Abstract
Skin trait variation impacts quality-of-life, especially for females from the viewpoint of beauty. To investigate genetic variation related to these traits, we conducted a GWAS of various skin phenotypes in 11,311 Japanese women and identified associations for age-spots, freckles, double eyelids, straight/curly hair, eyebrow thickness, hairiness, and sweating. In silico annotation with RoadMap Epigenomics epigenetic state maps and colocalization analysis of GWAS and GTEx Project eQTL signals provided information about tissue specificity, candidate causal variants, and functional target genes. Novel signals for skin-spot traits neighboured AKAP1/MSI2 (rs17833789; P = 2.2 × 10−9), BNC2 (rs10810635; P = 2.1 × 10−22), HSPA12A (rs12259842; P = 7.1 × 10−11), PPARGC1B (rs251468; P = 1.3 × 10−21), and RAB11FIP2 (rs10444039; P = 5.6 × 10−21). HSPA12A SNPs were the only protein-coding gene eQTLs identified across skin-spot loci. Double edged eyelid analysis identified that a signal around EMX2 (rs12570134; P = 8.2 × 10−15) was also associated with expression of EMX2 and the antisense-RNA gene EMX2OS in brain putamen basal ganglia tissue. A known hair morphology signal in EDAR was associated with both eyebrow thickness (rs3827760; P = 1.7 × 10−9) and straight/curly hair (rs260643; P = 1.6 × 10−103). Excessive hairiness signals’ top SNPs were also eQTLs for TBX15 (rs984225; P = 1.6 × 10−8), BCL2 (rs7226979; P = 7.3 × 10−11), and GCC2 and LIMS1 (rs6542772; P = 2.2 × 10−9). For excessive sweating, top variants in two signals in chr2:28.82-29.05 Mb (rs56089836; P = 1.7 × 10−11) were eQTLs for either PPP1CB or PLB1, while a top chr16:48.26–48.45 Mb locus SNP was a known ABCC11 missense variant (rs6500380; P = 6.8 × 10−10). In total, we identified twelve loci containing sixteen association signals, of which fifteen were novel. These findings will help dermatologic researchers better understand the genetic underpinnings of skin-related phenotypic variation in human populations.
Similar content being viewed by others
Introduction
Skin phenotypes such as freckles, hairiness, and excessive sweating can be serious problems for some individuals. Investigations among Japanese women (n~854 to 4345) for particular beauty problems revealed the following concerns broken down by individuals in particular age ranges and in order of significance: 20s) dry skin, large pores, and acne; 30s) freckles, dry skin, and large pores; 40s) freckles, wrinkles, and sagging skin; and 50s and older) wrinkles, freckles, and sagging skin1,2. These concerns often impact an individual’s Quality-of-Life (QOL), and there have been major developments in medical cosmetic treatment and the cosmetic industry in recent years to address these problems.
Review of patient family histories has suggested that genetic factors may be related to some of these phenotypes, and a number of studies have revealed potential causative genes for certain skin-related phenotypes. The thickness of human hair fibers is very differentiated across world-wide populations, and almost a decade ago, researchers identified its association with a non-synonymous variant in the ectodysplasin A receptor gene (EDAR)3,4. Recently, it was shown that the A481T and H615R alleles of OCA2, which is a causal gene for oculocutaneous albinism, are correlated with the skin color of Japanese people5,6,7. Several genome-wide association studies (GWAS) and candidate gene analyses in European ancestry population samples have identified variants associated with skin pigmentation in or near ASIP, HERC2, IRF4, MC1R, OCA2, SLC24A4, TYR, and BNC28,9,10, while IRF4, MC1R, ASIP, and BNC2 were also found to be associated with freckles and facial pigmented spots11,12. Associations were also identified for MC1R and ASIP with red hair color, TCHH and WNT10A with hair curl, and OCA2, IRF4, SLC45A2, SLC24A4, and MC1R with blond versus brown hair color. A more recent 23andMe GWAS report of 42 human phenotypes found multiple loci associated with skin-related traits such as male-pattern baldness, unibrow, chin dimples, and nose size13. However, most large-scale association studies have been performed in population samples with predominantly European ancestry. Analysis of traits in multiple ethnicities is an important part of modern GWAS analyses14,15,16, and a GWAS analysis of multiple skin-related traits in an East Asian population sample should help identify new loci and confirm and or refine previously reported association signals from those studies. If new genes and variants related to these phenotypes can be identified, it may be possible to decrease cosmetic problems and medical care costs by providing instructions on lifestyle habits, diet, and makeup, as well as preventive measures such as use of custom-made cosmetics and therapeutic intervention in the initial stage of symptom development.
Here, we report on a GWAS of skin-related phenotypes that analyzed over eleven-thousand Japanese females and identified both novel loci and refined known association signals. Use of epigenetic state data combined with colocalization analysis of GWAS and eQTL signals allowed us to narrow down the lists of candidate causal variants, identify likely target genes, and better understand the biologic functions of a number of genes with important effects that were detected in this study.
Results
We performed a genome-wide association study (GWAS) of Japanese female subjects who answered a questionnaire about various phenotypes and provided DNA for genetic analysis of those traits. The data was collected in two study-stages, termed LL01 and LL02 (LL01 = 5750, LL02 = 5628), with a detailed description of the dataset and methods used for genotyping, imputation, and annotation available in a recently published GWAS report of self-reported food reactions that used the same set of samples17. For the current report, we analyzed up to 11311 subjects who fulfilled quality-control (QC) criteria and provided self-reported phenotype information on the presence of age spots, freckles, double-edged eyelids, eyebrow thickness (thick vs. thin eyebrows), hair morphology (straight vs. curly hair), excessive hairiness, and excessive sweating (Supplementary Table S1; plot of principal component analysis in Supplementary Fig. S1). For several phenotypes (double-edged eyelid, eyebrow thickness, hair morphology, and excessive sweating), trait data was only available from the LL02 study-stage. Meta-analysis of the two GWAS stages was performed using 536506 QC+ variants from a custom Affymetrix Axiom array, for which there was negligible inflation of genome-wide statistics observed across the seven phenotypes (λGC: 1.0031–1.043; Supplementary Fig. S2). A summary-statistics based approach18 was used to impute genome-wide missing genotypes in order to produce the Manhattan plot (Fig. 1) and to allow for analysis of data from previous reports, but for the purposes of the association study analyses, we defined associated genomic regions as those with genotyped SNPs that achieved a multiple-testing adjusted P < 1.73 × 10−8 (see Methods). Within each associated region, we then performed genotype-based imputation19,20,21 and step-wise conditioning on top SNPs to identify independent secondary signals (P conditioned < 1 × 10−5). Across the seven skin-related phenotypes, we identified twelve separate loci made up of sixteen independent association signals. Fifteen signals were novel associations for the current phenotype being examined (Table 1). The following sections will describe each of the signals in the context of functional annotations and eQTL signals (see Methods) in order to help identify candidate causal variants and target genes.
Meta-analysis Manhattan plots for skin-related phenotypes. Manhattan plots of −log10(P value) for skin phenotype GWAS analyses after summary statistics-based imputation using 1000 Genomes Project Phase 1 Release 3 reference data. Top association signals were labeled with up to two annotated genes from Supplementary Worksheet S1 for variants with r2 > 0.8 to the top SNP. Peaks with more than two genes overlap more than one independent association signal. The horizontal red line denotes the multiple testing corrected P-value cutoff of 1.73 × 10−8 that was used for identifying association signals.
Age spots and freckles association signals
Previously reported freckles/facial pigmented spots association signals identified in two GWAS studies using Northern European ancestry samples11,12 displayed only weak evidence for replication in our Japanese dataset (Table 2; Supplementary Table S2). Both studies’ top SNPs in the basonuclin 2 gene (BNC2) locus were only nominally significant in our data (Prs2153271 = 0.0044; Prs62543565 = 0.0351), and in the interferon regulatory factor 4 (IRF4), melanocortin 1 receptor (MC1R), and agouti signaling protein (ASIP) loci, their top SNPs were predominantly monomorphic in 1000G East Asian samples (Supplementary Table S2) or were only weakly significant in our data (Table 2; P IRF4,rs9405675 = 0.0113; P MC1R:rs8060934 = 0.0027). Two previous reports12,22 had also constructed a compound heterozygosity score using six MC1R missense variants: rs1805005: Val60Leu; rs1805007: Arg151Gly; rs1805008: Arg160Trp; rs1805009: Asp294His; rs2228479: Val92Met; rs885479: Arg163Gln. The first four listed SNPs were monomorphic in Japanese, and one of the two polymorphic variants was nominally associated with freckles in our dataset (Prs2228479 = 0.00044; Prs885479 = 0.61053), but neither was associated with age-spots.
In the current study, we further identified five loci associated with age-spots and/or freckles (Table 1). Two were associated with both phenotypes at genome-wide significance, and three were genome-wide significant only for the freckles phenotype (Fig. 1). To examine whether variants in those GWAS signals had evidence as regulatory variants, we performed colocalization analysis of GTEx Portal23 eQTL statistics from their single-tissue and multi-tissue Metasoft modified random effects (RE2)24,25 analyses using the Approximate Bayes Factor (ABF)26 and Summary-data based Mendelian Randomization (SMR)27 methods (see Methods). GWAS/eQTL signal pairs that were at least nominally significant by the SMR test-of-linkage (P SMR < 0.05), non-significant for SMR’s HEIDI test-of-pleiotropy (HEIDI: heterogeneity in dependent instruments; P HEIDI ≥ 0.05) and had posterior probability of colocalization from the ABF test PP H4.ABF > 0.3 were considered as having nominal evidence for colocalization, while pairs with PP H4.ABF > 0.5 and PP H4.ABF > 0.9 were considered as having moderate and strong support, respectively. Lower values of P SMR and higher values of P HEIDI also provide stronger evidence of colocalization/pleiotropy of causal variants at a GWAS and an eQTL signal.
High LD SNPs in the top chr9:16.79–16.81 Mb locus (rs10810635; P freckles = 2.09 × 10−22; Supplementary Fig. S3a) overlapped introns of BNC2, with some overlapping RoadMap Epigenomics28,29 predicted enhancer activity as well as DNase hypersensitive sites (DHS) and Transcription factor binding sites (TFBS) identified in the ReMap 2018 database30,31 70 kb upstream of an alternative promoter (Supplementary Worksheet S3; Supplementary Fig. S3c). Visualization of the 25-state chromatin state model in 127 reference epigenomes also found that for three SNPs (rs10810635, rs67920508, rs16935073) active enhancer function was predicted in skin-spots relevant tissue samples such as foreskin fibroblasts and foreskin melanocytes (http://egg2.wustl.edu/roadmap/web_portal/imputed.html#chr_imp). The colocalization tests showed nominal support for antisense RNA RP11-62F24.2 eQTLs in only a single-tissue (Supplementary Fig. S3b). That ncRNA overlaps the alternative BNC2 promoter, suggesting that these SNPs may indirectly regulate BNC2 expression in a tissue or cell-type restricted manner.
The age-spots/freckles signal in the Peroxisome proliferator-activated receptor gamma coactivator 1 beta (PPARGC1B) gene at chr5:149.19–149.23 Mb (Supplementary Fig. S4; Supplementary Worksheet S1; top SNP rs251468; P age spots = 6.24 × 10−12; P freckles = 1.27 × 10−21) had four of five high LD (r2 equiv > 0.8; Supplementary Worksheet S3) freckles variants overlapping predicted enhancer function, and the top variant had function predicted in twelve tissue/cell types while also overlapping TFBS for multiple TFs. No SNPs were eQTLs in GTEx Portal data, so the functional impact of these SNPs on PPARGC1B could not be confirmed from in silico analysis.
The chr10:119.55–119.60 Mb region downstream of the RAB11 family interacting protein 2 (RAB11FIP2) contained three independent freckles and two independent age-spots signals (Supplementary Figs S5a–c and S6a,b; Table 1). We observed overlap between high LD variants in these signals and predicted epigenetic activity (Supplementary Figs S5d and S6c; Supplementary Worksheets S2 and S3), but there was no indication from eQTL data that linked putative regulatory function to target genes.
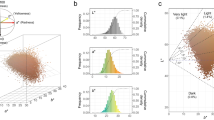
A nearby unlinked freckles association signal at chr10:118.45–118.48 Mb overlapped the heat shock protein family A (Hsp70) member 12A (HSPA12A) gene (Fig. 2a; top SNP rs12259842; P = 7.08 × 10−11). We observed two groups of moderate/high LD SNPs based on effect-allele frequency (EAF ~0.34 vs ~0.25, respectively) and haplotype cluster analysis. The two haplotype clusters (HAPS1 and HAPS3) overlapped the effect-allele (set to minor allele in Japanese) at those SNPs in combination or by HAPS1 alone (Supplementary Fig. S7). HAPS1 versus HAPS1-3 SNPs showed only slight differences in the association statistics (P HAPS1 ~ 2 × 10−10 vs P HAPS1-3 ~ 8 × 10−10). Similarly, they clustered together as strongly associated with HSPA12A expression in the two freckles pertinent GTEx skin tissue samples (Fig. 2b), but only HAPS1-3 SNPs were strongly significant eQTLs in non-skin tissue samples (bottom panels of Fig. 2b; Supplementary Fig. S8b). Both ABF and the SMR test-of-linkage were strongly supportive of colocalization of GWAS and eQTL signals, but HEIDI was only non-significant (supporting pleiotropy) in the sun un-exposed skin tissue, for which both HAPS1 and HAPS1-3 SNPs clustered closest together. In epigenetics data, four peaks overlapped the GWAS SNPs, with all four overlapping HAPS1-3 SNPs and two overlapping HAPS1 SNPs (Fig. 2c; Supplementary Worksheet S3; approx. peak centers: peak #1 = 118.448 Mb, peak #2 = 118.451 Mb, peak #3 = 118.454 Mb, and peak #4 = 118.458 Mb). In peak #1, two HAPS1-3 SNPs (rs1900500, rs2921967), overlapped promoter and enhancer function predicted in large numbers of tissue/cell types. In a 1 kb region of peak #2, enhancer function was predicted in a larger number of tissue/cell types (from 60–70 tissue/cell types) for seven HAPS1-3 variants, while in peak #3, HAPS1-3 SNP rs2907231 overlapped enhancer elements identified in forty tissue/cell types and lay in a region bound by forty TFs. Given that HAPS1 eQTLs were restricted to skin tissue samples, we expected a causal regulatory SNP to have epigenetic function predicted only in a restricted set of tissues that included skin cell types. Of the top five GWAS HAPS1 SNPs, two did not overlap any predicted epigenetic function and one SNP overlapped enhancer function predicted in a large number of tissues, although skin tissue samples were not among them. However, the other two top HAPS1 SNPs (rs3010484, rs2907229), which lay in peak #4, had enhancer function predicted in only a limited number of tissue types (skin, brain) including one foreskin melanocytes tissue sample. Those two SNPs were also the strongest eQTLs in both GTEx skin tissues, resided right next to each other, and appeared to be perfectly linked with each other in each 1000 G population based on allele frequencies (Supplementary Worksheet S3). Our results suggest that at least three HAPS1-3 variants regulate HSPA12A expression in a multi-tissue fashion, while a pair of HAPS1 SNPs have regulatory activity at least partially restricted to skin tissues.
Chr10:118.45–118.48 Mb (HSPA12A) freckles locus. Freckles associated SNPs possess strong evidence for colocalization with single-tissue and multi-tissue HSPA12A eQTL signals. (a) Regional association plots of −log10(P-values) around the Chr10:118.45–118.48 Mb (HSPA12A) freckles association locus. Top sub-panel presents points sized by r2 to the top GWAS SNP and coloured either black or by haplotype cluster HAPS1 or HAPS1-3 assignment (legend in the upper left corner). The bottom sub-panel shows SNPs with and without conditioning on the top SNP. (b) Shows −log10(P-values) for HSPA12A eQTL data for single-tissue and multi-tissue Metasoft RE2 analyses, with labels at the right-hand side. Points are sized and coloured the same as in the top (a) sub-panel. Colocalization statistics from ABF and SMR methods and the percent of mod. LD GWAS SNPs overlapping mod. LD eQTL SNPs are shown at the left of each sub-panel as an inlayed table. GENCODE gene models are shown below the eQTL plots. (c) presents output from the WashU EpiGenome Browser of an epilogos plot of the Roadmap Epigenomics 25-state imputed model of epigenetic states along with tracks of high LD candidate causal variants divided by assigned haplotype cluster(s) and GENCODE transcript models in the region. All panels are plotted on the same x-axis coordinates.
The final signal (top SNP rs17833789; P freckles = 2.19 × 10−9) lay 2–3 kb downstream of the A-kinase anchoring protein 1 (AKAP1) gene and 7–10 kb upstream of the musashi RNA binding protein 2 (MSI2) gene in the chr17:55.21−55.25 Mb region (Supplementary Fig. S9a). High LD SNPs overlapped enhancer function predicted in only a small number of tissue samples (Supplementary Fig. S9b; Supplementary Worksheet S3), and enhancer activity for three of those (rs17833789, rs62060349, rs9907841) was only identified in melanocyte skin tissue samples. No SNPs were eQTLs for either AKAP1 or MSI2, but analysis of FANTOM5 CAGE expression data32,33 pointed to a functional difference for MSI2 compared to AKAP1. AKAP1 displayed similar RLE expression values in keratinocytes and both dark and light-skin subjects’ melanocytes (mean Keratinocytes = 17.6; mean dark.melanocytes = 9.1; mean light.melanocytes = 14.0). In contrast, MSI2 exhibited low expression levels in both keratinocytes and dark skin melanocytes (mean Keratinocytes = 3.8; mean dark.melanocytes = 4.5), but values in light-skin subjects’ melanocytes were approximately six-times higher (mean light.melanocytes = 27.8).
The seven top SNPs across the skin-spots loci accounted for 5.2% of freckles phenotypic variance (calculated as a “pseudo-R2”), while the three age-spots signals explained 1.36% of its variance (Supplementary Table S5). We observed a large degree of overlap between the two phenotypes (Supplementary Table S6), with 47.8% (3553/7438) of age-spots cases also being freckles cases, and 88.1% (3553/4034) of freckles cases being age-spots cases. We constructed a genetic risk score (GRS) from the regression model coefficients and found that the proportion of freckles (any versus strong freckling) increased linearly with respect to GRS bin (Supplementary Fig. S10).
Double eyelid association signals
The chr10:119.25–119.35 Mb locus contained two novel independent signals associated with the double eyelid phenotype (Fig. 3a,b; Supplementary Worksheet S4), and SNPs in both signals (Signal #1: chr10:119.25–119.32 Mb: rs12570134: P = 8.15 × 10−15; Signal #2: chr10:119.33–119.35 Mb: rs1415425: Pcond. = 1.90 × 10−7) were within or near the empty spiracles homeobox 2 (EMX2) gene or the antisense long non-coding RNA (lncRNA) gene EMX2OS. Two signal #1 SNPs overlapped predicted promoter activity (rs12777466, rs12777755: Supplementary Worksheet S4), were conserved in both GERP and SiPhy annotations, lay within the EMX2 5′-UTR, and both were predicted in HaploReg to cause a TF motif change. Notably, promoter activity was predicted in tissues pertinent to facial development34,35, such as human embryonic stem cell (ESC) derived CD56+ mesoderm cells. Signal #2 displayed weak functional evidence, with only two of sixty-five high LD intergenic SNPs overlapping any epigenetic annotation (Fig. 3e). GWAS/eQTL analysis using brain putamen (basal ganglia) tissue samples identified moderate to strong support for colocalization/pleiotropy of signal #1 with EMX2 (Fig. 3c) and EMX2OS eQTLs (Fig. 3d), respectively.
Chr10:119.25–119.32 Mb (EMX2/EMX2OS2) double-edged eyelid locus. Two independent signals were identified in the chr10:119.25–119.32 (EMX/EMX2OS2) double-edged eyelid association locus, and Signal 1 SNPs displayed moderate and strong evidence for colocalization with eQTL signals for EMX2 and the anti-sense RNA gene EMX2OS, respectively. (a,b) Present regional association plots of −log10(P-values) for Signal 1 and 2, respectively. (c,d) Show −log10(P-values) from Brain_Putamen_basal_ganglia tissue samples for association with EMX2 and EMX2OS expression, respectively. GWAS and eQTL panels are configured as described in the Fig. 2 legend, but eQTL panel point colours denote different independent eQTL signals, with the top signals in a region in green and orange. GENCODE gene models are shown below the eQTL plots. (e) Presents output from the WashU EpiGenome Browser of an epilogos plot of the Roadmap Epigenomics 25-state imputed model of epigenetic states along with tracks of high LD signal 1 and signal 2 variants and GENCODE transcript models in the region.
Replication analysis of previous hair-related trait associations
We looked for replication/overlap with known signals involving similar hair-related traits. We mapped previously analyzed GWAS phenotypes to one from the current study (phenotype mapping of previous = current phenotypes: hair shape = hair morphology; eyebrow thickness, unibrow, or beard thickness = excessive hairiness; eyebrow thickness or unibrow = eyebrow thickness) and extracted any SNPs achieving FDR < 0.1 (Table 2). The strongest replication was for a previous association with hair shape in the EDAR gene (Table 2). However, in general, previous associations displayed little overlap with our results. None of four strong associations with beard thickness (P < 5 × 10−8) were validated in our excessive hairiness data, but one of four suggestive associations achieved P = 2.9 × 10−6 in our data (rs11121667 in C1orf127; P reported = 3.8 × 10−7; Table 2 and Supplementary Table S3a)36. Only one of ten previous eyebrow thickness associated SNPs was validated in our eyebrow thickness and excessive hairiness analyses (rs3827760; P reported = 1.2 × 10−7; Table 2 and Supplementary Table S3b,c), while 19 of 57 SNPs previously associated with unibrow were low frequency/monomorphic in Japanese or not present in 1000G, and only one of seven suggestive and six of 50 strong unibrow associations could be replicated in our data. Five of the replicated unibrow signals were associated with both eyebrow thickness and hairiness traits (Table 2; Supplementary Table S3d,e). The strongest of the non-EDAR signals that replicated lay in the chromosome 3 SOX2-ATP11B gene region, and our top hairiness SNP in the region matched that reported by Pickrell et al.13 for unibrow (P rs1345417 = 7.57 × 10−9).
Pan-hair related phenotype association signals at chr2:108.93–109.57 Mb
For the three hair-related traits (hair morphology, eyebrow thickness, and excessive hairiness), we identified overlapping association signals in and around the EDAR gene on chromosome 2 (Supplementary Fig. S11a–c). The eyebrow thickness and hair morphology associations almost completely shared high LD SNPs (Supplementary Worksheets S5 and S6), and a known missense variant (rs3827760) that has been associated with multiple traits3,4,37,38,39,40 ranked first and second across the two signals, respectively (rs3827760: P eyebrows = 1.71 × 10−9; P straight hair = 3.14 × 10−103). As noted above, our results replicate a previously reported association between that variant and straight vs. curly hair in Asian population samples37 (Table 2; Supplementary Tables S3 and S4).
In contrast, top imputed excessive hairiness variants in the chr2:108.93–109.57 Mb locus (r2 > 0.8: n = 230; top SNP rs6542772: P = 2.16 × 10−9; Supplementary Worksheet S7) were only moderately linked to rs3827760 (r2 = 0.507) and did not overlap EDAR. Rather, they resided within or near the LIM zinc finger domain containing 1 (LIMS1), the GRIP and coiled-coil domain-containing protein 2 (GCC2), or RAN binding protein 2 (RANBP2) genes (Supplementary Fig. S11d). Many of those variants were GTExPortal eQTLs for LIMS1 and GCC2 as well as the antisense RNAs LIMS1-AS1 (AC010095.5) and GCC2-AS1 (AC012487.2) (Supplementary Worksheets S7). Colocalization analysis identified strong support for LIMS1-AS1 in multi-tissue RE2 data (Supplementary Fig. S12b) and for GCC2 in a single tissue (Supplementary Fig. S12d), although the HEIDI tests did not support pleiotropy with GCC2. Moderate support for colocalization/pleiotropy was also found for GCC2-AS1 in both single-tissue and RE2 analyses (Supplementary Fig. S12c) and for LIMS1 in several tissues (Supplementary Fig. S12e; Supplementary Worksheet S11,S12); only LIMS1 eQTLs were in very high LD (r2 > 0.9) to both GWAS and eQTL top variants.
Hair density related traits associated loci
In addition, we identified two novel association signals for the excessive hairiness trait in the chr1:119.45–119.77 Mb and chr18:60.92–60.94 Mb regions.
Chr1:119.45–119.77 Mb locus (Fig. 4a) SNPs lay within the T-box 15 (TBX15) gene or between TBX15 and the Tryptophanyl-tRNA synthetase, mitochondrial (WARS2) gene (Supplementary Worksheet S7); overlap with promoter and intronic enhancer annotations within TBX15 (Fig. 4c) supported it being the functional gene. Although very high LD GWAS variants (r2 > 0.9) were strong eQTLs for WARS2 expression in multiple GTExPortal tissues (Supplementary Worksheet S7), they were actually lowly ranked within WARS2 multi-tissue eQTL statistics. In contrast, colocalization/pleiotropy analysis identified strong support for TBX15 eQTLs across several tissues as well as multi-tissue RE2 data (Fig. 4b).
Chr1:119.45–119.77 Mb (TBX15/WARS2) excessive hairiness locus. Excessive hairiness associated SNPs in the Chr1:119.45–119.77 Mb locus possessed strong evidence for colocalization with single-tissue and multi-tissue TBX15 eQTL signals. (a) Shows regional association plots of −log10(P-values). (b) Shows −log10(P-values) for TBX15 eQTL data for single-tissue and multi-tissue Metasoft RE2 analyses. GWAS and eQTL panels are configured as described in the Figs 2 and 3 legends. (c) Presents output from the WashU EpiGenome Browser of an epilogos plot of the Roadmap Epigenomics 25-state imputed model of epigenetic states along with tracks of high LD (r2 equiv > 0.8) and very high LD (r2 equiv > 0.9) variants and GENCODE transcript models in the region.
In the final excessive hairiness locus at chr18:60.92−60.94 Mb (Fig. 5a; rs7226979; P = 7.26 × 10−11), all top linked SNPs lay within BCL2 gene introns and overlapped predicted enhancer activity (Supplementary Worksheet S7; Fig. 5c). Colocalization/pleiotropy of GWAS and BCL2 eQTL variants was strongly supported by both methods in multiple tissues (Fig. 5b; Supplementary Worksheet S7).
Chr18:60.92–60.94 Mb (BCL2) excessive hairiness locus. Excessive hairiness associated SNPs in the Chr18:60.92–60.94 Mb (BCL2) locus possess strong evidence for colocalization with single-tissue and multi-tissue BCL2 eQTL signals. (a) Regional association plots of −log10(P-values) around the Chr18:60.92–60.94 (BCL2) excessive hairiness locus. (b) Shows −log10(P-values) for BCL2 eQTL data for single-tissue and multi-tissue Metasoft RE2 P-values. GWAS and eQTL panels are configured as described in the Figs 2 and 3 legends. (c) Presents output from the WashU EpiGenome Browser of an epilogos plot of the Roadmap Epigenomics 25-state imputed model of epigenetic states along with a track of variants with mod./high LD in both GWAS and eQTL data (r2 > 0.7 in GWAS and eQTL analyses for subcutaneous adipose, tibial artery, and tibial nerve tissues) and GENCODE transcript models in the region.
Excessive sweating association signals
For the excessive sweating phenotype, we identified two novel loci, with one locus at chr2:28.82−29.05 Mb containing two independent association signals and another locus with a single signal at chr16:48.26−48.45 Mb.
The chr2 region’s primary association signal (Signal #1: chr2:28.83−29.05 Mb; top SNP rs56089836: P = 1.70 × 10−11) lay upstream of the protein phosphatase 1 catalytic subunit beta (PPP1CB) gene and downstream of the Phospholipase B1 (PLB1) gene, while the secondary association signal (signal #2: chr2:28.82–28.85 Mb; top SNP rs1534480: P unconditioned = 3.28 × 10−7; P Cond. SNP 1 = 7.74 × 10−6) resided within the PLB1 gene (Fig. 6a,b; Supplementary Worksheet S8). Both signals had variants that overlapped predicted enhancer and/or promoter elements (Fig. 6e). One missense SNP was found in the second signal (rs7601771; H/D; Cac/Gac) but was predicted by all dbNSFP41 consequence predictions in the UCSC Genome Browser42 to be tolerated/benign. Colocalization/pleiotropy analysis identified strong support for overlap of signal #2 variants with PLB1 eQTLs from multiple tissues (Fig. 6c and Supplementary Fig. 13e), with AC097724.3 multi-tissue eQTL data (Supplementary Fig. 13d), and with AC074011.2 eQTLs from a single-tissue sample (Supplementary Fig. S13c). A secondary PPP1CB eQTL signal in tibial nerve tissue had strong support for colocalization with Signal #1 variants (Supplementary Fig. S13f) in ABF and basic SMR tests but had no support of pleiotropy from the HEIDI test.
Chr2:28.82–29.05 Mb (PLB1/PPP1CB) excessive sweating locus. Of two independent GWAS signals in the chr2:28.82–29.05 Mb (PLB1/PPP1CB) excessive sweating locus, Signal #1 colocalizes with PPP1CB eQTL from a single tissue, while Signal #2 colocalizes with a PLB1 eQTL signal identified in multiple tissues. (a,b) Show regional association plots of the Chr2:28.82–29.05 Mb (PLB1/PPP1CB) locus for the two independent signals. (c) Colocalization analyses of Signal #2 SNPs with PLB1 subcutaneous adipose tissue and multi-tissue Metasoft RE2 eQTLs. (d) Colocalization analysis of Signal #1 SNPs with PPP1CB tibial nerve eQTLs. GWAS and eQTL panels are configured as described in the Figs 2 and 3 legends. (e) Presents output from the WashU EpiGenome Browser of an epilogos plot of the Roadmap Epigenomics 25-state imputed model of epigenetic states along with tracks of high LD signal 1 and signal 2 variants and GENCODE transcript models in the region.
Chr16:48.26−48.45 Mb locus variants (Supplementary Fig. S14a; top SNP rs6500380; P = 6.84 × 10−10; Supplementary Worksheet S8) lay within or near the Lon peptidase 2, peroxisomal (LONP2), ATP binding cassette subfamily C member 11 (ABCC11), and siah E3 ubiquitin protein ligase 1 (SIAH1) genes. One high LD SNP was a known ABCC11 missense variant (rs17822931; P = 2.02 × 10−9; ABCC11 c.538 G > A (p.Gly180Arg)). All linked variants were also LONP2 eQTLs in multiple tissues (Supplementary Worksheet S8), but no significant colocalization with eQTLs was identified. Our results suggest the likely causal variant in this locus is the known coding SNP, rs17822931.
Discussion
In this study, we performed GWAS analyses of seven skin-related phenotypes (age spots, freckles, single vs. double eyelids, hairiness, straight vs. curly hair, eyebrow thickness, and excessive sweating) and identified twelve loci containing sixteen association signals, of which fifteen were novel discoveries. The single previously reported association signal involved the association of hair morphology phenotypes (straight vs. curly hair, eyebrow thickness) with SNPs in EDAR (top SNPs rs260643 and rs3827760, respectively). The following will briefly discuss our findings for each of these phenotypes and place the putative causal genes in the context of current biological and medical knowledge.
Skin-spot phenotypes
Japanese women suffer in early adulthood from pigmented spots, followed by facial wrinkles in their thirties, forties, and fifties2. Freckles, which are pigmented macules generally between 1–3 mm in diameter, characteristically occur on the face as well as the back of the hands, shoulders, and neck43. Age spots (lentigines), which are a notable outward manifestation of skin aging, have been reported to occur earlier and be more conspicuous in skin of Asians compared to that of Caucasians44. It is well known that pigmented spots are associated with chronic ultraviolet exposure45, but the genetics related to this phenotype has been less well understood, especially for individuals of Asian ancestry. Motokawa et al. first reported that MC1R (melanocortin-1-receptor) gene variants were associated in Asians with melanogenic phenotypes and thus identified both environmental factors and genetic factors as involved in the production of age spots and freckles46. Two previous reports’ top SNPs in the IRF4 (rs12203592), MC1R (rs12931267, rs35063026), and RALY/ASIP (rs619865, rs6059655) regions were monomorphic in 1000G ASN and had MAF <= 0.01 in 1000G AFR samples, and therefore likely represent population specific causal variants11,12. Our study identified a novel association signal in the known BNC2 gene locus and found novel loci in and around the genes PPARGC1B, RAB11FIP2, HSPA12A, and AKAP1/MSI2.
BNC2 is a zinc-finger transcription factor that appears to be necessary for development of craniofacial and long bones as well as ectodermal appendages (hair follicles, salivary glands, palatal rugae)47,48, and it has previously been associated with skin pigmentation and pigmented spots11,12,49. Epigenetic and eQTL data indicate the most likely function for the signal’s variants are as an enhancer for an alternative promoter near BNC2 exon 310. PPARGC1B codes the Peroxisome proliferator-activated receptor gamma coactivator 1-beta, the target of which (PPARγ) has previously been associated with melanogenesis. Various earlier reports support this gene’s involvement in pigmentation related phenotypes, with 2,4,6-Octatrienoic acid reported to act via PPARγ activation to promote melanogenesis and antioxidant defense in melanocytes50, and ciglitazone, a PPARγ agonist, observed to cause an increase of melanin production in cells and cultured skin51. Our analysis suggests that variants at this locus may regulate PPARGC1B expression through modification of enhancer elements. RAB11FIP2 codes for the Rab11 family-interacting protein 2, which acts in regulating vesicular transport from the endosomal recycling compartment to the plasma membrane (http://www.uniprot.org/uniprot/Q7L804). Depletion of Rab11 causes accumulation of pigment in melanocytes52, the Rab11b isoform mediates melanin exocytosis from melanocytes53, and interaction between RAB11FIP2 and myosin 5b (MYO5B) was reported to regulate movement of vesicles containing RAB11a54. In the AKAP1/MSI2 locus, both genes possess support for putative function, but support for MSI2 appears strongest. Response elements for another novel skin spot associated gene, PPARGC1B, have been reported to help regulate AKAP1 expression55, but we found MSI2 to be differentially expressed in light-skin melanocytes versus dark-skin melanocytes and keratinocytes from FANTOM5. MSI2 was also previously found necessary to maintain the resting state of hair follicle stem cells56, and the hair follicle bulge is also a main location of melanocytic stem cells57.
In contrast to those four loci, for which SNPs lacked eQTL activity for protein-coding genes, skin-spots variants around HSPA12A were associated with gene expression. HSPA12A is a member of the 70-kDa heat shock protein (HSP70) family, which is a highly-conserved family of chaperone genes that help facilitate many aspects of proteostasis such as protein folding, multi-protein complex assembly, and transport58. HSPA12A’s sub-cellular compartment has been localized to exosomes in urine samples59,60 and specifically localized to exosomes that express markers of podocytes, which are similar to the filopodia that form between melanocytes and keratinocytes61. That suggests that HSPA12A may play a role in the transfer of melanin from/along melanocytic filipodia.
To understand how our top skin-spots associations related to skin-pigmentation phenotypes, we examined overlap with GWAS summary statistics for two UK Biobank (UKBB)62 variables (Ease of skin tanning, Skin colour) for which GWAS analyses were performed by the Gene ATLAS (http://geneatlas.roslin.ed.ac.uk; see Methods)63,64,65. Across the seven primary and secondary skin-spot signals’ top SNPs, we observed co-localization with top skin-pigmentation SNPs for PPARGC1B chr5:149.19–149.23 Mb (rs251468: P freckles = 1.27 × 10−21, P tanning = 3.32 × 10−50, P skin colour = 2.52 × 10−41) and RAB11FIP2 chr10:119.56–119.58 Mb (rs10444039: P freckles = 5.62 × 10−21, P tanning = 3.78 × 10−71, P skin colour = 4.93 × 10−53). That suggests that overall skin-pigmentation and acquisition of pigmented spots at least partially share mechanisms for their development.
In addition, the overlap between age-spots and freckles cases/controls in our sample suggests that the definition/translation of age-spots and freckles in Japan (shimi and sobakasu) may differ from the United States and European sense, with freckles (sobakasu) in Japan perhaps representing more of an extreme end of a continuum of skin spot accumulation due to sun exposure. Based on a genetic risk score analysis (Supplementary Fig. S10), we found that the presence of multiple effect-alleles imparts a greater pre-disposition for acquisition of freckles. Further GRS analysis in independent cohorts would be needed to confirm that, and in the future, as direct to consumer genetic testing becomes more prevalent, it may be warranted to recommend to individuals with multiple effect alleles for these association signals that they reduce sun exposure to avoid the appearance of pigmented skin spots.
Double versus single eyelid phenotype
The human eyelid is a complex craniofacial structure consisting of interconnecting and layered substructures made up mostly of skin, adipose, muscular, and nerve tissues that help protect the eye from injury and insult due to external environmental threats such as temperature and humidity variation, wind, and dust and airborne objects66. The structure and appearance of the human upper eyelid shows variation in the amount of upper eyelid crease, with individuals having some degree of upper eyelid crease or those who lack such a crease termed double and single eyelids (or mono-lids), respectively67. In Asians, individuals with either single eyelids or eyelids with intermediate features make up approximately 44% of the population68, but they are very uncommon in most other ethnic populations67. Such eyelid variation has been ascribed to anatomical differences for the location at which the orbital septum fuses with the levator aponeurosis69, as well as to underlying differences in the shape and structure of the orbital socket70. Because individuals who possess double eyelids are sometimes considered more attractive compared to those having single eyelids, double eyelid surgery (blepharoplasty) has become a popular medical procedure67,71,72, with more than 40,000 Japanese women receiving eyelid modification surgery every year.
In this study, we identified double eyelid associated SNPs that were also eQTLs for EMX2 and the antisense RNA EMX2OS, the latter of which has been reported to regulate EMX2 expression73,74. EMX2 is a homeobox-containing transcription factor gene that appears important in development of skeletal and neural structures75,76,77. In light of recent reports of EMX2 in the context of craniofacial development35, our results suggest that these SNPs may regulate EMX2 expression at embryonic stages important for human facial structure development that lead to concomitant upper eyelid differences.
Hair morphology related phenotypes
Some women are affected by the presence of dense eyebrows, which may necessitate additional time spent on processing their eyebrows. In addition, scalp and body hair can assume different morphological shapes, both in terms of shaft thickness and shaft straightness/curl. In our study, the only genome-wide significant signals associated with dense eyebrows or straight/curly hair phenotypes replicated a known association of the EDAR missense variant rs38277603,4,36,37,78. EDAR codes for the ectodysplasin A receptor, which is a membrane-bound receptor of the TNF-R family, mutations of which have also been linked with Mendelian diseases associated with skin such as Ectodermal dysplasia79,80,81 as well as with facial and ear characteristics38,39, and dental traits40.
Excessive hairiness (hirsutism) phenotype
Excessive hairiness, which is medically termed hirsutism, represents excess hair growth in females, and can generally be divided into sub-types corresponding to: (1) individuals with high circulating androgen levels or high sensitivity to androgen, or (2) idiopathic hirsutism, for which an individual possesses normal androgen levels and no other discernible cause for the excess hair growth82. From examination of previous hair-density related trait analyses, we identified overlap with both eyebrow thickness or hirsutism signals (Table 2), which suggests some shared mechanisms between these phenotypes. In addition to validation of SNPs associated with these previous traits, our study identified three novel hirsutism loci that were also eQTLs for BCL2, GCC2 and LIMS1, and TBX15.
BCL2 acts as an anti-apoptotic regulatory protein that blocks cell death and has a known role in the cycle of life-and-death related to hair follicle growth83. Since regulation of catagen (apoptosis-driven involution of hair follicles) stage timing is an extremely important part of the process84, our analysis suggests that differences in BCL2 expression due to variants at this locus may shift the timing or length of catagen and thereby lead to variation in hair-density between individuals.
In the GCC2/LIMS1 locus, our analysis suggests that SNPs may impact expression of both genes. GCC2 localizes to the trans-Golgi network and functions in the tethering and capture of inbound vesicles in endosomal transport85,86,87. LIMS1 localizes to focal adhesion plaques to regulate cell adhesion and spreading through interaction with integrin-linked kinase (ILK)88,89,90 and Parvins as part of the ILK, PINCH, and Parvin complex (IPP)91. ILK was reported to be necessary for hair morphogenesis92, and deletion of LIMS1 from mouse keratinocytes was shown to impair hair follicle growth93. In addition, LIMS1 contributes to BCL2-dependent survival signaling and acts to inhibit JNK-mediated apoptosis94.
TBX15 is known to be involved in development, particularly in chondrocyte hypertrophy and skeletal limb development95, and Tbx15 was previously shown to be involved in hair pigmentation and hair length in mice96. It has also been related to the determination of muscle fiber-type97 and metabolic subtypes of adipocytes98,99. One of our signal’s top variants (rs984222) was previously associated with BMI and Waist-hip ratio (WHR) in individuals of European ancestry100, which was later confirmed in a trans-ethnic meta-analysis of European and African-American ancestry population samples101. Using the Gene ATLAS Region PheWAS tool, we also identified rs984222 as positively associated with WHR (β = 0.00147, P = 6.95 × 10−26), but while it was also associated with whole-body/leg/arm bioimpedance measurements (β whole body = 1.2651, P whole body = 3.29 × 10−21; β leg(left) = 0.56755, P leg(left) = 3.79 × 10−18; β arm(left) = 0.6744, P arm(left) = 1.57 × 10−17; β arm(right) = 0.63848, P arm(right) = 1.64 × 10−16; β leg(right) = 0.52175, P leg(right) = 1.58 × 10−15), it was not significantly associated with Body fat percentage (β = −0.0030593 P = 0.81055). In addition, it was positively associated with Standing height (β = 0.070321, P = 1.05 × 10−9) but negatively associated with BMI (β = −0.04843, P = 5.00 × 10−7) and Hip circumference (β = −0.081466, P = 1.52 × 10−5). For our analysis, we had included BMI as a covariate (Supplementary Table S1), suggesting that the association with excessive hairiness was also not related to adiposity. Taken together, these findings suggest that these TBX15 regulating variants act in a pleiotropic manner that impacts skeletal development, fat store distribution, and hair follicle density and/or activity.
Excessive sweating phenotype
Sweating ability plays an important function in regulating body temperature and is performed by eccrine and apocrine sweat glands present on the skin. Eccrine glands secrete a clear fluid made up mostly of water, NaCl, and other salts, and compared to apocrine glands, they are present in the greatest number across the human body (~2–4 million) and provide for most of sweat’s cooling properties102. Eccrine gland secretion is mainly controlled through cholinergic and to a lesser extent adrenergic signaling originating from the pre-optic region of the hypothalamus, but other signaling molecules appear to act at a local level103.
Hyperhidrosis, the clinical term for excessive sweating, can be caused by various pathologies104,105, and the presence of increased sweating can lead to embarrassment in social situations and sometimes impairs quality of life (QOL)106. In the absence of known causes, idiopathic hyperhidrosis does not appear to be a result of morphological changes in eccrine gland numbers or size but rather a complex dysfunction of autonomic nervous system central control107. In studies of hyperhidrosis patients, reports have identified other autonomic nervous system abnormalities, such as differences for cardiovascular stress responses, that point to over-functioning of sympathetic108 and possibly parasympathetic nervous system fibers109. Previously, a genome-wide linkage analysis of primary palmar hyperhidrosis in Japanese identified a significant region at 14q11.2-q13, but that region did not show significant associations in our analysis110. In the current study of excessive sweating, we found associations with SNPs in two gene regions on chromosome 2 and chromosome 16.
In the chromosome 2 PLB1/PPP1CB region, our analyses identified associated variants in two independent signals as either PLB1 or PPP1CB eQTLs. Previous reports lend support for both PLB1 and PPP1CB as plausible genes for the association with sweating. PLB1 codes for phospholipase B1 protein (PLB), which is an enzyme that has both phospholipase A1 and A2 enzymatic activities. PLB1 was originally identified in humans by its expression in the epidermis and suggested to function by promoting the skin barrier function by breakdown of lipids into free fatty acids111. In addition, based on its role in promoting acrosome exocytosis in sperm112,113,114, there is a possibility that it could function to modulate secretory processes in other situations such as sweating. On the other hand, PPP1CB encodes the beta-subunit of the Serine/threonine-protein phosphatase PP1, which is a key enzyme involved in the regulation of a large number of cellular processes115. Based on previous reports that phosphorylation of the water-specific channel aquaporin-5 (AQP5) regulates its ability to mediate water flow116,117 we hypothesize that PPP1CB eQTL SNPs may effect sweat production by modulating the amount of PPP1CB that is present and thereby influencing phosphorylation levels of AQP5 or other proteins necessary for sweat gland function.
The single signal on chromosome 16 encompassed a region including ABCC11 and LONP2 genes. After eQTL and LD structure analysis, we concluded that a known missense SNP in ABCC11 (rs17822931) was the likely causal variant among the five tightly linked SNPs in this locus. ABCC11 encodes ATP binding cassette subfamily C member 11, which is a member of the multi-drug resistance protein (MRP) sub-family of the ATP-binding cassette gene family. The missense SNP has previously been associated with dry versus wet earwax types118 and axillary ozmidrosis (body odor)119,120,121. The current study is the first report that shows that this SNP is also associated with hyperhidrosis.
Conclusions
In this report, we identified a dozen loci encompassing 16 association signals for the dermatological phenotypes of skin-spots, double vs. single eyelids, eyebrows (thick/thin), hair (straight/curly), excessive hairiness, and excessive sweating. For skin-spot phenotypes (freckles and age-spots), we found four novel loci encompassing the PPARGC1B, RAB11FIP2, HSPA12A, and AKAP1/MSI2 genes, along with a novel East Asian signal in the known BNC2 gene locus. Future research should combine these signals with association signals identified in other ethnic groups and confirm the preliminary genetic risk score that we calculated using the current dataset. Such a score may help women identify their risk for skin-spot acquisition and take appropriate actions to mediate their impact. For the double eyelid phenotype, EMX2 represents the likely regulatory target of the two independent signals, but understanding their actual functional impact will require analyzing the variants’ effect on EMX2OS and EMX2 expression in the context of particular developmental stages. For hair-related phenotypes, we replicated previous reports that implicated the rs3827760 EDAR missense variant with hair morphology and hair straightness/curliness, but additionally, we identified that SNPs near EDAR are associated with excessive hairiness (hair density) and are eQTLs for two neighboring protein-coding genes, namely GCC2 and LIMS1, as well as for neighboring lncRNAs. The LD structure in the EDAR region in Japanese makes it difficult to confirm the findings in further Japanese population samples, so future research should examine the relationship of these eQTL variants with the trait in other population samples. In contrast to the EDAR missense variant rs3827760, these high LD SNPs are common alleles in other world-wide population samples, and therefore, they should be amenable to validation. For the excessive sweating phenotype (hyperhidrosis), previous GWAS analyses have not been performed, and our current analysis implicated variants in one region that may regulate PLB1 and/or PPP1CB expression, and in another region, identified a known missense variant in ABCC11. An excessive rate of sweating may affect QOL in an individual’s social life, and therefore, cosmetic procedures to reduce hyperhidrosis has demand in the medical beauty-care industry. Further experimental research about these genes in the context of excessive sweating will be necessary and will hopefully lay a foundation for identifying systemic or topical agents to ameliorate their effects.
Methods
Details about the subject, sample, and phenotype data collection, sample processing and genotype Quality Control (QC) procedures, statistical analysis, linkage disequilibrium (LD) statistics calculation, in silico functional annotation of variants, and eQTL analyses can be found in a recent report that used the same underlying set of samples17. The following sections will include brief versions of the fully described methods, methods that were not included in that report, or for which previous methods differ (different procedures or upgraded datasets) or require further explanation.
Subject, sample, and phenotype data collection
Subjects were voluntarily enrolled in a study investigating the genetics of various human traits run by the MTI (http://www.mti.co.jp/eng/) subsidiary EverGene. Questionnaires soliciting trait information were filled-out by subjects online. Subjects were collected in two stages, denoted as LL01 and LL02, with 11378 individuals completing the questionnaires and providing DNA samples (LL01 = 5750, LL02 = 5628). The Institutional Review Board at the Tsukuba International Clinical Pharmacology Clinic approved the study design, such as the consent form, general questionnaire topics, and genotyping, and the study was performed in accordance with applicable regulations and guidelines. Informed consent was obtained from each patient for sample collection, genotyping, trait questionnaire, and trait analysis using genome-wide association study analysis.
Sample processing, genotyping and quality control
There were 607857 total variants assayed using the custom Axiom array EverGene1 chip. For downstream analyses, the following QC criteria had to be fulfilled in both stages: 1) ≥99% call-rate, 2) MAF ≥ 0.01, 3) HWE P-value ≥ 1 × 10−6, and 4) concordance-rate >90%. After QC, there were 536506 variants.
Principal component analysis (PCA)
We performed principal component analysis (PCA)122 with PLINK2 v1.90pVer.b3.42 (release date 16 Aug 2016)123,124 to identify population structure. A PCA plot of samples used in the current analysis is shown in Supplementary Fig. S1, while a complete description of the PCA based process for sample filtering can be seen in Supplementary Fig. S1 of the earlier Khor SS, et al. report17.
Identification of duplicated samples
Using LD-pruned data, we ran identify-by-descent (IBD) analysis with PLINK2 ver. 1.90p’s and filtered data as previously described. For the current study, there were 11311 subjects who passed QC procedures and who also answered at least some of the skin phenotype questions on the questionnaire. After running the current GWAS analysis, we checked and found that a very small number of sample pairs (n = 14) were identified with PI_HAT > 0.1875 and PI_HAT < 0.8, generally suggestive of second to first-degree relatives within the sample-set. Since the percent of total samples in that PI_HAT range was very low (~0.12%), their presence should have little impact on the overall GWAS statistics.
Definition of skin phenotype cases and controls
Certain constitutional phenotypes were asked as the question “Please describe your constitution, ease, strength, etc. [with respect to certain possible traits]”). The possible answers corresponded to the English phrases “Very applicable”, “Slightly true”, and “Not applicable”. For these traits, cases were considered as those answering “Very applicable” or “Slightly true” and controls as those answering “Not applicable”. For this paper, the traits included age spots (Japanese = Shimi), freckles (Japanese = Sobakasu), and hairiness (Japanese = Kebukasa).
Another set of constitutional questions were queried in the form “For each of the following, are you closer to A or B? Please answer subjectively.” Phenotypes queried in such a manner included thick vs. thin eyebrows, straight vs. curly hair, and excessive sweating. For thick/thin eyebrows, responses were A: Thick eyebrows (Japanese: Mayuge ga koi) and B: Thin eyebrows (Japanese: Mayuge ga usui). For straight/curly hair, responses were A: Straight hair (Japanese: Kami ga sutorēto) and B: Curly hair (Japanese: Kami ga kusekke). For excessive sweating, responses were A: It is easy to sweat (Japanese: Ase o kaki yasui) and B: It is hard to sweat (Japanese: Ase o kaki nikui). For each of those phenotypes, A responders were set as cases and B responders set as controls.
The final constitutional question was a multiple-choice question that asked “Please choose all that apply to yourself from the following (Multiple selections possible)”. One possible choice was for the double-edged eyelid phenotype (Japanese: Futaemabuta). Subjects checking that box were considered as responding affirmatively to the presence of the phenotype and thus used as cases, while those that did not mark the selection were considered as responding negatively to the presence of the phenotype and thus were used as controls.
Statistical analysis and genotype imputation
The R 3.4.1 statistical environment was used for management of data, statistical analyses, and figure plotting125. The primary association analyses for LL01 or LL02 datasets was run using PLINK2’s logistic regression analysis method (-logistic flag). For each phenotype analyzed, we included PC1 and PC2 (from the stage 3 PCA described above) as covariates, and then Age or BMI as covariates if a test of them against the phenotype in a regression model showed that they were significant (P < 0.05). The specific covariates that were included in each regression analysis are shown in Supplementary Table S1. Meta-analysis statistics combined LL01 and LL02 data using the inverse-variance weighting method with beta-coefficients and standard errors from the regression analyses126. Our assumption for the effective number of SNPs (ME) for the current genotyping platform was previously described17, resulting in a single GWAS P-value cut-off of P meta < 1.21 × 10−7 (0.05/411,521). Based on the number of phenotypes analysed in the current analysis, we defined primary association signals as those with genotyped variants that achieved a multiple-testing adjusted P-value cut-off of P meta < 1.73 × 10−8 (P meta < 1.21 × 10−7/7 skin phenotypes).
For plotting the Manhattan plot and to allow comparison/validation with data from previous reports, we performed a genome-wide summary statistics based imputation of the meta-analysis data using the program DISTMIX18 and the 1000 Genomes Phase 1 Release 3 reference data127. For primary association signals, we analyzed imputed genotype data that we produced using a more accurate genotype-based imputation method that was described in the Khor et al. report. Variants were filtered for allelic R2 > 0.7. In each region of imputed data, we performed step-wise logistic regression analysis conditioning on the top imputed variant in each signal until no further variants with P < 1 × 10−5 were identified; top variants achieving that significance level after a conditioning step were considered as secondary association signals.
Allele frequencies and LD statistics were calculated using PLINK 1.9 and then imported into R for analysis. For variance explained calculations, we imported the genotype data into R and used generalized linear model (glm) for logistic (family = ‘binomial’) or linear (family = ‘gaussian’) regression analysis using all of the top SNPs with PC1, PC2, and Age or BMI as covariates in a full regression model of the phenotype of interest. Using the R rms package128, we performed a likelihood ratio test of the full model (glm1) against the model without the associated variants (glm0) and then calculated a pseudo-R2129 as a surrogate for the proportion of variance explained. With n being the number of samples and LR the χ2 statistic from the likelihood ratio test comparison of glm1 and glm0 (lrtest(glm0, glm1)), we calculated pseudo-R2 as (1 − exp(−LR/n))/(1 − exp(2logLik(glm0)/n)).
For the freckles case-control phenotype, we calculated a genetic risk score (GRS) using the riskScore function from the R package PredictABEL130 with extracted components from the glm models. Within each GRS value, we then summarized the proportion of individuals who were positive for either any freckles (Freckles = Very applicable or Slightly applicable) or strong freckles (Freckles = Very applicable).
LD measures
We used PLINK 1.9 to calculate traditional LD r2 and D’ measures from the imputed genotype data. Additionally, we calculated an alternative measure that we term r2 equiv , which is based on conditional regression analysis to measure the signal decrease at a SNP B relative to a top SNP A and was described in detail in the Khor et al. report. Moderate LD SNPs were generally considered as those with r2 equiv > 0.5 and high LD SNPs as those with r2 equiv > 0.8.
In silico functional analysis of associated variants
We attempted to use the current dbSNP147 rsID as much as possible. The RsMergeArch.bcp.gz table was imported into R from NCBI’s ftp site and we identified the current rsID for SNPs present in the various annotation sources used below.
We annotated genotyped and 1000G variants using HaploReg 4.1131, including dbSNP gene function annotation and evolutionary conservation scores (GERP), and GWAS and eQTL results from the Genome-Wide Repository of Associations Between SNPs and Phenotypes (GRASP)132. Since HaploReg and 1000G used different dbSNP versions, we identified both the current rsID and all previously used rsIDs for each SNP, passed all rsIDs to HaploReg for annotation, and then processed the output to resolve the current rsID with the one actually used by HaploReg.
Since HaploReg is no longer maintained or updated, we added/updated data from several sources. A current version of the NHGRI/EBI GWAS Catalog was downloaded on February 6, 2018 from the UCSC Genome Browser and used for annotation of SNPs for previous GWAS results133,134. For more comprehensive eQTL associations, we downloaded multi-tissue eQTL Metasoft results (file: GTEx_Analysis_v7.metasoft.txt.gz) from the GTExPortal dataset page (https://www.gtexportal.org/home/datasets) along with files for variant and gene meta-information (variants: GTEx_Analysis_2016-01-15_v7_WholeGenomeSeq_635Ind_PASS_AB02_GQ20_HETX_MISS15_PLINKQC.lookup_table.txt.gz, genes: gencode.v19.genes.v7.patched_contigs.gtf)135. That data was imported into R and used to annotate SNPs with P-values from multi-tissue analysis using fixed-effects (FE), random effects (RE), and Metasoft modified random effects (RE2) models. That data was also used in the colocalization analysis that is described in the next section. We also labelled each SNP in our output with the minimum single-tissue P-value along with a text string of any gene/tissue pairs and their P-values that surpassed a multiple testing corrected 0.05/#tissues. Overlap of variants in each associated region with experimentally defined transcription factor binding sites (TFBS) was done using the ReMap 2018 annotation tool30,31.
In addition, we summarized the RoadMap Epigenomics data using our own scripts. We downloaded the 25-state/12-mark imputed chromosome segment model of epigenetic states from http://egg2.wustl.edu/roadmap/data/byFileType/chromhmmSegmentations/ChmmModels/compressedStateTracks/hg19_chromHMM_imputed25.gz, and we used bedtools (command flags “intersect -wb -a”) to extract segment state data for genomic regions associated in the GWAS. We then added meta-information for samples (https://docs.google.com/spreadsheet/ccc?key=0Am6FxqAtrFDwdHU1UC13ZUxKYy1XVEJPUzV6MEtQOXc&usp=sharing#gid=15) and information about imputed marks (http://egg2.wustl.edu/roadmap/data/byFileType/chromhmmSegmentations/ChmmModels/imputed12marks/jointModel/final/annotation_25_imputed12marks.txt) to be able to aggregate across tissue samples from the same anatomical class and collapse information from similar epigenetic states. Epigenetic state counts shown in Supplementary Worksheets S1–S8 summarize Promoter overlap counts from four Promoter states (TssA, PromU, PromD1, Prom D2), Enhancer overlap from ten states (TxReg, TxEnh5′, TxEnh3′, TxEnhW, EnhA1, EnhA2, EnhAF, EnhW1, EnhW2, EnhAc), Transcription overlap from four states (Tx5′, Tx, Tx3′, TxWk, TxReg), and bivalent/poised from PromP and PromBiv states. Other reported states only had a single state in the dataset.
Variant overlap with coding gene models was done using the UCSC Genome Browser GENCODE Ver. 24 tracks (wgEncodeGencodeBasicV24lift37.txt.gz and wgEncodeGencodeAttrsV24lift37.txt.gz)42,136, with gene transcripts merged to identify gene coding start and stop coordinates, and then overlaps with SNPs identified using the R Bioconductor GenomicRanges packages. We labeled SNPs with four categories of genic overlap/nearness: 1) “within” = SNP between start and stop coordinates of a gene’s coding region, 2) “upstream” = SNP < 100 kb upstream of the gene start position, 3) “downstream” = SNP < 40 kb downstream of the gene stop position, 4) “closest” = for SNPs with no genes fulfilling the first three rules, we picked the closest gene to the SNP. “Closest” gene is not provided as a separate column but listed in a column “Genes (all)” that either contains the union of within/upstream/downstream genes for a SNP, or if those are missing, contains the closest gene. The 100 kb upstream and 40 kb downstream cutoffs were chosen based on previous reports that analyzed the general distance from Transcription Start Site (TSS) and Transcription End Site (TES) within which most eQTL SNPs were identified137,138.
Analysis of colocalization of GWAS and eQTL signals
For colocalization testing of GWAS and GTEx eQTL association signals, we used the Approximate Bayes Factor (ABF) method in the R coloc (ver. 2.3–7) package’s coloc.abf function26 and the Summary data-based Mendelian Randomization method from the SMR program (ver 0.702); SMR is available on the CNS Genomics web-site (http://cnsgenomics.com/software/smr/#Overview)27. For each GWAS signal, we determined a list of pertinent genes from the gene and eQTL annotations in Supplementary Worksheets S1–S8 and then checked the GTEx Portal’s Gene eQTL Visualizer to identify relevant tissues in which association with those genes was observed. For a subset of the relevant tissues, we downloaded the complete GTEx Version 7 single-tissue eQTL files found on the datasets page under “Tissue Specific All SNP Gene Associations” for those tissues and processed the data in R.
Since we observed multiple independent eQTL signals for certain genes, we first processed a locus’ single-tissue or multi-tissue data for a particular gene into groups of SNPs that were in LD to a particular unassigned top eQTL variant. Briefly, a gene’s eQTL data was sorted by association statistics, and SNPs that had LD r2 > 0.01 to the top unlinked SNP were assigned to that SNP. If a SNP had LD r2 > 0.01 but the signal was very strong compared to what one would expect based on the LD, then it was left unassigned. That was determined by calculating the proportion of signal strength at a SNP B relative to the top SNP A (−log10(B)/−log10(A)) and then dividing by the r2. If that value was large (>10), then it was considered that the signal at SNP B was not due to just LD and SNP B was not assigned to SNP A’s signal. LD was calculated using PLINK2 across either EUR or AFR samples’ data from 1000 Genomes Project Phase 3. Although the GTEx samples were mostly of European ancestry, it was also apparent from top SNPs’ MAFs that some signals were coming from the African ancestry data, so heuristic MAF cutoffs were used to determine which 1000G population sample should be used for the LD calculation: 1) If a top SNP had MAFEUR < 0.01 and MAFAFR < 0.01, then we could not calculate LD statistics and a SNP was not assigned to any signal, 2) if MAFEUR < 0.01 and MAFAFR ≥ 0.01, then the AFR data was used for LD calculation, 3) if MAFEUR ≥ 0.01 then EUR data was used for LD. The MAFEUR < 0.01 and MAFAFR ≥ 0.01 SNPs were often monomorphic in EAS samples, so including putative AFR ancestry specific signals in our data was incorporated mostly to produce figures that included all eQTL variants that were visible in the GTEx Portal browser.
One important difference existed between the ABF and SMR tests in terms of data input: the coloc.abf test could use either beta-coefficients and standard errors or P-values as input, but the SMR test could use only the former. In both cases, we considered the use of beta and SE as preferable. However, while beta-coefficients and standard errors were available for single-tissue data, they were not available for the multi-tissue Metasoft RE2 analysis, which was preferable since the RE2 analysis was reported to be more powerful for identifying eQTLs that act across multiple tissues in certain instances. Therefore, to pick the most powerful choice at a particular signal, we used the FE beta and SE for multi-tissue data if the FE statistics were strongly correlated with those from the RE2 analysis, but otherwise, we used the RE2 P-values as input to the ABF test along with required MAF and sample-size values. If that was done, then we would expect that the SMR results, which had to use the FE statistics as input, would differ from the ABF results. The status for whether FE beta was used for the ABF analysis is reported in Supplementary Worksheets S9 and S10. Since GTEx eQTL analyses were performed using standardized expression values, we assumed a value of 1.0 for the standard deviation of expression trait values in the analysis using coloc.abf.
The ABF method outputs posterior probabilities for five states H0–H4: PP H0.ABF = no causal variant, PP H1.ABF = only trait 1 has causal variant, PP H2.ABF = only trait 2 has causal variant, PP H3.ABF = two different causal variants, PP H4.ABF = a single common causal variant (i.e., colocalized). The SMR test outputs two P-values: one for the SMR test (P SMR ) and the other for their HEIDI test (heterogeneity in dependent instruments; P HEIDI ). The SMR test by itself tests for linkage between a causal GWAS variant and a causal eQTL variant but does not provide evidence that the same causal variant impacts both traits. On the other-hand, HEIDI tests for heterogeneity across linked SNPs in a region and tries to determine whether pleiotropy exists in addition to simply linkage. For the purposes of this report, we considered a GWAS/eQTL signal pair as having at least nominal support for colocalization/pleiotropy if it was nominally significant for the SMR test (P SMR < 0.05), non-significant for the HEIDI test (P HEIDI ≥ 0.05), and had PP H4.ABF > 0.3. If a signal had PP H4.ABF > 0.9 or PP H4.ABF > 0.5, then a signal was described as having strong or moderate support, respectively. Additionally, lower values of P SMR combined with higher P HEIDI values provided more evidence for colocalization.
Haplotype cluster analysis
For one association signal, we performed haplotype cluster analysis to examine the genetic structure between putative causal variants. For the chr10:118.45–118.48 Mb locus that was examined, we calculated the maximum effect allele frequency (EAF max ) across linked SNPs with r2 equiv > 0.8 (EAF max = 0.35) and extracted associated variants EAF > 0.01, EAF < = EAF max , P meta-analysis < 1 × 10−5, r2 > 0.5, and D’ > 0.98. Then, we determined a set of non-redundant SNPs by calculating the Canberra distance between SNPs in the imputed phased haplotypes using the R dist function, performed complete hierarchical clustering using the hclust function, and identified clusters of SNPs with less than 1% dissimilarity from one another. From each cluster, we then extracted a single exemplar SNP. To perform haplotype clustering, we then extracted haplotype data for the non-redundant set of SNPs, calculated the Euclidean distance between each haplotype, and performed complete hierarchical clustering. Labels were assigned to haplotypes using the cutree function with a tree height cutoff determined from examining the plotted tree and picking a height that yielded a sensible number of haplotype clusters.
FANTOM5 CAGE peak analysis
Regional filtering of FANTOM5 CAGE data33,139 was performed using the ZENBU browser140, the CAGE peak data table downloaded, and summaries produced around appropriate sample types.
Browser link for: AKAP1 analysis: http://fantom.gsc.riken.jp/zenbu/gLyphs/#config=ONHzqgf2E5Xtmnpsh2gURB;loc=hg19::chr17:55162544.55162663.
Browser link for MSI2 analysis: http://fantom.gsc.riken.jp/zenbu/gLyphs/#config=ONHzqgf2E5Xtmnpsh2gURB;loc=hg19::chr17:55333785.55334472.
Analysis of Gene ATLAS/UK Biobank GWAS data
We examined SNPs in certain association signals for whether they were also associated with UK Biobank traits that had been analyzed by the Gene ATLAS (http://geneatlas.roslin.ed.ac.uk)63,64,65. For skin-spots signals, we merged imputed GWAS data for two traits (“Ease of skin tanning” and “Skin colour”) from the Gene ATLAS download page with our freckles GWAS summary statistics and examined the overlap between the top freckles or UKBB SNPs. In the case of the excessive hairiness phenotype, we used the Gene ATLAS PheGWAS function to identify associations with other traits in the TBX15 gene region. The top SNP for multiple anthropometric traits was also our top genotyped TBX15 variant (rs984222), for which we then extracted beta-coefficients and association P-values for several traits using their Phewas function.
Figure plotting
Figures that were not associated with outside software/web-services were plotted using self-written R programs. Figures of epigenetic states were produced from RoadMap Epigenomics data using the Washington University Epigenome Browser (http://epigenomegateway.wustl.edu/browser/)141 using their “Screenshot” function to produce publication quality images. Genes models shown come from GENCODE V19142. BED files of top SNPs in association signals were added as custom tracks. Custom tracks were added for imputed 25-state model from the RoadMap Epigenomics Project as compressed state tracks and as epilogos visualization. The custom track for 25-state model: http://egg2.wustl.edu/roadmap/data/byFileType/chromhmmSegmentations/ChmmModels/compressedStateTracks/hg19_chromHMM_imputed25.gz. Custom track for epilogos visualization: http://egg2.wustl.edu/roadmap/data/byFileType/chromhmmSegmentations/ChmmModels/epilogos/imputed/qcat.gz.
Data availability
Due to a concern for subject privacy and restrictions in the study consent form, the genotype data for this study is not publicly available to outside researchers. However, we do make the genome-wide summary statistics (β-coefficient and SE, P-value, effect-allele frequency) available as Supplementary Datasets S1–S7; a description and legend for each dataset is available in the Supplementary Information. Reasonable requests for other data should be addressed to Dr. Todd A. Johnson (todd.johnson@stagen.co.jp).
References
Nishiyama, S. & Takahashi, M. Revaluation of skin feature for 10s and 20s generation and development of cosmetic products suitable for them. Fragrance Journal 19, 60–66 (1991).
Naganuma, M. The Damage on the Skin Induced by UV Exposure in Sunlight. Oleoscience 7, 347–355 (2007).
Fujimoto, A. et al. A replication study confirmed the EDAR gene to be a major contributor to population differentiation regarding head hair thickness in Asia. Hum Genet 124, 179–185 (2008).
Fujimoto, A. et al. A scan for genetic determinants of human hair morphology: EDAR is associated with Asian hair thickness. Hum Mol Genet 17, 835–843 (2008).
Suzuki, T., Miyamura, Y. & Tomita, Y. High frequency of the Ala481Thr mutation of the P gene in the Japanese population. Am J Med Genet A 118A, 402–403 (2003).
Yuasa, I. et al. OCA2 481Thr, a hypofunctional allele in pigmentation, is characteristic of northeastern Asian populations. J Hum Genet 52, 690–693 (2007).
Abe, Y., Tamiya, G., Nakamura, T., Hozumi, Y. & Suzuki, T. Association of melanogenesis genes with skin color variation among Japanese females. J Dermatol Sci 69, 167–172 (2013).
Han, J. et al. A genome-wide association study identifies novel alleles associated with hair color and skin pigmentation. PLoS Genet 4, e1000074 (2008).
Liu, F. et al. Genetics of skin color variation in Europeans: genome-wide association studies with functional follow-up. Hum Genet 134, 823–835 (2015).
Visser, M., Palstra, R. J. & Kayser, M. Human skin color is influenced by an intergenic DNA polymorphism regulating transcription of the nearby BNC2 pigmentation gene. Hum Mol Genet 23, 5750–5762 (2014).
Eriksson, N. et al. Web-based, participant-driven studies yield novel genetic associations for common traits. PLoS Genet 6, e1000993 (2010).
Jacobs, L. C. et al. A Genome-Wide Association Study Identifies the Skin Color Genes IRF4, MC1R, ASIP, and BNC2 Influencing Facial Pigmented Spots. J Invest Dermatol 135, 1735–1742 (2015).
Pickrell, J. K. et al. Detection and interpretation of shared genetic influences on 42 human traits. Nat Genet 48, 709–717 (2016).
Li, Y. R. & Keating, B. J. Trans-ethnic genome-wide association studies: advantages and challenges of mapping in diverse populations. Genome Med 6, 91 (2014).
Keller, M. F. et al. Trans-ethnic meta-analysis of white blood cell phenotypes. Hum Mol Genet 23, 6944–6960 (2014).
Golder, V. et al. Frequency and predictors of the lupus low disease activity state in a multi-national and multi-ethnic cohort. Arthritis Res Ther 18, 260 (2016).
Khor, S. S. et al. Genome-wide association study of self-reported food reactions in Japanese identifies shrimp and peach specific loci in the HLA-DR/DQ gene region. Sci Rep 8, 1069 (2018).
Lee, D. et al. DISTMIX: direct imputation of summary statistics for unmeasured SNPs from mixed ethnicity cohorts. Bioinformatics 31, 3099–3104 (2015).
Loh, P. R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat Genet 48, 1443–1448 (2016).
Loh, P. R., Palamara, P. F. & Price, A. L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat Genet 48, 811–816 (2016).
Browning, B. L. & Browning, S. R. Genotype Imputation with Millions of Reference Samples. Am J Hum Genet 98, 116–126 (2016).
Liu, F. et al. The MC1R Gene and Youthful Looks. Curr Biol 26, 1213–1220 (2016).
GTEx, C. et al. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Han, B. & Eskin, E. Random-effects model aimed at discovering associations in meta-analysis of genome-wide association studies. Am J Hum Genet 88, 586–598 (2011).
Duong, D. et al. Applying meta-analysis to genotype-tissue expression data from multiple tissues to identify eQTLs and increase the number of eGenes. Bioinformatics 33, i67–i74 (2017).
Giambartolomei, C. et al. Bayesian test for colocalisation between pairs of genetic association studies using summary statistics. PLoS Genet 10, e1004383 (2014).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat Genet 48, 481–487 (2016).
Zhou, X. et al. Epigenomic annotation of genetic variants using the Roadmap Epigenome Browser. Nat Biotechnol 33, 345–346 (2015).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Chèneby, J., Gheorghe, M., Artufel, M., Mathelier, A. & Ballester, B. ReMap 2018: an updated atlas of regulatory regions from an integrative analysis of DNA-binding ChIP-seq experiments. Nucleic Acids Res 46, D267–D275 (2018).
Griffon, A. et al. Integrative analysis of public ChIP-seq experiments reveals a complex multi-cell regulatory landscape. Nucleic Acids Res 43, e27 (2015).
A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Lizio, M. et al. Gateways to the FANTOM5 promoter level mammalian expression atlas. Genome Biol 16, 22 (2015).
Square, T. et al. A gene expression map of the larval Xenopus laevis head reveals developmental changes underlying the evolution of new skeletal elements. Dev Biol 397, 293–304 (2015).
Askary, A. et al. Genome-wide analysis of facial skeletal regionalization in zebrafish. Development (2017).
Adhikari, K. et al. A genome-wide association scan in admixed Latin Americans identifies loci influencing facial and scalp hair features. Nat Commun 7, 10815 (2016).
Tan, J. et al. The adaptive variant EDARV370A is associated with straight hair in East Asians. Hum Genet 132, 1187–1191 (2013).
Adhikari, K. et al. A genome-wide association study identifies multiple loci for variation in human ear morphology. Nat Commun 6, 7500 (2015).
Adhikari, K. et al. A genome-wide association scan implicates DCHS2, RUNX2, GLI3, PAX1 and EDAR in human facial variation. Nat Commun 7, 11616 (2016).
Park, J. H. et al. Effects of an Asian-specific nonsynonymous EDAR variant on multiple dental traits. J Hum Genet 57, 508–514 (2012).
Liu, X., Jian, X. & Boerwinkle, E. dbNSFPv2.0: A Database of Human Non-synonymous SNVs and Their Functional Predictions and Annotations. Hum Mutat 34, E2393–402 (2013).
Speir, M. L. et al. The UCSC Genome Browser database: 2016 update. Nucleic Acids Res 44, D717–25 (2016).
Wilson, P. D. & Kligman, A. M. Do freckles protect the skin from actinic damage. Br J Dermatol 106, 27–32 (1982).
Vierkötter, A. et al. Development of lentigines in German and Japanese women correlates with variants in the SLC45A2 gene. J Invest Dermatol 132, 733–736 (2012).
Hillebrand, G. G. et al. Quantitative evaluation of skin condition in an epidemiological survey of females living in northern versus southern Japan. J Dermatol Sci 27(Suppl 1), S42–52 (2001).
Motokawa, T., Kato, T., Hashimoto, Y. & Katagiri, T. Effect of Val92Met and Arg163Gln variants of the MC1R gene on freckles and solar lentigines in Japanese. Pigment Cell Res 20, 140–143 (2007).
Vanhoutteghem, A. et al. The importance of basonuclin 2 in adult mice and its relation to basonuclin 1. Mech Dev 140, 53–73 (2016).
Vanhoutteghem, A. et al. The zinc-finger protein basonuclin 2 is required for proper mitotic arrest, prevention of premature meiotic initiation and meiotic progression in mouse male germ cells. Development 141, 4298–4310 (2014).
Jacobs, L. C. et al. Comprehensive candidate gene study highlights UGT1A and BNC2 as new genes determining continuous skin color variation in Europeans. Hum Genet 132, 147–158 (2013).
Flori, E. et al. 2,4,6-Octatrienoic acid is a novel promoter of melanogenesis and antioxidant defence in normal human melanocytes via PPAR-γ activation. Pigment Cell Melanoma Res 24, 618–630 (2011).
Lee, J. S., Choi, Y. M. & Kang, H. Y. PPAR-gamma agonist, ciglitazone, increases pigmentation and migration of human melanocytes. Exp Dermatol 16, 118–123 (2007).
Beaumont, K. A. et al. The recycling endosome protein Rab17 regulates melanocytic filopodia formation and melanosome trafficking. Traffic 12, 627–643 (2011).
Tarafder, A. K. et al. Rab11b mediates melanin transfer between donor melanocytes and acceptor keratinocytes via coupled exo/endocytosis. J Invest Dermatol 134, 1056–1066 (2014).
Schafer, J. C. et al. Rab11-FIP2 interaction with MYO5B regulates movement of Rab11a-containing recycling vesicles. Traffic 15, 292–308 (2014).
Rodriguez-Cuenca, S. et al. Peroxisome proliferator-activated receptor γ-dependent regulation of lipolytic nodes and metabolic flexibility. Mol Cell Biol 32, 1555–1565 (2012).
Ma, X. et al. Msi2 Maintains Quiescent State of Hair Follicle Stem Cells by Directly Repressing the Hh Signaling Pathway. J Invest Dermatol 137, 1015–1024 (2017).
Lang, D., Mascarenhas, J. B. & Shea, C. R. Melanocytes, melanocyte stem cells, and melanoma stem cells. Clin Dermatol 31, 166–178 (2013).
Radons, J. The human HSP70 family of chaperones: where do we stand. Cell Stress Chaperones 21, 379–404 (2016).
Prunotto, M. et al. Proteomic analysis of podocyte exosome-enriched fraction from normal human urine. J Proteomics 82, 193–229 (2013).
Gonzales, P. A. et al. Large-scale proteomics and phosphoproteomics of urinary exosomes. J Am Soc Nephrol 20, 363–379 (2009).
Ando, H. et al. Melanosomes are transferred from melanocytes to keratinocytes through the processes of packaging, release, uptake, and dispersion. J Invest Dermatol 132, 1222–1229 (2012).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med 12, e1001779 (2015).
Canela-Xandri, O., Law, A., Gray, A., Woolliams, J. A. & Tenesa, A. A new tool called DISSECT for analysing large genomic data sets using a Big Data approach. Nat Commun 6, 10162 (2015).
Canela-Xandri, O., Rawlik, K., Woolliams, J. A. & Tenesa, A. Improved Genetic Profiling of Anthropometric Traits Using a Big Data Approach. PLoS One 11, e0166755 (2016).
Canela-Xandri, O., Rawlik, K. & Tenesa, A. An atlas of genetic associations in UK Biobank. bioRxiv (2017).
Patel, B. C. K., Gupta, R. & Joos, Z. P. Medscape Eyelid Anatomy. http://reference.medscape.com/article/834932-overview (April 25).
Chen, W. P. Asian Blepharoplasty and the Eyelid Crease E-Book (2015).
Tochihara, K., Saito, K., Mizuguchi, A. & Ikeda, K. Studies on the Relation between Properly Dressed Clothes and Morphological Factors of Face (I). Journal of the Nagoya Women’s College 25, 1–12 (1979).
Doxanas, M. T. & Anderson, R. L. Oriental eyelids. An anatomic study. Arch Ophthalmol 102, 1232–1235 (1984).
Fakhro, A., Yim, H. W., Kim, Y. K. & Nguyen, A. H. The Evolution of Looks and Expectations of Asian Eyelid and Eye Appearance. Semin Plast Surg 29, 135–144 (2015).
Cho, M. & Glavas, I. P. Anatomic properties of the upper eyelid in Asian Americans. Dermatol Surg 35, 1736–1740 (2009).
Kakizaki, H., Malhotra, R. & Selva, D. Upper eyelid anatomy: an update. Ann Plast Surg 63, 336–343 (2009).
Spigoni, G., Gedressi, C. & Mallamaci, A. Regulation of Emx2 expression by antisense transcripts in murine cortico-cerebral precursors. PLoS One 5, e8658 (2010).
Noonan, F. C., Goodfellow, P. J., Staloch, L. J., Mutch, D. G. & Simon, T. C. Antisense transcripts at the EMX2 locus in human and mouse. Genomics 81, 58–66 (2003).
Fukuchi-Shimogori, T. & Grove, E. A. Emx2 patterns the neocortex by regulating FGF positional signaling. Nat Neurosci 6, 825–831 (2003).
Feenstra, J. M. et al. Detection of genes regulated by Lmx1b during limb dorsalization. Dev Growth Differ 54, 451–462 (2012).
Weimer, K., Theobald, J., Campbell, K. S., Esser, K. A. & DiMario, J. X. Genome-wide expression analysis and EMX2 gene expression in embryonic myoblasts committed to diverse skeletal muscle fiber type fates. Dev Dyn 242, 1001–1020 (2013).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am J Hum Genet 94, 559–573 (2014).
Chassaing, N., Bourthoumieu, S., Cossee, M., Calvas, P. & Vincent, M. C. Mutations in EDAR account for one-quarter of non-ED1-related hypohidrotic ectodermal dysplasia. Hum Mutat 27, 255–259 (2006).
Mégarbané, H. et al. Unusual presentation of a severe autosomal recessive anhydrotic ectodermal dysplasia with a novel mutation in the EDAR gene. Am J Med Genet A 146A, 2657–2662 (2008).
Stecksén-Blicks, C., Falk Kieri, C., Hägg, D. & Schmitt-Egenolf, M. Hair shaft structures in EDAR induced ectodermal dysplasia. BMC Med Genet 16, 79 (2015).
Piérard-Franchimont, C. & Piérard, G. E. Alterations in hair follicle dynamics in women. Biomed Res Int 2013, 957432 (2013).
Botchkareva, N. V., Ahluwalia, G. & Shander, D. Apoptosis in the hair follicle. J Invest Dermatol 126, 258–264 (2006).
Paus, R. Principles of hair cycle control. J Dermatol 25, 793–802 (1998).
Reddy, J. V. et al. A functional role for the GCC185 golgin in mannose 6-phosphate receptor recycling. Mol Biol Cell 17, 4353–4363 (2006).
Cheung, P. Y. & Pfeffer, S. R. Transport Vesicle Tethering at the Trans Golgi Network: Coiled Coil Proteins in Action. Front Cell Dev Biol 4, 18 (2016).
Brown, F. C., Schindelhaim, C. H. & Pfeffer, S. R. GCC185 plays independent roles in Golgi structure maintenance and AP-1-mediated vesicle tethering. J Cell Biol 194, 779–787 (2011).
Tu, Y., Li, F., Goicoechea, S. & Wu, C. The LIM-only protein PINCH directly interacts with integrin-linked kinase and is recruited to integrin-rich sites in spreading cells. Mol Cell Biol 19, 2425–2434 (1999).
Chiswell, B. P., Zhang, R., Murphy, J. W., Boggon, T. J. & Calderwood, D. A. The structural basis of integrin-linked kinase-PINCH interactions. Proc Natl Acad Sci USA 105, 20677–20682 (2008).
Fukuda, T., Chen, K., Shi, X. & Wu, C. PINCH-1 is an obligate partner of integrin-linked kinase (ILK) functioning in cell shape modulation, motility, and survival. J Biol Chem 278, 51324–51333 (2003).
Wickström, S. A., Lange, A., Montanez, E. & Fässler, R. The ILK/PINCH/parvin complex: the kinase is dead, long live the pseudokinase. EMBO J 29, 281–291 (2010).
Lorenz, K. et al. Integrin-linked kinase is required for epidermal and hair follicle morphogenesis. J Cell Biol 177, 501–513 (2007).
Karaköse, E. et al. The focal adhesion protein PINCH-1 associates with EPLIN at integrin adhesion sites. J Cell Sci 128, 1023–1033 (2015).
Montanez, E., Karaköse, E., Tischner, D., Villunger, A. & Fässler, R. PINCH-1 promotes Bcl-2-dependent survival signalling and inhibits JNK-mediated apoptosis in the primitive endoderm. J Cell Sci 125, 5233–5240 (2012).
Singh, M. K. et al. The T-box transcription factor Tbx15 is required for skeletal development. Mech Dev 122, 131–144 (2005).
Candille, S. I. et al. Dorsoventral patterning of the mouse coat by Tbx15. PLoS Biol 2, E3 (2004).
Lee, K. Y. et al. Tbx15 controls skeletal muscle fibre-type determination and muscle metabolism. Nat Commun 6, 8054 (2015).
Lee, K. Y. et al. Tbx15 Defines a Glycolytic Subpopulation and White Adipocyte Heterogeneity. Diabetes 66, 2822–2829 (2017).
Yamamoto, Y. et al. Adipose depots possess unique developmental gene signatures. Obesity (Silver Spring) 18, 872–878 (2010).
Heid, I. M. et al. Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat Genet 42, 949–960 (2010).
Liu, C. T. et al. Multi-ethnic fine-mapping of 14 central adiposity loci. Hum Mol Genet 23, 4738–4744 (2014).
D, B. The human eccrine sweat gland: Structure, function and disorders. Journal of Local and Global Health Science 5 (2015).
Shibasaki, M. & Crandall, C. G. Mechanisms and controllers of eccrine sweating in humans. Front Biosci ( Schol Ed ) 2, 685–696 (2010).
Romero, F. R., Haddad, G. R., Miot, H. A. & Cataneo, D. C. Palmar hyperhidrosis: clinical, pathophysiological, diagnostic and therapeutic aspects. An Bras Dermatol 91, 716–725 (2016).
Schlereth, T., Dieterich, M. & Birklein, F. Hyperhidrosis–causes and treatment of enhanced sweating. Dtsch Arztebl Int 106, 32–37 (2009).
Eisenach, J. H., Atkinson, J. L. & Fealey, R. D. Hyperhidrosis: evolving therapies for a well-established phenomenon. Mayo Clin Proc 80, 657–666 (2005).
Shibasaki, M., Secher, N. H., Selmer, C., Kondo, N. & Crandall, C. G. Central command is capable of modulating sweating from non-glabrous human skin. J Physiol 553, 999–1004 (2003).
Shih, C. J., Wu, J. J. & Lin, M. T. Autonomic dysfunction in palmar hyperhidrosis. J Auton Nerv Syst 8, 33–43 (1983).
Birner, P., Heinzl, H., Schindl, M., Pumprla, J. & Schnider, P. Cardiac autonomic function in patients suffering from primary focal hyperhidrosis. Eur Neurol 44, 112–116 (2000).
Higashimoto, I. et al. Primary palmar hyperhidrosis locus maps to 14q11.2-q13. Am J Med Genet A 140, 567–572 (2006).
Maury, E. et al. Human epidermis is a novel site of phospholipase B expression. Biochem Biophys Res Commun 295, 362–369 (2002).
Asano, A., Nelson, J. L., Zhang, S. & Travis, A. J. Characterization of the proteomes associating with three distinct membrane raft sub-types in murine sperm. Proteomics 10, 3494–3505 (2010).
Asano, A., Nelson-Harrington, J. L. & Travis, A. J. Phospholipase B is activated in response to sterol removal and stimulates acrosome exocytosis in murine sperm. J Biol Chem 288, 28104–28115 (2013).
Asano, A., Nelson-Harrington, J. L. & Travis, A. J. Membrane rafts regulate phospholipase B activation in murine sperm. Commun Integr Biol 6, e27362 (2013).
Bradshaw, R. A. & Dennis, E. A. Handbook of Cell Signaling (Academic Press, 2009).
Koffman, J. S., Arnspang, E. C., Marlar, S. & Nejsum, L. N. Opposing Effects of cAMP and T259 Phosphorylation on Plasma Membrane Diffusion of the Water Channel Aquaporin-5 in Madin-Darby Canine Kidney Cells. PLoS One 10, e0133324 (2015).
Kitchen, P. et al. Plasma Membrane Abundance of Human Aquaporin 5 Is Dynamically Regulated by Multiple Pathways. PLoS One 10, e0143027 (2015).
Yoshiura, K. et al. A SNP in the ABCC11 gene is the determinant of human earwax type. Nat Genet 38, 324–330 (2006).
Toyoda, Y., Gomi, T., Nakagawa, H., Nagakura, M. & Ishikawa, T. Diagnosis of Human Axillary Osmidrosis by Genotyping of the Human ABCC11 Gene: Clinical Practice and Basic Scientific Evidence. Biomed Res Int 2016, 7670483 (2016).
Toyoda, Y., Takada, T., Miyata, H., Ishikawa, T. & Suzuki, H. Regulation of the Axillary Osmidrosis-Associated ABCC11 Protein Stability by N-Linked Glycosylation: Effect of Glucose Condition. PLoS One 11, e0157172 (2016).
Ren, Y. et al. A missense variant of the ABCC11 gene is associated with Axillary Osmidrosis susceptibility and clinical phenotypes in the Chinese Han Population. Sci Rep 7, 46335 (2017).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38, 904–909 (2006).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81, 559–575 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
R_Core_Team. R: A Language and Environment for Statistical Computing, https://www.R-project.org/.
de Bakker, P. I. et al. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum Mol Genet 17, R122–8 (2008).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
FE, H. rms: Regression Modeling Strategies. R package version 5.0-1., https://CRAN.R-project.org/package=rms.
Steyerberg, E. W. Clinical prediction models: a practical approach to development, validation, and updating (Springer, New York, NY, 2009).
Suman, K., Yurii, S. A. & A., C. J. W. J. PredictABEL: Assessment of Risk Prediction Models 2014.
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 40, D930–4 (2012).
Eicher, J. D. et al. GRASPv2.0: an update on the Genome-Wide Repository of Associations between SNPs and phenotypes. Nucleic Acids Res 43, D799–804 (2015).
Burdett, T. et al. The NHGRI-EBI Catalog of published genome-wide association studies. http://www.ebi.ac.uk/gwas (Accessed February 2 2017).
Welter, D. et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res 42, D1001–6 (2014).
Consortium, G. T. E. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
UCSC. UCSC Genome Browser, http://genome.ucsc.edu.
Veyrieras, J. B. et al. Exon-specific QTLs skew the inferred distribution of expression QTLs detected using gene expression array data. PLoS One 7, e30629 (2012).
Veyrieras, J. B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet 4, e1000214 (2008).
FANTOM_Consortium_and_the_RIKEN_PMI_and_CLST_(DGT). et al. A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014).
Severin, J. et al. Interactive visualization and analysis of large-scale sequencing datasets using ZENBU. Nat Biotechnol 32, 217–219 (2014).
Zhou, X. et al. The Human Epigenome Browser at Washington University. Nat Methods 8, 989–990 (2011).
Harrow, J. et al. GENCODE: The reference human genome annotation for The ENCODE Project. Genome Res 22, 1760–1774 (2012).
Acknowledgements
The authors would like to acknowledge the Genotype-Tissue Expression (GTEx) Project and Portal, which was essential to this study’s analyses. Support for the GTEx Project was provided by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS. GTEx data used in this study were obtained in September 2017-February 2018 and corresponds to GTEx Release V7.
Author information
Authors and Affiliations
Contributions
C.E. and T.A.J. wrote the manuscript. T.A.J. performed genetic and statistical analyses and made the figures and tables. K.N. wrote programs and performed analyses. S.K. supervised the statistical and bioinformatics analyses. R.M. and Mai.Ka. performed genotyping. R.M. also performed genotype QC. M.Ko. and N.Ku. prepared the questionnaires. T.Y., M.T. and A.T. performed management of DNA specimens and ID tracking. M.A. and M.H. performed study design. A.K., Y.H. and K.I. recruited subjects. Y.T., N.Ka. and Mak.Ka. supervised the analyses and writing of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
T.A.J., S.K., N.Ka., S.K. are employees of StaGen Co. Ltd. R.M., M.A., Mai.Ka., T.Y., A.K., M.T., Y.H., A.T., K.I., M.H., M.Ko., and N.Ku. are employees of EverGene/MTI. EverGene/MTI has applied for patents related to some of the skin-related phenotype associations described in this report. N.Ka. is also Chairman of StaGen Co. Ltd. and a Director of the Tsukuba International Clinical Pharmacology Clinic. C.E., Y.T. and Mak.Ka. declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Endo, C., Johnson, T.A., Morino, R. et al. Genome-wide association study in Japanese females identifies fifteen novel skin-related trait associations. Sci Rep 8, 8974 (2018). https://doi.org/10.1038/s41598-018-27145-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-27145-2
This article is cited by
-
A study of genetic variants associated with skin traits in the Vietnamese population
BMC Genomics (2024)
-
Comparisons between wrinkles and photo-ageing detected and self-reported by the participant or identified by trained assessors reveal insights from Chinese individuals in the Singapore/Malaysia Cross-sectional Genetics Epidemiology Study (SMCGES) cohort
Journal of Physiological Anthropology (2024)
-
Genome-wide association studies for economically important traits in mink using copy number variation
Scientific Reports (2024)
-
Commentary: Facial Aesthetic Dermatological Procedures and Photoprotection in Chinese Populations
Dermatology and Therapy (2023)
-
Automatic landmarking identifies new loci associated with face morphology and implicates Neanderthal introgression in human nasal shape
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.