Abstract
Clinical and animal studies have demonstrated beneficial effects of early consumption of dairy lipids and a probiotic, Lactobacillus fermentum (Lf), on infant gut physiology. The objective of this study was to investigate their long-term effects on gut microbiota and host entero-insular axis and metabolism. Piglets were suckled with a milk formula containing only plant lipids (PL), a half-half mixture of plant lipids and dairy lipids (DL), or this mixture supplemented with Lf (DL + Lf). They were weaned on a standard diet and challenged with a high-energy diet until postnatal day 140. DL and DL + Lf modulated gut microbiota composition and metabolism, increasing abundance of several Clostridia genera. Moreover, DL + Lf specifically decreased the faecal content of 2-oxoglutarate and lysine compared to PL and 5-aminovalerate compared to PL and DL. It also increased short-chain fatty acid concentrations like propionate compared to DL. Furthermore, DL + Lf had a beneficial effect on the endocrine function, enhancing caecal GLP-1 and GLP-1 meal-stimulated secretion. Correlations highlighted the consistent relationship between microbiota and gut physiology. Together, our results evidence a beneficial programming effect of DL + Lf in infant formula composition on faecal microbiota and entero-insular axis function.
Similar content being viewed by others
Introduction
According to the developmental origins of health and disease (DOHaD), the early life environment may have long-lasting effects on health1,2,3. Underlying mechanisms to such a programming remain largely unknown but gut microbiota has recently been identified as a potential key actor, able to induce long-lasting changes in host intestinal functions4. Gut microbiota dysbiosis has therefore been associated with an increased susceptibility to metabolic disorders5.
The early postnatal period is a crucial window for the interplay between diet, gut microbiota and host metabolism6,7,8. Neonatal nutrition may modulate early implantation of gut microbiota and therefore exert long-lasting metabolic and physiological effects in adulthood9. Differences in gut microbiota diversity and composition have been demonstrated from the first week of life between breastfed and formula-fed infants10, the last ones being at higher risk for later insulin resistance and type 2 diabetes11,12. The structure and composition of the lipid matrix and microbiota of breast milk could partly explain its long-term protective effects.
Indeed, human milk contains high concentrations of medium-chain fatty acids associated with a complex structure, named milk fat globules13,14. In contrast, the lipid matrix of most formulas on the market is exclusively made of vegetable oils containing mostly long-chain fatty acids13. We demonstrated that the addition of dairy lipids in formulas had short-term benefits on digestion, gut physiology and microbiota in a piglet model15. Moreover, mimicking the milk fat globule size and coating was associated with long-term beneficial effects in a murine model, with a decreased fat accumulation and improved metabolic profile16.
Besides, recent data emphasize that breast milk has a diversified bacterial ecosystem with over 200 different genera and 700 species17,18,19. These bacteria would account for almost one-third of total bacteria present in the gut of breastfed infants during the first month of life20. The addition of probiotics in infant formulas may, therefore, be of great interest. Among probiotics, Lactobacillus fermentum CECT 5716 (Lf), a lactic acid bacteria originally isolated from human milk21, has been proven to exert beneficial short-term effects, in particular on immediate gut microbiota composition22, prevention of infections23 and barrier dysfunction24.
If short-term benefits of dairy lipids and Lf supplied separately have previously been demonstrated on gut physiology and microbiota composition, their long-term outcomes on gut microbiota and glucose metabolism remain unknown. Moreover, the effects of dairy lipids and Lf supplied together have not been evaluated yet. We hypothesized that dairy lipids and Lf effects on gut microbiota and physiology in the neonate could have long-term consequences in adulthood on gut microbiota, host entero-insular axis and metabolism. We used a diet-induced overweight Yucatan minipig model25 to reveal a potential latent programming effect induced by the infant formula composition.
Results
The study design and the composition of the infant formulas used in this study are described in Fig. 1 and Table 1 (see Methods for details).
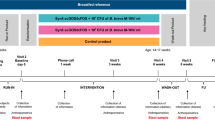
Study design from PND2 to PND140. Piglets were allowed to suckle the dam until PND2. They were then separated from their dam and fed with an automatic formula feeder until weaning, at PND 28. Formulas contained as lipids either: only plant lipids (PL), a half-half mixture of plant and dairy lipids (DL) or a half-half mixture of plant and dairy lipids supplemented with Lf (DL + Lf). From PND28 to 56, pigs were fed commercial starter diets and then challenged with a HE diet until euthanasia, at PND140. Glucose homeostasis was evaluated at PND133 via an IVGTT and fasting plasma was used for metabolomics analysis. Faeces (*) were collected at PND33 (after weaning), PND55 (before HE transition), PND61 (after HE transition), PND77, PND98 and PND135, 3, 6 and 12 weeks after HE transition, respectively. At euthanasia, faecal content was collected for metabolomics and microbiota analysis and the entero-insular axis was investigated. PND, postnatal day; HE, high-energy; IVGTT, intravenous glucose tolerance test.
Effects of dairy lipids and Lf on adult gut microbiota
Diversity
Alpha diversity (species richness, Chao1, Simpson, Shannon and inverse Simpson indices) measured on postnatal day (PND) 140 faecal samples was not affected by the composition of the infant formula (Fig. 2A and Supplementary Table 1), nor was beta diversity (weighted and unweighted Unifrac, Jaccard and Bray-Curtis distances) (Supplementary Table 1). To evaluate the contribution of diet to the pig microbiota, a permutational multivariate analysis of variance was applied using the distance matrices. The infant formula composition accounted for 9.8 to 12.1% of the total variation between samples.
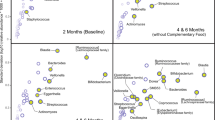
Composition and metabolism of gut microbiota. (A) α diversity indices in faecal samples of adult PL, DL and DL + Lf pigs. (B) Bacteria composition at phylum and (C) genus levels (relative abundance) in faecal samples of adult PL, DL and DL + Lf pigs. (D) Discriminating genera in the faeces of adult PL, DL and DL + Lf pigs. (E) Two-dimensional PLS-DA score plot of integrated 1H-NMR spectra of adult PL, DL and DL + Lf faecal samples. Each dot represents an animal, projected onto first (horizontal axis) and second (vertical axis) PLS-DA variables. The ellipse determines the 95% confidence interval, which is drawn using Hotelling’s T2 statistic. PLS-DA model parameters: four latent variables included, R2Y (cum) = 90.4% (% of variance explained by the model), Q2 = 0.66 (predictive ability of the model) and p-value = 0.02. (F) Discriminating faecal metabolites between adult PL, DL and DL + Lf pigs. Formulas contained as lipids either: only plant lipids (PL, n = 6–9), a half-half mixture of plant and dairy lipids (DL, n = 7–8) or a half-half mixture of plant and dairy lipids supplemented with Lf (DL + Lf, n = 7–9).
Composition
Eight phyla, 50 families and 171 genera were identified in faecal contents. The relative abundances of predominant bacterial phyla and genera are shown in Fig. 2B,C. Firmicutes constituted the dominant phylum in all groups, accounting for an average of 55% of the total sequences (Fig. 2B). Other bacterial phyla were Bacteroidetes (37%), Proteobacteria (4.6%), Fusobacteria (1.8%), Actinobacteria (1.0%), Tenericutes (0.30%), Fibrobacteres (0.12%), and Deferribacteres (0.08%). Genus-level analysis showed that the top six genera were Prevotellaceae NK3B31 group, unknown genus (Bacteroidales S24-7 group), Bacteroides, Christensenellaceae R-7 group, Rikenellaceae RC9 gut group and Ruminococcaceae UCG-005 (Fig. 2C).
482 operational taxonomic units (OTUs) allocated between seven phyla, 36 families and 123 genera contributed to at least 0.1% of the microbial community in at least 5% of the samples. The fat matrix and the addition of the probiotic Lf had multiple effects (a fat matrix effect, PL vs. DL and PL vs. (DL and DL + Lf); a probiotic effect, DL vs. DL + Lf and (DL and PL) vs. DL + Lf; and a synergetic effect, PL vs. DL + Lf) on the composition of the faecal microbiota (Fig. 2D).
Among the 78 discriminating OTUs, 25 (32%) were specific to the lipid matrix whereas 26 (33%) were specific to the probiotic Lf. OTUs within the Firmicutes phylum were most affected by the formula composition (67% of the discriminating OTUs) (Supplementary Table 1). At phylum level, the addition of dairy lipids was associated with increased abundance of Fusobacteria and decreased abundances of Firmicutes and Tenericutes. Probiotic Lf did not have any impact on phyla abundances. At the genus level, 11 genera were affected by the formula composition, among which five belonged to Clostridiales order. The addition of dairy lipids was associated with higher abundances of Hungatella, Enterococcus and Fusobacterium (DL and DL + Lf vs. PL) (Fig. 2D). The addition of Lf stimulated a significant expansion of Lachnospiraceae AC2044 group and reduction of Prevotella 2 and two unknown genera belonging to p-2534-18B5 gut group and Porphyromonadaceae families (DL + Lf vs. PL and DL). Dairy lipids and probiotic Lf addition had synergetic effects on four genera, increasing three genera belonging to Clostridiales order (Peptoclostridium, Flavonifractor and Lachnoclostridium), and decreasing Fibrobacter.
Metabolic activity
Short-chain fatty acid (SCFA) concentration was measured at different time points on faecal samples and at euthanasia on caecal and colon contents. Total faecal SCFA production did not differ with age whatever the formula (p = 0.15) and was similar between groups from weaning until PND77 (Table 2). At PND98, six weeks after the beginning of the high-energy (HE) diet, total faecal SCFAs, acetate and valerate concentrations tended (p = 0.09, 0.07 and 0.07, respectively) to increase and isobutyrate concentration was significantly increased (p = 0.02) in DL + Lf compared to DL. Besides, isovalerate was significantly (p = 0.03) increased in both PL and DL + Lf compared to DL. At PND133, faecal propionate level tended (p = 0.08) to increase in DL + Lf compared to DL. No differences were observed at euthanasia for total SCFAs in caecal and colonic contents (Table 2).
High-resolution 1H-NMR spectra were recorded from faecal contents at PND140. A valid and robust PLS-DA model was constructed (Fig. 2E). The DL + Lf group was clearly separated from the two other groups which shared an overlapping area. Among the 40 identified metabolites, five were statistically different between groups (Fig. 2F). Glycerol was increased in DL and DL + Lf compared to PL, and 2-oxoglutarate and L-lysine were decreased in DL + Lf compared to PL. 5-aminovalerate was decreased in DL + Lf compared to PL and DL whereas propionate was increased in DL + Lf compared to DL.
The correlations between the microbial composition and metabolites were analysed using Spearman correlations (Supplementary Table 2). Several discriminating genera were correlated to each other. For example, Lachnoclostridium was positively associated with Flavonifractor (R = 0.77, p < 0.01) and Hungatella (R = 0.57, p = 0.01) and negatively associated with Lachnospiraceae AC2044 group (R = −0.61, p < 0.01). Several discriminating genera were also correlated to discriminating metabolites or SCFA concentrations. For instance, Hungatella was positively associated with caecal acetate (R = 0.52, p = 0.01) whereas Prevotella 2 was negatively associated with faecal propionate (R = −0.54, p = 0.01) and Peptoclostridium was negatively associated with 5-aminovalerate (R = −0.51, p = 0.01).
Effects of dairy lipids and Lf on adult GLP-1 secretion
Enteroendocrine and glucagon-like peptide-1 (GLP-1)-secreting cell densities, as well as the percentage of GLP-1-secreting cells amongst all enteroendocrine cells were similar between groups in the caecum and the colon (Supplementary Table 3). However, caecal GLP-1, but not colon GLP-1, was increased in DL + Lf pigs compared to PL and DL groups (Fig. 3A,B). In addition, plasma GLP-1 secretory response to meal stimulation was increased in DL + Lf group compared to PL and DL groups (Fig. 3D), although fasting plasma GLP-1 was not different between groups (Fig. 3C). Even though no significant difference was observed between groups for the endocrine pancreas anatomy and its insulin content (Supplementary Table 3), caecal GLP-1 secreting L-cell density was positively correlated to caecal GLP-1, to plasma GLP-1 secretory response to meal stimulation, and to the endocrine pancreas (% endocrine tissue) (Supplementary Table 4). Similarly, colon GLP-1 secreting L-cell density was positively correlated to GLP-1 concentration in the colon and fasting plasma GLP-1 (Supplementary Table 4).
Intestinal endocrine function. (A) Caecal and (B) colon GLP-1 of adult PL, DL and DL + Lf pigs. (C) Fasting plasma GLP-1 and (D) plasma GLP-1 secretory response to meal stimulation of adult PL, DL and DL + Lf pigs. Formulas contained as lipids either: only plant lipids (PL, n = 7–9), a half-half mixture of plant and dairy lipids (DL, n = 6–8) or a half-half mixture of plant and dairy lipids supplemented with Lf (DL + Lf, n = 4–9). Data are shown as Means ± SEM. GLP-1, Glucagon-like peptide-1. A significant diet × sex effect was observed for caecal GLP-1, with the DL + Lf females having a higher concentration than DL and PL females.
Independently of formula composition, free fatty acid receptor (FFAR) 2 expression was significantly increased in the caecum compared to the colon (Fig. 4A) whereas FFAR3 expression was significantly increased in the colon compared to the caecum (Fig. 4B). FFAR3 expressions in both the caecum and the colon were positively correlated to the percentage of GLP-1-secreting cells amongst enteroendocrine cells (Supplementary Table 4).
Relative caecal and colonic expression of FFAR2 (A) and FFAR3 (B) in adult PL, DL and DL + Lf pigs. Formulas contained as lipids either: only plant lipids (PL, n = 9), a half-half mixture of plant and dairy lipids (DL, n = 7) or a half-half mixture of plant and dairy lipids supplemented with Lf (DL + Lf, n = 8). Data represent the ratio of the relative expression of groups to colon PL group (±SEM). FFAR, Free Fatty Acid Receptor.
To investigate the role of the gut microbiota as an actor of nutritional programming, its composition and metabolic activity were correlated to the entero-insular axis-related parameters (Supplementary Table 5 and Fig. 5). Faecal SCFA concentrations measured after six weeks of HE diet were positively correlated to caecal GLP-1 secreting L-cell density (total SCFAs, acetate, isobutyrate), caecal GLP-1 concentration (propionate) and plasma GLP-1 secretory response to meal stimulation (total SCFAs, acetate, butyrate). Plasma GLP-1 secretory response to meal stimulation was also positively correlated to several SCFAs concentrations measured at PND140 (total SCFAs, acetate, propionate, butyrate, isobutyrate, valerate and isovalerate, colon butyrate) and to three discriminating genera (Flavonifractor, Hungatella and Lachnoclostridium). Caecal GLP-1 concentration was positively correlated to butyrate concentration (faeces at PND77 and colon at PND140) (Supplementary Table 5). Lachnospiraceae AC2044 group was positively correlated to colon GLP-1 secreting L-cell density and fasting plasma GLP-1 (Supplementary Table 5).
Significant correlations between gut microbiota composition, luminal metabolites and host entero-insular axis parameters. Within the gut microbiota, the three genera Flavonifractor, Lachnoclostridium and Hungatella were positively correlated. They all belong to the Clostridia class, including SCFA-producing bacteria. These genera were also positively correlated to plasma GLP-1 secretory response to meal stimulation. Plasma GLP-1 secretory response to meal stimulation was, as expected, positively associated with luminal SCFA concentrations, specifically propionate. The percentage of caecal GLP-1 secreting-cells amongst enteroendocrine cells was positively correlated to the caecal GLP-1 content, to the plasma GLP-1 secretory response to meal stimulation and to the pancreatic endocrine tissue, in accordance with the well-known trophic effect of GLP-1 on β cells. These results highlight potential mechanisms between bacteria composition and metabolism, and host metabolism. GLP-1, Glucagon-like peptide-1; SCFA, short-chain fatty acid; C3, propionate. R > |0.50| and p < 0.05. In bold: significant effect of the infant formula composition.
Effects of dairy lipids and Lf on adult metabolic adaptation to a three-months HE diet challenge
Growth, food intake and body composition
Even if formula composition had an impact on weight gain during the suckling period, no significant difference in growth and food intake was observed between the three groups after weaning and throughout the experiment (Supplementary Fig. 1). Three months of HE diet induced a substantial increase in backfat (reflect of adiposity), regardless of the formula received in early life and all groups had similar body and relative organ weights at euthanasia (PND140) (Supplementary Table 6).
Inflammation and metabolic disorders
Three months of HE diet led to a weak inflammation as indicated by a plasma level of haptoglobin slightly above 2.5 g/L. However, the composition of infant formulas received in early life had no significant long-term impact on plasma haptoglobin and TNFα levels, and on caecal and colon alkaline phosphatase activity (Supplementary Table 7). Only plasma IL-1β was higher in DL + Lf compared to PL pigs (Supplementary Table 7). Cytokine secretions (IFNγ, TNFα, IL-10) of peripheral blood mononuclear cells (PBMCs) stimulated with concanavalin A (ConA) mitogen or lipopolysaccharide (LPS) were not significantly different between groups (Supplementary Table 7). Finally, plasma IL-1β was negatively correlated to an unknown genus belonging to the Porphyromonadaceae family (R = −0.53, p = 0.02).
The composition of infant formulas had no significant long-term impact on lipid profile and plasma fasting glucose and insulin concentrations at PND133 (Table 3). Besides, the intravenous glucose tolerance test (IVGTT), aiming at challenging glucose homeostasis, did not reveal significant differences between groups in glucose and insulin profiles as well as in calculated indexes (Table 3 and Supplementary Fig. 2). Even though infant formulas had no significant impact on metabolism and liver sensitivity to insulin with regards to genes related to insulin action (INSR and GLUT2) and PEPCK, the expression of liver glucose-6-phosphatase (G6Pase) was decreased in DL and DL + Lf groups compared to PL (Table 4). In addition, fasting insulin was positively correlated to Lachnospiraceae AC2044 group (R = 0.53, p = 0.02) (Supplementary Table 5). Plasma GLP-1 secretory response to meal stimulation was negatively correlated to INSR expression in the liver (R = −0.66, p = 0.01) (Supplementary Table 4), itself negatively correlated to Flavonifractor (R = −0.52, p = 0.02) (Supplementary Table 5).
Discussion
The objective of this study was to investigate the potential long-term programming effect of early infant formula consumption containing dairy lipids with or without probiotic Lf on gut microbiota and host entero-insular axis and metabolism. Our hypothesis was that early consumption of dairy lipids and Lf may program gut microbiota composition and metabolism and therefore affect the long-term response of adult pigs facing a HE diet. The HE diet was administered to highlight a potential latent programming effect of the formula composition. Our results demonstrated that both the addition of dairy lipids and probiotic Lf in infant formulas had an impact on gut microbiota composition. However, only the addition of Lf had a noticeable long-term promoting effect on SCFA production (especially propionate) and GLP-1 secreting function.
The first and main result of this study is that the composition of the infant formula fed in early life had a long-term programming effect on gut microbiota composition (from phylum to genus) and metabolic activity. The nutritional intervention limited to the suckling period was sufficient to permanently affect gut microbiota. The infant formula composition impacted genera mainly belonging to Clostridia class (Lachnospiraceae, Ruminococcaceae and Peptostreptococcaceae families): Hungatella, Lachnospiraceae AC2044 group, Peptoclostridium, Flavonifractor and Lachnoclostridium were increased in DL and/or DL + Lf pigs compared to PL pigs. In our study, strong positive correlations (R > 0.70, p ≤ 0.01) were observed between OTUs, especially between the ones belonging to Ruminococcaceae and Lachnospiraceae families. These are the most abundant families from the phylum Firmicutes (and order Clostridiales) found in the mammalian gut environment: 10–20% of the total bacteria in the human colon26 and 18.7% in faecal samples from weaned pigs27. The dominant genus in the large intestine of six-month-old pigs also belongs to Clostridiales28. Lachnospiraceae and Ruminococcaceae have been associated with gut health maintenance through SCFA production29. An increase in Clostridia class (butyrate-producing bacteria) was associated with a protective microbiota and with gut function and immune homeostasis maintenance in both mice and humans30,31,32 although no change in immune parameters was observed in our study. In addition, Clostridia class was shown to be significantly decreased in diabetic patients33,34 and in rats fed a high-fat high-sucrose diet35 although some studies demonstrated an enhancement of Clostridiales order in response to an adverse diet or obesity36,37,38. However, in our study, increased abundance of these genera in DL and/or DL + Lf pigs compared to PL pigs had no consequences on glucose tolerance after three months of HE diet challenge. All three groups of animals displayed an insulin-resistance state (elevated HOMA-IR values) and an impaired glucose tolerance (IVGTT) but not diabetes, as demonstrated by fasting glycaemia within the physiological range.
The increase of Lachnoclostridium in DL + Lf compared to PL pigs is of particular interest since it can ferment mono- and disaccharides and produce SCFAs39. Some members of Lachnoclostridium were found to be close to Clostridium lactatifermentans, a strain that ferment lactate to SCFAs (mainly acetate and propionate and also traces of butyrate and isovalerate)40 and Lachnoclostridium abundance was decreased in mice fed a high-fat diet41. In our study, Lachnoclostridium was positively correlated with Hungatella and Flavonifractor (Fig. 5). Lachnoclostridium was positively associated with caecal isovalerate and Hungatella with caecal total SCFAs (R = 0.46) and acetate. SCFA production was also increased in DL + Lf compared to DL after six weeks (PND98, total SCFAs, acetate, isobutyrate, valerate and isovalerate) and three months (PND133 and 140 for faecal propionate) of HE diet. The infant formula containing dairy lipids and Lf compared to PL was therefore associated with both a long-term increased abundance of genera belonging to Clostridia class and an increased SCFAs concentration. These results suggest a beneficial effect on microbiota composition and metabolic activity since a decreased SCFA production has generally been associated with obesity42 and adverse diets43,44.
The infant formula composition also affected the faecal concentration of 4 metabolites: lysine, 5-aminovalerate, 2-oxoglutarate and glycerol. Lysine, 5-aminovalerate and 2-oxoglutarate were decreased in DL + Lf compared to PL (and also compared to DL for 5-aminovalerate) whereas glycerol was increased in DL and DL + Lf compared to PL. 5-aminovalerate is thought to be a metabolite of protein catabolism (such as lysine) by anaerobic bacteria45,46. It can further be degraded to acetate, propionate, ammonia and valerate and is a substrate of 4-aminobutyrate-2-oxoglutarate aminotransferase. Higher expression of microbial genes involved in the metabolism of essential amino acids (lysine) was observed in mice transplanted with obese twin microbiomes42 suggesting that such an increased microbial protein metabolism was associated with altered host health. Thus, in our study, the decrease in lysine and 5-aminovalerate in DL ± Lf would be beneficial for the host. Besides, 2-oxoglutarate, an intermediate of the Krebs cycle, was decreased in the urine of rats fed a high-fat/high-sucrose diet47 and negatively correlated with the faecal abundance of SCFAs in control rats48 suggesting once again that its decrease in DL + Lf was consistent with the increased SCFA production. The colonic level of glycerol, a trihydroxy sugar alcohol intermediate in carbohydrate and lipid metabolism, is partly dependent on in situ microbial synthesis and fermentation processes. Indeed, some fermentation products such as butyrate may stimulate glycerol absorption49 whereas several alcohols such as 1,3-propanediol inhibit it50. It was demonstrated that a slower glycerol metabolism resulted in higher levels of propionate51 which seems in agreement with the increased levels of glycerol and propionate observed in DL + Lf. Even if the interpretation of modulation of gut microbiota is not an easy task due to substantial discordance between studies owing to the population or animal model chosen, the methods used and the hypervariable region of the 16S gene examined, all the effects observed in our study suggest a beneficial effect of the addition of dairy lipids with Lf in infant formula on later gut microbiota composition (increased abundance of Clostridiales genera) and metabolism (increased SCFAs and decreased oxoglutarate).
Gut microbiota has been associated with several host metabolic pathways such as the stimulation of the intestinal endocrine function, mainly via SCFAs52,53,54. In our study, the three discriminating genera, Flavonifractor, Lachnoclostridium and Hungatella, increased in DL + Lf compared to PL, were positively correlated to plasma GLP-1 secretory response to meal stimulation, suggesting that they indeed participated to enhanced intestinal endocrine function observed in DL + Lf animals (Fig. 5). Besides, Lachnospiraceae AC2044 genus (more abundant in DL + Lf than in PL and DL) was positively correlated to colonic GLP-1 density, fasting plasma GLP-1 and fasting insulinemia. Plasma GLP-1 secretory response to meal stimulation was, as expected, positively associated with all SCFA concentrations measured at PND133 (including acetate, propionate and butyrate). These results highlight potential mechanisms between bacteria composition and metabolism, and host metabolism. Among SCFAs, propionate, which was constantly increased in DL + Lf, would be the most potent agonist of FFAR2 and FFAR3 for GLP-1 secretion55. FFAR3 expression in both the caecum and the colon was positively correlated to the percentage of caecal GLP-1 secreting-cells amongst enteroendocrine cells suggesting mechanistic links between SCFAs, FFAR expression and intestinal endocrine function.
Accordingly, caecal GLP-1 concentration (basal condition) and plasma secretory response to meal stimulation (challenged condition) were significantly increased in DL + Lf group compared to PL and DL groups. This reflects an improved enteroendocrine cell function in DL + Lf animals, which may help them to better cope with the HE-diet challenge. Indeed, obesity and metabolic changes occurring with the development of type-2 diabetes have generally been associated in humans with an impaired postprandial secretion of GLP-156. In addition, high-fat diet consumption has been associated with a decreased number of GLP-1 positive cells in the colon of mice, and with a decreased response of primary small intestinal L-cell cultures to nutrient stimuli, suggesting an impairment of the enteroendocrine cell function by a chronic high-fat diet57.
An improved intestinal endocrine function may be beneficial to glucose homeostasis by stimulating insulin secretion and preventing glucose intolerance56. Even if the infant formula composition had no long-term effect on global metabolism and glucose tolerance in our study, caecal GLP-1 L-cell density was positively correlated to the pancreatic endocrine tissue in accordance with the well-known trophic effect of GLP-1 on β cells58 (Fig. 5). Furthermore, GLP-1 secretory response to meal stimulation was negatively correlated to INSR expression in the liver, reinforcing the hypothesis that increased GLP-1 secretion may be an adaptive mechanism to slow down the establishment of glucose intolerance59. Besides, the decreased hepatic expression of neoglucogenic enzyme G6P in DL and DL + Lf liver suggests better insulin sensitivity in these groups compared to PL. Several hypotheses can be made to explain the lack of clear long-term outcomes of infant formula composition on metabolism. The objective of the HE diet provided in adulthood was to challenge the metabolic adaptation capabilities of the animals, to reveal a potential programming effect of the infant formula composition. Our model was successful, since we were able to demonstrate a programming effect on microbiota and its metabolism, as well as on the intestinal endocrine function but the challenge duration might have been too short to reveal their consequences on metabolism adaptation to the HE diet. Besides, pigs received the HE diet during puberty which could have challenged more drastically their metabolism60. It is however plausible that their increase in GLP-1 secreting capacity would help DL + Lf pigs to metabolically cope with the HE diet on a longer term.
To conclude, our study highlighted a programming effect of the infant formula composition, i.e. the fat matrix and the addition of probiotic Lf, on gut microbiota composition and metabolism. However, only the addition of Lf had a long-term promoting effect on the growth of intestinal SCFA-producing bacteria from the Clostridiales order (such as Lachnoclostridium) accompanied by an increase in SCFA production (especially propionate) and GLP-1 secreting function under a HE-diet challenge (Fig. 5). The addition of dairy lipids and Lf in infant formulas would, therefore, be safe on the long-term and may represent a clever way to better approach the long-term physiological effects of breast milk.
Methods
Ethical Approval
The present study was designed and conducted in compliance with the current ethical standards of the European and French guidelines (directive 2010/63/EU and decree 2013-118, respectively). The local Ethics Committee CREEA (Rennes Committee of Ethics in Animal Experimentation) approved the entire protocol (authorization #2016011111546978). Animals were observed daily throughout the experimental protocol to ensure their welfare and they received no medication or antibiotic treatment.
Animals and study design
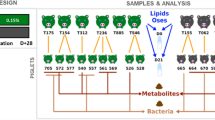
A total of 27 female and male Yucatan piglets (Saint-Gilles, France) were used in three replications. One animal was excluded because of medical issues. Piglets were separated from their dam at PND2 and housed in individual stainless-steel metabolic cages. They were fed one of the three experimental formulas with an automatic milk feeder as previously described61 until weaning, at PND28. To account for litter-to-litter variation, three piglets with a body weight (BW) close to the mean birth weight of the litter were selected from each litter and assigned to one of the three formulas. Allocation to formulas was balanced between groups for birth weight, BW at PND2 and sex. Birth weight was similar in all groups (791 ± 32 g). Formulas were rehydrated each day at 20% dry extract in water before distribution. The ration was allocated in ten meals automatically distributed during the day. BW was measured twice a week and feeding schedules were adjusted accordingly. The daily net energy offered was 1450 kJ/kg BW0.75. Formula intake was automatically recorded daily. At weaning, pigs were fed commercial starter diets (Cooperl Arc Atlantique) until PND56 and then a HE diet until euthanasia at PND140 (Fig. 1). The HE diet was formulated to provide 37.8% of digestible energy from lipids and 45.2% from carbohydrates -mainly sucrose-, to mimic an obesogenic environment (vs. 2.2% from lipids and 75.2% from carbohydrates -mainly fibers- in a standard minipig diet). Pigs had ad libitum access to water and food from PND28 to PND140. Solid food intake was recorded weekly (and daily one week before and two weeks after the administration of the HE diet) and BW was measured weekly. At PND130, plasma lipid profile was analysed and glucose tolerance was assessed in vivo by an intravenous glucose tolerance test (IVGTT). Pigs were then euthanized at PND140, after 12 weeks of HE diet, and tissues collected and weighed.
Diets
The three infant formulas were separately manufactured by Lactalis (Retiers, France) under the same conditions, using the same raw materials except for some plant oils and dairy lipids that were different between the PL formula and the two formulas that contained dairy lipids (DL and DL + Lf). Probiotic Lf was added at the end of the process, directly in the dry powder. Each infant formula was manufactured in one batch before the start of the experiment and was used for the three replications. The formulas were adapted to meet piglet energy and protein requirements. The three formulas had the same energy, protein, lipid and carbohydrate levels. They differed by the fat origin [only plant lipids (PL) vs. half-half plant lipids and dairy lipids (DL) and the supplementation with Lf (DL + Lf)] (Table 1). Compared to a standard infant formula, the experimental formulas contained higher amounts of proteins and lipids and a lower amount of lactose. However, lipids:proteins and linoleic acid:α-linolenic acid (ω6:ω3 = 6) ratios were kept similar to those found in infant formulas. The formulas were made of a mixture of skim milk powder to reach a casein:whey proteins ratio of 30:70 w/w. The HE diet, used to reveal a nutritional programming, contained 20% fat and 20% sucrose (Supplementary Table 8).
Body composition
The adiposity was evaluated with the back fat thickness before the administration of the HE diet (PND56) and at the end of the experiment (PND140). Back fat thickness was measured ultrasonically (Sonolayer SAL-32B, Toshiba, Tokyo, Japan) on both sides at the P2-position.
Faecal sample collection
Faeces were collected at different stages of development: at PND 33, 55, 61, 77, 98 and 133 to determine the fermentative activity of the gut microbiota (Fig. 1). They were mixed with 0.5% ortho-phosphoric acid solution, centrifuged at 1700 g for 15 min at 4 °C and supernatants collected and stored at −20 °C until further analysis of SCFA content by gas chromatography62.
Blood sampling and intravenous glucose tolerance test (IVGTT)
At PND125, a catheter was inserted under general anaesthesia into one external jugular vein of the pigs. Blood was collected in 9 mL heparin-coated tubes before culture of the PBMCs, or in tubes containing K2-EDTA (0.8 M, 10 µL/mL of blood) to assay plasma cytokines. The IVGTT was performed two to five days after surgery, after an overnight fast. It consisted in multiple blood samplings before (t = −15, 0 min, basal samples) and after (t = 2, 4, 6, 8, 10, 15, 20, 25, 30, 40, 50, 60 and 75 min) an intravenous glucose injection (0.3 g/kg BW). After the IVGTT, a meal test was performed. 100 g of the HE diet were given to pigs and blood samples were collected 30 min after the meal. Blood was collected in tubes containing K2-EDTA for glucose, insulin, inflammatory markers and lipid profile and in tubes containing K2-EDTA plus an anti-dipeptidylpeptidase-IV (DPP-IV, 10 µL/mL of blood) for GLP-1 (Millipore, Billerica, MA, USA). Tubes containing “Choay” heparin were used for LPS assay. After centrifugation (10 min, 2500 g, 4 °C), plasma samples for glucose, insulin, inflammatory markers and lipid assays were stored at −20 °C and the ones for GLP-1 assay at −80 °C. Glucose and insulin assays were performed on all plasma samples whereas haptoglobin and lipid profile were only performed on basal samples. GLP-1 assays were performed on basal samples and 30 minutes after the meal test.
Post Mortem Tissue Collection
At PND140, after an overnight fast, pigs were euthanized in the experimental slaughterhouse by electrical stunning immediately followed by exsanguination. Liver, pancreas, visceral adipose tissue, caecum and colon (full and empty) were weighed. Digestive contents from caecum, colon and faeces were collected, weighed, and 200–300 mg were immediately frozen in liquid nitrogen and stored at −80 °C for microbiota analysis and metabolomics. In addition digestive contents were mixed with 1 ml of 0.5% ortho-phosphoric acid solution per g of digesta for quantification of SCFAs. Around 1 g of tissues were frozen in liquid nitrogen and stored at −80 °C for further analysis. Adjacent pieces of caecum and colon were fixed in 4% paraformaldehyde for immunohistochemistry. For determination of gene expression, liver, caecal and colon samples were kept in a RNA later solution (Thermo Fisher Scientific, Villebon-sur-Yvette, France) for 24 h at 4 °C and stored at −20 °C. One cm3 sample of the body of pancreas was directly frozen and stored at −80 °C for insulin extraction and another cm3 was fixed in 4% paraformaldehyde for immunohistochemistry. Finally, caecal and colon tissues were collected, rinsed with cold PBS and stored at −80 °C until GLP-1 extraction and assay.
Faecal microbiota analyses
DNA extraction and sequencing
Extraction of total bacterial DNA from rectal contents was performed using Guanidium Thiocyanate and the mechanical bead-beating disruption method as previously described63. DNA concentration was determined with DeNovix spectrophotometer (Wilmington, USA). The V3-V4 region of the 16S rRNA gene was amplified for 30 amplification cycles with an annealing temperature of 65 °C by using the primers F343 (CTTTCCCTACACGACGCTCTTCCGATCTACGGRAGGCAGCAG) and R784 (GGAGTTCAGACGTGTGCTCTTCCGATCTTACCAGGGTATCTAATCCT). The amplicon lengths were about 450 bp (depending on the species). Because MiSeq sequencing enables paired 250 bp reads, the ends of each read overlap and can be stitched together to generate extremely high-quality, full-length reads covering the entire V3-V4 region. Single multiplexing was performed using a home-made 6 bp index, which was added during a second PCR with 12 cycles using the forward primer (AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGAC) and the modified reverse primer (CAAGCAGAAGACGGCATACGAGAT-index-GTGACTGGAGTTCAGACGTGT). The resulting PCR products were purified and loaded onto the Illumina MiSeq cartridge according to the manufacturer instructions. The quality of the run was checked internally using PhiX, and for further analysis, each paired sequence was assigned to its sample using the previously integrated index.
Bioinformatics and statistical analysis
The 16rRNA raw sequences were analysed using the bioinformatic pipeline FROGS (Find Rapidly OTU with Galaxy Solution)64. After performing the quality control depletions, 458,295 total sequences (out of the 674,981 initial sequences) with a mean of 20,831 sequences per sample were used for the analyses. Reads were classified into 909 OTUs. OTUs were assigned using the silva123 16S database. Multi-affiliation was manually checked: all sequences with a low pintail quality in the silva database were removed; a single affiliation was accepted when corresponding to more than 90% of all OTU sequences. The phylogenetic tree was constructed using Phy_tree. After multi-affiliation management, the resulting OTU table (unrarefied raw data) was used for subsequent statistical analyses. Data were filtered to keep only those that contributed to at least 0.1% of the microbial community in at least 5% of the samples. Differences in phyla, genera and OTUs, were assessed with pairwise comparisons, after aggregation at the desired taxonomic rank (phyloseq’ tax_glom function) by using EdgeR package (Bioconductor)65, correcting for a sex effect. Multiple testing corrections (False Discovery Rate) were used to avoid false positives (significance threshold = 0.05).
To investigate within- and between-sample diversities (α and β, respectively), all samples were rarefied to the same depth (15,000 sequences) before analysis with phyloseq R package. α diversity was studied using species richness, Chao1, Simpson index, Shannon index, and inverse Simpson index. Significant differences between formulas and their interactions with sex were assessed using ANOVA (aov function). Compositional β diversities were studied using Jaccard and Bray-Curtis distances whereas phylogenetic β diversities were studied using Unifrac and Weighted Unifrac distances. Group differences in β diversity measurements were evaluated with principal co-ordinate analysis (PCoA) and permutational multivariate analysis (PERMANOVA) of variance using distance matrices. Significance was assessed with adonis and anosim statistical tests (vegan package of R, infant formula composition × sex model) to evaluate the distances at 9999 permutations between groups.
Metabolomics
Sample preparation and 1H-Nuclear Magnetic Resonance (NMR) spectroscopy
Faecal water extraction protocol was adapted from Zhao et al.66. Briefly, 500 µL of phosphate buffer (pH 7.0) prepared in D2O were added to 100 mg of rectal content, homogenized using a FastPrep (MP Biomedicals), centrifuged at 15,000 g for 10 min at 4 °C and supernatants were kept aside. 500 µL of phosphate buffer (pH 7.0) prepared in D2O were added to the pellets and processed as above. The two supernatants were pooled and centrifuged (15 min, 4 °C, 15,000 g) before being transferred into 5 mm RMN tubes. All 1H-NMR spectra were obtained on a Bruker Avance III HD NMR spectrometer operating at 600.13 MHz for 1H resonance frequency using an inverse detection 5 mm 1H-13C-15N-31P cryoprobe attached to a CryoPlatform (the preamplifier cooling unit). 1H NMR spectra were acquired at 300 K using the Carr-Purcell-Meiboom-Gill (CPMG) spin-echo pulse sequence with presaturation, with a total spin-echo delay of 240 ms to attenuate broad signals from proteins and lipoproteins for faecal samples. A total of 512 scans were collected, using a spectral width of 20 ppm and a relaxation delay of 2 s. All spectra were Fourier transformed, manually phased, baseline corrected and referenced to TMSP using Bruker TopSpin 3.5 software (Bruker, GMBH, Karlsruhe, Germany). The spectral assignment was based on matching data to reference spectra in a home-made reference database.
Data reduction and multivariate statistical analysis
Data were reduced using the AMIX software (version 3.9.11, Bruker, Rheinstetten, Germany) to integrate 0.01 ppm regions corresponding to the δ 9–0.5 ppm region. To account for differences in sample amount, each integrated region was normalized to the total spectral area.
SIMCA-P + software (V13, Umetrics AB, Umea, Sweden) was used to perform the multivariate analyses. Before multivariate analysis, Orthogonal Signal Correction (OSC) filtering67 was used to remove variation not linked to the infant formula composition. Filtered data were mean-centered and Pareto scaled. Principal Component Analysis was firstly applied to detect outliers and intrinsic clusters. Then, Partial Least Squares – Discriminant Analysis (PLS-DA) was used to assess the relationship between diet (infant formula) and metabolomic profiles. PLS-DA is a supervised method that maximizes the separation between groups. The R2 (percentage of explained variance) and Q2 (predictive capacity of the model) criteria were used to assess model performance. Models were considered to be valid and robust when R2 (percentage of explained variance) > 50% and Q2 (predictive capacity of the model) > 40%68. Permutation tests involving 200 iterations were also used to assess the robustness of the models. The number of latent variables and Q² value were calculated using seven-fold cross validation. Discriminant variables were determined using VIP (Variable Importance in the Projection) and Kruskal-Wallis test was applied to assess which variables were significantly different. Multiple testing corrections (False Discovery Rate) were used to avoid false positives (significance threshold = 0.05).
Immunohistochemical analysis
Fresh caecum, colon and body of the endocrine pancreas samples were fixed in 4% paraformaldehyde for 72 h at room temperature. They were then placed at 4 °C in PBS containing 30% sucrose and embedded in the Tissue-Tek Optimum Cutting Temperature compound (Sakura Finetek Europe B. V., Zoeterwoude, The Netherlands), frozen in isopentane and sectioned (10 μm for pancreas and 7 µm for caecum and colon) using a cryostat-microtome. Caecum and colon sections (n = 6 per group) were rehydrated with PBS and incubated with 4% normal horse serum, Triton X-100 (0.5%) and sodium azide (0.1%) in PBS for 30 min at room temperature. Sections were then exposed overnight to a rabbit anti-GLP-1 antibody (1:200; Abcam, Cambridge, UK). After washing with PBS, they were incubated with an anti-rabbit antibody conjugated with Alexa Fluor 555 (1:500; Cell Signaling Technology, Leiden, The Netherland) for 2.5 h. Sections were then washed with PBS, incubated with a mouse anti-Chromogranin A antibody (1:1000; Abcam, Cambridge, UK) for 3 h, washed with PBS, incubated with an anti-mouse antibody conjugated with Alexa Fluor 488 (1:500; Cell Signaling Technology, Leiden, The Netherlands) for 2.5 h, washed again with PBS and cover-slipped with Vectashield. Sections were examined by using a fluorescence microscope (Eclipse E400, Nikon Instruments France, Champigny-Sur-Marne, France) attached to a digital camera (Digital Still DXM 1200, Nikon Instruments France). The whole section of the tissue was scanned using digital slide scanner (Nanozoomer 2.0-RS, Hamamatsu Photonics France SARL, Massy, France). Recorded files were analysed using the software NDPI (Hamamatsu Photonics) for determination of the number of enteroendocrine (chromogranin A-labelled) cells and GLP-1-secreting cells per area of mucosa. Immunohistochemical analysis of pancreas (n = 7 per group) was processed as previously described69 to obtain the percentage of endocrine tissue and the number and diameter of islets.
Hormone, glucose, lipid and inflammatory marker assays
Plasma glucose, non-esterified fatty acids (NEFA), triglycerides, total cholesterol, High-density lipoprotein (HDL)-cholesterol and haptoglobin were assessed by an automated spectrophotometric method (Konelab 20i, Thermo Fisher Scientific, Illkirsh, France) using specific commercial kits (Biomérieux, Bruz, France). The intra-assay coefficient of variation was <5%.
GLP-1 content was extracted from caecum and colon by homogenisation (Polytron 3100, Kinematica, 25,000 rpm, 2 × 20 s) of 1 g of tissue in 5 ml of ethanol acid solution (1% HCl 12 M, 74% absolute ethanol, 25% H2O). After 24 h at 4 °C, samples were centrifuged (20 min, 2000 g, 4 °C) and supernatants diluted (1:300 and 1:250 for caecum and colon, respectively). Intestinal and plasma GLP-1 concentrations were measured using a GLP-1 active ELISA kit (Millipore). Non-responding pigs to meal-stimulation were removed from the secretory response analysis.
Insulin content was extracted from the pancreas by homogenisation in 10 mL of an ethanol acid solution (75% absolute ethanol, 23.5% ultrapure H2O, 1.5% HCl 12 N) (Polytron 3100, Kinematica, 25,000 rpm, 2 × 20 s). After an overnight storage at −20 °C, samples were centrifuged (30 min, 190 g, 4 °C) and supernatants stored at −20 °C. Ethanol acid solution (10 mL) was added to pellets for a second extraction, stored overnight at −20 °C and centrifuged (30 min, 190 g, 4 °C). Supernatants were collected and pooled to the ones from the first extraction, diluted in a PBS/BSA solution (1:3000) and pancreas insulin concentration and plasma insulin were measured by a radioimmunoassay (RIA) method, using iodinated porcine insulin as a tracer (INSULIN-CT, CisbioInternational, Gif sur Yvette, France). The intra- and inter-assay CV were 15 and 11% respectively for a concentration of 35 µIU/mL.
Plasma cytokines (IL-1β and TNFα) were measured using Cloud-Clone Corp kits (USA, SEA563Po and SEA133Po, respectively) according to manufacturer’s instructions.
Gene expression in liver, caecum and colon tissues
Total RNA extraction, integrity and quantitative RT-PCR
Total RNA was isolated from liver, caecal and colon samples stored in RNA later solution according to the TRIzol method (Invitrogen, France). Extracted RNAs were quantified using a DeNovix spectrophotometer (Wilmington, USA). The RNA quality and integrity were confirmed using the Agilent RNA 6000 Nano kit utilizing Agilent 2100 Bioanalyzer (Agilent Technologies France, Massy, France). After a DNase treatment (Ambion), cDNA was prepared by reverse transcription of 2 μg total RNA using a High Capacity Complementary DNA Reverse Transcription Kit according to the manufacturer’s instructions (Applied Biosystems). Quantitative real-time PCR was performed on a StepOnePlus real-time PCR machine using Fast SyberGreen master mix (Applied Biosystems) for detection.
RPL4, GAPDH and YWHAZ were used as housekeeping genes in the liver while TBP and ACTB were used as housekeeping genes in the caecum and colon according to their stability (Supplementary Table 9). Relative expressions of the target genes were determined using the 2−ΔΔCt method. Target genes in the liver were genes encoding gluconeogenic enzymes: G6Pase and PEPCK (phosphoenolpyruvate carboxykinase), and genes related to insulin action: GLUT2 (glucose transporter 2) and INSR (insulin receptor). Target genes in the caecum and the colon were FFAR 2 and 3.
Insulin resistance and minimal model
Insulin resistance was evaluated using the Homeostasis Model Assessment Insulin Resistance Index (HOMA-IR = (G0 (mmol.L−1) × I0 (µIU.mL−1)/22.5) where G0 and I0 are fasting glycemia and insulinemia, respectively. Glucose tolerance was estimated through the following indexes: the incremental area under the curve (AUC) over 30 and 75 min for glucose and the rate of glucose disappearance (KG, slope of the function f(t) = Ln(G), in %/min). Beta-cell secretory capacity was estimated through the calculation of the AUC over 30 and 75 min for insulin, the acute insulin response (AIR, mean insulin concentration above basal values for the first 6 min) and the insulinogenic index (AUCinsulin/AUCglucose) at 30 and 75 min.
The glucose and insulin responses were integrated using the minimal model described by Bergman70 to obtain the glucose efficiency index SG and the insulin sensitivity index SI (SAAM Institute Inc, Seattle, WA, USA). The S2 index, described to be well-correlated to the insulin sensitivity index obtained by the gold standard euglycemic hyperinsulinemic clamp71 was also calculated.
Alkaline phosphatase activity in caecum and colon
Bioactive phosphatase alkaline concentration was determined in caecum and colon with a commercial kit (Sensolyte®, Anaspec, San Jose, USA) and expressed per mg of soluble protein.
Culture and stimulation of PBMCs
Peripheral blood was diluted with an equal volume of Hank’s Balanced Saline Solution (HBSS) supplemented with 2% foetal calf serum (FCS), 100 IU/mL penicillin and 100 µg/mL streptomycine sulphate. Diluted blood (6 mL) was then layered over 2.5 mL of Ficoll-Hypaque (Histopaque-1077, Sigma, #10771) and centrifuged at 400 g for 30 min at room temperature without brake. Mononuclear cells were collected at the interface between Ficoll and plasma. HBSS-FCS was added to mononuclear cells up to a total volume of 50 mL and centrifuged 10 min at 600 g and 4 °C. Then the pellet was suspended in 10 mL of room temperature ACK lysing buffer to lyse red blood cells, if any. After 5 min, reaction was stopped by adding 10 mL of ice-cold complete RPMI medium (supplemented with 10% of FCS). PBMCs (1.25 × 106 cells per well in flat-bottomed 96 well plates) were cultured for 72 h at 37 °C under an atmosphere containing 5% CO2. Three different conditions of culture were performed: unstimulated condition (only with RPMI complete medium), in the presence of 5 µg/ml ConA (Sigma, C5275, Saint Quentin Fallavier, France), or in the presence of 10 µg/ml LPS-EB ultrapure from Escherichia coli 0111:B4 strain (tlrl-3pelps, InvivoGen, San Diego, USA). Finally, supernatants were collected and frozen at −20 °C for later cytokine analysis by ELISA. Indeed, IL-10, IFNγ and TNFα were measured in culture supernatants of PBMCs using capture sandwich ELISA (R&D Systems, USA: DY693B, DY985 and DY690B, respectively). Secretion levels (pg/mL) were obtained by subtracting basal condition level to stimulated condition levels.
Statistical analysis
Statistical analyses of microbiota and metabolomics data were described above. For all other data, statistical analyses were performed using R Core Team (2016; R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria; http://www.R-project.org/). Normality was tested with Shapiro and Wilk test. For parameters with unequal variances between groups, Box-Cox transformations were used. Differences between groups were assessed using a two-way ANOVA (lm function) testing formula, replication, sex and interactions between diet and sex, and between diet and replication, followed by Tukey post-hoc test (Tukey HSD).
BW, energy intake, glucose and insulin responses to IVGTT, FFAR expression across sites were subjected to repeated measures ANOVA (lme function) including diet, time (age or min after glucose injection), site (caecum and colon for FFAR expression only), replication, sex and interactions between diet and all other factors, followed by Tukey post-hoc test (lsmeans package). Sex effect and interaction diet × sex were not significant unless otherwise mentioned.
Spearman’s r-test was used to evaluate correlations between parameters for all groups. Correlations were considered when p < 0.05 and |R| > 0.5.
Data are presented as mean values with their standard errors (SEM). Differences were considered significant at p < 0.05 and a trend at p < 0.1.
Data Availability
The datasets generated during and/or analysed during the current study, which are not already included in this published article (and its Supplementary Information files), are available from the corresponding author on reasonable request.
References
Barker, D. J. The origins of the developmental origins theory. J. Intern. Med. 261, 412–417, https://doi.org/10.1111/j.1365-2796.2007.01809.x (2007).
Mcmillen, I. C. & Robinson, J. S. Developmental Origins of the Metabolic Syndrome: Prediction, Plasticity, and Programming. Physiological Reviews 85, 571–633, https://doi.org/10.1152/physrev.00053.2003 (2005).
Suzuki, K. The developing world of DOHaD. J. Dev. Orig. Health Dis. 1–4, https://doi.org/10.1017/s2040174417000691 (2017).
Cox, L. M. et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 158, 705–721, https://doi.org/10.1016/j.cell.2014.05.052 (2014).
Cani, P. D. et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 57, 1470–1481, https://doi.org/10.2337/db07-1403 (2008).
Wang, M., Ivanov, I., Davidson, L. A., Chapkin, R. S. & Donovan, S. M. In Nutrigenomics and Proteomics in Health and Disease. 220–257 (John Wiley & Sons, Ltd, 2017).
Nauta, A. J., Ben Amor, K., Knol, J., Garssen, J. & van der Beek, E. M. Relevance of pre- and postnatal nutrition to development and interplay between the microbiota and metabolic and immune systems. The American journal of clinical nutrition 98, 586S–593S, https://doi.org/10.3945/ajcn.112.039644 (2013).
Thompson, A. L. Developmental origins of obesity: early feeding environments, infant growth, and the intestinal microbiome. Am. J. Hum. Biol. 24, 350–360, https://doi.org/10.1002/ajhb.22254 (2012).
Chatelais, L. et al. The level of protein in milk formula modifies ileal sensitivity to LPS later in life in a piglet model. PLoS One 6, e19594, https://doi.org/10.1371/journal.pone.0019594 (2011).
Davis, E. C., Wang, M. & Donovan, S. M. The role of early life nutrition in the establishment of gastrointestinal microbial composition and function. Gut Microbes, 1–29 https://doi.org/10.1080/19490976.2016.1278104 (2017).
Horta, B. L., Loret de Mola, C. & Victora, C. G. Long-term consequences of breastfeeding on cholesterol, obesity, systolic blood pressure and type 2 diabetes: a systematic review and meta-analysis. Acta Paediatr. 104, 30–37, https://doi.org/10.1111/apa.13133 (2015).
Hui, L. L. et al. The association of breastfeeding with insulin resistance at 17 years: Prospective observations from Hong Kong’s “Children of 1997” birth cohort. Matern. Child Nutr. https://doi.org/10.1111/mcn.12490 (2017).
Nejrup, R. G., Licht, T. R. & Hellgren, L. I. Fatty acid composition and phospholipid types used in infant formulas modifies the establishment of human gut bacteria in germ-free mice. Sci. Rep. 7, 3975, https://doi.org/10.1038/s41598-017-04298-0 (2017).
Lopez, C., Cauty, C. & Guyomarc’h, F. Organization of lipids in milks, infant milk formulas and various dairy products: role of technological processes and potential impacts. Dairy Sci. Technol. 95, 863–893, https://doi.org/10.1007/s13594-015-0263-0 (2015).
Le Huerou-Luron, I. et al. A mixture of milk and vegetable lipids in infant formula changes gut digestion, mucosal immunity and microbiota composition in neonatal piglets. Eur. J. Nutr. 57, 463–476, https://doi.org/10.1007/s00394-016-1329-3 (2018).
Baars, A. et al. Milk fat globule membrane coating of large lipid droplets in the diet of young mice prevents body fat accumulation in adulthood. Br. J. Nutr. 115, 1930–1937, https://doi.org/10.1017/s0007114516001082 (2016).
Jost, T., Lacroix, C., Braegger, C. & Chassard, C. Impact of human milk bacteria and oligosaccharides on neonatal gut microbiota establishment and gut health. Nutr. Rev. 73, 426–437, https://doi.org/10.1093/nutrit/nuu016 (2015).
Cabrera-Rubio, R. et al. The human milk microbiome changes over lactation and is shaped by maternal weight and mode of delivery. Am. J. Clin. Nutr. 96, 544–551, https://doi.org/10.3945/ajcn.112.037382 (2012).
Murphy, K. et al. The Composition of Human Milk and Infant Faecal Microbiota Over the First Three Months of Life: A Pilot Study. Sci. Rep. 7, 40597, https://doi.org/10.1038/srep40597 (2017).
Pannaraj, P. S. et al. Association between breast milk bacterial communities and establishment and development of the infant gut microbiome. JAMA Pediatrics, https://doi.org/10.1001/jamapediatrics.2017.0378 (2017).
Martin, R. et al. Probiotic potential of 3 Lactobacilli strains isolated from breast milk. J. Hum. Lact. 21, 8–17; quiz 18–21, 41 https://doi.org/10.1177/0890334404272393 (2005).
Maldonado-Lobon, J. A. et al. Long-term safety of early consumption of Lactobacillus fermentum CECT5716: A 3-year follow-up of a randomized controlled trial. Pharmacol. Res. https://doi.org/10.1016/j.phrs.2015.01.006 (2015).
Lopez-Huertas, E. Safety and efficacy of human breast milk Lactobacillus fermentum CECT 5716. A mini-review of studies with infant formulae. Benef. Microbes 6, 219–224, https://doi.org/10.3920/bm2014.0091 (2015).
Vanhaecke, T. et al. L. fermentum CECT 5716 prevents stress-induced intestinal barrier dysfunction in newborn rats. Neurogastroenterol. Motil. 29, https://doi.org/10.1111/nmo.13069 (2017).
Roura, E. et al. Critical review evaluating the pig as a model for human nutritional physiology. Nutr. Res. Rev. 29, 60–90, https://doi.org/10.1017/s0954422416000020 (2016).
Vital, M., Howe, A. C. & Tiedje, J. M. Revealing the Bacterial Butyrate Synthesis Pathways by Analyzing (Meta)genomic Data. MBio 5, https://doi.org/10.1128/mBio.00889-14 (2014).
Frese, S. A., Parker, K., Calvert, C. C. & Mills, D. A. Diet shapes the gut microbiome of pigs during nursing and weaning. Microbiome 3, 28, https://doi.org/10.1186/s40168-015-0091-8 (2015).
Zhao, W. et al. The Dynamic Distribution of Porcine Microbiota across Different Ages and Gastrointestinal Tract Segments. PLoS One 10, e0117441, https://doi.org/10.1371/journal.pone.0117441 (2015).
Biddle, A., Stewart, L., Blanchard, J. & Leschine, S. Untangling the Genetic Basis of Fibrolytic Specialization by Lachnospiraceae and Ruminococcaceae in Diverse Gut Communities. Diversity 5, 627 (2013).
Lopetuso, L. R., Scaldaferri, F., Petito, V. & Gasbarrini, A. Commensal Clostridia: leading players in the maintenance of gut homeostasis. Gut Pathogens 5, 23–23, https://doi.org/10.1186/1757-4749-5-23 (2013).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450, https://doi.org/10.1038/nature12721 (2013).
Atarashi, K. et al. Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500, 232–236, https://doi.org/10.1038/nature12331 (2013).
Larsen, N. et al. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS One 5, e9085, https://doi.org/10.1371/journal.pone.0009085 (2010).
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60, https://doi.org/10.1038/nature11450 (2012).
Etxeberria, U. et al. Shifts in microbiota species and fermentation products in a dietary model enriched in fat and sucrose. Benef. Microbes 6, 97–111, https://doi.org/10.3920/bm2013.0097 (2015).
Hildebrandt, M. A. et al. High Fat Diet Determines the Composition of the Murine Gut Microbiome Independently of Obesity. Gastroenterology 137, 1716-1724.e1711–1712, https://doi.org/10.1053/j.gastro.2009.08.042 (2009).
de La Serre, C. B. et al. Propensity to high-fat diet-induced obesity in rats is associated with changes in the gut microbiota and gut inflammation. Am. J. Physiol. Gastrointest. Liver Physiol. 299, G440–448, https://doi.org/10.1152/ajpgi.00098.2010 (2010).
Pedersen, R. et al. Characterisation of Gut Microbiota in Ossabaw and Göttingen Minipigs as Models of Obesity and Metabolic Syndrome. PLoS One 8, e56612, https://doi.org/10.1371/journal.pone.0056612 (2013).
Yutin, N. & Galperin, M. Y. A genomic update on clostridial phylogeny: Gram-negative spore-formers and other misplaced clostridia. Environmental microbiology 15, 2631–2641, https://doi.org/10.1111/1462-2920.12173 (2013).
Stanley, D., Hughes, R. J., Geier, M. S. & Moore, R. J. Bacteria within the Gastrointestinal Tract Microbiota Correlated with Improved Growth and Feed Conversion: Challenges Presented for the Identification of Performance Enhancing Probiotic Bacteria. Frontiers in Microbiology 7, 187, https://doi.org/10.3389/fmicb.2016.00187 (2016).
Bally, P. Microbiote intestinal et développement de l’obésité: une approche par métagénomique et métabolomique du concept de répondeur et non-répondeur, thesis (2015).
Ridaura, V. K. et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 341, 1241214, https://doi.org/10.1126/science.1241214 (2013).
Agus, A. et al. Western diet induces a shift in microbiota composition enhancing susceptibility to Adherent-Invasive E. coli infection and intestinal inflammation. Sci. Rep. 6, 19032, https://doi.org/10.1038/srep19032 (2016).
Heinritz, S. N. et al. Impact of a High-Fat or High-Fiber Diet on Intestinal Microbiota and Metabolic Markers in a Pig Model. Nutrients 8, https://doi.org/10.3390/nu8050317 (2016).
Barker, H. A. Amino acid degradation by anaerobic bacteria. Annu. Rev. Biochem. 50, 23–40, https://doi.org/10.1146/annurev.bi.50.070181.000323 (1981).
Barker, H. A., D’Ari, L. & Kahn, J. Enzymatic reactions in the degradation of 5-aminovalerate by Clostridium aminovalericum. J. Biol. Chem. 262, 8994–9003 (1987).
Yang, Y. et al. Effects of high fructose and salt feeding on systematic metabonome probed via (1) H NMR spectroscopy. Magn. Reson. Chem. 53, 295–303, https://doi.org/10.1002/mrc.4198 (2015).
Swann, J. R. et al. Variation in antibiotic-induced microbial recolonization impacts on the host metabolic phenotypes of rats. J. Proteome Res. 10, 3590–3603, https://doi.org/10.1021/pr200243t (2011).
Fujimoto, N., Inoue, K., Ohgusu, Y., Hayashi, Y. & Yuasa, H. Enhanced uptake of glycerol by butyrate treatment in HCT-15 human colon cancer cell line. Drug Metab. Pharmacokinet. 22, 195–198 (2007).
Kato, T., Hayashi, Y., Inoue, K. & Yuasa, H. Glycerol absorption by Na+-dependent carrier-mediated transport in the closed loop of the rat small intestine. Biol. Pharm. Bull. 28, 553–555 (2005).
De Weirdt, R. et al. Human faecal microbiota display variable patterns of glycerol metabolism. FEMS Microbiol. Ecol. 74, 601–611, https://doi.org/10.1111/j.1574-6941.2010.00974.x (2010).
den Besten, G. et al. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. Journal of Lipid Research 54, 2325–2340, https://doi.org/10.1194/jlr.R036012 (2013).
Layden, B. T., Angueira, A. R., Brodsky, M., Durai, V. & Lowe, W. L. Jr. Short chain fatty acids and their receptors: new metabolic targets. Transl. Res. 161, 131–140, https://doi.org/10.1016/j.trsl.2012.10.007 (2013).
Ríos-Covián, D. et al. Intestinal Short Chain Fatty Acids and their Link with Diet and Human Health. Frontiers in Microbiology 7, 185, https://doi.org/10.3389/fmicb.2016.00185 (2016).
Le Poul, E. et al. Functional characterization of human receptors for short chain fatty acids and their role in polymorphonuclear cell activation. J. Biol. Chem. 278, 25481–25489, https://doi.org/10.1074/jbc.M301403200 (2003).
Bodnaruc, A. M., Prud’homme, D., Blanchet, R. & Giroux, I. Nutritional modulation of endogenous glucagon-like peptide-1 secretion: a review. Nutrition & Metabolism 13, 92, https://doi.org/10.1186/s12986-016-0153-3 (2016).
Richards, P. et al. High fat diet impairs the function of glucagon-like peptide-1 producing L-cells. Peptides 77, 21–27, https://doi.org/10.1016/j.peptides.2015.06.006 (2016).
Mudaliar, S. & Henry, R. R. Effects of incretin hormones on beta-cell mass and function, body weight, and hepatic and myocardial function. Am. J. Med. 123, S19–27, https://doi.org/10.1016/j.amjmed.2009.12.006 (2010).
Nakajima, S., Hira, T. & Hara, H. Postprandial glucagon-like peptide-1 secretion is increased during the progression of glucose intolerance and obesity in high-fat/high-sucrose diet-fed rats. Br. J. Nutr. 113, 1477–1488, https://doi.org/10.1017/s0007114515000550 (2015).
Cho, Y. H., Craig, M. E. & Donaghue, K. C. Puberty as an accelerator for diabetes complications. Pediatr. Diabetes 15, 18–26, https://doi.org/10.1111/pedi.12112 (2014).
Morise, A. et al. Impact of intrauterine growth retardation and early protein intake on growth, adipose tissue, and the insulin-like growth factor system in piglets. Pediatr. Res. 65, 45–50, https://doi.org/10.1203/PDR.0b013e318189b0b4 (2009).
Jouany, J. P. et al. Role of the rumen ciliate protozoa Polyplastron multivesiculatum, Entodinium sp. and Isotricha prostoma in the digestion of a mixed diet in sheep. Reprod. Nutr. Dev. 21, 871–884 (1981).
Godon, J.-J., Zumstein, E., Dabert, P., Habouzit, F. & Moletta, R. Microbial 16S rDNA diversity in an anaerobic digester. Water Science and Technology 36, 49–55, https://doi.org/10.1016/S0273-1223(97)00506-4 (1997).
Escudie, F. et al. FROGS: Find, Rapidly, OTUs with Galaxy Solution. Bioinformatics 34, 1287–1294, https://doi.org/10.1093/bioinformatics/btx791 (2017).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, https://doi.org/10.1093/bioinformatics/btp616 (2010).
Zhao, Y. et al. Gut microbiota composition modifies fecal metabolic profiles in mice. J. Proteome Res. 12, 2987–2999, https://doi.org/10.1021/pr400263n (2013).
Wold, S., Antti, H., Lindgren, F. & Öhman, J. Orthogonal signal correction of near-infrared spectra. Chemometrics and Intelligent Laboratory Systems 44, 175–185, https://doi.org/10.1016/S0169-7439(98)00109-9 (1998).
McCombie, G. et al. ω-3 oil intake during weight loss in obese women results in remodelling of plasma triglyceride and fatty acids. Metabolomics 5, 363–374, https://doi.org/10.1007/s11306-009-0161-7 (2009).
Blat, S. et al. The protein level of isoenergetic formulae does not modulate postprandial insulin secretion in piglets and has no consequences on later glucose tolerance. Br. J. Nutr. 108, 102–112, https://doi.org/10.1017/s0007114511005253 (2012).
Bergman, R. N. Minimal model: perspective from 2005. Horm. Res. 64(Suppl 3), 8–15, https://doi.org/10.1159/000089312 (2005).
Christoffersen, B., Ribel, U., Raun, K., Golozoubova, V. & Pacini, G. Evaluation of different methods for assessment of insulin sensitivity in Gottingen minipigs: introduction of a new, simpler method. Am. J. Physiol. Regul. Integr. Comp. Physiol. 297, R1195–1201, https://doi.org/10.1152/ajpregu.90851.2008 (2009).
Acknowledgements
The authors acknowledge all the technical staff of Nutrition Gut Brain team and all the staff of Rennes porcine experimental facilities (UEPR, especially Francis Le Gouevec, Julien Georges and Mickaël Genissel) for their professionalism, help, expert assistance and animal care and feeding. They also thank the Nutrition, chemical food safety and consumer behaviour Division of the INRA for financing the metabolomics study and the French National Infrastructure of Metabolomics and Fluxomics (MetaboHUB-ANR-11-INBS-0010) for their support. The authors thank Servane Le Page for her help in PCR analysis, Roselyne Gautier for her help in preparing the samples for metabolomics analyses, Laure France for infant formula manufacturing, Georges Guillemois and Jean-François Rouaud for HE diet manufacturing and Thibaud Le Mouel for SCFA assay.
Author information
Authors and Affiliations
Contributions
P.L.R., C.B., I.L.H.-L. and S.B. conceived the project and M.L., P.L.R., C.B., P.G., I.L.H.-L. and S.B. designed the experiment. I.C., P.L.R. and C.B. provided the infant formulas. M.L., A.C., M.F., L.L.N., V.R., I.N., S.F.-B., M.R. C.C., M.T.-F., I.L.H.-L. and S.B. performed analysis. M.L., S.D., C.C., M.T.-F., I.L.H.-L. and S.B. analysed the data. M.L., I.L.H.-L. and S.B. wrote the draft manuscript. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The present work received funding from Lactalis Recherche et Développement (Retiers, France) and from the Association Nationale de la Recherche et de la Technologie (ANRT) (grant CIFRE n°2014/0580). M.L., I.C., P.L.R. and C.B. are employees of Lactalis. The other authors declare no potential conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lemaire, M., Dou, S., Cahu, A. et al. Addition of dairy lipids and probiotic Lactobacillus fermentum in infant formula programs gut microbiota and entero-insular axis in adult minipigs. Sci Rep 8, 11656 (2018). https://doi.org/10.1038/s41598-018-29971-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-29971-w
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.