sequence alignment (Fig. S1) for major differences between the structurally aligned residues reveals ~14 positions with drastically different charged residues. In this study, a drastically variable residue substitution is defined as a specific position that is occupied by oppositely charged amino acids in the two proteins. The full list of amino acid residues is presented in the supplemental information (Table S1). The differences are predominantly located on the exteriors of the proteins and the P450BSβ surface is characterized by more number of positively charged residues than the OleTJE surface.
Another interesting structural difference is the length of the loop between the F and G helices, with P450BSβ having a F-G loop shorter by about 3 amino acid residues as compared to OleTJE (Fig. 1 right). This difference has been recently highlighted to be an important factor enabling the accommodation of the long C20 fatty acid arachidic acid39. The impact of shortening this loop in OleTJE was evaluated to result in a loss of decarboxylation in favor of hydroxylation40.
Apo state dynamics reveal differences in flexibility and active site channels
The simulations of the 2 enzymes in the apo state reveals significant differences in flexible regions of the protein. While the secondary structure domains are fairly stable, there are significant differences in the looped regions. The loops between H and I helices and residues 340 and 350 are more flexible in OleTJE while loops between β1 and β2 sheets and G and K helices are more flexible in P450BSβ. The F-G loops are flexible in both enzymes with OleTJE F-G loop being slightly more flexible. This could be due to its extended length as compared to P450BSβ.
Reactions catalyzed by P450s involve the turnover of different entities including activators, reactants, active site water molecules and products. This has resulted in a well characterized description in literature of channels leading from the surface of the P450s to the active site with an established convention to identify tunnels based on their positioning relative to the specific secondary structure elements (specific α-helices and β-sheets)31. While this study is the first to discuss the active site tunnels observed in MD simulations of P450BSβ, our observations of tunnels in OleTJE agree with the recently published results of Du et al.16 A comparison reveals mostly similar active site channels between OleTJE and P450BSβ (Fig. 2) in the apo state. The 2e and 2b channels are described as channels that enable access from the surface to the active site between the B-C loop and the B-B’ loop respectively. Two different orientations are observed for the 2e loop in both enzymes (2e1 and 2e2). The 5 channel is also observed in both enzymes with access afforded between the K and K′ helices. A clear difference between the two enzymes is the observation of the solvent tunnel, S, in OleTJE which is absent in P450BSβ. However, P450BSβ is characterized by channel 1 that enables access between the C, H helices and close to the G-H loop.
Apo state dynamics: (Top) Root Mean Square Fluctuations (RMSFs) enable comparisons of flexible regions in P450BSβ and OleTJE. The important structural domains of the protein are highlighted and correspond to the color-coded domains depicted in Fig. 1. (Below: Left and Right) CAVER analysis reveals dominant protein channels connecting the enzyme surface to the active site in P450BSβ (Left) and OleTJE (Right).
Dynamics at the intermediate state (compound II and substrate radical) reveals differences in substrate mobility
A molecular level description of compound II and substrate (radical) dynamics has thus far been elusive and has hindered us from exploring the specific actors that enable catalytic bifurcation. Differences observed in P450BSβ and OleTJE at this crucial catalytic bifurcation stage are likely to play a role in enabling the different characteristic reactivities. A comparison reveals key differences in both flexibility and channels leading to the active site, Fig. 3. The RMSFs reveal differences in flexibility at the loops on either side of the I helix. The I helix is the largest α-helix in both enzymes and straddles from one end of the enzyme across the active site over to the other end. This helix also accommodates the active site arginine (Arg245 in OleTJE and Arg242 in P450BSβ) that is known to be crucial for the catalytic activity in both enzymes. The H-I loop is more flexible in OleTJE while the I-K loop is more flexible in P450BSβ. Other loops in P450BSβ that are slightly more flexible include the F-G loop and the loops closer to the C-terminal.
Enzyme dynamics in the intermediate state (compound II and substrate radical): (Top) Root Mean Square Fluctuations (RMSFs) enable comparisons of flexible regions in P450BSβ and OleTJE. The important structural domains of the protein are highlighted and correspond to the color-coded domains depicted in Fig. 1. (Bottom: Left and Right) CAVER analysis reveals dominant protein channels connecting the enzyme surface to the active site in P450BSβ (Left) and OleTJE (Right).
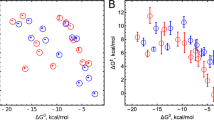
The CAVER analysis indicates a key difference between the enzymes in this substrate bound state. Unlike OleTJE that is characterized by 2e2 active site channel as the sole dominant channel, P450BSβ is characterized by tunnels not only in the B-C loop (2e1 and 2e2) but also along the substrate binding groove (2b and 2d). This is indicative of a more open and accessible substrate binding groove in P450BSβ. This trend of a more accessible substrate binding groove is also evident in the reactant state (Fig. S2), where in OleTJE is characterized by channels 2e2 and 2e1 while P450BSβ enables access to the active site via channels 2a, 2c, 2e1 and 2b. In order to investigate if this more open active site binding groove has any impact on substrate dynamics, the mobility of the substrate was evaluated using Root Mean Square Deviations (RMSDs) and Fluctuations. The average and, more prominently, the standard deviations in RMSDs for the substrate are estimated to be higher in P450BSβ (1.34 ± 0.15 Å) as compared to OleTJE (1.28 ± 0.09 Å). A closer look at the RMSD trend (Fig. 4 Left) throughout the 150 ns trajectory reveals subtle but consistently higher RMSDs throughout the simulation time for P450BSβ. The RMSFs of the oxygen and carbon atoms of the substrate (Fig. 4 Right) also indicate a consistently greater flexibility of the substrate across its length with the tail being significantly more flexible in P450BSβ. These differences in substrate mobility are further exaggerated in the reactant state (Fig. S3) with average RMSD values of 2.14 ± 0.30 Å for P450BSβ and 1.84 ± 0.25 Å for OleTJE and the RMSF of all substrate atoms in P450BSβ significantly higher than in the case of OleTJE.
Substrate dynamics in the intermediate state: (Left) Trends for the Root Mean Square Deviations of the substrate center of mass over the 150 ns simulation trajectory. The darker lines depict the moving average for the RMSD value while the light regions indicate instantaneous values. (Right) Comparison of the flexibility of substrate oxygen and carbon atoms- C1 indicates the carboxylate Carbon and C14 the terminal carbon on the substrate tail. Values for P450BSβ are indicated in red and for OleTJE in black.
Recent reports have alluded to the significance of the substrate mobility at the active site as an important factor in determining the catalytic fate of the substrate in OleTJE. Matthews et al. noted an absence of large scale structural changes in their H85Q and F79A OleTJE mutants, but observed major changes in their product slate suggesting that subtle changes in mobility of the substrates might be important factor determining catalytic activity15. Another experimental study exploring the impact of amino acid mutations to the F-G loop in OleTJE also suggests that coordination of the substrate and its stability at the active site might be important regulatory factors for determination of its catalytic activity40. Our observations of the difference in substrate mobility in P450BSβ as compared to OleTJE corroborate the above hypothesis.
Arginine mobility correlates with substrate mobility
An analysis of the important active site residues that coordinate with the substrate reveals the active site arginine (Arg242 in P450BSβ and Arg245 in OleTJE) to be an important contributor to substrate binding with strong H-bonds with the carboxylate group on the substrate. Considering the differences in flexibility of the loops adjoining the I-helix containing this arginine residue, we evaluated if the difference in substrate mobility could be attributed to the mobility of this residue. Figures 5 and S4 illustrate a dynamic cross correlation analysis between atoms of the arginine, compound II (compound I in S4) and the substrate carboxylate atoms. Positive correlation coefficients obtained from this analysis indicate the movements of a subset of atoms in the system correlates well with the movement of the other subset of atoms41. As is evident in both P450BSβ and OleTJE, the movement in substrate carboxylate atoms correlates very well with the arginine nitrogen atoms. Correlation coefficients close to zero for the Heme atoms indicate that its movement with the substrate is not correlated.
Dynamic cross-correlation analysis indicates substrate correlation with the Arginine residue. The H-bonding nitrogen atoms in Arg242/245, the carboxylate atoms on the substrate and the iron-hydroxo atoms on compound II are considered. Positively correlated motions are shown in red while bluish regions indicate no correlation.
Enzyme-substrate interactions mediated by specific active-site residue contacts can also limit substrate mobility. The list of residues within close contact of the substrate was compiled for each enzyme over the 150 ns trajectory. The most important contributors to substrate binding are listed in Fig. 6 with Arginine being the highest contributor. It is also interesting to note that the arginine in OleTJE binds the substrate better than in P450BSβ. This is further supported by hydrogen bond occupancies for the arginine residue in the two enzymes. The arginine is observed to coordinate with the two carboxylate oxygens for 48% and 35% of the simulation time in OleTJE as compared to 28% and 23% in P450BSβ. Another notable difference in the coordination of the arginine residue is the presence of a methionine residue in OleTJE which coordinates with the arginine backbone NH via its backbone C=O. This is observed to bind ~53% of the time as compared to the corresponding neighbor in P450BSβ that is observed to coordinate ~34% of the time. While the lower RMSDs for arginine in OleTJE (0.26 ± 0.07 Å) when compared to P450BSβ (0.20 ± 0.04 Å) might be responsible for limiting substrate mobility, the stronger coordination of the arginine with the substrate carboxylate atoms also suggest of an increased capacity to effect decarboxylation in OleTJE.
A closer look at the active site residues aiding substrate binding (Fig. 7) and their contributions reveals an important difference that could explain the increased flexibility of the substrate tail in P450BSβ. The substrate tail is stabilized by Pro293 and Pro296 in OleTJE on either side while in P450BSβ the substrate tail is stabilized by Pro291 and Phe292 both of which are present only on one side substrate. Furthermore, Pro296 in OleTJE is substituted by Gly294 in P450BSβ which does not contribute to binding. All of these factors result in a much less favorable total MMGBSA binding energy for the myristiate in P450BSβ (−46.6 ± 4 kcal/mol) as compared to OleTJE (−53.7 ± 3.1 kcal/mol).
Residue-wise binding energy decomposition for P450BSβ (Left) and OleTJE (Right). The values indicate binding energy contributions and standard deviations calculated over the 150 ns trajectory in kcal/mol. Only residues contributing >0.5 kcal/mol are considered. All contributing residues are in conserved positions except for Phe287 in P450BSβ and Pro293 in OleTJE.
Water occupancies suggest heme occlusion in the OleTJE active site
Water molecules are known to play a functional role in cytochrome P450s both directly by impacting catalysis as a proton transport enabler and in more subtle ways by impacting protein structure and substrate binding42,43,44,45,46. Fig. 8 compares water occupancies observed in our 150 ns MD trajectories of OleTJE and P450BSβ in the apo, reactant (compound I and substrate) and intermediate (compound II and substrate radical) states. It is evident that the region surrounding the heme oxygen atom is characterized by greater water occupancy in OleTJE as compared to P450BSβ. The difference is particularly stark in substrate bound (reactant and intermediate) states where in the region between the His85 and the Heme oxygen is occupied by at least 2 water molecules in OleTJE as compared to an occasional occupation of a water molecule in P450BSβ. These observations are consistent with Falpone et al.’s wherein they consider the presence of three water molecules at the OleTJE active site in their QM/MM calculations evaluating the decarboxylation mechanism10. While they did not assign a catalytic role for these water molecules at the active site, their sustained presence at the active site might suggest their role in occluding the rebound mechanism in OleTJE.
Active site water occupancies: Volumetric occupancies of water calculated within 10 Å of heme in various states-apo, compound I and compound II-are depicted as wire frame structures in P450BSβ (red) and OleTJE(green). The wireframe volumes signify atleast a 75% occupancy of water calculated using 15000 frames over 150 ns trajectories in each state. The arrows point to key water occupancies that might occlude substrate rebound onto the heme oxygen.
One of the apparent active site differences between P450BSβ and OleTJE has been the presence of a Glutamine in the former and a Histidine in the latter leading to initial assignment of a catalytic role for His85 as a proton donor in OleTJE4. The observation of decarboxylation activity on the H85Q mutant eliminated its role as the primary determinant of catalytic fate15. From our observations, it is apparent that His85 might play a more subtle role in enabling decarboxylation. In the intermediate state, the active site histidine seems to coordinate the waters close to compound II’s hydroxo group, which plausibly form a shield to obstruct substrate access, thereby enabling the substrate to evade rebound. However, it must be noted that these water molecules are not solely coordinated by His85 and are also stabilized by the active site arginine along with other residues and hence the Q85H mutant in P450BSβ and H85Q mutant in OleTJE don’t result in absolute reversal of their characteristic wild-type reactivities.
Conclusions
Hydrocarbons have energized mankind’s rapid development and fueled our technological advancement for the past century. While they shall continue to do so for the near future, their lack of sustainability is driving us towards developing more sustainable alternatives. Biological routes to producing our fuels have been a significant focus area for research towards this goal and have resulted in the nascent commercialization of technologies for the production of alternative oxygenated fuels such as ethanol and fatty acid methyl esters. However, the biological production of hydrocarbons has thus far been elusive. The P450 OleTJE enzyme, that bears significant similarities with the P450 hydroxylase, presents a promising platform for the direct biological production of hydrocarbons from fatty acids. Recent enzyme engineering efforts to improve OleTJE performance have mostly resulted in shifting OleTJE activity from decarboxylation towards hydroxylation leading to the current hypothesis that OleTJE’s decarboxylation activity is a fragile reaction that seems to predominantly occur by the evasion of oxygen rebound, which is the dominant route to hydroxylation14,15,40.
In this study, we elucidate differences in the dynamics of wild-type P450BSβ and OleTJE during different stages of the mechanism leading up to the bifurcation of the catalytic pathways in the crucial intermediate state with (compound II and the substrate radical). We develop point charges to enable molecular simulations and provide parameters for the community to perform compound II simulations. A comparison of the apo-states reveals differing flexible regions in the two enzymes which might be effected by the charge variable substitutions observed in the structural comparison. Active site channels reveal mostly similar channels in both enzymes except for the rare 1 channel in P450BSβ and the S (solvent) channel in OleTJE. Simulations in the intermediate state highlight differences in substrate mobility and active site water occupancies between the two enzymes and provides the first evidence of their plausible role in determining the catalytic fate of the substrate. This corroborates proposed experimental hypotheses that attribute OleTJE’s hydroxylating activity to increased mobility at the active site39,40. Our simulations also demonstrate the importance of the active site arginine in not only limiting substrate mobility in OleTJE but might also be crucial in the increased capacity to decarboxylate the substrate. Furthermore, these insights also point towards future directions for enzyme engineering strategies to improve OleTJE performance will likely involve controlling substrate mobility at the active site and effect it by controlling water occupancies and substrate stabilizing enzyme interactions.
Data availability
The parameter (.frcmod and.prep) files for compound I and compound II are provided as text within the supplemental information included along with the manuscript. The data that support the findings of this study are available from the corresponding author on reasonable request.
References
Meunier, B., de Visser, S. P. & Shaik, S. Mechanism of Oxidation Reactions Catalyzed by Cytochrome P450 Enzymes. Chem. Rev. 104, 3947–3980, https://doi.org/10.1021/cr020443g (2004).
Rude, M. A. et al. Terminal Olefin (1-Alkene) Biosynthesis by a Novel P450 Fatty Acid Decarboxylase from Jeotgalicoccus Species. Appl. Environ. Microbiol. 77, 1718–1727, https://doi.org/10.1128/AEM.02580-10 (2011).
Choi, Y. J. & Lee, S. Y. Microbial production of short-chain alkanes. Nature 502, 571, https://doi.org/10.1038/nature12536 (2013).
Belcher, J. et al. Structure and Biochemical Properties of the Alkene Producing Cytochrome P450 OleTJE (CYP152L1) from the Jeotgalicoccus sp. 8456 Bacterium. J. Biol. Chem. 289, 6535–6550, https://doi.org/10.1074/jbc.M113.527325 (2014).
Fujishiro, T. et al. Crystal Structure of H2O2-dependent Cytochrome P450SPα with Its Bound Fatty Acid Substrate Insight into the Regioselective Hydroxylation of Fatty Acids at the α Position. J. Biol. Chem. 286, 29941–29950, https://doi.org/10.1074/jbc.M111.245225 (2011).
Matsunaga, I., Ueda, A., Fujiwara, N., Sumimoto, T. & Ichihara, K. Characterization of the ybdT gene product of Bacillus subtilis: Novel fatty acid β-hydroxylating cytochrome P450. Lipids 34, 841–846, https://doi.org/10.1007/s11745-999-0431-3 (1999).
Grant, J. L., Hsieh, C. H. & Makris, T. M. Decarboxylation of Fatty Acids to Terminal Alkenes by Cytochrome P450 Compound I. J. Am. Chem. Soc. 137, 4940–4943, https://doi.org/10.1021/jacs.5b01965 (2015).
Grant, J. L., Mitchell, M. E. & Makris, T. M. Catalytic strategy for carbon−carbon bond scission by the cytochrome P450 OleT. PNAS 113, 10049–10054, https://doi.org/10.1073/pnas.1606294113 (2016).
Ji, L. et al. Drug Metabolism by Cytochrome P450 Enzymes: What Distinguishes the Pathways Leading to Substrate Hydroxylation Over Desaturation? Chem. Eur. J. 21, 9083–9092, https://doi.org/10.1002/chem.201500329 (2015).
Faponle, A. S., Quesne, M. G. & de Visser, S. P. Origin of the Regioselective Fatty-Acid Hydroxylation versus Decarboxylation by a Cytochrome P450 Peroxygenase: What Drives the Reaction to Biofuel Production? Chem. Eur. J. 22, 5478–5483, https://doi.org/10.1002/chem.201600739 (2016).
Wang, J.-b, Lonsdale, R. & Reetz, T. M. Exploring substrate scope and stereoselectivity of P450 peroxygenase OleT JE in olefin-forming oxidative decarboxylation. Chemical Communications 52, 8131–8133, https://doi.org/10.1039/C6CC04345C (2016).
Fang, B. et al. Mutagenesis and redox partners analysis of the P450 fatty acid decarboxylase OleTJE. Scientific Reports 7, https://doi.org/10.1038/srep44258 (2017).
Dennig, A. et al. Oxidative Decarboxylation of Short-Chain Fatty Acids to 1-Alkenes. Angew. Chem. Int. Ed. 54, 8819–8822, https://doi.org/10.1002/anie.201502925 (2015).
Hsieh, C. H. et al. The Enigmatic P450 Decarboxylase OleT Is Capable of, but Evolved To Frustrate, Oxygen Rebound Chemistry. Biochemistry 56, 3347–3357, https://doi.org/10.1021/acs.biochem.7b00338 (2017).
Matthews, S. et al. Catalytic Determinants of Alkene Production by the Cytochrome P450 Peroxygenase OleTJE. J. Biol. Chem. 292, 5128–5143, https://doi.org/10.1074/jbc.M116.762336 (2017).
Du, J., Liu, L., Guo, L. Z., Yao, X. J. & Yang, J. M. Molecular basis of P450 OleTJE: an investigation of substrate binding mechanism and major pathways. J Comput Aided Mol Des 31, 483–495, https://doi.org/10.1007/s10822-017-0013-x (2017).
Liu, Y. et al. Hydrogen peroxide-independent production of α-alkenes by OleTJE P450 fatty acid decarboxylase. Biotechnology for Biofuels 7, 28, https://doi.org/10.1186/1754-6834-7-28 (2014).
Lee, D.-S. et al. Substrate Recognition and Molecular Mechanism of Fatty Acid Hydroxylation by Cytochrome P450 from Bacillus subtilis Crystallographic, Spectroscopic, and Mutational Studies. J. Biol. Chem. 278, 9761–9767, https://doi.org/10.1074/jbc.M211575200 (2003).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25, 1605–1612, https://doi.org/10.1002/jcc.20084 (2004).
Case, D. A. et al. Amber 16 & AmberTools17. AMBER 2017, University of California, San Francisco (2017).
Shahrokh, K., Orendt, A., Yost, G. S. & Cheatham, T. E. Quantum mechanically derived AMBER-compatible heme parameters for various states of the cytochrome P450 catalytic cycle. J Comput Chem 33, 119–133, https://doi.org/10.1002/jcc.21922 (2012).
Becke, A. D. Density-functional thermochemistry. III. The role of exact exchange. The Journal of Chemical Physics 98, 5648–5652 (1993).
Dunning, T. H. & Hay, P. J. In Methods of Electronic Structure Theory Modern Theoretical Chemistry 1–27 (Springer, Boston, MA, 1977).
Dupradeau, F.-Y. et al. R.E.DD.B.: A database for RESP and ESP atomic charges, and force field libraries. Nucleic Acids Res 36, D360–D367, https://doi.org/10.1093/nar/gkm887 (2008).
Maier, J. A. et al. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713, https://doi.org/10.1021/acs.jctc.5b00255 (2015).
Wang, J. M., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general amber force field. Journal of Computational Chemistry 25, 1157–1174, https://doi.org/10.1002/jcc.20035 (2004).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. The Journal of Chemical Physics 79, 926–935, https://doi.org/10.1063/1.445869 (1983).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. Journal of Molecular Graphics and Modelling 25, 247–260, https://doi.org/10.1016/j.jmgm.2005.12.005 (2006).
Anandakrishnan, R., Aguilar, B. & Onufriev, A. V. H+ +3.0: automating pK prediction and the preparation of biomolecular structures for atomistic molecular modeling and simulations. Nucleic Acids Res 40, W537–W541, https://doi.org/10.1093/nar/gks375 (2012).
Ryckaert, J.-P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. Journal of Computational Physics 23, 327–341, https://doi.org/10.1016/0021-9991(77)90098-5 (1977).
Cojocaru, V., Winn, P. J. & Wade, R. C. The ins and outs of cytochrome P450s. Biochimica et Biophysica Acta (BBA) - General Subjects 1770, 390–401, https://doi.org/10.1016/j.bbagen.2006.07.005 (2007).
Chovancova, E. et al. CAVER 3.0: A Tool for the Analysis of Transport Pathways in Dynamic Protein Structures. PLOS Computational Biology 8, e1002708, https://doi.org/10.1371/journal.pcbi.1002708 (2012).
Miller, B. R. et al. MMPBSA.py: An Efficient Program for End-State Free Energy Calculations. J. Chem. Theory Comput. 8, 3314–3321, https://doi.org/10.1021/ct300418h (2012).
Bharadwaj, V. S., Dean, A. M. & Maupin, C. M. Insights into the Glycyl Radical Enzyme Active Site of Benzylsuccinate Synthase: A Computational Study. J. Am. Chem. Soc. 135, 12279–12288, https://doi.org/10.1021/ja404842r (2013).
Schutt, T. C., Bharadwaj, V. S., Granum, D. M. & Mark Maupin, C. The impact of active site protonation on substrate ring conformation in Melanocarpus albomyces cellobiohydrolase Cel7B. Physical Chemistry Chemical Physics 17, 16947–16958, https://doi.org/10.1039/C5CP01801C (2015).
Roe, D. R. & Cheatham, T. E. PTRAJ and CPPTRAJ: Software for Processing and Analysis of Molecular Dynamics Trajectory Data. J. Chem. Theory Comput. 9, 3084–3095, https://doi.org/10.1021/ct400341p (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. Journal of Molecular Graphics 14, 33–38, https://doi.org/10.1016/0263-7855(96)00018-5 (1996).
Hasemann, C. A., Kurumbail, R. G., Boddupalli, S. S., Peterson, J. A. & Deisenhofer, J. Structure and function of cytochromes P450:a comparative analysis of three crystal structures. Structure 3, 41–62, https://doi.org/10.1016/S0969-2126(01)00134-4 (1995).
Munro, A. W., McLean, K. J., Grant, J. L. & Makris, T. M. Structure and function of the cytochrome P450 peroxygenase enzymes. Biochemical Society Transactions 46, 183–196, https://doi.org/10.1042/BST20170218 (2018).
Amaya, J. A., Rutland, C. D., Leschinsky, N. & Makris, T. M. A Distal Loop Controls Product Release and Chemo- and Regioselectivity in Cytochrome P450 Decarboxylases. Biochemistry 57, 344–353, https://doi.org/10.1021/acs.biochem.7b01065 (2018).
Hünenberger, P. H., Mark, A. E. & van Gunsteren, W. F. Fluctuation and cross-correlation analysis of protein motions observed in nanosecond molecular dynamics simulations. J. Mol. Biol. 252, 492–503, https://doi.org/10.1006/jmbi.1995.0514 (1995).
Rydberg, P., Rod, T. H., Olsen, L. & Ryde, U. Dynamics of Water Molecules in the Active-Site Cavity of Human Cytochromes P450. J. Phys. Chem. B 111, 5445–5457, https://doi.org/10.1021/jp070390c (2007).
Fishelovitch, D., Shaik, S., Wolfson, H. J. & Nussinov, R. How Does the Reductase Help To Regulate the Catalytic Cycle of Cytochrome P450 3A4 Using the Conserved Water Channel? J. Phys. Chem. B 114, 5964–5970, https://doi.org/10.1021/jp101894k (2010).
Oprea, T. I., Hummer, G. & García, A. E. Identification of a functional water channel in cytochrome P450 enzymes. PNAS 94, 2133–2138, https://doi.org/10.1073/pnas.94.6.2133 (1997).
Madrona, Y., Hollingsworth, S. A., Khan, B. & Poulos, T. L. P450cin Active Site Water: Implications for Substrate Binding and Solvent Accessibility. Biochemistry 52, 5039–5050, https://doi.org/10.1021/bi4006946 (2013).
Sen, K. & Thiel, W. Role of Two Alternate Water Networks in Compound I Formation in P450eryF. J. Phys. Chem. B 118, 2810–2820, https://doi.org/10.1021/jp411272h (2014).
Acknowledgements
This work was authored in part by Alliance for Sustainable Energy, LLC, the manager and operator of the National Renewable Energy Laboratory for the U.S. Department of Energy (DOE) under Contract No. DE-AC36-08GO28308. Funding provided by the U.S. Department of Energy Bioenergy Technologies Office of under award number No. DE-AC36-08GO28308 and by the National Renewable Energy Laboratory’s (NREL) Laboratory Directed Research and Development program. We are also grateful for the access and use of NREL’s Computational Sciences’ resources (Peregrine) supported by the DOE Office of EERE under contract number DE-AC36-08GO28308. The views expressed in the article do not necessarily represent the views of the DOE or the U.S. Government. The U.S. Government retains and the publisher, by accepting the article for publication, acknowledges that the U.S. Government retains a nonexclusive, paid-up, irrevocable, worldwide license to publish or reproduce the published form of this work, or allow others to do so, for U.S. Government purposes.
Author information
Authors and Affiliations
Contributions
V.S.B. conceived of the presented idea. S.K. performed QM & RESP calculations for point charge calculations. V.S.B. set-up, performed and analyzed the MD simulations, wrote the manuscript and prepared the figures. M.F.C. verified the methods and analysis. All authors including M.T.G. provided critical feedback and helped shape the research, analysis and manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bharadwaj, V.S., Kim, S., Guarnieri, M.T. et al. Different Behaviors of a Substrate in P450 Decarboxylase and Hydroxylase Reveal Reactivity-Enabling Actors. Sci Rep 8, 12826 (2018). https://doi.org/10.1038/s41598-018-31237-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31237-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.