Abstract
Mechanisms of injury and repair in alveolar epithelial cells (AECs) are critically involved in the progression of various lung diseases including idiopathic pulmonary fibrosis (IPF). Homeobox only protein x (HOPX) contributes to the formation of distal lung during development. In adult lung, alveolar epithelial type (AT) I cells express HOPX and lineage-labeled Hopx+ cells give rise to both ATI and ATII cells after pneumonectomy. However, the cell function of HOPX-expressing cells in adult fibrotic lung diseases has not been investigated. In this study, we have established a flow cytometry-based method to evaluate HOPX-expressing cells in the lung. HOPX expression in cultured ATII cells increased over culture time, which was accompanied by a decrease of proSP-C, an ATII marker. Moreover, HOPX expression was increased in AECs from bleomycin-instilled mouse lungs in vivo. Small interfering RNA-based knockdown of Hopx resulted in suppressing ATII-ATI trans-differentiation and activating cellular proliferation in vitro. In IPF lungs, HOPX expression was decreased in whole lungs and significantly correlated to a decline in lung function and progression of IPF. In conclusion, HOPX is upregulated during early alveolar injury and repair process in the lung. Decreased HOPX expression might contribute to failed regenerative processes in end-stage IPF lungs.
Similar content being viewed by others
Introduction
The lung is a complex organ which is directly exposed to numerous external stimuli, such as cigarette smoke, pathogens, air pollution, or particles from the environment1. The distal lung is composed of alveoli which are covered by alveolar epithelial type (AT) I cells and ATII cells. ATI cells are squamous, large, and flat cells, which cover 95% of the internal surface area of the alveoli2. ATI cells play an indispensable role for gas exchange and alveolar-capillary barrier function2. Thus, it is critically important to preserve and regenerate ATI cells for proper respiratory function. ATII cells maintain alveolar homeostasis as a “caretaker” of alveoli3 to produce and secrete surfactant proteins4, to keep the fluid balance of the alveoli5, and to serve as progenitor cells for ATI/ATII cells during lung turnover and repair6. Mechanisms of alveolar epithelial injury and repair have been proposed to be critically involved in the progression of various lung diseases, including idiopathic pulmonary fibrosis (IPF), a fatal interstitial lung disease1.
Homeobox only protein x (HOPX) is a Homeobox protein important for anterior-posterior patterning during organ development7,8. In the developing lung, HOPX contributes to the formation of distal lung9. In adult murine lung, ATI cells express HOPX10 and lineage-labeled Hopx+ cells, which are further positive for ATI cell markers, give rise to both ATI and ATII cells after pneumonectomy11. Thus, HOPX might be involved in the bidirectional trans-differentiation of ATII to ATI or ATI to ATII during adult alveolar repair. It is also known that HOPX-expressing cells serve as stem cells in intestinal epithelium12 or hair follicle13. Furthermore, HOPX is further described as a tumor suppressor gene in the lung that regulates cellular proliferation and metastasis14,15. However, the function of HOPX-expressing cells in the adult fibrotic lung has not yet been clarified. In the present study, we aimed to gain a detailed insight into the dynamics of HOPX-expressing alveolar epithelial cells (AECs) during lung injury and repair and established a flow cytometry (FCM)-based method to quantify the HOPX-expressing cells in AECs. We then investigated HOPX expression in an ATII-ATI trans-differentiation culture model in vitro, in bleomycin-induced mouse model of pulmonary fibrosis in vivo, and by database analyses of expression profiles of IPF lungs in silico.
Results
ProSP-C and HOPX expression during ATII-ATI trans-differentiation
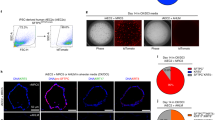
To investigate the potential role of HOPX in adult lung injury/repair, we first investigated the changes of HOPX expression in primary mouse (pm) ATII cells (95+/−3% of proSP-C expression) from wildtype mice during 2D-culture in vitro. This in vitro culture system is well-established to study trans-differentiation of ATII cells into ATI-like cells16,17,18, a process known to be implicated in lung repair. Hopx expression was increased (Fig. 1A) over 5 days of culture along with decreased Sftpc expression, as evaluated by qRT-PCR (Fig. 1B) and FCM (Fig. 1C and D, mean fluorescent intensity (MFI) for Fig. 1E), which is consistent with previous reports10,16. Moreover, after an initial increase, the proliferation marker Mki67 was decreased during the time course (Fig. 1F). We next optimized the FCM-based quantification of HOPX expression in pmATII cells using the same trans-differentiation system (Fig. 1G,H). HOPX expression was gradually increased during the culture (Fig. 1H). In addition, we evaluated the co-expression of proSP-C and HOPX within freshly isolated pmATII cells over culture time (Fig. 1I). We found that HOPX+/proSP-C+ cells as well as HOPX+/proSP-C− cells were increased while HOPX−/proSP-C+ cells were decreased during trans-differentiation (Fig. 1J).
Expression of HOPX and proSP-C during ATII-ATI cell trans-differentiation in vitro. Quantitative (q) RT-PCR analysis of (A) Hopx (B) Sftpc, (C,D) FCM analysis of proSP-C expression over culture, (E) mean fluorescent intensity (MFI) of proSP-C-Alexa-fluor 488, and (F) qRT-PCR analysis of Mki67 during 5 days culture of pmATII cells (n = 3). (G,H) FCM-based quantification of HOPX expression during the culture. (I) and (J) FCM-based quantification of HOPX/proSP-C co-expression during the culture (n = 3). *p < 0.05, **p < 0.01, ***p < 0.005.
HOPX expression was increased in the alveolar epithelium in bleomycin (BLM)-instilled lungs
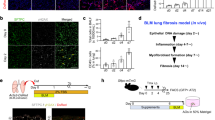
Disturbed ATII to ATI cell trans-differentiation has been linked to lung fibrosis16,17. Thus, we next sought to investigate the expression changes of HOPX in fibrotic lung diseases using the FCM analysis. We evaluated Hopx expression in the mouse model of pulmonary fibrosis induced by intra-tracheal BLM instillation. We isolated ATII cells from phosphate buffered saline (PBS)-instilled lungs (PBS-pmATII cells) as a control group and BLM-instilled mouse lungs (BLM-pmATII cells) after 14 days of the initial PBS/BLM instillation. We found that mRNA expression of Hopx (Fig. 2A) and T1α (Fig. 2B) was significantly upregulated whereas Sftpc (Fig. 2C) was significantly downregulated in BLM-pmATII cells compared with PBS-pmATII cells. Next, we evaluated the expression of proSP-C and HOPX by FCM (Fig. 2D). Quantification of the FCM analysis revealed a significant decrease of HOPX−/proSP-C+ cells while HOPX+/proSP-C+ cells were significantly increased in BLM-pmATII cells compared to in PBS-pmATII cells (Fig. 2E). Importantly, further immunofluorescence (IF) also revealed that HOPX expression was increased in BLM-instilled lungs compared with PBS-instilled lungs in situ (Fig. 2F and G). The cells which co-expressed both HOPX and proSP-C were increased in BLM-lungs (Fig. 2G, shown in white arrows, and Fig. 2I, shown in green dots) compared to PBS-lungs (Fig. 2F and H). The co-expression of proSP-C and HOPX was also confirmed by IF of cytospun pmATIIs which were freshly isolated from PBS- (Fig. 2J) or BLM-instilled lungs (Fig. 2K).
Expression of HOPX/proSP-C lung epithelial cell subpopulations in BLM-induced pulmonary fibrosis model in vivo. Quantitative RT-PCR analysis of (A) Hopx (B) T1α, and (C) Sftpc in EpCAM+ cells from PBS or BLM-instilled lungs (n = 4). (D) FCM-based evaluation of HOPX/proSP-C expression in pmATII cells from PBS or BLM-instilled lungs (representative images from n = 3). Quantification of (E) HOPX−/proSP-C+, HOPX+/proSP-C−, and HOPX+/proSP-C+ cells in PBS and BLM-instilled lungs (n = 3). Immunofluorescence staining (IF) of HOPX (white) and proSP-C (red) in the sections from (F) PBS and (G) BLM-instilled lungs. Visualization of the co-expression of HOPX and proSP-C from (H) PBS and (I) BLM-instilled lungs using ZEN2009 software. IF of HOPX (white) and proSP-C (red) in cytospun EpCAM+ cells from (J) PBS and (K) BLM-instilled lungs. *p < 0.05, **p < 0.01, ***p < 0.005.
HOPX knockdown activated cellular proliferation
To investigate whether HOPX is involved in the proliferation of lung epithelial cells, we silenced Hopx by siRNA in the murine alveolar epithelial cell line MLE12, which endogenously expresses HOPX. We found that HOPX/Hopx expression was efficiently reduced in the cells as assessed by qRT-PCR (Fig. 3A) and western blotting (Fig. 3B, quantification in Fig. 3C). Next, we performed an EdU-based proliferation assay using the siRNA-transfected MLE12 cells with co-staining of HOPX by FCM. We found that Hopx knockdown (siHopx) increased the HOPX−/EdU+ population while reducing the HOPX+/EdU+ population (Fig. 3D,E). Further, we observed an increase in proliferation using a scratch assay in siHopx-transfected MLE12 cells (Fig. 3F,G). In support of these observations, we found that the knockdown of Hopx significantly increased Mki67 expression as well as ATII marker Sftpc (Fig. 3H and I), and increased net metabolic activity as assessed by WST1 assay (Fig. 3J). In addition, we evaluated Ki67+ cells with and without HOPX in the BLM-instilled lung by IF. The number of Ki67+ cells within the HOPX+ fraction was significantly lower than that within the HOPX- fraction (Fig. 3K). Altogether, these results suggest that HOPX is involved in suppression of AEC proliferation.
Effect of HOPX on proliferation and differentiation in MLE12 epithelial cells in vitro. Evaluation of si-RNA-based Hopx knockdown in MLE12 lung epithelial cells with (A) qRT-PCR of Hopx, (B) Western blotting of HOPX, (C) quantification of Western blotting data. (D) EdU assay of MLE12 lung epithelial cells co-stained with HOPX antibody in vitro. Alexa Fluor 647-EdU+ cells and Alexa Fluor 488-HOPX+ cells were evaluated by FCM (Representative image of n = 2). (E) Quantification of HOPX+ EdU+ cells and HOPX-EdU+ cells in MLE12 cells (n = 2). (F,G) Cellular scratch assay, qRT-PCR analysis of (H) Mki67 and (I) Sftpc, and (J) WST assay of MLE12 cells transfected with control siRNA and Hopx siRNA. (J) The ratio of Ki67 positive/negative cells with HOPX co-expression in IF. *p < 0.05, **p < 0.01, ***p < 0.005.
HOPX expression in IPF lungs
Thus far, we have shown that HOPX contributes to adult lung injury/repair process in mouse lungs. Next, we sought to clarify whether HOPX might act as a modulator of lung injury/repair in human lungs. To this end, we first analyzed microarray datasets in silico to investigate the expression changes of HOPX in IPF lungs. In contrast to the mouse data, we found that HOPX expression was significantly decreased in whole lung homogenate from IPF lungs compared with control lungs (Fig. 4A) and the lung function parameter DLCO was significantly correlated with HOPX expression (Fig. 4B), as indicated in the dataset from Lung Genomics Research Consortium (LGRC) (Gene Expression Omnibus Series (GSE) accession number: 47460; National Center for Biotechnology Information, Bethesda, MD, USA; https:// www. ncbi. nlm. nih. gov/ geo/). As shown in Fig. 4C by the expression of matrix metalloprotease (MMP7), a biomarker for progression of IPF19, HOPX expression was significantly lower in later stage than in earlier stage. Next, we confirmed decreased expression of HOPX (Fig. 4D) and SFTPC (Fig. 4E) transcripts in our own cohort (lung tissue from IPF explant and Donor lungs), while ACTA2, a mesenchymal marker, was increased (Fig. 4F).
Expression and localization of HOPX in control (donor) and IPF lungs. In silico analysis from Lung Genomics Research Consortium (LGRC) of (A) HOPX mRNA expression in organ donor (control) and IPF lungs, (B) correlation between HOPX mRNA expression and DLco (blue dots: control, red dots: IPF). (C) Correlation between HOPX mRNA and MMP7 mRNA in control (blue dots) and IPF (red dots) in silico. QRT-PCR analysis of (D) HOPX mRNA, (E) SFTPC mRNA, and (F) ACTA2 mRNA from whole lung homogenate of control and IPF lungs in our cohort. (G) Immunohistochemical staining of serial sections of control and IPF lung tissue for KRT5, proSP-C, HOPX, KRT7, and Ki67. In IPF lungs, proSP-C+/KRT7+ cells revealed cytoplasmic or in part nuclear expression of HOPX without nuclear Ki67 (Indicated by arrows in panels 1–4). ProSP-Clow/KRT7+ AECs of IPF-lungs also indicated robust HOPX-immunoreactivity with nuclear Ki67 expression (asterisks in panel 2 and 4). Some proSP-C+/KRT7+ cells without HOPX immunoreactivity were observed in the area of relatively preserved alveolar septa (white arrowheads in panel 5). KRT5+/KRT7+ bronchiolar basal cells did not express HOPX, and indicated nuclear expression of Ki67 (Indicated by black arrowheads in panel 2). (H) Immunohistochemical staining of serial sections of control donor lung tissues for KRT5, proSP-C, HOPX, KRT7, and Ki67. In donor lungs, HOPX expression was present in proSP-C+/KRT7+ ATII (arrows) and proSP-Clow/KRT7+ AECs (asterisks) (panels 2 and 3 in Fig. 4O), and faint in normal bronchiolar epithelium. ProSP-Clow/KRT7+ AECs expressing HOPX also revealed nuclear Ki67 expression (indicated by asterisks). In silico single cell RNA sequence analysis of (I) HOPX, (J) SFTPC, (K) KRT7, (L) CTNNB1, and (M) CTND1 in EpCAM+/HTII280+ cells. *p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.0001.
Next, we stained serial lung tissue sections of organ donor and IPF patients’ lungs with cytokeratin 5 (KRT5, a marker for airway basal cells), proSP-C, HOPX, and cytokeratin 7 (KRT7, a marker for general epithelia), and Ki67, to show the expression pattern of HOPX and proSP-C as well as their proliferation status in IPF lungs (Fig. 4G and H). In IPF, KRT7+ cells are more abundant than in control lungs as previously shown in the study from our laboratory20. We observed that proSP-C+/KRT7+ cells of IPF lungs frequently exhibited an intense expression of HOPX without nuclear Ki67 expression (arrows in panel 1–4 in Fig. 4G). Whereas proSP-C+/KRT7+ cells without HOPX positivity were often observed within preserved alveolar septae (white arrowheads in panel 5, Fig. 4G), proSP-C+/KRT7+ cells within fibrotic regions were mostly HOPX positive (black arrows, Fig. 4G).
In addition, proSP-Clow/KRT7+ AECs of IPF lungs also exhibited robust HOPX immunoreactivity with nuclear Ki67 expression (asterisks in panels 2 and 4 in Fig. 4G). Finally, KRT5+/KRT7+ basal cells in abnormal bronchiolar structures of IPF lungs did not express HOPX, and exhibited strong nuclear expression of Ki67 (panel 2 in Fig. 4G, indicated by black arrowheads). The data suggest that in the fibrotic area, there are HOPX+/proSP-C+/KRT7+ cells, HOPX+/proSP-Clow/−/KRT7+ cells, and HOPX−/proSP-C+/KRT7+ cells as observed in the BLM-instilled mouse lungs. Intriguingly, HOPX+/proSP-C+/KRT7+ cells and HOPX+/proSP-Clow/−/KRT7+ cells were less in the donor lungs and preserved alveolar septa, compared to fibrotic area, as observed in the BLM lungs. In addition, there are few HOPX−/proSP-C+/KRT7+ cells in the fibrotic area.
To further corroborate our immunohistochemistry study, we analyzed a recently published single cell RNA sequence analysis of EpCAM+/HTII280+ cells from control and IPF patients’ lungs21. Importantly, EpCAM+/HTII280+ cells from IPF patients exhibited decreased HOPX (Fig. 4I) and SFTPC expression (Fig. 4J), and increased expression of KRT7 (Fig. 4K). Furthermore, CTNNB1 (Fig. 4L) and CCND1 (Fig. 4M) were significantly upregulated in EpCAM+/HTII280+ cells from IPF lungs. These data further support the idea that decreased HOPX expression in IPF contributes to failure of AEC regeneration and excessive/aberrant epithelial proliferation.
Discussion
Temporal and spatial heterogeneity is one of the histological characteristics of IPF. Several stages of IPF, including initiation, progression, and the terminal stage of the disease, exist at the same time in the same lung. This heterogeneity suggests an ongoing dynamic process with different stages of alveolar injury, repair, and failed regeneration. There are various reported biomarkers of IPF22. Of these, serum levels of surfactant protein A (SP-A) and D (SP-D), which are synthesized and secreted by ATII, have been reported to predict mortality23, exacerbation24, or progression25 of IPF. The increase of these ATII-associated proteins may reflect alveolar dysfunction induced by alveolar injury processes in IPF. On the other hand, MMP7 and MMP1, which degrade extracellular matrix, are increased in plasma from IPF patients. The increase of MMPs may reflect the excessive tissue repair and remodeling22,26. We showed that HOPX is increased in relatively early injury and decreases along with the progression of the disease. In the present study, HOPX is shown to be upregulated in pmATII cells in BLM model of pulmonary fibrosis in vivo. In contrast, we found that HOPX mRNA was decreased in whole lung homogenate and human EpCAM+/HTII280+ cells from end-stage IPF lungs. These results may be explained by several potential areas which warrant further study. BLM lung injury is known to not fully recapitulate all aspects of clinical IPF and may represent a relatively “early” time point of regeneration in alveolar epithelium after injury whereas analyzed IPF patient samples derived from explants represent rather later disease stages. It is well-known that animals can recover and regenerate in the BLM model of fibrosis when the level of injury is not too severe. Our data allow the hypothesis that at the initial stage of regeneration, expression of HOPX is still low to allow ATII cells to undergo self-renewal, whereas at the later stage, HOPX-expressing cells are increased to suppress the aberrant/excessive proliferation and remodeling of alveoli. Thus, it is entirely plausible that a HOPX-expressing population survived the initial injury and participates in epithelial regeneration. It will be interesting in future studies to explore the levels of HOPX in different fibrotic animal models (e.g. adeno-TGF-β27) or models with varying degrees of severity. Importantly, we recently observed loss of HOPX in an ex vivo human tissue model of early IPF using precision cut lung slices28, and while HOPX expression increased in BLM-derived pmATII cells, human EpCAM+/HTII280+ cells in IPF lungs decreased21. These findings support the hypothesis that EpCAM+/HTII280+ cells in IPF have already lost their regenerative capability, while pmATII cells in BLM might still exhibit regenerative capacity in the relatively “early” repair period. Taken together, HOPX may be a marker for the extent of alveolar injury or a marker of the progression of repair, and failed regeneration.
Using the FCM-based technique and IF, we describe 3 different AEC phenotypes in terms of HOPX and proSP-C expression in murine and human alveolar epithelial cells: HOPX−/proSP-C+ ATII cells, HOPX+/proSP-C− ATI cells, and HOPX+/proSP-C+ AECs. Of note, we observed an increase in the number of HOPX+/proSP-C+ cells in both BLM-treated mouse lungs and human IPF lungs. These HOPX+/proSP-C+ AECs could be intermediate cell types between ATII-ATI during trans-differentiation, or bipotent progenitor cells capable to differentiate either ATII or ATI cells, which have been described in the developing lung29 and also in alveolar epithelial progenitor lineage from adult human lung in the recent study30. Our data support the notion that either intermediate or bipotent progenitor cell type could emerge during alveolar injury and repair process.
HOPX is also known to be a tumor suppressor gene in several organs31,32,33,34. In the lung, HOPX expression inhibited lung adenocarcinoma formation and metastasis14,15. In our study, we demonstrate that HOPX is involved in suppression of epithelial cell proliferation in vitro. IHC on IPF lungs also indicated that HOPX+/proSP-C+/KRT7+ AECs lacked nuclear Ki67 expression. In addition, KRT5+/proSP-C−/HOPX−/KRT7+ abnormal bronchiolar basal cells in IPF showed more nuclear Ki67 expression. End-stage IPF lungs indicate fibrotic scarring as well as increased bronchiolization, i. e. enhanced bronchiolar proliferation and replacement of alveoli by bronchiolar epithelium, HOPX expression may decrease during this process along with the progression of IPF.
In IPF, alveolar epithelial regeneration is incomplete or ineffective35. This may be due to the persistence of “repetitive triggers” caused by genetic susceptibility36,37,38,39 or epigenetic changes40,41, resulting in permanent and persistent epithelial injury and apoptosis42. In a variety of studies, “maladaptive” pro-apoptotic endoplasmic reticulum stress, DNA damage as well as senescence have been well documented in the AECII of patients with sporadic and familial IPF20,43. These conditions might be responsible for the reduction of genes involved in alveolar maintenance, such as HOPX. In addition, IPF is characterized by a “hyper-responsive” regenerative loop due to re-activation of developmental pathways including sonic hedgehog44, TGF-β45, and Wnt signaling46, resulting in inappropriate proliferation of alveolar epithelial precursors, which are also prone to die due to replicative senescence, and presumably also due to loss of HOPX. Impaired alveolar regeneration in IPF is paralleled by the abnormal proliferation of bronchiolar basal cells, which initiate bronchiolization, leading to the migration and invasion of these bronchiolar progenitors into distal alveolar spaces, and finally replacement of alveoli. Indeed, bronchiolar basal cells which do not express HOPX, can be found in abnormal luminal position directly near ATII in alveoli47. Therefore, we suggest that the loss of HOPX due to increased bronchiolization of alveoli aggravate the impaired alveolar regeneration. Our data support the hypothesis that HOPX expression maintains AEC quiescence during alveolar homeostasis, whereas HOPX induction limits aberrant AEC proliferation during normal lung injury/repair. Furthermore, aberrant HOPX expression may contribute to increased alveolar senescence in IPF.
Conclusion
HOPX contributes to alveolar injury/repair process in fibrotic lung diseases, but fails to regenerate alveolar epithelium in IPF, due to loss of HOPX. HOPX may thus be potential indicator of the progression of fibrosis.
Methods
Human Samples
The study protocol was approved by the Ethics Committee of the Justus-Liebig-University Giessen (No. 111/08 and 58/15), and informed consent was obtained in written form from each subject. All IPF diagnoses were made on the basis of the recent IPF consensus guidelines48 and a usual interstitial pneumonia (UIP) pattern was consistently observed in all used lung tissues. All methods, including those involving human samples, were performed in accordance with the relevant guidelines and regulations.
Animals
Six to eight-week-old female wildtype C57/BL6N mice were obtained from Charles River and housed in rooms with constant humidity and temperature with 12 h light cycles and free access to water and rodent chow. All animal studies were performed under the strict governmental and international guidelines and approved by the local government for the administrative region of Upper Bavaria (Project 55.2-1-54-2532-88-12). All methods were performed in accordance with the relevant guidelines and regulations.
Isolation and culture of primary mouse AECs
Primary mouse (pm) ATII cells were isolated as previously described with some modifications49. Briefly, lungs were perfused with ice-cold phosphate buffer saline (PBS) (Invitrogen, Thermo Fischer Scientific, Waltham, MA USA) via right ventricle. Then dispase (Roche Applied Science, Mannheim, Germany) and agarose (low gelling temperature) (Sigma Aldrich, St. Louis, MO, USA) were intratracheally instilled into the mouse lungs. CD45 positive white blood cells were magnetically depleted by CD45 microbeads (Miltenyi Biotec, Teterow, Germany) and pmATII cells were magnetically isolated by epithelial cell adhesion molecule (EpCAM) microbeads (Miltenyi Biotec). pmAECs were cultured on plastic plates with DMEM/F12 medium supplemented with 10% fetal bovine serum (FBS, Invitrogen, Thermo Fischer Scientific), 100 mg/l streptomycin, and 100 U/ml penicillin (Sigma Adrich) for 5 days and harvested on days 2, 3 and 5.
Evaluation of HOPX and proSP-C expressing cells by flow cytometry
HOPX and proSP-C expression within epithelial cells was evaluated by flow cytometry (FCM). Magnetically-isolated cells were fixed and permeabilized with IntraPrepTM Permeabilization Reagent (Beckman Coulter, Brea, CA, USA) and stained with HOPX mouse monoclonal antibody (sc-398703, 1:200, Santa Cruz Biotechnology, Dallas, TX, USA) and proSP-C rabbit polyclonal antibody (1:1000, Merck Millipore, Billerica, MA, USA). Normal mouse IgG (Santa Cruz Biotechnology) and rabbit serum (Life Technologies Corp.) were used for the isotype controls. Alexa Fluor 647 goat anti-mouse IgG and Alexa Fluor 488 goat anti-rabbit IgG (Life Technologies Corp.) were used for secondary antibodies. Intracellular expression of HOPX and proSP-C was detected using a FACS LSR II flow cytometer (BD Biosciences, San Jose, CA, USA). The HOPX and pro-SP-C positive populations were quantified by FlowJo software (Tomy Degital Biology Co., Ltd., Tokyo, Japan).
HOPX expression in bleomycin-induced lung injury/repair model
Eight-week-old female wildtype C57/BL6N mice were instilled with PBS or 2 units (as a single dose of 2 U/kg body weight in 50 µl PBS) of bleomycin (BLM, Bleomycin sulfate, Almirall, Barcelona, Spain) in vivo. After 14 days of instillation, fixed lung tissue was used for IF. pmATII cells from PBS/BLM mouse lungs were separated as above and the cells were used for quantitative (q) RT-PCR and FCM analysis.
Hopx siRNA transfection
Murine lung epithelial cells (MLE12) were purchased from ATCC (CRL-2110) and cultured in RPMI medium (Invitrogen, Thermo Fischer Scientific) supplemented with 10%FBS, 100 mg/l streptomycin, and 100 U/ml penicillin.MLE12 cells were grown until 70–80% confluency in six-well cluster plates and transfected with 150 pmol/l of specific double-stranded siRNA targeted against the Hopx transcript (sc-38672; Santa Cruz Biotechnology, Inc.) in serum-free Opti-MEM medium (Life Technologies) in combination with Lipofectamine 2000 transfection reagent (Life Technologies). Control transfections were performed using 150 pmol/l non-silencing control siRNA (SC-37007; Santa Cruz Biotechnology,Inc.). After 24 h of transfection, the cells were collected for RNA isolation, metabolic assay, FCM analysis, or lysed for western blotting.
WST-1 metabolic assay
WST-1 metabolic assay was performed as previously described50.
EdU assay
EdU-based proliferation assay was performed using Click-iT® EdU Alexa Fluor® 647 Imaging Kit (Invitrogen, Thermo Fischer Scientific) as manufacturer’s instructions with some modifications. Briefly, MLE12 cells were incubated with EdU solution for 24 hours, fixed with 3.7% formaldehyde, and permeabilized with 0.5% TritonX-100. Then the cells were intracellularly stained with mouse HOPX antibody as described above. Alexa Fluor 647 secondary dye was used to visualize EdU positive cells and Alexa Fluor 488 goat anti-mouse IgG was used to evaluate HOPX expression. The number of EdU positive/HOPX positive, or EdU positive/HOPX negative cells were measured and quantified by a FACS LSR II flow cytometer (BD Biosciences).
Scratch assay
Scratch assay was performed using Culture-Insert 2 well in μ-Dish 35 mm (Nippon Genetics Co. Ltd., Tokyo, Japan) according to the manufacturer’s instructions. The gap between the cells were measured using ImageJ software (from NIH, MD, USA).
Total RNA isolation and qRT-PCR
Total RNA from pmAECs and whole lung homogenate was isolated using Total RNA Kit, peqGOLD (VWR International, Radnor, PA, USA) according to the manufacturer’s instructions. cDNA was synthetized as previously described28. QRT-PCR was performed using SYBR green and the LightCycler 480 system (Roche, Mannheim, Germany). Expression changes of alveolar epithelial genes (Hopx, Sftpc, T1α), proliferation markers (Mki67, Ctnnb1) were evaluated by qRT-PCR. Primer sequences were as follows; mHopx forward: 5′TCTCCATCCTTAGTCAGACGC3′, reverse: 5′GGGTGCTTGTTGACCTTGTT3′, mSftpc forward: 5′AGCAAAGAGGTCCTGATGGA3′, reverse: 5′GAGCAGAGCCCCTACAATCA 3′, mT1α forward: 5′ACAGGTGCTACTGGAGGGCTT3′, reverse: 5′TCCTCTAAGGGAGGCTTCGTC3′, mHprt: 5′ CCTAAGATGAGCGCAAGTTGAA3′, 5′ CCACAGGACTAGAACACCTGCTAA3′.
Immunofluorescence staining
Immunofluorescence staining (IF) of the lung sections was performed as previously described49 with some modifications. Briefly, after deparaffinized, the lung sections were boiled in citrate buffer (10 mM Citric Acid, pH 6.0) for 10 minutes at 90 °C to retrieve antigens. Samples were blocked by 5% BSA with 0.1% TritonX-100 (company) in PBS for 2 hours at room temperature, and then stained with primary antibodies overnight at 4 °C. We used the following primary antibodies; HOPX mouse monoclonal antibody (1:200, Santa Cruz Inc., sc-398703) and proSP-C rabbit polyclonal antibody (1:1000, Merck Millipore, Billerica, MA, USA, AB3786). Normal mouse IgG (Santa Cruz Biotechnology) and rabbit serum (Life Technologies Corp.) were used for the isotype controls. Alexa Fluor 647 goat anti-mouse IgG and Alexa Fluor 488 goat anti-rabbit IgG (Life Technologies Corp.) were used for secondary antibodies. On the following day, the sections were washed with 1% BSA/PBS and stained with secondary antibodies for 1 hour at room temperature. Samples were visualized using a Zeiss LSM710 confocal microscope (Carl Zeiss, Oberkochen, Germany). Images were obtained, and co-localization analysis was performed with ZEN2009 software (Carl Zeiss).
Immunohistochemistry
Immunohistochemistry (IHC) of the lung sections was performed as previously described in detail20. The primary antibodies used for IHC are listed as follows: proSP-C rabbit polyclonal antibody (1:750, Merck Millipore, AB3786), rabbit monoclonal antibody for human cytokeratin-5 (KRT5) (1:200, Abcam, Cambridge, United Kingdom, ab75869), rabbit monoclonal antibody for human cytokeratin-7 (KRT7) (1:200, Abcam, ab68459), HOPX mouse monoclonal antibody (1:50, Santa Cruz Inc., sc-398703), and mouse monoclonal antibody for human Ki67 (1:50, Dianova, DIA-670-P05). Immunostained lung sections were scanned with a scanning device (Nano-Zoomer, Hamamatsu), and examined histopathologically using the ‘NDP.view2 software’ at 200×, 400× and 800× original magnification. IHC for mentioned antibodies was undertaken in 12 IPF- and 5 control-donor lung samples.
Western blotting
Western blotting was performed as previously described20
In silico analysis
HOPX expression was evaluated using the microarray dataset from Lung Genomics Research Consortium (LGRC) and the dataset from the previous reports21.
Data presentation and statistical analysis
Unless otherwise described, all data are presented as means ± standard deviation (SD). Statistical analyses were performed using the Graph Pad Prism 6 software.
For the statistical comparison of differences between two groups, the unpaired t-test was applied. For the statistical comparison of differences between three groups, one-way ANOVA with Bonferroni’s multiple comparisons test as post-test was applied. Statistical significance was defined as p < 0.05.
Data availability
The datasets analyzed during the current study are available from the corresponding author on reasonable request.
References
Knudsen, L., Ruppert, C. & Ochs, M. Tissue remodelling in pulmonary fibrosis. Cell Tissue Res 367, 607–626 (2017).
Williams, M. C. Alveolar type I cells: molecular phenotype and development. Annu Rev Physiol 65, 669–695 (2003).
Ward, H. E. & Nicholas, T. E. Alveolar type I and type II cells. Aust N Z J Med 14, 731–734 (1984).
Kuroki, Y. & Voelker, D. R. Pulmonary surfactant proteins. J Biol Chem 269, 25943–25946 (1994).
Fehrenbach, H. Alveolar epithelial type II cell: defender of the alveolus revisited. Respir Res 2, 33–46 (2001).
Barkauskas, C. E. et al. Type 2 alveolar cells are stem cells in adult lung. J Clin Invest 123, 3025–3036 (2013).
Shin, C. H. et al. Modulation of cardiac growth and development by HOP, an unusual homeodomain protein. Cell 110, 725–735 (2002).
Chen, F. et al. Hop is an unusual homeobox gene that modulates cardiac development. Cell 110, 713–723 (2002).
Yin, Z. et al. Hop functions downstream of Nkx2.1 and GATA6 to mediate HDAC-dependent negative regulation of pulmonary gene expression. Am J Physiol Lung Cell Mol Physiol 291, L191–199 (2006).
Liebler, J. M. et al. Combinations of differentiation markers distinguish subpopulations of alveolar epithelial cells in adult lung. Am J Physiol Lung Cell Mol Physiol 310, L114–120 (2016).
Jain, R. et al. Plasticity of Hopx(+) type I alveolar cells to regenerate type II cells in the lung. Nat Commun 6, 6727, https://doi.org/10.1038/ncomms7727 (2015).
Takeda, N. et al. Interconversion between intestinal stem cell populations in distinct niches. Science 334, 1420–1424 (2011).
Takeda, N. et al. Hopx expression defines a subset of multipotent hair follicle stem cells and a progenitor population primed to give rise to K6+niche cells. Development 140, 1655–1664 (2013).
Cheung, W. K. et al. Control of alveolar differentiation by the lineage transcription factors GATA6 and HOPX inhibits lung adenocarcinoma metastasis. Cancer Cell 23, 725–738 (2013).
Pacheco-Pinedo, E. C. et al. Wnt/beta-catenin signaling accelerates mouse lung tumorigenesis by imposing an embryonic distal progenitor phenotype on lung epithelium. J Clin Invest 121, 1935–1945 (2011).
Mutze, K., Vierkotten, S., Milosevic, J., Eickelberg, O. & Konigshoff, M. Enolase 1 (ENO1) and protein disulfide-isomerase associated 3 (PDIA3) regulate Wnt/beta-catenin-driven trans-differentiation of murine alveolar epithelial cells. Dis Model Mech 8, 877–890 (2015).
Flozak, A. S. et al. Beta-catenin/T-cell factor signaling is activated during lung injury and promotes the survival and migration of alveolar epithelial cells. J Biol Chem 285, 3157–3167 (2010).
Demaio, L. et al. Characterization of mouse alveolar epithelial cell monolayers. Am J Physiol Lung Cell Mol Physiol 296, L1051–1058 (2009).
Bauer, Y. et al. MMP-7 is a predictive biomarker of disease progression in patients with idiopathic pulmonary fibrosis. ERJ Open Res 3, https://doi.org/10.1183/23120541.00074-2016 (2017).
Lehmann, M. et al. Senolytic drugs target alveolar epithelial cell function and attenuate experimental lung fibrosis ex vivo. Eur Respir J 50 (2017).
Xu, Y. et al. Single-cell RNA sequencing identifies diverse roles of epithelial cells in idiopathic pulmonary fibrosis. JCI Insight 1, e90558 (2016).
Drakopanagiotakis, F., Wujak, L., Wygrecka, M. & Markart, P. Biomarkers in idiopathic pulmonary fibrosis. Matrix Biol 68-69, 404–421 (2018).
Takahashi, H. et al. Monitoring markers of disease activity for interstitial lung diseases with serum surfactant proteins A and D. Respirology 11(Suppl), S51–54 (2006).
Collard, H. R. et al. Plasma biomarker profiles in acute exacerbation of idiopathic pulmonary fibrosis. Am J Physiol Lung Cell Mol Physiol 299, L3–7 (2010).
Wang, K., Ju, Q., Cao, J., Tang, W. & Zhang, J. Impact of serum SP-A and SP-D levels on comparison and prognosis of idiopathic pulmonary fibrosis: A systematic review and meta-analysis. Medicine (Baltimore) 96, e7083 (2017).
Rosas, I. O. et al. MMP1 and MMP7 as potential peripheral blood biomarkers in idiopathic pulmonary fibrosis. PLoS Med 5, e93 (2008).
Ask, K. et al. Progressive pulmonary fibrosis is mediated by TGF-beta isoform 1 but not TGF-beta3. Int J Biochem Cell Biol 40, 484–495 (2008).
Alsafadi, H. N. et al. An ex vivo model to induce early fibrosis-like changes in human precision-cut lung slices. Am J Physiol Lung Cell Mol Physiol 312, L896–L902 (2017).
Desai, T. J., Brownfield, D. G. & Krasnow, M. A. Alveolar progenitor and stem cells in lung development, renewal and cancer. Nature 507, 190–194 (2014).
Zacharias, W. J. et al. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 555, 251–255 (2018).
Ren, X. et al. HOPX hypermethylation promotes metastasis via activating SNAIL transcription in nasopharyngeal carcinoma. Nat Commun 8, 14053, https://doi.org/10.1038/ncomms14053 (2017).
Kikuchi, M. et al. Epigenetic silencing of HOPX contributes to cancer aggressiveness in breast cancer. Cancer Lett 384, 70–78 (2017).
Yap, L. F. et al. HOPX functions as a tumour suppressor in head and neck cancer. Sci Rep 6, 38758, https://doi.org/10.1038/srep38758 (2016).
Yamashita, K., Katoh, H. & Watanabe, M. The homeobox only protein homeobox (HOPX) and colorectal cancer. Int J Mol Sci 14, 23231–23243 (2013).
Konishi, K. et al. Gene expression profiles of acute exacerbations of idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 180, 167–175 (2009).
Seibold, M. A. et al. A common MUC5B promoter polymorphism and pulmonary fibrosis. N Engl J Med 364, 1503–1512 (2011).
Nogee, L. M. et al. A mutation in the surfactant protein C gene associated with familial interstitial lung disease. N Engl J Med 344, 573–579 (2001).
Lawson, W. E. et al. Genetic mutations in surfactant protein C are a rare cause of sporadic cases of IPF. Thorax 59, 977–980 (2004).
Tsakiri, K. D. et al. Adult-onset pulmonary fibrosis caused by mutations in telomerase. Proc Natl Acad Sci USA 104, 7552–7557 (2007).
Yang, I. V. et al. Relationship of DNA methylation and gene expression in idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 190, 1263–1272 (2014).
Huang, S. K., Scruggs, A. M., McEachin, R. C., White, E. S. & Peters-Golden, M. Lung fibroblasts from patients with idiopathic pulmonary fibrosis exhibit genome-wide differences in DNA methylation compared to fibroblasts from nonfibrotic lung. PLoS One 9, e107055 (2014).
Barbas-Filho, J. V. et al. Evidence of type II pneumocyte apoptosis in the pathogenesis of idiopathic pulmonary fibrosis (IFP)/usual interstitial pneumonia (UIP). J Clin Pathol 54, 132–138 (2001).
Korfei, M. et al. Epithelial endoplasmic reticulum stress and apoptosis in sporadic idiopathic pulmonary fibrosis. Am J Respir Crit Care Med 178, 838–846 (2008).
Bolanos, A. L. et al. Role of Sonic Hedgehog in idiopathic pulmonary fibrosis. Am J Physiol Lung Cell Mol Physiol 303, L978–990 (2012).
Jenkins, R. G. et al. Ligation of protease-activated receptor 1 enhances alpha(v)beta6 integrin-dependent TGF-beta activation and promotes acute lung injury. J Clin Invest 116, 1606–1614 (2006).
Konigshoff, M. et al. WNT1-inducible signaling protein-1 mediates pulmonary fibrosis in mice and is upregulated in humans with idiopathic pulmonary fibrosis. J Clin Invest 119, 772–787 (2009).
Korfei, M. et al. Comparative proteomic analysis of lung tissue from patients with idiopathic pulmonary fibrosis (IPF) and lung transplant donor lungs. J Proteome Res 10, 2185–2205 (2011).
American Thoracic Society. Idiopathic pulmonary fibrosis: diagnosis and treatment. International consensus statement. American Thoracic Society (ATS), and the European Respiratory Society (ERS). Am J Respir Crit Care Med 161, 646–664 (2000).
Ota, C. et al. Histone deacetylase inhibitor restores surfactant protein-C expression in alveolar-epithelial type II cells and attenuates bleomycin-induced pulmonary fibrosis in vivo. Exp Lung Res 41, 422–434 (2015).
Uhl, F. E. et al. Preclinical validation and imaging of Wnt-induced repair in human 3D lung tissue cultures. Eur Respir J 46, 1150–1166 (2015).
Acknowledgements
The authors are thankful to all the members in the Königshoff Laboratory for experimental suggestions and discussions. The authors also appreciate Nadine Adam and Rabea Imker for bleomycin experiments and histological assistance, Anastasia van den Berg and Julia Kipp for the assistance of ATII isolation, and Maria Magdalena Stein for qRT-PCR and western blotting of the human samples. The authors are grateful to the technical help of Biomedical Research Unit of Tohoku University Hospital. This study is supported by a post-doctoral research fellowship from the European Respiratory Society (LTRF-2015-4822, Ota), Grant-in-Aid for Scientific Research (C), Japan Society for the Promotion of Science (16K10056, Ota), and EC | European Research Council (ERC-2010-StG 261302, Königshoff). Human samples were provided through the European IPF Registry and Biobank (AG).
Author information
Authors and Affiliations
Contributions
C. Ota and M. Königshoff designed and planned research, analyzed and organized data, and wrote the manuscript. J.P. Ng-Blichfeldt, H.N. Alsafadi, M. Lehmann, W. Skronska-Wasek, and M.M. de-Santis planned and performed ATII isolation, FACS analysis, and qRT-PCR. M. Korfei performed immunohistochemistry. A. Guenther contributed tissue specimens. D.E. Wagner analyzed in silico data. D.E. Wagner, J.P. Ng-Blichfeldt, M. Korfei, M. Lehmann, and M. Königshoff edited manuscript, and all authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ota, C., Ng-Blichfeldt, JP., Korfei, M. et al. Dynamic expression of HOPX in alveolar epithelial cells reflects injury and repair during the progression of pulmonary fibrosis. Sci Rep 8, 12983 (2018). https://doi.org/10.1038/s41598-018-31214-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31214-x
This article is cited by
-
Lung Organoids: Systematic Review of Recent Advancements and its Future Perspectives
Tissue Engineering and Regenerative Medicine (2024)
-
A Maverick Review of Common Stem/Progenitor Markers in Lung Development
Stem Cell Reviews and Reports (2022)
-
TRIM72 promotes alveolar epithelial cell membrane repair and ameliorates lung fibrosis
Respiratory Research (2020)
-
Differential effects of Nintedanib and Pirfenidone on lung alveolar epithelial cell function in ex vivo murine and human lung tissue cultures of pulmonary fibrosis
Respiratory Research (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.