Abstract
The opinion expressed by Eriksson and colleagues’ fails to recognise that there are no standard experimental designs for academic investigations involving omics analyses of genetically modified crops and that the only valid comparator to determine the effect of the process of transgenesis is a near isogenic variety grown at the same time and location, as was the case in our investigation of NK603 maize. Eriksson does not acknowledge that the quality of the rat liver tissues in our chronic Roundup toxicity study has neither been questioned nor branded as unsuitable for further investigation. In addition, Eriksson fails to appreciate that the statistical methods we used to analyse the liver metabolomics dataset are recognised as appropriate as some of a number of approaches that can be taken. Moreover, Eriksson neglects to mention that the proteomics analysis of the liver tissues highlights structural and functional damage from Roundup exposure. Thus our results are sound and the claims by Eriksson and colleagues of experimental flaws are unfounded.Replying to: Eriksson et al. Sci Rep 8 (2018); https://doi.org/10.1038/s41598-018-30440-7.
Similar content being viewed by others
Introduction
NK603 study1
Eriksson criticizes us2 for not including spatially separated biological replicates and for analysing only 2 temporal replicates. However, our study aims were strictly restricted to identifying potential metabolic differences between NK603 genetically modified (GM) maize and a near-isogenic control grown under agricultural conditions. While some requirements for GM crop field trials performed by industry for regulatory purposes are set down in EU law, there are no standard experimental designs for academic investigations involving omics analyses; some use the suggested randomised block field design3,4, most do not5,6,7. Moreover, we assessed the consequences of Roundup herbicide application, which has not previously been undertaken3. Most GM vs non-GM omics investigations use one environmental and temporal replicate to test equivalence4,8,9,10,11,12,13. We acknowledged further experiments are needed under different environmental conditions to determine the full range of GM process effects on this maize type. However, this does not invalidate our results.
Eriksson complains we did not include evidence to show that the non-GM control variety is near-isogenic to NK603. GM material available in the marketplace is always backcrossed with other varieties and the pure isogenic is never available for independent research. Furthermore, no international guidance defines ‘near-isogenic’. Therefore this is a judgment call. We used a type of NK603 (Monsanto/Dekalb DKC 26–78) and the nearest available isogenic line (Monsanto/Dekalb DKC 26–75). The use of other near isogenic non-GM lines, if available, would constitute additional valid comparators, and provide useful information about variability between related strains. However, for GM vs non-GM comparative studies conducted both by those based in academia as well as those in industrial settings for regulatory purposes, the usual method is to employ a single near-isogenic non-GM comparator. For example, in the study of NK603 GM maize conducted by the developer company (Monsanto), only a single near-isogenic non-GM comparator was used, although no information as to its exact nature was provided14. Thus our experimental design of using only one near isogenic non-GM comparator is the norm within the field and the degree of detail we provide of the non-GM near isogenic maize variety used1 exceeds that disclosed by industry14.
Eriksson states that there is a lack of reference to expected natural variation in the levels of proteins and metabolites among different maize cultivars in our investigation. However, the point of our study was to assess the effect of the GM process on the composition of NK603. Thus the only scientifically valid comparator is the non-GM closest (isogenic) relative. Comparisons with different maize varieties that do not constitute near-isogenic strains, and which are grown at different locations and times, would only serve to increase variation in the dataset and thus mask rather than highlight the effects of the GM process, negating the purpose of the study. Additional valid comparisons could be made between other maize strains harbouring the NK603 event (produced by out-crossing of the original genetically engineered parent) and their near-isogenic maize variety(s), always with the caveat that they must be grown at the same time and location. Such an investigation would determine if our findings are unique to the NK603 strain we have studied or occur generally with this genetic modification event.
We agree that the cadaverine and putrescine content of maize varies; this is covered in our Discussion.
Roundup study
Rat liver tissues were obtained15 from animals described in a previous report16. Eriksson tries to discredit the findings of our study by association with this previous investigation16, which sparked much controversy17,18,19,20,21,22,23. Livers were freshly excised from euthanized animals, snap frozen and appropriately stored to maintain integrity. No evidence exists suggesting that these tissues are unsuitable for experimental use. The influence of age and the presence of tumours was not a concern. This can be independently verified as the raw data (age, tumour presence) are available15,16,24.
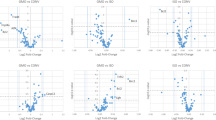
We did not overlook the consequences of the metabolomics Benjamini-Hochberg false discovery rate. We provide careful interpretation and highlight limitations in the Discussion (Lines 5–12, page 6; Lines 17–26, page 9). Numerous statistical methods have been applied to extract biologically meaningful information from metabolomics3,5,6. We employed methods that are recognised as appropriate by experts in the field. Surprisingly, Eriksson neglects to acknowledge our proteomics analysis, which provides data of high statistical significance revealing damage from Roundup ingestion. Therefore our bioinformatics and statistical analyses of both proteomics and metabolomics are sound and when taken together provide a consistent pattern of liver structural and functional defects.
Thus the claims by Eriksson and colleagues of experimental flaws in our investigation of both NK603 maize composition and chronic ultra-low dose toxicity of Roundup are unfounded.
References
Mesnage, R. et al. An integrated multi-omics analysis of the NK603 Roundup-tolerant GM maize reveals metabolism disturbances caused by the transformation process. Sci Rep. 6, 37855 (2016).
Eriksson, D., Ammann, K., Chassy, B., Chawade, A. Comments on two recent publications on GM maize and Roundup. Sci. Rep. 8, https://doi.org/10.1038/s41598-018-30440-7 (2018).
Harrigan, G. G. et al. Evaluation of metabolomics profiles of grain from maize hybrids derived from near-isogenic GM positive and negative segregant inbreds demonstrates that observed differences cannot be attributed unequivocally to the GM trait. Metabolomics. 12, 82 (2016).
Coll, A. et al. Lack of repeatable differential expression patterns between MON810 and comparable commercial varieties of maize. Plant Mol Biol 68, 105–117 (2008).
Leon, C. et al. Metabolomics of transgenic maize combining Fourier transform-ion cyclotron resonance-mass spectrometry, capillary electrophoresis-mass spectrometry and pressurized liquid extraction. J Chromatogr A. 1216, 7314–7323 (2009).
Barros, E. et al. Comparison of two GM maize varieties with a near-isogenic non-GM variety using transcriptomics, proteomics and metabolomics. Plant Biotechnol J. 8, 436–451 (2010).
Albo, A. G. et al. Proteomic analysis of a genetically modified maize flour carrying Cry1Ab gene and comparison to the corresponding wild-type. Maydica. 52, 443–455 (2007).
Ruebelt, M. C. et al. Application of two-dimensional gel electrophoresis to interrogate alterations in the proteome of genetically modified crops. 2. Assessing natural variability. J Agric Food Chem. 54, 2162–2168 (2006).
Islam, N., Campbell, P. M., Higgins, T. J., Hirano, H. & Akhurst, R. J. Transgenic peas expressing an alpha-amylase inhibitor gene from beans show altered expression and modification of endogenous proteins. Electrophoresis. 30, 1863–1868 (2009).
Coll, A. et al. Gene expression profiles of MON810 and comparable non-GM maize varieties cultured in the field are more similar than are those of conventional lines. Transgenic Res. 18, 801–808 (2009).
Arruda, S. C., Barbosa, H. S., Azevedo, R. A. & Arruda, M. A. Comparative studies focusing on transgenic through cp4EPSPS gene and non-transgenic soybean plants: an analysis of protein species and enzymes. J Proteomics. 93, 107–116 (2013).
Agapito-Tenfen, S. Z. et al. Effect of stacking insecticidal cry and herbicide tolerance epsps transgenes on transgenic maize proteome. BMC Plant Biol. 14, 346 (2014).
Wang, L. et al. Comparative proteomics of Bt-transgenic and non-transgenic cotton leaves. Proteome Sci. 13, 15 (2015).
Hammond., B., Dudek, R., Lemen, J. & Nemeth, M. Results of a 13 week safety assurance study with rats fed grain from glyphosate tolerant corn. Food Chem Toxicol 42, 1003–1014 (2004).
Mesnage, R., Renney, G., Séralini, G. E., Ward, M. & Antoniou, M. N. Multiomics reveal non-alcoholic fatty liver disease in rats following chronic exposure to an ultra-low dose of Roundup herbicide. Sci Rep. 7, 39328 (2017).
Séralini, G. E. et al. Republished study: long-term toxicity of a Roundup herbicide and a Roundup-tolerant genetically modified maize. Environ Sci Eur. 26, 14 (2014).
Séralini, G. E. et al. RETRACTED: Long term toxicity of a Roundup herbicide and a Roundup-tolerant genetically modified maize. Food Chem Toxicol. 50, 4221–4231 (2012).
Fagan, J., Traavik, T. & Bøhn, T. The Seralini affair: degeneration of Science to Re- Science? Environ Sci Eur. 27, 19 (2015).
Portier, C. J., Goldman, L. R. & Goldstein, B. D. Inconclusive findings: now you see them, now you don’t! Environ Health Perspect. 122, A36 (2014).
Roberfroid, M. Letter to theeditor: retraction of the Séralini et al. article. Food Chem Toxicol. 65, 390 (2014).
endsciencecensorship.org. https://web.archive.org/web/20151024050433/http://www.endsciencecensorship.org/en/page/Statement#signed-by (2014).
Baum, Hedlund, Aristei, Goldman. Monsanto Papers|Secret Documents|Page Two. https://www.baumhedlundlaw.com/toxic-tort-law/monsanto-roundup-lawsuit/monsanto-secret-documents-page-two/ (2017).
Horel, S. & Foucart, S. L’affaire Séralini ou l’histoire secrète d’un torpillage. Le Monde, 5 October. http://www.lemonde.fr/planete/article/2017/10/05/l-affaire-seralini-ou-l-histoire-secrete-d-un-torpillage_5196526_3244.html#psfHj6spOPZ9ahyF.99 (2017).
Mesnage, R. et al. Transcriptome profile analysis reflects rat liver and kidney damage following chronic ultra-low dose Roundup exposure. Environ Health. 14, 70 (2015). Erratum in: Environ Health. 16, 28, 2017.
Author information
Authors and Affiliations
Contributions
M.N.A. led the drafting of the article with technical input from R.M., S.A.-T. and G.-E.S.
Corresponding author
Ethics declarations
Competing Interests
M.A. declares that research in his laboratory is supported by the Sustainable Food Alliance (USA), Breast Cancer UK, The Sheepdrove Trust (UK) and the Safe Food Institute (Australia). He has served as an expert witness on behalf of the State of Vermont (U.S.A.) in a case involving the labelling of food products containing ingredients from genetically engineered organisms (although this case did not come to court). R.M. declares that he has no financial competing interests but is supported by grants from the Sustainable Food Alliance (USA). G.-E.S. declares that he has no financial competing interests. M.N.A., R.M. and G.-E.S. are members of the scientific council of CRIIGEN (Comité de recherche et d’information indépendantes sur le génie génétique). M.N.A. and G.-E.S. are members of ENSSER (European Network of Scientists for Social and Environmental Responsibility). S.A.-T. declares that she has no financial or non-financial competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Antoniou, M.N., Mesnage, R., Agapito-Tenfen, S. et al. Reply to ‘Comments on two recent publications on GM maize and Roundup’. Sci Rep 8, 13339 (2018). https://doi.org/10.1038/s41598-018-30751-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30751-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.