Abstract
The transcription factor PROX1 is essential for development and cell fate specification. Its function in cancer is context-dependent since PROX1 has been shown to play both oncogenic and tumour suppressive roles. Here, we show that PROX1 suppresses the transcription of MMP14, a metalloprotease involved in angiogenesis and cancer invasion, by binding and suppressing the activity of MMP14 promoter. Prox1 deletion in murine dermal lymphatic vessels in vivo and in human LECs increased MMP14 expression. In a hepatocellular carcinoma cell line expressing high endogenous levels of PROX1, its silencing increased both MMP14 expression and MMP14-dependent invasion in 3D. Moreover, PROX1 ectopic expression reduced the MMP14-dependent 3D invasiveness of breast cancer cells and angiogenic sprouting of blood endothelial cells in conjunction with MMP14 suppression. Our study uncovers a new transcriptional regulatory mechanism of cancer cell invasion and endothelial cell specification.
Similar content being viewed by others
Introduction
The transcription factor PROX1 is involved in the development of the central nervous system, lens, heart, liver and pancreas1,2,3,4,5,6. PROX1 is also necessary and sufficient for the differentiation of lymphatic endothelial cells (LECs)7,8. The role of PROX1 in cancer is context and tumour type-dependent since it has been shown to have both oncogenic and tumour-suppressive properties9. In agreement with the concept that during oncogenesis an aberrant developmental program is activated, altered PROX1 expression is often found in malignant cells of organs, whose normal development depends on PROX19. Glioma, esophageal carcinoma and colon cancer display high PROX1 levels10,11,12,13 indicative of an oncogenic role, while in hepatocellular carcinoma (HCC) PROX1 expression is reduced, suggesting a tumour-suppressive role14,15,16. Moreover, high expression of PROX1 was recently reported to associate to better survival in gastric cancer17.
PROX1 expression was also recently investigated in Kaposi’s sarcoma (KS), an angiogenic tumour of endothelial origin causally linked to KS herpesvirus (KSHV) infection, and which is the second most common malignancy among AIDS patients (AIDS-associated KS)18. In this study, PROX1 was expressed in the large majority (93.3%) of the cases analysed19. Interestingly, we and others have demonstrated that infection of LECs with KSHV reduces PROX1 expression20,21,22. Since our previous work showed that the PROX1 downregulation in KSHV-infected LECs reprogrammed the LECs into a more invasive cell type that was dependent on the membrane type 1 matrix metalloproteinase MMP1420, we have sought to investigate whether PROX1 regulates the MMP14 levels.
Here we report that PROX1 and MMP14 expressions are inversely correlated and that PROX1 binds and represses transcription from the MMP14 promoter. Moreover, by manipulating PROX1 expression we could regulate MMP14 expression in an in vivo mouse model and change the invasive properties of cancer and blood endothelial cells in vitro. Our work reveals that PROX1 regulates MMP14 expression in several cellular contexts, and thus represents an important modulator of normal and cancer cell behaviour.
Results
PROX1 and MMP14 expression are inversely correlated
PROX1 is expressed in KS tumours as shown in several publications19,21,22,23. Given our previous observations on MMP14 expression in KS biopsies20, we decided to investigate the possible cross-regulation of PROX1 and MMP14 in KS tumours. Sections from ten AIDS-associated KS patients were double-stained for PROX1 and MMP14. The expression of PROX1 and MMP14 appeared inversely correlated throughout the lesions (Fig. 1a,b). In particular, about 98% of the cells with the highest MMP14 intensity had the lowest levels of PROX1, and 95% of the cells presenting the highest PROX1 signal showed low or no MMP14 expression.
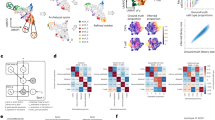
PROX1 and MMP14 expressions are inversely correlated. (a) Representative image of a KS section from an AIDS patient stained with MMP14 (green) and Prox1 (magenta) specific antibodies. Nuclei were counterstained with Hoechst 33342. (b) MMP14 and Prox1 signal intensities for each cell were quantified from 3 images/section of 10 different patients. Each dot represents a cell and the different colours represent the different patients. (c) 57 specimens from TMA were stained as in (a), representative images are shown for one benign hyperplasia (left) and one papillary thyroid cancer (right). White arrows indicate cells with Prox1 cytoplasmic staining. (d) Transcript levels of Prox1 and MMP14 in different tissues/organs derived from FANTOM5 database. Tags per million for MMP14 (Blue) and Prox1(red) are shown for each tissue/organ. Pearson´s correlation coefficient (r): −0.4515. Significance of the correlation p = 0.0065.
A decrease in PROX1 expression was recently observed in thyroid cancer when compared to adjacent healthy tissue, and reintroduction of PROX1 in papillary thyroid cancer (PTC) cell lines abolished the malignant properties (growth, adhesion and invasiveness) of these cells24. Therefore, to investigate if reciprocal levels of PROX1 and MMP14 would be observed also in thyroid cancer, we performed double-staining IHC for PROX1 and MMP14 on a tissue microarray (TMA) including 57 specimens of which 44 originated from different thyroid cancers and 13 from benign hyperplasias. We found TMA specimens where only PROX1 (12/57) or MMP14 (14/57) was expressed and samples where both proteins were expressed (22/57) (Fig. 1c). Similar to KS results, in case of PROX1 and MMP14 co-expression in the same tissue section, nuclear PROX1 and cytoplasmic/membrane-associated MMP14 were not found in the same cells, but interestingly, in some specimens cytoplasmic PROX1 and MMP14 were co-localized (Fig. 1c, white arrows). The inversely correlated expression pattern was independent of the stage and the type of the tumour as illustrated in Fig. 1c, where specimens from a benign thyroid hyperplasia and a malignant PTC are shown. Cytoplasmic localization of PROX1 has also been described during lens development25 and in thyroid and gastric cancer cells17,24. Although PROX1 function in the cytoplasm is not known, it is evident that there it cannot act as a transcriptional repressor.
To assess if the inversely correlated expression of PROX1 and MMP14 occurs also in healthy tissues, we investigated their mRNA levels reported in the FANTOM5 database (www.humanproteinatlas.com). The levels of MMP14 and PROX1 were inversely correlated in the majority of the analysed, normal tissues, except in the spleen, where both PROX1 and MMP14 mRNA were expressed at intermediate levels (Fig. 1d). Taken together, observations across different cancer types suggest that PROX1 negatively regulates MMP14 expression.
PROX1 binds to MMP14 promoter and represses its transcription
To test if PROX1 directly suppresses MMP14 transcription, we initially performed a luciferase-based reporter assay using plasmids harboring 0.4, 1.2 and 7.2 kb fragments of the 5′-flanking region of the MMP14 gene upstream of the closest transcription start site (TSS), linked to a firefly luciferase gene (described in26 and depicted in the schematic in Fig. 2a, upper panel). The results revealed that Prox1 wild-type (WT) significantly reduced the luciferase activity of the 7.2 kb and of the 1.2 kb MMP14 promoter fragments (Fig. 2a, lower panel). Notably, a PROX1 mutant (MUT) with point mutations in the Prospero region, responsible in Drosophila for the DNA binding and lacking transcriptional activity27, had no effect on the MMP14 reporter activity of any of the constructs tested. Next, we assessed whether PROX1 was negatively regulating MMP14 promoter activity by direct binding to DNA, as suggested by the lack of effect in the presence of the PROX1 MUT. To this end, we performed ChIP following ectopic expression of PROX1 in iLECs. The samples were then subjected to qPCR using primers recognizing different regions of the MMP14 promoter (from −1340 to −36 bp upstream of MMP14 TSS) (diagram in Fig. 2b, upper panel). The ChIP results revealed that PROX1 binds to the MMP14 promoter in the regions designated as b and c (Fig. 2b) that correspond to sequences previously identified as negative regulatory regions26. In silico analysis of these sequences showed that both b and c fragments were harboring putative PROX1-binding sites28. The fragment b contains one PROX1-binding site from 11239 to 11223 bp upstream of MMP14 TSS (PROX1 BS1, Fig. 2c, left panel); whereas the fragment c contains four consecutive PROX1 binding sites from 1020 to 963 bp upstream of MMP14 TSS (PROX1 BS2, Fig. 2c, left panel). To study the contribution of these putative binding sites to PROX1 transcriptional activity, we generated the ΔBS1 and ΔBS2 mutants, lacking the PROX1 binding sites in the b and c fragment, respectively, as well as ΔBS1-2, devoid of all putative PROX1 binding sites within the b and c fragments of the MMP14 promoter. The luciferase activity of the ΔBS1 and ΔBS2 was still suppressed by approximately 50% in the presence of WT PROX1 (Fig. 2c, right panel). However, by combining the two deletions (ΔBS1-2) the repression of MMP14 promoter activity by PROX1 was abolished.
PROX1 binds to the MMP14 promoter and regulates its expression. (a) Upper panel: schematic diagram of the MMP14 promoter fragments, numbers indicate the bp upstream (−) or downstream (+) of the MMP14 transcription start site (TSS, where bp = 0 is MMP14 TSS), indicated by the black arrow. Bottom panel: luciferase reporter assay using either the pGL3 backbone or its indicated derivatives depicted above. HeLa cells were co- transfected with the indicated reporter plasmids (0.1 µg) and expression vectors for PROX1 WT or MUT (1 µg) in duplicates. An empty pAMC vector was used as a control (mock). 32 h after transfection, cell extracts were collected and luciferase activities were measured. The graph includes the data from three independent experiments. Error bars represent SD. (b) Upper panel: schematic representation of the MMP14 promoter region spanning from bp −1340 to −36. Letters denote the fragments amplified by different sets of primers (a–h) used in the ChIP-qPCR below. Bottom panel: Chromatin immunoprecipitation using either PROX1 or control IgG antibodies followed by qPCR with primers amplifying the indicated regions of the 1.2 kb MMP14 promoter region. Average fold enrichment over the IgG is shown for three independent experiments; error bars represent SD. (c) Left: schematic representation of the 1.2 kb MMP14 promoter reporter plasmid, the b and c regions immunoprecipitated in (b) are marked as red boxes. PROX1 binding site containing regions (BS1 and BS2) are illustrated below as red dotted lines, numbers indicate their position upstream of the MMP14 TSS. These sequences were deleted in the 1.2 kb MMP14 promoter reporter plasmid (white boxes), generating the 1.2 kb MMP14 promoter mutants ΔBS1, ΔBS2, ΔBS1-2. Right: Luciferase assay performed as in (a) using 1.2 kb MMP14 promoter either full length (FL) or the indicated mutants co-transfected with either a pAMC control vector (mock) or a PROX1 WT expression vector. *p < 0.05; **p > 0.01; ***p > 0.001.
Together, these data indicate that PROX1 is a negative transcriptional regulator of MMP14 and that the repression occurs through direct PROX1 binding to the MMP14 promoter regions in the positions from 1139 to 1123 (BS1) and from 1020 to 963 (BS2) bp upstream of MMP14 TSS.
PROX1 depletion increases MMP14 levels in murine lymphatic vessels and in human cells
Prompted by these observations (Figs 1 and 2) implicating PROX1 as a transcriptional regulator of MMP14, we tested if altering the PROX1 levels leads to changes in MMP14 expression in vivo. To address this in an in vivo physiological setting, we took advantage of the Prox1flox mouse29, crossed with the Cdh5-CreERT2 mouse30 to delete Prox1 specifically in endothelial cells by treatment with 4OH-Tamoxifen (4OHT). The Prox1flox/flox; Cdh5-CreERT2 mice and the Cre-negative Prox1flox/flox littermate controls were treated with 4OHT at 3 weeks of age. Efficient Prox1 deletion in dermal lymphatic vessels was confirmed by whole mount immunofluorescence of the ear one week after the treatment, in particular the lymphatic vessels of the Prox1flox/flox; Cdh5-CreERT2 mouse, identified with LYVE-1 staining, were negative for PROX1 (Fig. 3a). Furthermore, we observed no relevant changes in the organization and distribution of PECAM-positive (blood vascular) and LYVE-1 positive (lymphatic) vessels. The other ear was used for paraffin sections, which were stained with endomucin and podoplanin, markers for blood and lymphatic vessels, respectively (Fig. 3b) to further demonstrate that Prox1 inactivation did not modify the identity or organization of blood and lymphatic vessels in Prox1flox/flox; Cdh5-CreERT2 mice (Fig. 3b). Subsequently, to assess the expression of MMP14 in the PROX1-negative lymphatic vessels, mouse ears sections were co-stained with LYVE-1 and MMP14 (Fig. 3c). LYVE-1-positive lymphatic vessels in the Prox1flox/flox; Cdh5-CreERT2 ears showed significantly stronger MMP14 staining when compared to the Cre-negative littermate control ears. The expression of higher levels of MMP14 in the lymphatic vessels of the Prox1flox/flox; Cdh5-CreERT2 ears was also verified by staining for podoplanin, another lymphatic-specific marker (Fig. S1a). Moreover, no change in MMP14 expression in the endomucin-positive blood vessels was observed the in Prox1flox/flox; Cdh5-CreERT2 ears compared to Prox1flox/flox ears (Fig. S1b). These observations indicate that Prox1 modulates MMP14 levels in the murine LECs in vivo.
PROX1 depletion increases MMP14 expression in vivo and in vitro. (a) Whole-mount immunofluorescence of 4 weeks old Prox1flox; Cdh5-CreERT2 mice ear skin. Note efficient depletion of PROX1 in the Prox1flox/flox; Cdh5-CreERT2 ear compared to the control. (b) Representative images of IHC staining of the ear sections from mice described in (a) for podoplanin and endomucin. Nuclei were counterstained with Hoechst 33342. (c) Representative images of IHC staining for MMP14 and LYVE −1, a lymphatic endothelial marker, of the ear sections from mice described in (a). Nuclei were counterstained with Hoechst 33342. Pearsons’s co-localization index (PC) was calculated using 4 images of three Prox1flox/flox and four Prox1flox/flox; Cdh5-CreERT2. PC = 0.088 in control mice (N3); PC = 0.640 in Prox1-depleted mice (N = 4); p = 0.01 (t-test). (d,e) LECs were transfected with the indicated siRNAs and after 72 h whole cell extracts were analysed by RTqPCR (d) for the indicated targets with GAPDH as an internal control. Bars represent an average of three independent experiments, error bars show SD across the experiments. (e) Immunoblotting for the indicated proteins using γ-tubulin (TUBG1) as a loading control; numbers indicate the intensity of MMP14 band for each sample normalized to the corresponding loading control. Cropped membranes are shown, uncropped membranes can be found in Fig. S3a. (f,g) HEK293FT cells were transfected with the indicated siRNAs, 32 h later whole cell extracts were analysed by RTqPCR (f) for the indicated targets with ACT as an internal control. Bars indicate the average of two independent experiments, error bars show SD; and by immunoblotting (g) for the expression of the indicated proteins using γ-tubulin (TUBG1) as a loading control; numbers indicate the intensity of MMP14 band for each sample normalized to the corresponding loading control. Cropped membranes are shown, uncropped membranes can be found in Fig. S3b. **p < 0.01; ***p > 0.001.
We next analysed the effect of PROX1 manipulation on MMP14 expression in human cells. We depleted PROX1 in human LECs using two different PROX1-targeting siRNAs and measured MMP14 expression, comparing it to cells treated with control siRNA (Fig. 3d,e). Upon efficient silencing of PROX1, MMP14 mRNA and protein levels were increased about twofold at RNA and about two- to threefold at protein level, thus suggesting that PROX1 functions as a regulator of MMP14 expression in primary LECs.
PROX1 has been shown previously to regulate LEC fate in part via increased expression of VEGFR-3 and repression of blood endothelial genes, such as NRP131. To study whether upregulation of MMP14 in PROX1-depleted cells coincided with broad LEC fate changes, we analysed MMP14, VEGFR-3 and NRP2, a lymphatic-specific marker controlled by the Chicken Ovalbumin Upstream Promoter Transcription Factor 2 (COUP-TF II)32, as well as NRP1 expression in LECs 72 h post siRNA transfection and compared the expression of these genes in LECs and blood endothelial cells, such as BEC and HUVEC. Consistent with previous experiments, PROX1 depletion induced strong increase in MMP14 levels. However, VEGFR-3 and NRP2 expression levels were not changed and albeit there was a slight increase in NRP1, its levels were still significantly lower than in blood endothelial cells (Fig. S1c). Thus, this data supports the notion that PROX1 directly binds to and represses the MMP14 promoter, and that the induction of MMP14 in PROX1-depleted cells precedes broad changes of the LEC fate.
To provide further evidence for the existence of this signaling axis we silenced PROX1 expression in another PROX1-positive cell type, HEK 293FT (Fig. 3f,g). Similar to LECs, PROX1 transient silencing increased MMP14 expression in HEK293FT cells both at mRNA and protein levels.
PROX1 depletion increases MMP14 levels in cancer cells
We next asked if the PROX1-MMP14 regulation occurs in human cancer cells. Initially, we performed a ChIP-qPCR in HEPG2 hepatocellular carcinoma and SW620 colorectal carcinoma cell lines, transduced with Myc-tagged PROX1 expressing lentivirus. We found that PROX1 bound the same MMP14 promoter fragments as in iLEC (compare Fig. 2b with Fig. 4a). SW620 and HepG2 endogenously express PROX1 and silencing PROX1 in these cell lines led to four- (HEPG2) and threefold (SW620) increase of MMP14 mRNA and a two- to threefold increase of MMP14 protein (Fig. 4b,c).
PROX1 silencing increases MMP14 expression and MMP14-dependent 3D sprouting in cancer cells. (a) Chromatin immunoprecipitation in HepG2 and SW620 cells ectopically expressing Myg-tagged PROX1. Chromatin was immunoprecipitated using either an anti-Myc antibody or an irrelevant anti-HA antibody and subsequently amplified by qPCR for the indicated regions of the MMP14 promoter (illustrated in Fig. 2b). Average fold enrichment over the anti-HA control is shown for three independent experiments; error bars represent SD. (b) HepG2 and SW620 cells were transfected with the indicated siRNAs for 48 h and whole cell extracts were analysed by RTqPCR for the indicated targets with GAPDH as an internal control. Bars represent an average of two independent experiments, error bars show SD across the experiments. (c) HepG2 and SW620 cells were treated as in (b) and whole cell extract was analysed by immunoblot for the indicated proteins using actin as a loading control; numbers indicate the intensity of the MMP14 band normalized to the corresponding loading control. Cropped membranes are shown, uncropped membranes can be found in Fig. S3c,d. (d,e) 3D fibrin invasion assay with HepG2 cells transfected twice in consecutive days with the indicated siRNAs and embedded in 3D fibrin in the presence of either DMSO or 50 μM of the MMP14 inhibitor NSC405020. After 4 days, fibrin gels were fixed and stained with Phalloidin (Phall A488) and Hoechst 33342. (d) Representative images are shown. (e) Quantification of three images per condition from two independent experiments (n > 100 cells/condition). Bars represent the average area occupied by each cell cluster and normalized to the scr siRNA and DMSO-treated sample. n.s.: non-significant; **p > 0.01; ***p > 0.001.
MMP14 is involved in angiogenic sprouting of endothelial cells33 and invasiveness of several solid cancers34,35,36,37 through its ability to degrade several extracellular matrices38,39, including fibrin40. We have previously shown that the invasiveness of KSHV-infected LECs into a 3D fibrin matrix is MMP14-dependent20. In hepatocellular carcinoma, MMP14 expression has been associated with poor prognosis in HCC patients41 and, in HCC cell lines it plays an important role in invasiveness in vitro and metastasis in vivo42,43. Therefore, we next asked whether the observed MMP14 increase associated with PROX1 depletion would enhance the 3D invasiveness of HepG2 cells. HepG2 cells were transfected with either control siRNA or siRNAs targeting PROX1 and embedded in 3D fibrin matrix. After 5 days in 3D culture, PROX1 siRNA-treated cells displayed an increased sprouting when compared to the control siRNA-treated cells, which grew as round spheres (Fig. 4d, upper panels). To study if the observed sprouting was MMP14-dependent, the control and PROX1-depleted cells were treated with the MMP14 specific inhibitor NSC405020 during the 3D fibrin assay. The treatment with MMP14 inhibitor significantly diminished the sprouting of PROX1 siRNA treated cells, which, similarly to control cells, grew as round spheres in 3D matrix (Fig. 4d, bottom panels and Fig. 4e), thus corroborating the function of MMP14 in the increased invasiveness upon PROX1 depletion.
Taken together, these results indicate that PROX1 binds to the MMP14 promoter and thereby negatively regulates its expression in human tumour cells. Moreover, PROX1 depletion enhances HEPG2 MMP14-dependent invasiveness in 3D fibrin matrix.
Reintroduction of PROX1 inhibits 3D sprouting and invasiveness of endothelial and cancer cells
Since silencing of PROX1 increased MMP14 levels, the reintroduction of PROX1 into PROX1-negative cells should decrease MMP14 expression. To test this, we chose HuAR2T, an HuVEC cell line, and MDA-MB-321, an invasive breast cancer cell line, both expressing high levels of MMP14, but no PROX1. We transduced both cell lines with a lentivirus encoding either WT or DNA binding-deficient PROX1. MMP14 mRNA decreased dramatically upon ectopic expression of PROX1 WT, but not PROX1 MUT. The decrease in MMP14 protein upon PROX1 WT expression was 50% less compared to the control (Fig. 5a,b).
PROX1 reintroduction decreases MMP14 expression and reduces 3D invasiveness in BEC and cancer cells. (a) MDA-MB-231 and HuAR2T were treated with the indicated siRNAs for 72 h, and whole cell extracts were analysed by immunoblotting for the indicated proteins using actin as a loading control; numbers indicate the intensity of MMP14 band for each sample normalized to the corresponding loading control. Cropped membranes are shown, uncropped membranes can be found in Fig. S3e,f. (b) The samples in (a) were analysed by RTqPCR for the indicated targets with GAPDH as an internal control. Bars represent the average of three independent experiments, error bars show SD. (c) 3D fibrin invasion assay with MDA-MB-231 transduced with lentiviruses expressing PROX1 WT or MUT and stained with Phalloidin (Phall A488) and Hoechst. Empty vector was used as a control (mock). Representative images are shown. (d) Quantification of three images per condition described in (c) from three independent experiments (n > 200 cells/condition). Bars represent the average area occupied by each cell and normalized to the mock infected cells. (e) 3D sprouting assay of HuAR2T spheroids, treated and stained as in (c). Representative images are shown. (f) Quantification of three independent experiments described in (e). Bars represent the average of the total area of each spheroid divided by the area of the non-invading cells. Three spheroids per condition were quantified in three independent experiments. Error bars indicate SD. (g,h) MDA-MB-231 cells were transduced with PROX1 WT-expressing lentivirus for 24 h and subsequently transfected with a control vector (mock) or MMP14 expression vector for 24 h. Cells were then embedded in fibrin and treated as in (c). (g) Representative enlarged images are shown. (h) Quantification of three images/condition for two independent experiments (n > 100). Bars represent the average of the total area occupied by each cell and normalized to the mock transfected cells. Error bars indicate SD. **p < 0.005; ***p > 0.001.
MMP14 plays a pivotal role in BEC sprouting as well as in solid cancer invasiveness33,34,44. In particular in breast cancer, MMP14 expression is a marker of increased cell invasiveness35,45. Therefore, we tested if the PROX1-induced decrease in MMP14 affects the ability of endothelial and cancer cells to invade into 3D crosslinked fibrin. First, we confirmed that MMP14 silencing decreased the 3D sprouting growth of MDA-MB-231 cells in 3D fibrin (Fig. S2a). Next, MDA-MB-231 cells ectopically expressing PROX1 WT or MUT, as well as control vector-transduced cells were embedded in fibrin for 3 days. As shown in Fig. 5c,d the PROX1 WT expressing cells had a significantly decreased ability to invade into the matrix as compared to the PROX1 MUT or mock-treated cells.
MMP14 inhibition by the NSC405020 treatment reduced the 3D invasive sprouting of HuAR2T (Fig. S1b), demonstrating that the invasive sprouting of these cells is MMP14 dependent. Moreover, HuAR2T spheroids expressing ectopic PROX1 WT sprouted significantly less than the PROX1 MUT-expressing or untreated spheroids (Fig. 5e,f).
To confirm that the PROX1 expressing cells were less invasive in 3D fibrin, both HuAR2T spheroids and MDA-MB-231 single cells ectopically expressing PROX1 WT were stained with antibody specific for PROX1 and the invasiveness of the PROX1 positive cells was compared to that of PROX1 negative cells within the same fibrin gel. Results shown in Fig. S2c–f illustrate that the PROX1-expressing cells sprouted/invaded significantly less than the PROX1-negative cells. To test if the invasive phenotype of PROX1 WT expressing MDA-MB-231 cells could be rescued by MMP14, we reintroduced MMP14 by transient transfection. As shown in Fig. 5g,h the inhibition of invasiveness by PROX1 WT was restored by reintroduction of MMP14.
In conclusion, these results show that PROX1 acts as a negative regulator of MMP14 expression and MMP14-dependent 3D invasion in multiple cells types, such as BEC and breast cancer cells.
Discussion
MMP14 expression is controlled at different levels, and its promoter differs from those of other MMPs46. The transcription factors Sp126, E2F47, Egr148 and HIF249 have been identified as MMP14 transcriptional activators, and to our knowledge, no repressors of MMP14 have been found so far. Here we show that PROX1 suppresses MMP14 transcription. This discovery stems from our previous observation that KSHV-infection of LECs leads to a decrease in PROX1 expression with a concomitant increase in MMP14 expression and MMP14-dependent invasiveness20.
Our data further demonstrates increased levels of MMP14 in Prox1-depleted mouse dermal lymphatics suggesting that PROX1 suppresses MMP14 expression under physiological conditions. In the lymphatic vasculature, the role of MMP14 is controversial. In corneal lymphangiogenesis MMP14 degrades ECM components and activates MMP2 to promote migration and sprouting of the LECs50, and in LEC spheroids it stimulates cell sprouting in a 3D crosslinked matrix20. However, recent data shows that MMP14 depletion in LECs increased corneal lymphangiogenesis thus implicating MMP14 as an inhibitor of this process51. The physiological levels of PROX1 in LECs could thus contribute to maintain MMP14 expression at appropriate levels during lymphangiogenesis. This is supported by the phenotype of the Foxc2-deficient mice that fail to downregulate Prox1 in the developing lymphatic vessels. These Foxc2-deficient vessels display defective capillary sprouting and remain in an immature-capillary-like state52, which could be influenced by the lack of MMP14 expression due to the aberrant Prox1 expression. On the other hand, haploinsufficiency of PROX1 has also been reported to promote abnormal lymph leakage53, which could be due to untimely MMP14 expression. However, whether these phenotypes, linked to an increase or decrease in PROX1 expression, are induced by aberrant levels of MMP14, remains to be confirmed.
Activation of the TWIST-AKT2 axis and increased metastatic invasiveness have been linked to PROX1 downregulation in HCC16. As MMP14 has been also shown to enhance HCC metastasis42, it is possible that MMP14 upregulation and the concomitant increase in the MMP14-dependent invasive sprouting in HepG2 observed here in response to PROX1 downregulation, could contribute to the malignant phenotype of HCC.
A recent work reported PROX1 mRNA downregulation and the unexpected cytoplasmic mis-localization of the PROX1 in PTC24. In that study, reintroduction of nuclear PROX1 into a PTC cell line BCPAP suppressed the invasive phenotype of the cells implying that PROX1 inactivation contributes to thyroid carcinoma invasiveness. This, together with our data on the inversely correlated expression of PROX1 and MMP14 in the thyroid cancer biopsies, suggests that the increased invasiveness could be due to an increase in MMP14 expression.
Therefore, by governing the expression of MMP14, PROX1 might function as a gate-keeper that ensures appropriate MMP14 levels. This is also supported by the observation that PROX1 is expressed in mammary stem cells54, but not in breast cancer where PROX1 gene is silenced through hyper-methylation55 and MMP14 is highly expressed. The epigenetic silencing of PROX1 might be necessary for the MMP14-dependent invasiveness since here we show that PROX1 reintroduction in MDA-MB-231 significantly reduces MMP14 expression and 3D-invasiveness.
Taken together, demonstration that PROX1 negatively regulates MMP14 in normal tissues and in cancer supports the general nature of this signalling axis and its potential importance in physiological and pathological processes.
Material and Methods
Cell culture
HeLa (ATTC:CCL-2™) and HEK293FT (Thermofisher Scientific, R70007) cells were grown in Dulbecco modified Eagle’s medium (DMEM) supplemented with 10% Foetal Calf serum (FCS) and 1% L-Glutamine. HepG2 (ATTC:HB-8065), SW620 (ATCC:CCL-227™) and MDA-MB-321 (ATCC:CRM-HTB-26™) were cultured in RPMI 1640 supplemented with 20% FCS and 1% L-Glutamine. HuAR2T, human umbilical vein endothelial cells (HuVEC), conditionally immortalized with a doxycycline-inducible human telomerase reverse transcriptase (hTERT) and Simian virus 40 (SV40) large T antigen transgene expression, were grown in EGM2-MV medium supplemented with 200 ng/ml of Doxycycline. Human dermal LECs, HuVEC, BEC (purchased from Lonza), and iLECs, the intestine-derived LECs immortalised with the HPV E6 and E7 proteins56, were grown in EGM2-MV (Lonza). All cells were propagated at standard conditions (37 °C, 5% CO2).
Antibodies, plasmids, siRNAs, inhibitors
The following antibodies were used for immunoblots: mouse anti-actin (Santa Cruz Biotechnology; SC-8432), mouse anti-TBG1 (Sigma-Aldrich; T6557), goat anti-hPROX1 (R&D systems; AF2727), rabbit anti-MMP14 (R&D Systems; EP1264Y). Secondary antibodies HRP-conjugated were from Chemicon (Millipore) and Cell Signaling Technology.
For the staining of mouse ears the following antibodies were used: rat anti-mouse PECAM1 (BD-Biosciences; 553370), rabbit anti-mouse LYVE-1 (103-PA50AG, Reliatech), goat anti-PROX1 (R&D AF2727), rat anti mouse endomucin (Santa Cruz sc-65495), hamster anti mouse podoplanin (Developmental Studies hybridoma bank 8.1.1) and mouse monoclonal anti-MMP14 (LEM clone; Millipore). Secondary antibodies conjugated to Cy3, Alexa Fluor 488 or 647 were obtained from Jackson ImmunoResearch or Life Technologies (Thermofisher).
Reporter plasmids harbouring the MMP14 promoter fragments upstream of the firefly luciferase gene were generated using pGL3 backbone26. MMP14 promoter plasmids lacking the PROX1 binding sites (indicated as ΔBS1 and ΔBS2) were generated from the pGL3-1.2 kb MMP14 promoter using NEBuilder HiFi assembly kit. The backbone and the insert were amplified by PCR using the following primers (forward; reverse):
ΔBS1: TTGGATCTTAGATTTCGCTGATAGTCTAGTTTTC; GTGAGAGAGGACAGAGGTAGTGC.
ΔBS2: TTTTGTTCTGAGTCCAGTAAGTCCCTAAAG; TTTTGTTCTGAGTCCAGTAAGTCCCTAAAG.
and ligated using HiFi Assemby Master Mix (NEB) using the following ligation oligonucleotides:
ΔBS1: CTACCTCTGTCCTCTCTCACTTGGATCTTAGATTTCGCTG.
ΔBS2: CCAGACCTGTCTAGTTCCCATTTTGTTCTGAGTCCAGTAAG.
PROX1 gene was cloned in a pAMC plasmid and the mutation N624A and N626A in the PROX1 mutant (MUT) was introduced by site-directed mutagenesis8,27. Subsequently the PROX1 wild-type (WT) and PROX1 MUT were cloned into the pSIN lentiviral vector57. The inserts were verified by sequencing.
The pSport6-MMP14 expression plasmid, containing the MMP14 gene transcribed from CMV-IE promoter, was obtained from Genome-Biology Unit, University of Helsinki.
Two different Stealth RNAi™ targeting PROX1 (HSS 108596-7) and siRNA Negative Control (12935200) were purchased from Invitrogen. MMP14-targeting siRNAs (SI03648841; SI00071176) and the corresponding control siRNA (1027281) were purchased from Qiagen. The MMP14 inhibitor NSC405020 was obtained from Sigma-Aldrich (2044451).
Transfection, lentivirus production and transduction
DNA and RNA transfections were performed using Lipofectamine 2000 and Lipofectamine RNAiMAX, respectively according to the instructions provided by the manufacturer. Lentiviruses were produced in HEK293FT transfected with pLP1, pLP2, pVSVg and either an empty pSIN or pSIN-PROX1 WT or pSIN-PROX1 MUT. After 72 h, lentivirus particles were precipitated with PEG-IT (System Biosciences) according to the vendor’s instructions. Transduction of cells with lentiviruses was done by spinoculation (450 g, 30 min, RT) in the presence of 8 μg/ul of polybrene20.
RTqPCR
Total RNA was extracted from cells using Nucleospin Spin RNA extraction kit (Macherey Nagel) and reverse transcribed as described in20. Transcript levels were measured in triplicate using unlabelled primers and SYBR GREEN reaction mix (Fermentas). The following primer sequences were used (forward; reverse):
PROX1: TGTTCACCAGCACACCCGCC; TCCTTCCTGCATTGCACTTCCCG.
MMP14: GCAGAAGTTTTACGGCTTGCAA; CCTTCGAACATTGGCCTTGAT.
NRP1: GCATGAAGGCAGACAGAGATG; CTGTCGGCCATACTCATTGAA.
NRP2: GGGAACACCCAAGACAGTGA; TCAAACCTTCGGATGTCAGG.
VEGFR3: GACAGCTACAAATACGAGCATCTG; CTGTCTTGCAGTCGAGCAGA.
GAPDH: TCAACGACCCCTTCATTGAC; ATGCAGGGATGATGTTCTGG
Actin: TCACCCACACTGTGCCATCTACGA; CAGCGGAACCGCTCATTGCCAATGG.
Data were normalized to the cellular housekeeping genes GAPDH or ACT. Experiments were performed two or three times in three technical replicates, the graphs represent an average of the fold change and error bars indicate SD ± 95%CI across the different experiments.
Western blotting
Cells were lysed in RIPA buffer (150mM-NaCl; 1%-Igepal CA630-0.5% Na-deoxycholate-0.1% SDS-50mM; Tris-HCl-PH 8.0) supplemented with phosphatase (Pierce™ 88667) and protease (Pierce™ 88666) inhibitors. Pre-cleared cell extract was mixed with 5XLaemmli buffer and loaded on Criterion TGX precast gels (Bio-Rad). Gels were run for 40 min at 55 mA and transferred on nitrocellulose membranes using trans-blot Turbo Transfer system (Bio-Rad). Membranes were blocked in 2.5% non-fat dry milk in TBS-T (0, 1%Tween) for one hour at RT, gently rocking, and incubated O/N at 4 °C with the primary antibody diluted in blocking solution, membranes were then incubated with the appropriate HRP-conjugated secondary antibody (Chemicon International) for 1 h at RT. Luminescent signal was revealed using WesternBright Sirius detection kit (Advansta). Experiments were repeated at least two times.
Where indicated, band intensities were quantified using the Fiji software (https://imagej.net/Fiji). For each sample, MMP14 band intensity was normalized to the corresponding loading control, relative band intensities are shown.
Luciferase-based reporter assay
5000 HeLa cells/well were seeded in a 96-well plate. Next day cells were transfected in the presence of Lipofectamine2000 with 25 ng of the Renilla reporter, 50 ng of the indicated reporter plasmid, 1000 ng of either pAMC-PROX1 (WT or MUT) or 1000 ng of the corresponding control vector. 36 hours after transfection cells were lysed in 30 µl/well of 1X reporter lysis buffer (Promega). Luciferase activity was measured using a luciferase buffer according to the manufacturer’s instructions (Promega). The assay was repeated two or three times using two biological replicates and the error bars represent SD ± 95%CI across all experiments. For each sample, the firefly luciferase values were normalized to the Renilla luciferase values.
Chromatin immunoprecipitation (ChIP)
iLECs52,57 were transduced with PROX1-pSD4412 and 72 hrs later processed according to simpleChIP Kit (Cell Signalling Technology) instructions. Chromatin was precipitated using Goat anti-human PROX1 antibody (R&D Systems, AF2727) or normal goat IgG antibody (Santa Cruz Biotechnology, sc-2028).
HepG2 and SW620 were transduced with a pSIN-PROX1 WT encoding lentivirus, chromatin was precipitated with mouse monoclonal anti Myc-tag antibody (Cell Signaling Technology #2276) or a control HA-tag mouse monoclonal antibody (HA.11; MMs-101R-200, Biolegend).
After DNA purification (Macherey-Nagel PCR-purification Kit, 740609), the following primers were used to amplify different domains of the MMP14 promoter (forward; reverse):
a: AGGACCTGAAAAGCTTTTCAT; GGGTGGACAGAAATTAGGT
b: TCCTCTATTCCTTCCTTTGCT; AAGCACAACAGAAGCAGG
c: CTTTTCCAGACCTGTCTAGTTC; CACTGAAAAAGGAGGCAATTC
d: TAGAGGTGGAACTAAACCCC; CCTTTAATTGGAACTCGCTGG
e: TGCAGCCACATTACAAATGA; TGTCTATGTCCCTCCCTCTG
f: CATAGACAGTTGTCTACAGGG; TATTGGGGGCTATGTGGCTA
g: CCACATAGCCCCCAATAATT; TTGTGGTGCAGGCTGCCATC
h: AAGTCTCCCACATCCCGTCC; CCAGTGCCCTCCTTTCCTGGTTG
3D sprouting and invasion assays
4000 HuAR2T cells were seeded in the 0.5% low melting point agarose-coated round bottom 96-well plate. Spheroids were allowed to form for 36–48 hours, harvested and embedded in 3 mg/ml plasminogen-free fibrinogen (Calbiochem)20. Spheroids were grown in fibrin for 4 days. Alternatively, a single-cell suspension of 5000 MDA-MB-321 cells were embedded in fibrinogen and followed for 72 h. The fibrin droplets were fixed in 4% PFA and stained with Alexa488-conjugated Phalloidin (Life Technologies), nuclei were counterstained with Hoechst 33342 (1ug/ml). Images were taken using Zeiss LSM-780 as previously reported20.
Mice
Cdh5-CreERT2 and Prox1flox lines were described previously29,30. For induction of Cre recombination, 1 mg of 4-hydroxytamoxifen (4-OHT), dissolved in peanut oil (10 mg/ml), was administered to 3 weeks old mice by intraperitoneal injection. Mice were analysed 1 week after treatment.
Experimental procedures were approved by the Uppsala Laboratory Animal Ethical Committee (permit number:C130/15) and performed in accordance with relevant guidelines and regulations.
Tumour tissues
Paraffin embedded KS sections were kindly obtained from Dr. Justin Weir (Charing Cross Hospital, and the London Clinic, London), The research was covered by Riverside Research Ethics Committee (Study title: Kaposi’s Sarcoma Herpes Virus Infection And Immunity, REC reference: 04/Q0401/80); all patients gave written informed consent.
Tissue microarray of thyroid cancers was constructed from surgical specimens obtained from the archives of the Department of Pathology, Helsinki University Hospital, according to Finnish laws and regulations by permission of the director of the health care unit. The tissue samples were de-identified and analysed anonymously.
Immunohistochemistry
Tissue sections were deparaffinized and rehydrated. Antigen retrieval was performed by heating in 0.1 M citrate buffer (pH 6.0) prior to blocking with 5% normal donkey serum for 1 hour at RT. Sections were stained with the indicated primary antibody O/N at 4 °C in humidified chambers and with the appropriate secondary antibody for 1 hour at RT, nuclei were counterstained with Hoechst 33342 (1ug/ml).
For the whole mount immunofluorescence, the dorsal side of the ear was fixed in 4% paraformaldehyde (PFA), permeabilized in 0.3% Triton X-100 in PBS (PBST), blocked in PBST with 3% milk. Primary antibodies were incubated at 4 °C overnight in blocking buffer. After washing, the samples were incubated with fluorescence-conjugated secondary antibodies before further washing and mounting in Mowiol.
Samples were scanned in a Panoramic 250 viewer (Genome Biology Unit, Research Programs Unit, University of Helsinki). Alternatively, confocal Z-stack images were acquired as previously described20.
Analysis of gene expression database
To assess the reciprocal expression levels of PROX1 and MMP14 the FANTOM5 (http://www.proteinatlas.org/) was analysed. The data were obtained by CAP analysis of gene expression (CAGE) and reported by Tags per million.
Statistical analysis
Student T-test was used to assess whether the difference between the experimental groups was statistically significant. Analyses were performed using GraphPad PRISM6 and 7. Co-localization of proteins in the mouse ears was quantified using JACoP plug-in in the Fiji software (https://imagej.net/Fiji). KS sections were analysed using the ANIMA software58.
The sprouting of single invasive cells in fibrin was quantified with the Cell Profiler pipeline (http://cellprofiler.org/). For each image stack the area of each cell (visualized as Phalloidin-stained area related to one nucleus) was measured. Each stack included approximately 20–40 cells, and for each independent experiment three random stacks were quantified. The invasion of spheroids was quantified from the Phalloidin A488 labelled image by calculating the total area occupied by the spheroid and the area occupied by the central non-invasive area using the Image J software. The average ratio between the total area of the spheroids and the non-invasive body of the spheroid is indicated in the graphs.
Data availability
The gene expression data in Fig. 1d are freely available at http://www.proteinatlas.org. The data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Wigle, J. T., Chowdhury, K., Gruss, P. & Oliver, G. Prox1 function is crucial for mouse lens-fibre elongation. Nat Genet 21, 318–322, https://doi.org/10.1038/6844 (1999).
Burke, Z. & Oliver, G. Prox1 is an early specific marker for the developing liver and pancreas in the mammalian foregut endoderm. Mech Dev 118, 147–155 (2002).
Oliver, G. et al. Prox 1, a prospero-related homeobox gene expressed during mouse development. Mech Dev 44, 3–16 (1993).
Sosa-Pineda, B., Wigle, J. T. & Oliver, G. Hepatocyte migration during liver development requires Prox1. Nat Genet 25, 254–255, https://doi.org/10.1038/76996 (2000).
Lavado, A. & Oliver, G. Prox1 expression patterns in the developing and adult murine brain. Dev Dyn 236, 518–524, https://doi.org/10.1002/dvdy.21024 (2007).
Risebro, C. A. et al. Prox1 maintains muscle structure and growth in the developing heart. Development 136, 495–505, https://doi.org/10.1242/dev.030007 (2009).
Wigle, J. T. & Oliver, G. Prox1 function is required for the development of the murine lymphatic system. Cell 98, 769–778 (1999).
Petrova, T. V. et al. Lymphatic endothelial reprogramming of vascular endothelial cells by the Prox-1 homeobox transcription factor. EMBO J 21, 4593–4599 (2002).
Elsir, T., Smits, A., Lindstrom, M. S. & Nister, M. Transcription factor PROX1: its role in development and cancer. Cancer Metastasis Rev 31, 793–805, https://doi.org/10.1007/s10555-012-9390-8 (2012).
Elsir, T. et al. PROX1 is a predictor of survival for gliomas WHO grade II. Br J Cancer 104, 1747–1754, https://doi.org/10.1038/bjc.2011.162 (2011).
Petrova, T. V. et al. Transcription factor PROX1 induces colon cancer progression by promoting the transition from benign to highly dysplastic phenotype. Cancer Cell 13, 407–419, https://doi.org/10.1016/j.ccr.2008.02.020 (2008).
Ragusa, S. et al. PROX1 promotes metabolic adaptation and fuels outgrowth of Wnt(high) metastatic colon cancer cells. Cell Rep 8, 1957–1973, https://doi.org/10.1016/j.celrep.2014.08.041 (2014).
Skog, M. et al. Expression and prognostic value of transcription factor PROX1 in colorectal cancer. Br J Cancer 105, 1346–1351, https://doi.org/10.1038/bjc.2011.297 (2011).
Shimoda, M. et al. A homeobox protein, prox1, is involved in the differentiation, proliferation, and prognosis in hepatocellular carcinoma. Clin Cancer Res 12, 6005–6011, https://doi.org/10.1158/1078-0432.CCR-06-0712 (2006).
Laerm, A. et al. Prospero-related homeobox 1 (PROX1) is frequently inactivated by genomic deletions and epigenetic silencing in carcinomas of the bilary system. J Hepatol 46, 89–97, https://doi.org/10.1016/j.jhep.2006.07.033 (2007).
Chang, T. M. & Hung, W. C. Transcriptional repression of TWIST1 gene by Prospero-related homeobox 1 inhibits invasiveness of hepatocellular carcinoma cells. FEBS Lett 586, 3746–3752, https://doi.org/10.1016/j.febslet.2012.08.034 (2012).
Laitinen, A. et al. High PROX1 expression in gastric cancer predicts better survival. PLoS One 12, e0183868, https://doi.org/10.1371/journal.pone.0183868 (2017).
Yarchoan, R. & Uldrick, T. S. HIV-Associated Cancers and Related Diseases. N Engl J Med 378, 1029–1041, https://doi.org/10.1056/NEJMra1615896 (2018).
Benevenuto de Andrade, B. A. et al. Expression of PROX-1 in oral Kaposi’s sarcoma spindle cells. J Oral Pathol Med 43, 132–136, https://doi.org/10.1111/jop.12097 (2014).
Cheng, F. et al. KSHV-initiated notch activation leads to membrane-type-1 matrix metalloproteinase-dependent lymphatic endothelial-to-mesenchymal transition. Cell Host Microbe 10, 577–590, https://doi.org/10.1016/j.chom.2011.10.011 (2011).
Wang, H. W. et al. Kaposi sarcoma herpesvirus-induced cellular reprogramming contributes to the lymphatic endothelial gene expression in Kaposi sarcoma. Nat Genet 36, 687–693, https://doi.org/10.1038/ng1384 (2004).
Yoo, J. et al. Kaposin-B enhances the PROX1 mRNA stability during lymphatic reprogramming of vascular endothelial cells by Kaposi’s sarcoma herpes virus. PLoS Pathog 6, e1001046, https://doi.org/10.1371/journal.ppat.1001046 (2010).
Miettinen, M. & Wang, Z. F. Prox1 transcription factor as a marker for vascular tumors-evaluation of 314 vascular endothelial and 1086 nonvascular tumors. Am J Surg Pathol 36, 351–359, https://doi.org/10.1097/PAS.0b013e318236c312 (2012).
Choi, D. et al. Aberrant Activation of Notch Signaling Inhibits PROX1 Activity to Enhance the Malignant Behavior of Thyroid Cancer Cells. Cancer Res 76, 582–593, https://doi.org/10.1158/0008-5472.CAN-15-1199 (2016).
Duncan, M. K., Cui, W., Oh, D. J. & Tomarev, S. I. Prox1 is differentially localized during lens development. Mech Dev 112, 195–198 (2002).
Lohi, J., Lehti, K., Valtanen, H., Parks, W. C. & Keski-Oja, J. Structural analysis and promoter characterization of the human membrane-type matrix metalloproteinase-1 (MT1-MMP) gene. Gene 242, 75–86 (2000).
Shin, J. W. et al. Prox1 promotes lineage-specific expression of fibroblast growth factor (FGF) receptor-3 in lymphatic endothelium: a role for FGF signaling in lymphangiogenesis. Mol Biol Cell 17, 576–584, https://doi.org/10.1091/mbc.E05-04-0368 (2006).
Charest-Marcotte, A. et al. The homeobox protein Prox1 is a negative modulator of ERR{alpha}/PGC-1{alpha} bioenergetic functions. Genes Dev 24, 537–542, https://doi.org/10.1101/gad.1871610 (2010).
Martinez-Corral, I. et al. Nonvenous origin of dermal lymphatic vasculature. Circulation research 116, 1649–1654, https://doi.org/10.1161/CIRCRESAHA.116.306170 (2015).
Wang, Y. et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature 465, 483–486, https://doi.org/10.1038/nature09002 (2010).
Lee, S. et al. Prox1 physically and functionally interacts with COUP-TFII to specify lymphatic endothelial cell fate. Blood 113, 1856–1859, https://doi.org/10.1182/blood-2008-03-145789 (2009).
Lin, F. J. et al. Direct transcriptional regulation of neuropilin-2 by COUP-TFII modulates multiple steps in murine lymphatic vessel development. J Clin Invest 120, 1694–1707, https://doi.org/10.1172/JCI40101 (2010).
Genis, L., Galvez, B. G., Gonzalo, P. & Arroyo, A. G. MT1-MMP: universal or particular player in angiogenesis? Cancer Metastasis Rev 25, 77–86, https://doi.org/10.1007/s10555-006-7891-z (2006).
Iida, J. et al. Membrane type-1 matrix metalloproteinase promotes human melanoma invasion and growth. J Invest Dermatol 122, 167–176, https://doi.org/10.1046/j.0022-202X.2003.22114.x (2004).
Jiang, W. G. et al. Expression of membrane type-1 matrix metalloproteinase, MT1-MMP in human breast cancer and its impact on invasiveness of breast cancer cells. Int J Mol Med 17, 583–590 (2006).
Ranjan, A. & Kalraiya, R. D. Invasive potential of melanoma cells correlates with the expression of MT1-MMP and regulated by modulating its association with motility receptors via N-glycosylation on the receptors. Biomed Res Int 2014, 804680, https://doi.org/10.1155/2014/804680 (2014).
Tolde, O. et al. Neoplastic progression of the human breast cancer cell line G3S1 is associated with elevation of cytoskeletal dynamics and upregulation of MT1-MMP. Int J Oncol 36, 833–839 (2010).
Ohuchi, E. et al. Membrane type 1 matrix metalloproteinase digests interstitial collagens and other extracellular matrix macromolecules. J Biol Chem 272, 2446–2451 (1997).
d’Ortho, M. P. et al. Membrane-type matrix metalloproteinases 1 and 2 exhibit broad-spectrum proteolytic capacities comparable to many matrix metalloproteinases. Eur J Biochem 250, 751–757 (1997).
Hiraoka, N., Allen, E., Apel, I. J., Gyetko, M. R. & Weiss, S. J. Matrix metalloproteinases regulate neovascularization by acting as pericellular fibrinolysins. Cell 95, 365–377 (1998).
Miyoshi, A. et al. Snail accelerates cancer invasion by upregulating MMP expression and is associated with poor prognosis of hepatocellular carcinoma. Br J Cancer 92, 252–258, https://doi.org/10.1038/sj.bjc.6602266 (2005).
Ip, Y. C., Cheung, S. T., Leung, K. L. & Fan, S. T. Mechanism of metastasis by membrane type 1-matrix metalloproteinase in hepatocellular carcinoma. World J Gastroenterol 11, 6269–6276 (2005).
Murakami, K. et al. Invasiveness of hepatocellular carcinoma cell lines: contribution of membrane-type 1 matrix metalloproteinase. Neoplasia 1, 424–430 (1999).
Shay, G., Lynch, C. C. & Fingleton, B. Moving targets: Emerging roles for MMPs in cancer progression and metastasis. Matrix Biol 44-46, 200–206, https://doi.org/10.1016/j.matbio.2015.01.019 (2015).
Sugiyama, N. et al. EphA2 cleavage by MT1-MMP triggers single cancer cell invasion via homotypic cell repulsion. J Cell Biol 201, 467–484, https://doi.org/10.1083/jcb.201205176 (2013).
Turunen, S. P., Tatti-Bugaeva, O. & Lehti, K. Membrane-type matrix metalloproteases as diverse effectors of cancer progression. Biochim Biophys Acta 1864, 1974–1988, https://doi.org/10.1016/j.bbamcr.2017.04.002 (2017).
Johnson, J. L. et al. Regulation of matrix metalloproteinase genes by E2F transcription factors: Rb-Raf-1 interaction as a novel target for metastatic disease. Cancer Res 72, 516–526, https://doi.org/10.1158/0008-5472.CAN-11-2647 (2012).
Haas, T. L., Stitelman, D., Davis, S. J., Apte, S. S. & Madri, J. A. Egr-1 mediates extracellular matrix-driven transcription of membrane type 1 matrix metalloproteinase in endothelium. J Biol Chem 274, 22679–22685 (1999).
Petrella, B. L., Lohi, J. & Brinckerhoff, C. E. Identification of membrane type-1 matrix metalloproteinase as a target of hypoxia-inducible factor-2 alpha in von Hippel-Lindau renal cell carcinoma. Oncogene 24, 1043–1052, https://doi.org/10.1038/sj.onc.1208305 (2005).
Du, H. T. & Liu, P. Matrix metalloproteinase 14 participates in corneal lymphangiogenesis through the VEGF-C/VEGFR-3 signaling pathway. Exp Ther Med 12, 2120–2128, https://doi.org/10.3892/etm.2016.3601 (2016).
Wong, H. L. et al. MT1-MMP sheds LYVE-1 on lymphatic endothelial cells and suppresses VEGF-C production to inhibit lymphangiogenesis. Nat Commun 7, 10824, https://doi.org/10.1038/ncomms10824 (2016).
Norrmen, C. et al. FOXC2 controls formation and maturation of lymphatic collecting vessels through cooperation with NFATc1. J Cell Biol 185, 439–457, https://doi.org/10.1083/jcb.200901104 (2009).
Harvey, N. L. et al. Lymphatic vascular defects promoted by Prox1 haploinsufficiency cause adult-onset obesity. Nat Genet 37, 1072–1081, https://doi.org/10.1038/ng1642 (2005).
Sheridan, J. M. et al. A pooled shRNA screen for regulators of primary mammary stem and progenitor cells identifies roles for Asap1 and Prox1. BMC Cancer 15, 221, https://doi.org/10.1186/s12885-015-1187-z (2015).
Versmold, B. et al. Epigenetic silencing of the candidate tumor suppressor gene PROX1 in sporadic breast cancer. Int J Cancer 121, 547–554, https://doi.org/10.1002/ijc.22705 (2007).
Norrmen, C. et al. Liprin (beta)1 is highly expressed in lymphatic vasculature and is important for lymphatic vessel integrity. Blood 115, 906–909, https://doi.org/10.1182/blood-2009-03-212274 (2010).
Vart, R. J. et al. Kaposi’s sarcoma-associated herpesvirus-encoded interleukin-6 and G-protein-coupled receptor regulate angiopoietin-2 expression in lymphatic endothelial cells. Cancer Res 67, 4042–4051, https://doi.org/10.1158/0008-5472.CAN-06-3321 (2007).
Rantanen, V., Valori, M. & Hautaniemi, S. Anima: modular workflow system for comprehensive image data analysis. Front Bioeng Biotechnol 2, 25, https://doi.org/10.3389/fbioe.2014.00025 (2014).
Acknowledgements
We are grateful to Justin Weir (Charing Cross Hospital, and the London Clinic, London) for providing the KS tumour sections, Katri Koli (University of Helsinki) for providing the HepG2 cell line, Thomas F. Schulz (Hannover Medical School, Germany) for providing HuAR2T cell line and to Ralf Adams (Max Planck Institute for Molecular Biomedicine, Muenster) for providing the Cdh5-CreERT2 mice. Mikko Turunen (University of Helsinki) is acknowledged for his help in optimizing the ChIP. We thank also Raquel Diaz Martinez, Lotta Honkala, Nadezhda Zinovkina, Veronika Rezov, Sari Tynkkynen and Jenny Bärlund for the technical assistance. We acknowledge the Bioimaging Unit (University of Helsinki) and the Genome-Biology Unit (University of Helsinki) for the technical support in imaging. The study was supported by the Centre of Excellence grant (Translational Cancer Biology; P.M.O., K.A.) and a postdoctoral researcher grant (S.G.) from the Academy of Finland, Finnish Cancer Foundations (P.M.O, K.L.), Sigrid Juselius Foundation (P.M.O.), and University of Helsinki Foundations (P.M.O) and University of Helsinki Foundations (P.M.O). E.K. was supported by the Helsinki Biomedical Graduate Program (HBGP; University of Helsinki). We also acknowledge funding from the Karolinska Institutet (K.L.), the Swiss National Science Foundation (PPP0033-114898 to T.V.P.) and the Swedish Research Council (542-2014-3535 to T. M.).
Author information
Authors and Affiliations
Contributions
Conceptualization: S.G., K.L., P.M.O. Methodology: S.G., I.M.C., M.V.K., E.E., T.M., T.V.P. Software: S.G.,V.R., S.H. Validation: S.G., P.M.O. Formal analysis: S.G., J.C., V.R. Investigation: S.G., J.C., I.M.C., M.V.K., E.E., E.K., K.T. Resources: M.B., T.M., C.H., K.L., T.V.P., K.A., P.M.O. Writing –original draft: S.G., P.M.O. Writing-review and editing: S.G., E.E., M.B., T.M., T.V.P., C.H., K.A., S.H., K.L., P.M.O. Visualization: S.G., I.M.C. Supervision: S.G., P.M.O. Project administration: P.M.O. Funding acquisition: P.M.O.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gramolelli, S., Cheng, J., Martinez-Corral, I. et al. PROX1 is a transcriptional regulator of MMP14. Sci Rep 8, 9531 (2018). https://doi.org/10.1038/s41598-018-27739-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-27739-w
This article is cited by
-
The MMP14–caveolin axis and its potential relevance for lipoedema
Nature Reviews Endocrinology (2020)
-
Expression of Prox1 in Medullary Thyroid Carcinoma Is Associated with Chromogranin A and Calcitonin Expression and with Ki67 Proliferative Index, but Not with Prognosis
Endocrine Pathology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.