Abstract
Ultraviolet (UV)-sensitive visual pigment and its corresponding ability for UV vision was retained in early mammals from their common ancestry with sauropsids. Subsequently, UV-sensitive pigments, encoded by the short wavelength-sensitive 1 (SWS1) opsin gene, were converted to violet sensitivity or have lost function in multiple lineages during the diversification of mammals. However, many mammalian species, including most bats, are suggested to retain a UV-sensitive pigment. Notably, some cave-dwelling fruit bats and high duty cycle echolocating bats have lost their SWS1 genes, which are proposed to be due to their roosting ecology and as a sensory trade-off between vision and echolocation, respectively. Here, we sequenced SWS1 genes from ecologically diverse bats and found that this gene is also non-functional in both common vampire bat (Desmodus rotundus) and white-winged vampire bat (Diaemus youngi). Apart from species with pesudogenes, our evolutionary and functional studies demonstrate that the SWS1 pigment of bats are UV-sensitive and well-conserved since their common ancestor, suggesting an important role across major ecological types. Given the constrained function of SWS1 pigments in these bats, why some other species, such as vampire bats, have lost this gene is even more interesting and needs further investigation.
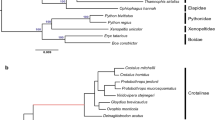
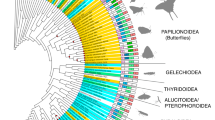
Similar content being viewed by others
Introduction
Ultraviolet (UV) light is perceived by diverse vertebrates to facilitate several behaviors, such as foraging1,2. The short wavelength-sensitive 1 (SWS1) visual pigment of ancestral vertebrates has been demonstrated to be UV-sensitive with a maximum absorption wavelength (λmax) of 361 nm3. UV sensitivity was then preserved in a variety of vertebrate taxa, with λmax values ranging from 355 to 371 nm4. In early mammals, the SWS1 pigment inherited from their amniote ancestor retained UV sensitivity (λmax = 359 nm)4. Subsequently, the SWS1 gene was lost in the ancestor of monotremes, the only mammalian group that possesses a functional short wavelength-sensitive 2 (SWS2) opsin gene, after their split from therians (marsupials and placentals)5. In contrast, the common ancestor of therians lost SWS2, but kept the SWS1 that encoded an UV-sensitive pigment4,5.
UV vision was initially thought to be rare in extant mammals, with only a few species, such as some bats, eulipotyphlans, rodents and marsupials, reported to have retained this sensitivity6,7,8,9,10. However, the recent finding, based on extensive genomic sequencing, suggests that UV-sensitive SWS1 pigments are more widely distributed in mammals than previously thought, with violet-sensitive pigments being derived multiple times11. Many mammals, however, have lost their SWS1 genes11,12, and thus UV/violet sensitivity, and is probably associated with having a nocturnal lifestyle12. Intriguingly, nocturnal activity itself cannot fully explain the losses of SWS1 as not all nocturnal species have lost this gene, suggesting other lineage-specific mechanisms potentially play roles12.
Bats (order Chiroptera) possess the second largest number of species, after rodents, within Mammals13 and are well-known for the special sensory adaptation, that is, advanced laryngeal echolocation14. Echolocation and ultrasonic hearing, processed using an enlarged auditory cortex in the brain, helps insectivorous bats detect and capture prey at night15. Old World fruit bats (family Pteropodidae), on the other hand, do not use laryngeal echolocation and rely largely on vision with enlarged eyes and visual cortex15. The pteropodid bats share a common ancestor with echolocating bats from superfamily Rhinolophoidea to form a suborder called Yinpterochiroptera16. Within the rhinolophoid species, the horseshoe and roundleaf bats (families Rhinolophidae and Hipposideridae) are insectivorous and use a sophisticated high duty cycle echolocation17. The other suborder of bats, Yangochiroptera, includes species (except Pteronotus parnellii) that use low duty cycle echolocation16,17, and have diverse diets, with many species from the family Phyllostomidae being frugivorous, nectarivorous, omnivorous and even sanguivorous18.
The majority of SWS1 genes from both echolocating and non-echolocating bat species were predicted to be UV-sensitive11,19,20,21,22. However, the exact λmax values (phenotypes) of bat SWS1 pigments have not been experimentally verified. Until now, 17 amino-acid sites have been identified as being critical for the spectral sensitivities of SWS1 pigments4. However, these sites have a limited ability to predict λmax values, based on gene sequences, for mammalian SWS1 pigments23. Interestingly, some species from the suborder Yinpterochiroptera, were reported to have independently lost their SWS1 genes11,20,24. Specifically, losses of SWS1 in the horseshoe and roundleaf bats were suggested to be sensory trade-offs with their sophisticated high duty cycle echolocation20. Individual losses of SWS1 genes were also noted in some Old World fruit bats, which were hypothesized to be related to their cave-roosting habit20. Here we sequenced the complete SWS1 coding regions from several ecologically diverse bat species, expressed these opsins, and then determined the spectral sensitivities of the visual pigments to study the roles of bat SWS1 pigments and also provide a functional supplement to the mammalian SWS1 pigments.
Results and Discussion
To investigate the spectral sensitivities of bat SWS1 pigments, we successfully amplified and sequenced complete SWS1 coding regions de novo for three species of bats and completed the missing 5′ and 3′ portions of sequences for an additional five species. Sequences from three (Cynopterus sphinx, Megaderma spasma and Rhinopoma hardwickii) of the eight species come from three different families of the suborder Yinpterochiroptera, while the other five species (Chaerephon plicatus, Miniopterus fuliginosus, Pipistrellus abramus, Leptonycteris yerbabuenae and Artibeus lituratus) are from four families of the suborder Yangochiroptera. Moreover, these eight species are ecologically diversified, representing non-echolocating Old World fruit bats (C. sphinx), echolocating New World fruit bats (L. yerbabuenae and A. lituratus) and insectivorous bats that use low duty cycle echolocation (M. spasma and R. hardwickii, C. plicatus, M. fuliginosus and P. abramus) (Supplementary Table 1). No frame-shift indels or premature stop codons were observed in the coding sequences from these eight species, suggesting that they have functional SWS1 pigments (Supplementary Fig. 1). Acquisition of the full-length coding sequences from these diverse bats enabled the subsequent evolutionary analyses and functional assays of the visual pigments.
In contrast to the above eight species, only partial coding regions could be amplified from two vampire bats, Desmodus rotundus and Diaemus youngi (Phyllostomidae) (Supplementary Table 1). Surprisingly, both of these sequences contained one or more deletions that lead to frame-shifts. Specifically, two of the four deletion events in the common vampire bat (Desmodus rotundus) result in frame-shifts, and there are a total of five premature stop codons in this gene. In the white-winged vampire bat (Diaemus youngi), there is one deletion, which is different from those in the common vampire bat, which causes a frame-shift that leads to a premature stop codon (Fig. 1). These results demonstrate that the two vampire bats independently lost their SWS1 genes after speciation. Previously, some Old World fruit bats from the family Pteropodidae and all high duty cycle echolocating bats from the families Rhinolophidae and Hipposideridae have been reported to have lost their UV pigment gene20. A sensory trade-off hypothesis, between vision and echolocation, was proposed for rhinolophid and hipposiderid bats that utilize specialized high duty cycle echolocation to explain the loss of the SWS1 gene in these species20.
Alignment of vampire bat SWS1 sequences. The nectar-feeding bat Leptonycteris yerbabuenae is a close relative of vampire bats and has an intact SWS1 coding region (“*” indicates the stop codon). Numbering of the alignment is according to the full-length L. yerbabuenae sequence. Only partial coding regions were sequenced from the two vampire bats, and are shown with a gray background. Deletion events in the vampire bat sequences are indicated by the red boxes and premature stop codons by the black boxes.
To examine selective pressure (ratio of non-synonymous and synonymous substitution rates, or ω) acting on bat SWS1 pigments, we first carried out an evolutionary analysis of the SWS1 sequences (no pseudogenes included) from both bats and other mammals (Supplementary Fig. 2A). Results from the clade model C analysis using PAML 4.9 show a subset of sites that are under different selective pressure in bats (ω = 0.21) compared to other mammalian species (0.32) (P < 0.001). Notably, even sites in SWS1 that are under purifying selection in both groups are more strongly conserved in bats. Additional comparisons between bats and species from single orders (i.e., Carnivora, Perissodactyla, Cetartiodactyla, Eulipotyphla, Primates and Rodentia) retain support the conclusion that the SWS1 genes in bats experienced stronger purifying selection than in other mammals, with the exception of eulipotyphlans (Table 1). A sliding window analysis was then conducted to calculate the ω values for species from each order. Results of this analysis show that the estimated ω values are higher in species from the orders Carnivora, Perissodactyla, Cetartiodactyla and Primates than in bats, while no obvious difference among bats, eulipotyphlans and rodents (Supplementary Fig. 2B), which is a pattern similar to that derived from clade model C. Previous findings, based on other evolutionary models in PAML, have shown purifying selection on bat SWS1 genes20,22. The results of our study further reveal that the negative selection acting on bat SWS1 is even stronger than on other mammals (except eulipotyphlan species), which suggests that the spectral sensitivities of SWS1 pigments from these bats have probably not shifted much.
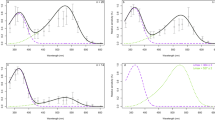
We used in vitro assays on purified recombinant SWS1 photopigments to determine their spectral sensitivities. Our results demonstrate that SWS1 from all eight bat species examined are UV-sensitive with λmax values ranging from 357 to 359 nm (Fig. 2). Moreover, the resurrected SWS1 pigment of the bat ancestor is also UV-sensitive (λmax = 358 nm), demonstrating a conserved function in bats with only minor λmax-shifts (Figs 2 and 3). Bat SWS1, therefore, is a new group of UV-sensitive pigments in mammals that has been functionally verified by in vitro assay, in addition to the previously characterized pigments from rodents (Mus musculus and Rattus norvegicus) and marsupials (Didelphis aurita and Sminthopsis crassicaudata), which have λmax values of 358–359 nm and 362–363 nm respectively25,26,27,28. All of the bat SWS1 pigments studied here are UV-sensitive, with differences of only 2 nm or less in spectral sensitivity, and are from species spanning various ecological types, including a non-echolocating fruit bat and low duty cycle echolocating bats feeding on fruits, nectar or insects (Fig. 3). The conserved UV pigments from the major clades of order Chiroptera examined in this study add further support to the hypothesis that UV sensitivity mediated by SWS1 pigment is advantageous and important for specific bats7,22.
Evolution of UV pigments in bats. λmax values of the UV pigments, type of echolocation, and diet are mapped onto the chiropteran phylogenetic tree. Absence of SWS1 (pseudogene) is indicate by Ψ. Amino-acid substitutions in the SWS1 opsins are indicated on the lineages, with only sites between transmembrane I and VII shown44. Numbering of the sites follows that of bovine rhodopsin. Losses of SWS1 genes in bats are indicated by gray branches in the tree. Gene loss events which have been published in previous studies are denoted by “*” after the species name11,20.
In contrast to the functional conservation of UV pigments in these species, some other bats have independently lost their SWS1 genes11,20,24. Notably, vampire bats, the only group of sanguivorous mammals29, have lost their SWS1 genes while other closely related phyllostomid species, such as L. yerbabuenae (λmax = 358 nm) and A. lituratus (λmax = 358 nm), retain functional copies. UV sensitivity through SWS1 pigment has been suggested to be useful for several activities such as detection of insects and flowers and for orientation7,20,21,22. While the reason underlying these losses of UV-sensitive pigments in vampire bats is not clear, it is possibly related to their changed foraging targets, which are mammals and birds30,31. In addition, the SWS1 gene from the Indian false vampire bat (Megaderma lyra) has also been reported to be a pseudogene, based on a low-coverage genome sequence11,32. We re-sequenced part of the SWS1 coding region from this species and confirmed that it is a pseudogene (Supplementary Table 1). However, a congener species, the lesser false vampire bat (Megaderma spasma), retains a functional UV pigment (λmax = 357 nm), with negligible functional difference from the bat ancestor (Fig. 3). Therefore the reasons for the independent losses of SWS1 genes in the vampire bats and also in the Indian false vampire bat are likely specifically related to their individual ecologies, and further studies are necessary to elucidate the causes.
Methods
Bat SWS1 sequencing
To examine the phenotypes of SWS1 pigments in bats of the order Chiroptera, complete coding sequences from representative species are required. Here we studied 11 species from the suborders Yinpterochiroptera, Cynopterus sphinx (family Pteropodidae), Rhinopoma hardwickii (Rhinopomatidae), Megamerda spasma and M. lyra (Megadermatidae), and Yangochiroptera, Chaerephon plicatus (Molossidae), Miniopterus fuliginosus (Miniopteridae), Pipistrellus abramus (Vespertilionidae), and four additional species, Leptonycteris yerbabuenae, Artibeus lituratus, Desmodus rotundus and Diaemus youngi from the family Phyllostomidae. Bat sample collection (wing membrane and eye tissues) for this study was approved by the Animal Ethics Committee of Shenyang Agricultural University, and the experiments were performed following the guidelines and regulations.
Eye tissue from C. sphinx was used for amplification of the SWS1 coding region. Total RNA extraction and first-strand cDNA synthesis were conducted using RNAiso Plus and PrimeScript II reagents (TaKaRa). Wing membrane tissues from the other ten species were collected for amplification of SWS1 exons, separately, from genomic DNA. The TIANamp Kit (Tiagen) was used for genomic DNA extraction. Eye cDNA from R. hardwickii was also synthesized and used as a PCR template, since specific amplification of SWS1 exons failed using genomic DNA for this species. Detailed information on primer pairs and corresponding annealing temperatures are provided in Supplementary Tables 1 and 2. PCR amplicons were purified, ligated into pGEM-T Easy cloning vector (Promega), transformed using DH5α competent cells (Tiagen), and then sequenced by an ABI 3730 machine (Applied Biosystems).
Evolutionary analyses on SWS1 genes
For the evolutionary analyses of the mammalian SWS1 genes, 73 additional complete coding sequences were obtained from NCBI (www.ncbi.nlm.nih.gov) and Ensembl (www.ensembl.org). These sequences represent species from 14 orders of mammals and enable comparisons of the selective pressures between bats and other groups (Supplementary Table 3).
A total of 81 mammalian sequences (no pseudogenes included) were used to determine whether the selective pressures acting on SWS1 genes were different in bats compared to other species. Sequences were aligned using Clustal W in MEGA 5 software33 and then checked by eye. Clade model C was implemented using CODEML in PAML 4.934,35. This model allows a site class with different ω values between the focal clade (ω3 assigned to the foreground) and the others (ω2 to the background). In contrast, the null model, called M1a, assumes nearly neutral evolution for this gene in all species. The superior model (clade model C or M1a) for the data was identified using a likelihood ratio test. The species tree for these 81 mammals used in this analysis was based on published phylogenies18,36,37,38,39,40,41. Clade model C was also applied to compare ω values between bats and species from each of the orders Carnivora, Perissodactyla, Cetartiodactyla, Eulipotyphla, Primates or Rodentia, separately.
To visualize ω values across the SWS1 gene sequence, a sliding window analysis was conducted using the software SWAAP 142. By using a window size of 90 bp and a step size of 18 bp, ω values for the SWS1 genes within each mammalian order (Chiroptera, Carnivora, Perissodactyla, Cetartiodactyla, Eulipotyphla, Primates or Rodentia) were calculated, respectively.
In vitro assays for bat SWS1 pigments
The 81 species dataset was used to infer the ancestral SWS1 sequence for bats. The best-fitting amino-acid model for the data is JTT + I + Γ, which was selected by ProtTest 3 software according to the corrected Akaike information criterion43. The ancestral sequence for bat SWS1 was reconstructed under the best-fitting model and the species topology by using CODEML34.
Complete coding sequences of eight bat SWS1 genes and the inferred sequence of the common ancestor of bats were synthesized and ligated into the pcDNA3.1 expression vector (Invitrogen) using the HindIII and XhoI restriction sites. A Kozak sequence “CCACC” was added in front of the start codon for each gene. A short sequence “ACA GAG ACC AGC CAA GTG GCG CCT GCC”, encoding “TETSQVAPA”, was added prior to the stop codon, and used as an epitope tag for protein purification. Plasmids containing the SWS1 genes were transfected into HEK293T cells individually using Xfect reagent (Clontech). Expressed SWS1 opsins were harvested from cells 48 h after transfection. Visual pigments were regenerated by incubating the cells together with 11-cis-retinal (Storm Eye Institute, Medical University of South Carolina). Visual pigments were then solubilized using n-Dodecyl β-D-maltoside and purified with the Rho 1D4 monoclonal antibody (University of British Columbia) following published procedures26. Absorbance spectra of the SWS1 pigments were measured by a spectrophotometer (U-3900, Hitachi) in the dark before and after sulfuric acid denaturation orderly26.
References
Hunt, D. M., Wilkie, S. E., Bowmaker, J. K. & Poopalasundaram, S. Vision in the ultraviolet. Cell Mol Life Sci 58, 1583–98 (2001).
Honkavaara, J., Koivula, M., Korpimaki, E., Siitari, H. & Viitala, J. Ultraviolet vision and foraging in terrestrial vertebrates. Oikos 98, 505–511 (2002).
Shi, Y. & Yokoyama, S. Molecular analysis of the evolutionary significance of ultraviolet vision in vertebrates. Proc Natl Acad Sci USA 100, 8308–13 (2003).
Yokoyama, S., Tada, T., Liu, Y., Faggionato, D. & Altun, A. A simple method for studying the molecular mechanisms of ultraviolet and violet reception in vertebrates. BMC Evol Biol 16, 64 (2016).
Davies, W. I., Collin, S. P. & Hunt, D. M. Molecular ecology and adaptation of visual photopigments in craniates. Mol Ecol 21, 3121–58 (2012).
Winter, Y., Lopez, J. & Von Helversen, O. Ultraviolet vision in a bat. Nature 425, 612–4 (2003).
Muller, B. et al. Bat eyes have ultraviolet-sensitive cone photoreceptors. PLoS One 4, e6390 (2009).
Glosmann, M., Steiner, M., Peichl, L. & Ahnelt, P. K. Cone photoreceptors and potential UV vision in a subterranean insectivore, the European mole. J Vis 8(23), 1–12 (2008).
Jacobs, G. H., Neitz, J. & Deegan, J. F. II Retinal receptors in rodents maximally sensitive to ultraviolet light. Nature 353, 655–6 (1991).
Arrese, C. A., Hart, N. S., Thomas, N., Beazley, L. D. & Shand, J. Trichromacy in Australian marsupials. Curr Biol 12, 657–60 (2002).
Emerling, C. A., Huynh, H. T., Nguyen, M. A., Meredith, R. W. & Springer, M. S. Spectral shifts of mammalian ultraviolet-sensitive pigments (short wavelength-sensitive opsin 1) are associated with eye length and photic niche evolution. Proc R Soc B 282, 20151817 (2015).
Jacobs, G. H. Losses of functional opsin genes, short-wavelength cone photopigments, and color vision–a significant trend in the evolution of mammalian vision. Vis Neurosci 30, 39–53 (2013).
Simmons, N. B. Order Chiroptera. In Mammal species of the world: a taxonomic and geographic reference (eds Wilson, D. E. & Reeder, D. M.) (Johns Hopkins University Press, Baltimore, 2005).
Jones, G. & Teeling, E. C. The evolution of echolocation in bats. Trends Ecol Evol 21, 149–56 (2006).
Speakman, J. R. The evolution of flight and echolocation in bats: another leap in the dark. Mammal Rev 31, 111–130 (2001).
Teeling, E. C. et al. A molecular phylogeny for bats illuminates biogeography and the fossil record. Science 307, 580–4 (2005).
Fenton, M. B., Faure, P. A. & Ratcliffe, J. M. Evolution of high duty cycle echolocation in bats. J Exp Biol 215, 2935–44 (2012).
Datzmann, T., von Helversen, O. & Mayer, F. Evolution of nectarivory in phyllostomid bats (Phyllostomidae Gray, 1825, Chiroptera: Mammalia). BMC Evol Biol 10, 165 (2010).
Wang, D. et al. Molecular evolution of bat color vision genes. Mol Biol Evol 21, 295–302 (2004).
Zhao, H. et al. The evolution of color vision in nocturnal mammals. Proc Natl Acad Sci USA 106, 8980–5 (2009).
Melin, A. D., Danosi, C. F., McCracken, G. F. & Dominy, N. J. Dichromatic vision in a fruit bat with diurnal proclivities: the Samoan flying fox (Pteropus samoensis). J Comp Physiol A 200, 1015–22 (2014).
Zhao, H., Xu, D., Zhou, Y., Flanders, J. & Zhang, S. Evolution of opsin genes reveals a functional role of vision in the echolocating little brown bat (Myotis lucifugus). Biochem Syst Ecol 37, 154–161 (2009).
Hauser, F. E., van Hazel, I. & Chang, B. S. Spectral tuning in vertebrate short wavelength-sensitive 1 (SWS1) visual pigments: can wavelength sensitivity be inferred from sequence data? J Exp Zool B 322, 529–39 (2014).
Muller, B., Goodman, S. M. & Peichl, L. Cone photoreceptor diversity in the retinas of fruit bats (Megachiroptera). Brain Behav Evol 70, 90–104 (2007).
Fasick, J. I., Applebury, M. L. & Oprian, D. D. Spectral tuning in the mammalian short-wavelength sensitive cone pigments. Biochemistry 41, 6860–5 (2002).
Yokoyama, S., Radlwimmer, F. B. & Kawamura, S. Regeneration of ultraviolet pigments of vertebrates. FEBS Lett 423, 155–8 (1998).
Cowing, J. A., Arrese, C. A., Davies, W. L., Beazley, L. D. & Hunt, D. M. Cone visual pigments in two marsupial species: the fat-tailed dunnart (Sminthopsis crassicaudata) and the honey possum (Tarsipes rostratus). Proc R Soc B 275, 1491–9 (2008).
Hunt, D. M. et al. Cone visual pigments in two species of South American marsupials. Gene 433, 50–5 (2009).
Zepeda Mendoza, M. L. et al. Hologenomic adaptations underlying the evolution of sanguivory in the common vampire bat. Nat Ecol Evol 2, 659–668 (2018).
Carter, G. G., Coen, C. E., Stenzler, L. M. & Lovette, I. J. Avian host DNA isolated from the feces of white-winged vampire bats (Diaemus youngi). Acta Chiropt 8, 255–274 (2006).
Bobrowiec, P. E. D., Lemes, M. R. & Gribel, R. Prey preference of the common vampire bat (Desmodus rotundus, Chiroptera) using molecular analysis. J Mammal 96, 54–63 (2015).
Parker, J. et al. Genome-wide signatures of convergent evolution in echolocating mammals. Nature 502, 228–31 (2013).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28, 2731–9 (2011).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24, 1586–91 (2007).
Bielawski, J. P. & Yang, Z. H. A maximum likelihood method for detecting functional divergence at individual codon sites, with application to gene family evolution. J Mol Evol 59, 121–132 (2004).
Fabre, P. H., Hautier, L., Dimitrov, D. & Douzery, E. J. A glimpse on the pattern of rodent diversification: a phylogenetic approach. BMC Evol Biol 12, 88 (2012).
Perelman, P. et al. A molecular phylogeny of living primates. PLoS Genet 7, e1001342 (2011).
Springer, M. S., Murphy, W. J., Eizirik, E. & O’Brien, S. J. Placental mammal diversification and the Cretaceous-Tertiary boundary. Proc Natl Acad Sci USA 100, 1056–61 (2003).
Miller-Butterworth, C. M. et al. A family matter: conclusive resolution of the taxonomic position of the long-fingered bats, miniopterus. Mol Biol Evol 24, 1553–61 (2007).
Hassanin, A. et al. Pattern and timing of diversification of Cetartiodactyla (Mammalia, Laurasiatheria), as revealed by a comprehensive analysis of mitochondrial genomes. C R Biol 335, 32–50 (2012).
Nyakatura, K. & Bininda-Emonds, O. R. Updating the evolutionary history of Carnivora (Mammalia): a new species-level supertree complete with divergence time estimates. BMC Biol 10, 12 (2012).
Pride, D. T. SWAAP - A tool for analyzing substitutions and similarity in multiple alignments. distributed by the author (2000).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–5 (2011).
Palczewski, K. et al. Crystal structure of rhodopsin: A G protein-coupled receptor. Science 289, 739–45 (2000).
Acknowledgements
We thank Yimeng Cui and Fangnan Liu for technical help in lab, and Dr. Rosalie K. Crouch (Storm Eye Institute, Medical University of South Carolina) and Dr. Lisa A. Neuhold (National Eye Institute, National Institutes of Health) for providing 11-cis-retinal. This work was supported by grants from the National Natural Science Foundation of China (No. 31601855) to Y.L. and (No. 31570382) to S.Z., and the Ministry of Science and Technology of the People’s Republic of China (No. 2016YFD0500300) to Y.L. and S.Z. and (2016YFC1200100) to S.Z.
Author information
Authors and Affiliations
Contributions
Y.L. designed the study; Y.L. and S.Z. contributed the experimental materials; L.L., H.C., H.L. and Y.X. conducted the experiments; L.L., H.C. and Y.L. analyzed the data; Y.L., D.M.I. and S.Z. wrote the paper.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, L., Chi, H., Liu, H. et al. Retention and losses of ultraviolet-sensitive visual pigments in bats. Sci Rep 8, 11933 (2018). https://doi.org/10.1038/s41598-018-29646-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-29646-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.