Abstract
The ACT domain (aspartate kinase, chorismate mutase and TyrA), an allosteric effector binding domain, is commonly found in amino acid metabolic enzymes. In addition to ACT domain-containing enzymes, plants have a novel family of ACT domain repeat (ACR) proteins, which do not contain any recognizable catalytic domain. Arabidopsis has 12 ACR proteins, whose functions are largely unknown. To study the functions of Arabidopsis ACR11, we have characterized two independent T-DNA insertion mutants, acr11-2 and acr11-3. RNA gel-blot analysis revealed that the expression of wild-type ACR11 transcripts was not detectable in the acr11 mutants. Interestingly, a lesion-mimic phenotype occurs in some rosette leaves of the acr11 mutants. In addition, high levels of reactive oxygen species (ROS), salicylic acid (SA), and callose accumulate in the mutant leaves when grown under normal conditions. The expression of several SA marker genes and the key SA biosynthetic gene ISOCHORISMATE SYNTHASE1 is up-regulated in the acr11 mutants. Furthermore, the acr11 mutants are more resistant to the infection of bacterial pathogen Pseudomonas syringae pathovar tomato DC3000. These results suggest that ACR11 may be directly or indirectly involved in the regulation of ROS and SA accumulation, which in turn modulates SA-associated defense responses and disease resistance in Arabidopsis.
Similar content being viewed by others
Introduction
Amino acids are essential organic compounds for all life forms. The synthesis of these important molecules is tightly regulated. It is well established that many key enzymes involved in amino acid biosynthesis are subject to feedback inhibition. These feedback-regulated enzymes are usually composed of catalytic domains and allosteric domains, which are responsible for catalyzing the reaction and allosteric regulation of the enzyme activity, respectively1,2,3,4. Interestingly, despite being feedback-regulated by different amino acids, the allosteric domain of these enzymes shares some common features in the primary sequence and tertiary structure, which has been named the ACT domain after bacterial aspartate kinase (AK), chorismate mutase (CM) and TyrA (prephenate dehydrogenase, PDH)5,6. AK catalyzes the first reaction of the biosynthesis of aspartate family amino acids, including lysine, methionine, and threonine. The activity of AK is feedback-regulated by lysine and threonine via the conserved ACT domain7. CM and PDH are involved in the biosynthesis of aromatic amino acids, which are feedback-regulated by phenylalanine and tyrosine through the regulatory ACT domain1,8.
Feedback regulation of amino acid biosynthetic enzymes has been extensively studied in bacteria. Interestingly, these enzymes are highly conserved from bacteria to plants. Most plant homologs have similar domain composition, e.g. a specific enzyme catalytic domain, followed by a general allosteric regulatory ACT domain2,3,7. It is likely that the ACT domain fused to the amino acid metabolic enzymes also serves as an allosteric ligand-binding domain in plants. Indeed, the Arabidopsis AK is feedback-inhibited by lysine and S-adenosylmethionine via the ACT domain9. The three committed enzymes in branched-chain amino acid BCAA biosynthesis, e.g. threonine deaminase, acetohydroxy acid synthase, and isopropylmalate synthase are feedback-regulated by branched-chain amino acids mainly via the ACT domains of these enzymes in Arabidopsis10. Furthermore, the Arabidopsis ACT domain-containing enzyme phosphoglycerate dehydrogenase is feedback regulated by its end product serine11. Thus, the ACT domain is a widespread allosteric regulatory domain that is highly conserved from bacteria to plants.
We previously identified a novel ACT domain repeat (ACR) protein family in Arabidopsis2,12. The ACR proteins contain only ACT domain repeats but not any recognizable catalytic domain. The functions of these plant ACR proteins are largely unknown. Arabidopsis has 12 ACR proteins, which are further divided into 3 different groups according to their ACT domain composition and sequence homology12. Group III ACR proteins, e.g. ACR11 and ACR12, are distinct in that they contain a non-conserved N-terminal transit peptide followed by two conserved ACT domains. Indeed, the Arabidopsis ACR11 and ACR12 proteins have been demonstrated to localize in the chloroplast12. The ACR11 gene is specifically expressed in green tissues, and is coordinately regulated with GLN2 encoding a chloroplastic glutamine synthetase 2 (GS2) in Arabidopsis12. Recently, the Arabidopsis ACR11 protein was shown to activate the activity of GS2 and levels of glutamine were significantly reduced in the acr11 mutant13. In addition, the Arabidopsis ACR11 protein was shown to interact with ferredoxin-dependent glutamine oxoglutarate aminotransferase 1 (Fd-GOGAT1) and the activity of Fd-GOGAT was reduced in the acr11 mutants14. It has been proposed that ACR11 may stabilize Fd-GOGAT and possibly modulates its activity14. Nevertheless, the molecular mechanisms of ACR11 have yet to be elucidated.
Here, we have characterized two independent acr11 T-DNA insertion mutants in Arabidopsis. Interestingly, spontaneous cell death occurs in the rosette leaves of the acr11 mutants. The lesion-mimic phenotype accompanied by increased levels of reactive oxygen species (ROS) and salicylic acid (SA)-associated defense responses make the acr11 mutants more resistant to the bacterial pathogen Pseudomonas syringae pathvar tomato DC3000 (Pst). The homeostasis of glutamine has been proposed to modulate SA-associated redox status and defense responses in Arabidopsis15. The functions of Arabidopsis ACR11 in the interconnections of GS/Fd-GOGAT cycle, glutamine homeostasis, redox balance, ROS accumulation, and SA-associated defense responses are discussed herein.
Results
Isolation and characterization of Arabidopsis acr11 mutants
The Arabidopsis ACR11 protein is predicted to contain a transit peptide with the cleavage site located at the 52nd residue (www.cbs.dtu.dk/services/TargetP/) followed by two ACT domains (Fig. 1a). To further characterize the functions of ACR11, we obtained two independent T-DNA insertion lines, SAIL_14H_10 and SALK_025722, from the Arabidopsis Biological Resource Center (ABRC). The acr11 homozygous mutant plants were isolated from these T-DNA insertion lines by PCR and confirmed by genomic Southern blot analysis (Supplementary Fig. S1). The T-DNA mutant of SAIL_14H_10 was previously named acr11-214. We have adopted the nomenclature and named the new allele derived from the SALK_025722 T-DNA line acr11-3. The positions of T-DNA insertion of acr11-2 and acr11-3 are shown in Fig. 1b. We used RNA gel-blot analysis to examine the expression of ACR11 in wild-type, acr11-2 and acr11-3 seedlings. Although the wild-type ACR11 transcript was not detectable in acr11-2, two faint bands, one higher and another lower than ACR11, were detected in the mutant (Fig. 1c). The identities of these two bands are unknown. By contrast, transcripts of ACR11 were not detectable in the acr11-3 mutant (Fig. 1c).
Molecular and phenotypic analyses of Arabidopsis acr11 mutants. (a) Schematic diagram of the Arabidopsis ACR11 protein. TP, transit peptide. (b) Schematic diagram of the ACR11 gene and the locations of T-DNA insertion in the acr11-2 and acr11-3 mutants. (c) RNA gel-blot analysis. Total RNA extracted from 2-week-old wild type (WT), acr11-2 and acr11-3 seedlings was used to detect the expression of ACR11. The full-length blot is shown in Supplementary Fig. S5. (d) Four-week-old Arabidopsis WT, acr11-2 and acr11-3 mutant plants grown in soil under normal conditions. Lesions appear in the rosette leaves of the acr11 mutants are indicated by white arrows.
The phenotypes of acr11-2 and acr11-3 are very similar. The mutant plants are smaller than wild type (Supplementary Fig. S2), and some lesions appear in the rosette leaves of acr11-2 and acr11-3 mutants when grown in soil under a 16-h light/8-h dark cycle (Fig. 1d). The lesion usually starts to develop in the rosette leaves of 3- to 4-week-old acr11 mutant plants (Fig. 1d). We used trypan blue to stain dead cells in the lesion-containing rosette leaves from acr11-2 and acr11-3. Compared with the wild type, the acr11 mutants possess more dead cells in the rosette leaves (Fig. 2). Thus, the lesions occur in some of the rosette leaves can be attributed to spontaneous cell death in the acr11 mutants.
Accumulation of ROS and callose in the acr11 mutants
It is known that accumulation of ROS commonly proceeds spontaneous cell death in plants16. We thus examined the accumulation of ROS, including H2O2, superoxide, and singlet oxygen species, in the rosette leaves of 5-week-old wild-type and acr11 mutant plants by diaminobenzidine tetrahydrochloride (DAB), nitroblue tetrazolium (NBT), and Singlet Oxygen Sensor Green (SOSG) staining, respectively. Levels of H2O2, superoxide, and singlet oxygen are significantly higher in the mutant leaves as compared with those of the wild-type (Fig. 3a–c). These results suggest that the acr11 mutants have accumulated excessive amounts of ROS when grown under normal conditions. In addition, we used aniline blue to stain callose in the rosette leaves from 5-week-old wild-type and acr11 mutant plants (Fig. 4a). The mutant leaves have accumulated significant amount of callose as compared with that of the wild type (Fig. 4a,b).
Comparison of reactive oxygen species levels in the rosette leaves of 5-week-old Arabidopsis wild-type (WT) and acr11 mutant plants. (a) Staining of hydrogen peroxide by 3,3′-diaminobenzidine. (b) Staining of superoxide radical by nitroblue tetrazolium. (c) Staining of singlet oxygen species by Singlet Oxygen Sensor Green (SOSG) fluorescent dye. Scale bars are 1 mm in (a) and (b), 200 μm in (c).
Callose deposition in the rosette leaves of 5-week-old Arabidopsis wild type (WT) and acr11 mutants. (a) Staining of callose deposits by aniline blue. Scale bar is 100 μm. (b) Quantification of callose deposits in WT and acr11 mutant leaves. Values shown are means ± SD per 0.97 mm2 from leaves of 5 independent plants. ***P < 0.001 represents the result of Student’s t test.
The expression of SA marker genes is induced in the acr11 mutants
The lesion-mimic phenotype and accumulation of ROS in the rosette leaves suggest that the SA-related signaling pathways may be activated in the acr11 mutants. To examine if the SA-associated responses are enhanced in the acr11 mutants, we used quantitative (q) RT-PCR analysis to measure the expression of SA marker genes in the rosette leaves of 5-week-old wild type and acr11 mutants. The selected SA maker genes include PATHOGENESIS-RELATED 1 (PR1), PR2, PR5, CALMODULIN BINDING PROTEIN 60 G (CBP60G) and two WRKY transcription factor genes, WRKY18 and WRKY53. Steady-state mRNA levels of PR1, PR2, PR5, CBP60, WRKY18, and WRKY53 were significantly higher in the acr11-2 and acr11-3 mutants as compared with those of the wild type (Fig. 5). In addition, RNA gel-blot analysis of PR1 and PR2 in Arabidopsis wild type and acr11-3 was shown in Supplementary Fig. S3. These results indicate that the SA-associated responses are constitutively activated in the acr11 mutant rosette leaves.
Quantitative RT-PCR analysis of salicylic acid-responsive genes. Total RNA extracted from rosette leaves of 5-week-old Arabidopsis wild-type (WT), acr11-2, and acr11-3 mutant plants was used for qRT-PCR analysis to detect the expression of PR1, PR2, PR5, CBP60G, WRKY18 and WRKY53. Relative expression indicates the fold-change of each gene as compared to that of WT. Results shown are means ± SD from three independent experiments. ***P < 0.001 represents the result of Student’s t test.
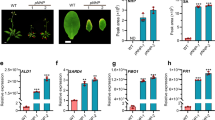
Accumulation of SA in the acr11 mutants
Perturbation of endogenous SA levels will directly affect the SA-associated responses in plants. To test if the constitutively activated SA responses in the acr11 mutants is caused by changes of SA levels, we measured the amount of SA in the rosette leaves of 5-week-old wild-type, acr11-2 and acr11-3 mutant plants. The results indicate that the SA levels in the mutant rosette leaves are significantly higher than those of the wild type (Fig. 6a). It is known that isochorismate synthase (ICS) is the major enzyme involved in SA biosynthesis in Arabidopsis17. There are two ICS genes, ICS1 and ICS2, in Arabidopsis. We used qRT-PCR analysis to examine the expression of ICS1 and ICS2 in the rosette leaves of wild-type and acr11 mutant plants. The expression levels of ICS1, but not ICS2, are much higher in the acr11-2 and acr11-3 mutants as compared to the wild type (Fig. 6b).
Enhanced salicylic acid (SA) accumulation is in the acr11 mutants. (a) Levels of free SA in the rosette leaves of 5-week-old Arabidopsis wild type (WT) and acr11 mutants. (b) Quantitative RT-PCR analysis of SA biosynthetic genes in the rosette leaves of 5-week-old Arabidopsis WT and acr11 mutants. The expression levels of ICS1 and ICS2 in WT were set at 1. Fold change indicates the relative expression of ICS1 and ICS2 as compared to that of WT. ICS, ISOCHORISMATE SYNTHASE. Results shown are means ± SD from three independent experiments. ***P < 0.001 represents the result of Student’s t test.
Enhanced disease resistance in the acr11 mutants
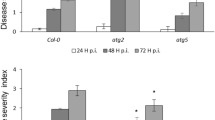
The phenotypes of lesion-mimic, enhanced SA-dependent responses, and accumulation of callose, ROS and SA suggest that the acr11 mutants may be more resistant to pathogen infection. To test this possibility, we used the virulence strain Pseudomonas syringae pathovar tomato DC3000 (Pst) to infect the rosette leaves of 5-week-old wild-type and acr11 mutant plants with syringe infiltration. The disease symptoms developed in the mutant leaves were significantly weaker than those of the wild type 3 days after inoculation (Fig. 7a). This phenotype was associated with bacterial growth in leaves infiltrated with Pst. The number of bacteria growing in the mutant leaves was significantly smaller than that of the wild type 1 to 3 days post inoculation (Fig. 7b).
Enhanced disease resistance in 5-week-old Arabidopsis acr11 mutants. (a) Symptoms of wild-type (WT), acr11-2 and acr11-3 rosette leaves 3 days after syringe infiltration with Pseudomonas syringae pv. tomato DC3000 (Pst). (b) Growth of Pst in Arabidopsis WT, acr11-2 and acr11-3 mutants. Bacterial titers were evaluated at 0 to 3 days post inoculation (dpi). Results are means ± SD from three independent experiments. Asterisks indicate significant differences (*P < 0.05; **P < 0.01; Student’s t test) compared to the WT.
In addition to syringe infiltration on individual rosette leaves, we also inoculated whole plants with Pst by dipping. Compared with the wild type, the acr11 mutant plants showed less severe disease symptoms 3 days after dip inoculation (Supplementary Fig. S4a). The rosette leaves of the acr11 mutants had less bacterial growth as compared with that of the wild type 1 to 3 days after dip inoculation (Supplementary Fig. S4b). These results suggest that the acr11 mutants are more resistant to Pst infection.
Discussion
The prediction that plant ACR11 homologs contain two ACT domains is reminiscent of the recent discovery of human arginine sensor CASTOR118,19. Unlike CASTOR1, the functions of plant ACR proteins are largely unknown. Nonetheless, these proteins share a common feature that they all contain multiple ACT domains, but not any recognizable catalytic domain. The human CASTOR1 and its homolog CASTOR2 are predicted to contain 2 ACT domains18. However, crystal structure analysis of the arginine-bound CASTOR1 reveals that it is in fact composed of 4 tandem ACT domains19,20,21. Therefore, the real composition of ACT domains in the ACR11 protein requires further studies on the crystal structure.
It is interesting that the Arabidopsis acr11 mutants have a lesion-mimic phenotype. Usually, the lesion first appears in the rosette leaves of 3- to 4-week-old acr11 mutant plants when grown in soil under a normal condition. Many plant lesion-mimic mutants are associated with disease resistance22,23. The spontaneous cell death phenotype and the induction of defense response are coordinated in the acr11 mutants. Furthermore, we have shown that the Arabidopsis acr11 mutants are more resistant to the bacterial pathogen Pst (Fig. 7 and Supplementary Fig. S4). These results suggest that the cell death pathway activated in the acr11 mutants is interconnected with the defense pathway against Pst.
The ACR11 protein is localized to the chloroplast12, which is also the site for SA biosynthesis and one of the major sites for ROS production inside the plant cell. The increased disease resistance of the acr11 mutants can be attributed to the accumulation of ROS and SA, which is well-documented to enhance plant immunity17,24,25. In the rosette leaves of Arabidopsis acr11 mutants, levels of ROS, SA and callose are significantly increased as compared to those of the wild type (Figs 3, 4 and 6). Furthermore, the expression of SA biosynthetic and responsive genes is dramatically induced in the acr11 mutant (Figs 5, 6). These results are consistent with the phenotype that the acr11 mutant is more resistant to pathogen infection. Loss-of-function in ACR11 will increase ROS levels and activate defense responses in the plant. However, if the amounts of ROS accumulated inside the cell are over the threshold, cell death will occur in the acr11 mutants. The molecular mechanisms of ACR11 in the chloroplast-triggered spontaneous cell death remain to be elucidated.
Alternatively, the increased disease resistance of the acr11 mutants can be attributed to shortage of nutrients, e.g. glutamine and its derivatives, for bacterial growth in the host plant. Pathogens have to obtain their nitrogen nutrients from the host plant. Thus, the nitrogen status of the host plant is tightly associated with pathogenesis26,27. Interestingly, the pathogenic Pst has been shown to selectively catabolize abundant amino acids, such as glutamine, glutamate, and aspartate from the host plant28. The Arabidopsis acr11 mutants have reduced levels of glutamine13. It is possible that decreased levels of glutamine may cause shortage of nutrients for bacterial growth, and thus confers pathogen resistance in the acr11 mutants.
In addition to its role in nutrition and metabolism, glutamine can also function as a signaling molecule in plants29,30. It has been shown that glutamine homeostasis can modulate SA-associated redox status and defense responses in Arabidopsis15. It is conceivable that ACR11 may be involved in the maintenance of Gln homeostasis in Arabidopsis. We propose a hypothetical working model that Gln-deficiency derived from a compromised GS/Fd-GOGAT cycle may affect redox balance and induce ROS production in the acr11 mutant (Fig. 8). This hypothesis may be partly supported by the following observations: (1) the ACR11 and GLN2 genes are coordinately expressed12; (2) ACR11 can activate GS2 and levels of glutamine are decreased in the acr11 mutant13; (3) ACR11 interacts with Fd-GOGAT1 and the activity of Fd-GOGAT is reduced in the acr11 mutant14. Arabidopsis Fd-GOGAT1 plays a major role in the assimilation of ammonium generated by photorespiration31. In addition to reduced levels of glutamine13, ammonium may also accumulate in the acr11 mutant. Excess amounts of ammonium are toxic to plants, which may activate ROS production and trigger the downstream SA-associated defense responses (Fig. 8).
A hypothetical working model of ACR11 function in modulating reactive oxygen species (ROS) production and salicylic acid (SA)-associated defense responses. ACR11 may integrate the information of glutamine homeostasis and redox balance to modulate the activities of glutamine synthetase (GS)/ferredoxin-dependent glutamine oxoglutarate aminotransferase (Fd-GOGAT) and ROS production in the chloroplast. Overproduction of ROS activates SA biosynthesis and enhances SA-associated defense responses. In addition, ACR11 may directly affect SA biosynthesis, which in turn affects ROS production and SA-associated defense responses.
The reactions catalyzed by GS/Fd-GOGAT require ATP and reducing power derived from photosynthesis. Thus, the GS/Fd-GOGAT cycle is a strong electron sink, which plays an important role in consuming and translocating reducing equivalents inside the plant cell. Perturbation of the GS/Fd-GOGAT cycle results in Gln-deficiency and redox imbalance, which may trigger the overproduction of ROS and enhance SA-associated defense responses in the acr11 mutant. Alternatively, ACR11 may have an effect on SA biosynthesis, and subsequently affect the accumulation of ROS. It is known that the interplay of SA and ROS can modulate the expression of defense genes32. Overproduction of SA may also result in increased levels of ROS and enhanced defense responses in the acr11 mutants. It will be interesting to investigate the role of ACR11 in the interconnection of Gln metabolism, ROS production, and SA-associated defense network in Arabidopsis.
Fd-GOGAT1 has been shown to interact and regulate the activities of UDP-sulfoquinovose synthase and serine hydroxymethyltransferase33,34. Interestingly, Fd-GOGAT1 plays a regulatory role when interacts with these enzymes, which is independent of its catalytic function33,34. Thus, it is possible that the ACR11/Fd-GOGAT complex may have additional functions independent of the assimilation of photorespiratory ammonium in Arabidopsis. We cannot exclude the possibility that the accumulation of ROS and SA, and the defense-related phenotypes observed in the acr11 mutants are not directly linked to the GS/Fd-GOGAT cycle. The ACR11 homologs are conserved from algae to land plants. Further studies on the functions of Arabidopsis ACR11 will provide insights into the molecular mechanism of ACR proteins in plants.
Methods
Plant materials and growth conditions
Arabidopsis thaliana ecotype Columbia-0 and T-DNA insertion mutants acr11-2 (Sail_14_H10) and acr11-3 (Salk_025722) were obtained from the Arabidopsis Biological Resource Center. Plants were grown in soil or on tissue culture plates in a controlled growth chamber on a 16-h light/8-h dark cycle at 23 °C as previously described35.
Trypan blue staining and detection of ROS
Rosette leaves from 5-week-old Arabidopsis wild-type and acr11 mutant plants were used for trypan blue, DAB, NBT, and SOSG staining as previously described with minor modifications36,37. The DAB (D5637, Sigma-Aldrich) staining solution, 1.25 mg/ml, was freshly prepared in sterilized water and adjusted to pH 3.8 with KOH. Detached rosette leaves were immersed and infiltrated under vacuum with DAB staining solution and then cleared in boiling 95% (v/v) ethanol for 10 min. For NBT staining, detached rosette leaves were immersed and infiltrated under vacuum with 3.5 mg/ml NBT (N6876, Sigma-Aldrich) staining solution in 10 mM potassium phosphate buffer containing 10 mM sodium azide. After vacuum infiltration, stained leaves were bleached in boiling 95% ethanol (v/v) for 10 min. The commercially available fluorescent dye SOSG (S36002, Thermo Fisher) was used to detect singlet oxygen. Rosette leaves from 5-week-old plants were infiltrated with a solution of 100 μM SOSG in 50 mM phosphate potassium buffer (pH 7.5). Plants were exposed to light for 30 min and infiltrated leaves were observed under a 510 META Zeiss confocal laser scanning microscope with excitation at 480 nm and emission at 530 nm.
Callose staining and microscopy
Aniline blue (415049, Sigma-Aldrich) was used to stain callose deposition. Rosette leaves of 5-week-old Arabidopsis were cleared overnight in 95% ethanol (v/v) at room temperature. The completely cleared leaves were rehydrated in sterilized water, and then immersed in aniline blue staining solution of 0.01% (w/v) in 0.15 M phosphate buffer, pH 9.5, for 30 min. The callose deposition was observed under a UV illumination using Zeiss Axio Scope A1 microscope. Callose deposits were quantified by the “analyze particles” function of ImageJ (http://rsb.info.nih.gov/ij/).
Quantitative (q) RT-PCR and RNA gel-blot analysis
Arabidopsis total RNA was isolated using a phenol extraction protocol as previously described38. Total RNA extracted from rosette leaves of 5-week-old Arabidopsis wild-type and acr11 mutant plants was digested with DNase I and used for qRT-PCR analysis. All qRT-PCRs were performed with three biological repeats and the expression data were normalized to the nuclear gene ACTIN2 (At3g18780). The following primers were used for qRT-PCR: PR1 (At2g14610), 5′-TTCACAACCAGGCACGAGGAG-3′, 5′-GCCAGACAAGTCACCGCTACC-3′; PR2 (At3g57260), 5′-CTTGAACGTCTCGCCTCCAGTC-3′, 5′-TCCAGAAACCGCGTTCTCGATG-3′; PR5 (At1g75040), 5′-CAATTGCCCTACCACCGTCTGG-3′, 5′-CTTAGACCGCCACAGTCTCCG-3′; CBP60G (At5g26920), 5′-CGGGCGTAACACTTCTCTTC-3′, 5′-AGCTTCGGCCTTTAATTGGT-3′; WRKY18 (At4g31800), 5′-CATACGAAGGGACGCATAAC-3′, 5′-CCTTTCGTTTTTCTCCAACG-3′; WRKY53 (At4g23810), 5′-GGCAGTGTTCCAGAATCTCC-3′, 5′-GCCTCTCTCTGGGCTTATTC-3′; ACTIN2 (At3g18780), 5′-GGTAACATTGTGCTCAGTGGTGG-3′, 5′-AACGACCTTAATCTTCATGCTGC-3′; ICS1 (At1G74710), 5′-TGGCGAGGAGAGTGAATTTG-3′, 5′-TGGGTCACTTCCAGCTACTA-3′; ICS2 (At1G18870), 5′-GTTTGCGGATGTCCAGTAGAA-3′, 5′-CCACCACCAAAGAACCCAATA-3′. For RNA gel-blot analysis, total RNA (10 μg) was separated in standard formaldehyde gel by electrophoresis and blotted onto a nylon membrane. To detect the transcripts of ACR11 and SA marker genes, digoxigenin (DIG)-labeled single-stranded DNA probes were generated by PCR using the following primers: ACR11 (At1g16880), 5′-ATGGCTATGGCCT CTGCTTC-3′, 5′-GAAACTTGACTCGTCAGTTG-3′; PR1 (At2g14610), 5′- ATGAATTTTACTGGCTATTCTCG-3′, 5′-TTAGTATGGCTTCTCGTTCAC-3′; PR2 (At3g57260), 5′-ATGTCTGAATCAAGGAGCTTAGC-3′, 5′-TTAGTTGAAATTAACTTCATACTTAG-3′. DIG probe labeling, pre-hybridization, hybridization, wash conditions and detection were performed according to the Boehringer-Mannheim Genius System User’s Guide: DIG Application Manual for Filter Hybridization.
Salicylic acid (SA) measurement
Rosette leaves from 5-week-old Arabidopsis wild-type and acr11 mutant plants were used for SA measurement. Sample extraction and quantitative analysis of free SA were performed as previously described39. The SA measurement was conducted by the Metabolomics Core at Academia Sinica using Ultra Performance Liquid Chromatography-High Definition Mass Spectrometry (Waters, http://www.waters.com).
Pathogen infection assays
Five-week-old Arabidopsis wild-type and acr11 mutant plants were used for Pst infection assays as previously described40. Arabidopsis plants were dipped in a bacterial suspension of 107 colony-forming units (cfu)/mL in 10 mM MgCl2 containing 0.01% (v/v) Silwet L-77 for 15 min. For inoculation by syringe infiltration, 3–4 leaves per plant were infiltrated with bacterial suspension of 105 cfu/ml using a 1-ml syringe without a needle. Plants were kept at 100% relative humidity for one day after infection, and symptoms were photographed 3 days post inoculation. For analysis of bacterial growth, 8 leaf discs with 0.5 cm diameter from 4 different plants sampled at 0 to 3 days after inoculation were used to measure bacterial growth as previously described40.
References
Chipman, D. M. & Shaanan, B. The ACT domain family. Curr. Opin. Struct. Biol. 11, 694–700 (2001).
Hsieh, M. H. & Goodman, H. M. Molecular characterization of a novel gene family encoding ACT domain repeat proteins in Arabidopsis. Plant Physiol. 130, 1797–1806 (2002).
Curien, G. et al. Amino acid biosynthesis: new architectures in allosteric enzymes. Plant Physiol. Biochem. 46, 325–339 (2008).
Lang, E. J. M., Cross, P. J., Mittelstädt, G., Jameson, G. B. & Parker, E. J. Allosteric ACTion: the varied ACT domains regulating enzymes of amino-acid metabolism. Curr. Opin. Struct. Biol. 29, 102–111 (2014).
Aravind, L. & Koonin, E. V. Gleaning non-trivial structural, functional and evolutionary information about proteins by iterative database searches. J. Mol. Biol. 287, 1023–1040 (1999).
Liberles, J. S., Thorolfsson, M. & Martinez, A. Allosteric mechanisms in ACT domain containing enzymes involved in amino acid metabolism. Amino Acids 28, 1–12 (2005).
Dumas, R., Cobessi, D., Robin, A. Y., Ferrer, J. L. & Curien, G. The many faces of aspartate kinases. Arch. Biochem. Biophys. 519, 186–193 (2012).
Zhang, S. et al. Chorismate mutase-prephenate dehydratase from Escherichia coli. Study of catalytic and regulatory domains using genetically engineered proteins. J. Biol. Chem. 273, 6248–6253 (1998).
Mas-Droux, C. et al. A novel organization of ACT domains in allosteric enzymes revealed by the crystal structure of Arabidopsis aspartate kinase. Plant Cell 18, 1681–1692 (2006).
Xing, A. & Last, R. L. A regulatory hierarchy of the Arabidopsis branched-chain amino acid metabolic network. Plant Cell 29, 1480–1499 (2017).
Okamura, E. & Hirai, M. Y. Novel regulatory mechanism of serine biosynthesis associated with 3-phosphoglycerate dehydrogenase in Arabidopsis thaliana. Sci. Rep. 7, 3533 (2017).
Sung, T. Y., Chung, T. Y., Hsu, C. P. & Hsieh, M. H. The ACR11 encodes a novel type of chloroplastic ACT domain repeat protein that is coordinately expressed with GLN2 in Arabidopsis. BMC Plant Biol. 11, 118 (2011).
Osanai, T., Kuwahara, A., Otsuki, H., Saito, K. & Hirai, M. Y. ACR11 is an activator of plastid-type glutamine synthetase GS2 in Arabidopsis thaliana. Plant and Cell Physiol. 58, 650–657 (2017).
Takabayashi, A., Niwata, A. & Tanaka, A. Direct interaction with ACR11 is necessary for post-transcriptional control of GLU1-encoded ferredoxin-dependent glutamate synthase in leaves. Sci. Rep. 6, 29668 (2016).
Liu, G. et al. Amino acid homeostasis modulates salicylic acid–associated redox status and defense responses in Arabidopsis. Plant Cell 22, 3845–3863 (2010).
Van Breusegem, F. & Dat, J. F. Reactive oxygen species in plant cell death. Plant Physiol. 141, 384–390 (2006).
Dempsey, D. A., Vlot, A. C., Wildermuth, M. C. & Klessig, D. F. Salicylic acid biosynthesis and metabolism. The Arabidopsis Book/American Society of Plant Biologists 9, e0156 (2011).
Chantranupong, L. et al. The CASTOR proteins are arginine sensors for the mTORC1 pathway. Cell 165, 153–164 (2016).
Saxton, R. A., Chantranupong, L., Knockenhauer, K. E., Schwartz, T. U. & Sabatini, D. M. Mechanism of arginine sensing by CASTOR1 upstream of mTORC1. Nature 536, 229–233 (2016).
Gai, Z. et al. Structural mechanism for the arginine sensing and regulation of CASTOR1 in the mTORC1 signaling pathway. Cell Discov. 2, 16051 (2016).
Xia, J., Wang, R., Zhang, T. & Ding, J. Structural insight into the arginine-binding specificity of CASTOR1 in amino acid-dependent mTORC1 signaling. Cell Discov. 2, 16035 (2016).
Lorrain, S., Vailleau, F., Balaque, C. & Roby, D. Lesion mimic mutants: keys for deciphering cell death and defense pathways in plants? Trends Plant Sci. 8, 263–271 (2003).
Bruggeman, Q., Raynaud, C., Benhamed, M. & Delarue, M. To die or not to die? Lessons from lesion mimic mutants. Front. Plant Sci. 30(6), 24 (2015).
Torres, M. A., Jones, J. D. G. & Dangl, J. L. Reactive oxygen species signaling in response to pathogens. Plant Physiol. 141, 373–378 (2006).
Dempsey, D. A. & Klessig, D. F. How does the multifaceted plant hormone salicylic acid combat disease in plants and are similar mechanisms utilized in humans? BMC Biology 15, 23 (2017).
Seifi, H. S., Van Bockhaven, J., Angenon, G. & Hofte, M. Glutamate metabolism in plant disease and defense: friend or foe? Mol. Plant Microbe Interact. 26, 475–485 (2013).
Fagard, M. et al. Nitrogen metabolism meets phytopathology. J. Exp. Bot. 65, 5643–5656 (2014).
Rico, A. & Preston, G. M. Pseudomonas syringae pv. tomato DC3000 uses constitutive and apoplast-induced nutrient assimilation pathways to catabolize nutrients that are abundant in the tomato apoplast. Mol. Plant Microbe Interact. 21, 269–282 (2008).
Chellamuthu, V. R. et al. A widespread glutamine-sensing mechanism in the plant kingdom. Cell 159, 1188–1199 (2014).
Kan, C. C., Chung, T. Y., Juo, Y. A. & Hsieh, M. H. Glutamine rapidly induces the expression of key transcription factor genes involved in nitrogen and stress responses in rice roots. BMC Genomics 16, 731 (2015).
Coschigano, K. T., Melo-Oliveira, R., Lim, J. & Coruzzi, G. M. Arabidopsis gls mutants and distinct Fd-GOGAT genes: implications for photorespiration and primary nitrogen assimilation. Plant Cell 10, 741–752 (1998).
Herrera-Vasquez, A., Salinas, P. & Holuigue, L. Salicylic acid and reactive oxygen species interplay in the transcriptional control of defense genes expression. Front. Plant Sci. 6, 171 (2015).
Jamai, A., Salomé, P. A., Schilling, S. H., Weber, A. P. & McClung, C. R. Arabidopsis photorespiratory serine hydroxymethyltransferase activity requires the mitochondrial accumulation of ferredoxin-dependent glutamate synthase. Plant Cell 21, 595–606 (2009).
Shimojima, M., Hoffmann-Benning, S., Garavito, R. M. & Benning, C. Ferredoxin-dependent glutamate synthase moonlights in plant sulfolipid biosynthesis by forming a complex with SQD1. Arch. Biochem. Biophys. 436, 206–214 (2005).
Hsieh, W. Y. et al. The Arabidopsis thiamin-deficient mutant palegreen1 lacks thiamin monophosphate phosphatase of the vitamin B1 biosynthesis pathway. Plant J. 91, 145–157 (2017).
Pogány, M. et al. Dual roles of reactive oxygen species and NADPH oxidase RBOHD in an Arabidopsis-Alternaria pathosystem. Plant Physiol. 151, 1459–1475 (2009).
Ramel, F., Sulmon, C., Bogard, M., Couee, I. & Gouesbet, G. Differential patterns of reactive oxygen species and antioxidative mechanisms during atrazine injury and sucrose-induced tolerance in Arabidopsis thaliana plantlets. BMC Plant Biol. 9, 28 (2009).
Tseng, C. C., Lee, C. J., Chung, Y. T., Sung, T. Y. & Hsieh, M. H. Differential regulation of Arabidopsis plastid gene expression and RNA editing in non-photosynthetic tissues. Plant Mol. Biol. 82, 375–392 (2013).
Pan, X., Welti, R. & Wang, X. Quantitative analysis of major plant hormones in crude plant extracts by high-performance liquid chromatography–mass spectrometry. Nat. Protoc. 5, 986–992 (2010).
Zimmerli, L., Jakab, G., Metraux, J. P. & Mauch-Mani, B. Potentiation of pathogen-specific defense mechanisms in Arabidopsis by beta-aminobutyric acid. Proc. Natl. Acad. Sci. USA 97, 12920–12925 (2000).
Acknowledgements
We thank Ching-Chih Tseng for technical assistance, Mei-Jane Fang for confocal microscopy, Nai-Chun Lin and Erh-Min Lai for the bacterial strain. This research was supported by a grant (NSC 99-2311-B-001-007-MY3) to M.H.H. from National Science Council, Taipei, Taiwan.
Author information
Authors and Affiliations
Contributions
M.H.H. conceived and designed the experiments. S.K.S., T.Y.S., T.Y.C., S.Y.L., S.C.L., J.C.L. and W.Y.H. conducted the experiments. S.K.S., T.Y.S., T.Y.C. and M.H.H. analyzed the data. M.H.H. wrote the manuscript. All authors read and approve the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, S.K., Sung, TY., Chung, TY. et al. ACR11 modulates levels of reactive oxygen species and salicylic acid-associated defense response in Arabidopsis. Sci Rep 8, 11851 (2018). https://doi.org/10.1038/s41598-018-30304-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-30304-0
This article is cited by
-
Proteome changes in pepper (Capsicum annuum L.) leaves induced by the green peach aphid (Myzus persicae Sulzer)
BMC Plant Biology (2021)
-
Interactions between ShPP2-1, an F-box family gene, and ACR11A regulate cold tolerance of tomato
Horticulture Research (2021)
-
Physiological and proteomic responses to drought stress in leaves of two wild grapevines (Vitis sylvestris): a comparative study
Plant Growth Regulation (2020)
-
The lineage and diversity of putative amino acid sensor ACR proteins in plants
Amino Acids (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.