Abstract
The obesity pandemic in the obstetrical population plus increased frequency of Cesarean delivery (CD) has increased vulnerability to surgical site infection (SSI). Here we characterized the microbiome at the site of skin incision before and after CD. Skin and relevant surgical sites were sampled before and after surgical antisepsis from obese (n = 31) and non-obese (n = 27) pregnant women. We quantified bacterial biomass by qPCR, microbial community composition by 16sRNA sequencing, assigned operational taxonomic units, and stained skin biopsies from incision for bacteria and biofilms. In obese women, incision site harbors significantly higher bacterial biomass of lower diversity. Phylum Firmicutes predominated over Actinobacteria, with phylotypes Clostridales and Bacteroidales over commensal Staphylococcus and Propionbacterium spp. Skin dysbiosis increased post-surgical prep and at end of surgery. Biofilms were identified post-prep in the majority (73%) of skin biopsies. At end of surgery, incision had significant gains in bacterial DNA and diversity, and obese women shared more genera with vagina and surgeon’s glove in CD. Our findings suggest microbiota at incision differs between obese and non-obese pregnant women, and changes throughout CD. An interaction between vaginal and cutaneous dysbiosis at the incision site may explain the a priori increased risk for SSI among obese pregnant women.
Similar content being viewed by others
Introduction
Obesity is a risk factor for Cesarean Delivery (CD)1,2 and is associated with a significantly higher risk of surgical site infection (SSI) including wound infections and post-partum endometritis3,4,5. Compared to women with a normal body mass index [(BMI) 18.5 to 25)], obese women (BMI > 30) are twice as likely to develop SSI post-CD6.
Medical, quality of care and financial incentives have generated pressure to reduce SSI rates for all surgical specialties7. Among universally effective strategies were the standardization of protocols for skin preparation and antisepsis and for use of preoperative antibiotics8,9. Despite their adoption in obstetrics, 6–10% of CDs continue to develop SSIs7,10,11.
Most species of bacteria that cause SSIs are part of the normal human microflora and form biofilms at sites where they exist as commensals12. Due to various factors that disturb local ecology biofilms that harbor bacteria they have been implicated as the cause of 65% of all microbial infections13,14. Microbiologic evidence the notion that biofilm bacterial communities demonstrate resistance to routine surgical preparation protocols15,16.Skin microbial community profiling and biofilm research have remained largely two separate scientific areas, but metagenomics has paved the road toward their convergence. Given the correlation between increasing body weight and higher risk for SSI, it was reasonable to propose that in obese pregnant women the anatomical characteristics of the panniculus fold create a unique microbiologic environment. To address this gap in knowledge we measured the bacterial biomass and microbial diversity and assessed for resence of biofilms in obese versus non-obese pregnant women. We further examined the effectiveness of a universally-recognized preoperative antisepsis protocol17 on commensal colonization at surgically relevant cutaneous sites prior to and after delivery of the newborn.
Results
Clinical Characteristics of the Study Population
For analysis purpose cases were separated into two groups: non-obese (BMI < 30, n = 27) and obese (BMI ≥ 3, n = 31, class 1: n = 11; class 2: n = 7; class 3: n = 13). Compared to normal BMI women, obese mothers had a higher pre-pregnancy BMI (P < 0.001), panniculus grade (P < 0.001) and a longer time period from showering before enrollment (P = 0.011). Within 14 days after Cesarean, 15% (9/58) of enrolled women developed an SSI (BMI < 30, n = 2 and BMI ≥ 30, n = 7, P = 0.111) Details about demographic and surgical characteristics of the groups are presented in Supplementary Table S1.
Topographical and Temporal Changes in Bacterial Biomass
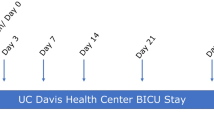
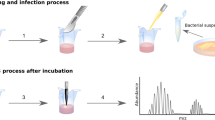
A schematic of the sampled sites relative to antisepsis and surgical times is provided in Fig. 1. Before surgical skin antisepsis, the highest bacterial loads for obese than for non-obese women were seen retro-auricular and in the Pfannenstiel area (Fig. 2A, P < 0.05). Compared to the mid-abdominal site, the average bacterial load at the Pfannenstiel site was 3.8-fold higher in obese group, but just 1.8-fold higher in the non-obese women.
Topographical and temporal changes in bacterial load in obese and non-obese women undergoing scheduled Cesarean delivery. Bacterial load quantified from samples collected from skin sites before prep (A), Pfannenstiel site before and after prep (B), mid-abdomen (C) and surgical scrub (D) before and after prep, vagina (E) and forearm (F) before procedure, and surgeon glove before and after surgery (G) and after prep and post-op at Pfannenstiel site (H).
Surgical antisepsis at Pfannenstiel site did not impact bacterial load in non-obese women (Fig. 2B, P = 0.917). Conversely, bacterial load at Pfannenstiel site was significantly reduced in obese women after prep (P = 0.006) to a point where there was no longer a difference between groups (P = 0.416). Obesity was characterized by an increase in bacterial load at the mid-abdominal site compared to non-obese women (Fig. 2C, P = 0.046). After skin prep, the bacterial load of the surgical scrub was significantly increased in both obese and non-obese women (Fig. 2D, P < 0.001). No differences were noted in vaginal bacterial load between obese and non-obese groups (Fig. 2E, P = 0.981). Vaginal bacterial load was notably higher (69-fold for non-obese and 24-fold for obese group vs. Pfannenstiel site) compared to the skin sites sampled at the same Time 1.
Sampling of the lower uterine segment after delivery of the baby showed this site is rich in bacteria in both obese and non-obese women with no differences between groups (Fig. 2F, P = 0.695). A similar pattern was observed on the surgical glove where the bacterial load was found to be significantly higher compared to pre-surgery (Fig. 2G, P < 0.001). The bacterial load of the glove and of the lower uterine segment at Time 3 were similar for both obese (P = 0.582) and non-obese women (P = 0.698). At the end of surgery and following wound closure, the edges of the Pfannenstiel incision had a very high bacterial load compared to the skin area immediately after scrubbing (Fig. 2H, P < 0.001). The level of added bacteria was similar for obese and non-obese groups (P = 0.872). Bacterial load at the Pfannenstiel incision after the procedure was determined by bacterial load of the lower uterine segment (multivariate regressions, R2 = 0.530, P < 0.001) but not by BMI, thickness of subcutaneous tissue, race or bacterial loads of the vagina or Pfannenstiel area after prep (P > 0.100).
Topographical and Temporal Changes in Microbial Communities at Phylum Level
Preprocessing of raw sequence data yielded a total of 1,998,254 high quality sequences which mapped to 452 genera. These converged taxonomically into 31 distinct phyla of which only one had status Candidatus.
Consistent with previous reports18, we identified at all skin sites a mixed representation of the phyla Actinobacteria, Firmicutes, Proteobacteria, Bacteroidetes and Fusobacterium with the remaining phyla (e.g. Tenericutes, Synergistetes) contributing <1% to any given sample. Actinobacteria and Firmicutes represented >80% of bacterial DNA (Fig. 3).
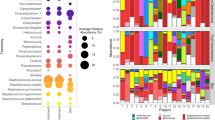
Topographical and temporal changes in microbial communities at the phylum level. As shown the sampled sites had a mixed representation of Actinobacteria, Firmicutes, Proteobacteria, Bacteroidetes and Fusobacterium with Tenericutes, Synergistetes contributing <1% to any given sample. Actinobacteria and Firmicutes represented >80% of bacterial DNA.
Time 1
Prior to skin prep, the most striking differences between obese and non-obese groups were observed at mid-abdomen and Pfannenstiel areas. At both these sites, obese women an increased representation of Firmicutes and Bacteroidetes with proportional decreases in Actinobacteria. The vaginal microbiota of obese women was characterized by reduced representation of Firmicutes and an increased frequency in Actinobacteria and Bacteroidetes.
Time 2
Surgical scrub was associated with a decreased proportion of Actinobacteria and Proteobacteria, but with an increased proportion of Firmicutes, Bacteroidetes and Fusobacteria.
Time 4
Examining the profile of phyla on the Pfannenstiel incision post-surgery, we noted an increased proportion of Proteobacteria, Bacteroidetes and of other less common phyla with further decrease in Actinobacteria. Post-surgery, in obese women the surgeon’s glove had higher abundance of Firmicutes, and Bacteroidetes. Additional analyses with interpretation in the context of current knowledge are included in Supplementary Appendix S2.
Topographical and Temporal Changes in Skin Phylotypes
The inter-personal variation in microbial community composition displayed as taxonomic phylotypes of Grice and Segre18 along with their phyla-level classification is shown in Fig. 4A. After adjustment for inter-personal variation, there were significant differences in microbial community composition between obese and non-obese women at mid-abdomen after prep and at Pfannenstiel site after prep and post-surgery (Supplementary Table S2). The phylotypes accounting for the differences between obese and non-obese women at these sites and times are listed in Supplementary Table S3. Among discriminating phylotypes, Clostridales showed consistent enrichment at sites of obese women.
Topographical and temporal changes in skin microbial phylotypes and skin dysbiosis at the Pfannenstiel site during planned Cesarean delivery. Individual (A) and average (B) changes among obese (n = 6) and non-obese (n = 6) patients at the indicated time points during Cesarean delivery. (C) Pfannenstiel site dysbiosis was observed in obese women throughout Cesarean delivery.
The phylotypes responsible for the differences in community composition before and after skin preparation of obese and non-obese women are presented in Supplementary Table S4 (temporal dissimilarity). At Pfannenstiel site, skin preparation of obese women resulted in increased representation of Clostridales (genera Anaerococus, Peptoniphilus, Finegoldia) and Bacteroidales (genera Prevotella, Porphyromonas) with decrease in the Corynebacterium and Staphylococcus. These changes partially reflected at mid-abdomen. In non-obese women, the phylotype maximally enriched at both Pfannenstiel and mid-abdomen, was Propionbacteria (member of Actinobacteria). Next, we analyzed the difference in skin microbiota between mid-abdomen and Pfannenstiel (topographical dissimilarity, Supplementary Table S4). After prep, the dissimilarity between Pfannenstiel and mid-abdomen narrowed for obese (before prep: 51.9% vs. after prep: 32.1%), but not for non-obese women (before prep: 57.0% vs. after: 57.9%). This can be further appreciated visually from Fig. 4B.
The dissimilarity in microbiota on the Pfannenstiel incision at the end of CD (Time 4) compared to prior to incision (Time 2, after prep) was underlined in obese women by an additional increase in representation of Clostridales (genera Peptoniphilus Anaerococus, Finegoldia and Ezakiella) with decrease in Staphylococci and Corynebacteria. At the end of CD, Clostridales became the most abundant incisional phylotype (35.6%). The rare phyla increased in representation were led by Synergystetes and were identified only in obese women (genera Jonquella and Pyramidobacter). In contrast, for non-obese women, the difference after surgery was accounted primarily by further increase in representation of Propionbacteria and Staphylococci (Supplementary Table S5). An increase in Propionbacterium was noted also in obese women but the representation of this phylotype at the end of CD was less than a third than in non-obese.
Temporal Changes in Skin Dysbiosis at Pfannenstiel Site
We estimated the level of dysbiosis at the surgical site based on the abundance of Clostridales and Bacteroidals phylotypes relative to skin the commensals Propionbacteria and Staphylococcus. The dysbiotic ratio between obese and non-obese women becomes significant after skin antisepsis and further increased after surgery (Fig. 4C).
Changes in Skin Genera and in Alpha-Diversity
Clinically relevant comparisons in alpha-diversity metrices for richness (Fisher’s alpha index), dominance (Berger-Parker d index) and diversity (Shannon-Weiner H index) at genera level are presented in Supplementary Fig. S1. In obese women, skin preparation of the Pfannenstiel site resulted in increased species richness and bacterial diversity, but a decreased of the dominance index which also remained lower post-op.
The Circos visualization of shared genera at the Pfannenstiel area before and after prep is presented in Fig. 5A. After prep, the Pfanneniel site harbored more genera compared to pre-op. The enrichment in new genera was higher in obese women (82 vs 69 new genera) with only 19 new genera in common implying a unique profile in obese women. The same type of analysis among Pfannenstiel incision post-op, vagina and surgeon’s glove demonstrated shared genera with the extent of overlap higher for obese women (11 genera common to all 3 sites for non-obese vs. 32 for obese women) Fig. 5B. Of sequences matching to genera uniquely shared in obese women, 87% matched to Clostridales and Bacteroidals phylotypes (leading genera: Anaerococcus, Peptoniphilus, Finegoldia, Porphyromonas).
Circos visualization of bacterial genera shared among pre- and post-op Pfannenstiel incision (A), and post-op Pfannenstiel site, vagina and surgeon’s glove (B) in obese and non-obese women undergoing scheduled Cesarean delivery. After prep, the Pfannenstiel site showed more genera compared to pre-op. The enrichment was higher in obese women. Shared genera were observed between Pfannenstiel incision post-op, vagina and surgeon’s glove, and the extent of overlap was higher for obese women.
Identification of biofilms on skin biopsy after surgical antisepsis
Biofilms represent a reservoir of hidden bacteria which can become mobilized upon scrubbing. Of the 17 skin biopsies that were both positive by Gram stain and also had associated keratinized epithelium, 13 (76%) were positive for integration host factor (IHF). IHF is one of two bacterial DNA-binding proteins of the DNABII family that are abundant in biofilms and, along with extracellular bacterial DNA, provide essential structural support to the biofilm19,20,21. Positive staining for IHF was independent of BMI. Representative micrographs of skin biopsies from 4 women are presented in Supplementary Fig. S2.
Discussion
Compared with non-obese women, obese mothers have significantly higher pre-operative bacterial biomass at the anticipated Pfannenstiel incision site. Skin pre-operative antisepsis with CHG decreases bacterial quantity at the future surgical site to a point where obese and non-obese women begin surgery with similar bacterial loads. Excess bacteria from the Pfannenstiel area of obese woen may have been displaced to the mid-abdomen since this site and surgical scrub showed a significant gain in bacteria immediately following antisepsis. We demonstrated a considerable surge in bacterial DNA load on the surgeon’s glove, and the Pfannenstiel incision at the end of surgery both in obese (2.4-fold) and non-obese (3.3-fold) women. We provided pragmatic evidence that despite strict aseptic techniques and maintenance of a safe environment in the operating room, elective CD is not a sterile procedure. We identified a significant gain in bacterial biomass at the end of surgery in both obese and non-obese women on the Pfannenstiel site which is expected to be essentially sterile. The significance of our findings is supported by previous studies concluding tissue bacterial load has independent predictive value for subsequent infection22,23,24,25,26.
The commensal, mutualistic, parasitic and saprophytic relationships established between host’s skin and its inhabiting microorganisms has been reviewed in numerous publications27,28,29,30,31. Our findings are in agreement with the notion that skin occlusion favoring an acidic pH and high humidity favors growth of Firmicutes, known to be associated with SSI in obese non-pregnant patients32. Examination of the phyla on the Pfannenstiel incision post-surgery revealed an increased proportion of Proteobacteria, and of less common phyla. Within these, two species (Jonquella and Pyramidobacter spp.) in phylum Synergystetes had the leading representation. Synergystetes is a newly classified gram-negative anaerobic phylum identified in periodontal, infections33. Interestingly, mice fed a high fat diet reported Synergistetes part of the “obese microbiota” with translocation resulting from loss of gut barrier34. Thus, we raise the possibility of Synergystetes as potential etiologic agent of polymicrobial nature of SSI in obese women. Collectively, our findings imply that obese pregnant women undergoing CD may have an increased risk of SSI, due to anatomical differences from the pannus, which creates a unique moist environment at the anticipated site of surgery. It further suggests that surgical antisepsis which is expected to disrupt skin biofilms may uncover a unique profile of microbiota skin biofilm resulting in increased bacterial richness and diversity, and decreased dominance in obese women. This intervention, in certain patients, may disrupt the equilibrium of the “good” commensal skin bacteria and the “bad” pathogenic bacteria, further increasing risk of infection.
The skin and genital tract are the most influential reservoirs for bacterial contamination35,36,37. Traditionally, minimizing the bioburden of cutaneous microflora on the surgical site prior to surgery or intra-operatively is fundamental for prevention of SSI. Our results suggest marked increased in bacterial DNA at Pfannenstiel incision post-surgery in both groups, questioning the overall sterility of the current surgical technique for CD. The lower uterine segment and the physician’s glove after delivery of the baby’s head demonstrated these sites were rich in bacterial DNA, implying they may be the source of bacterial DNA being deposited on the maternal skin at the end of surgery. Our inference is supported by finding shared common genera between Pfannenstiel incision post-op, vagina and surgeon’s glove, with more overlap in obese women. Recently, the concept and benefits (i.e. asthma, atopic disease) of neonatal vaginal seeding, immediately after CD, received increased attention38,39. Our analysis of the shared genera among Pfannenstiel incision, vagina and surgeon’s glove demonstrates this colonization process occurs in the context of the current surgical technique of CD thus implying vaginal seeding is unnecessary. However, routine cleansing the vagina with antiseptic solution, changing the entire surgical team’s gloves and surgical instruments intra-operatively, after delivery of the placenta, would reduce the rate of post-CD wound infections40,41.
Biofilms are described as microbial communities that attach to an interface or to each other, and are embed into a matrix of extracellular polymeric substances (EPS) aimed to protect microorganisms from outside perturbations14. Biofilms allows for microbial communication, enhanced virulence and breakdown of nutrients aiding microbial succession and development14 .Biofilms tolerate antimicrobial properties of the immune system, antiseptics and antibiotics42,43. Specifically, the antimicrobial effectiveness of ChloraPrepTM was reduced when challenged by S. epidermidis and S. aureus (Firmicutes), when in a biofilm formation15,16 .In our study we identified persistence of biofilms after surgical antisepsis. Our literature search suggests most microorganisms identified using metagenomic tools carry the ability to adhere and form biofilms44. Overall, by using sequencing technology, we identified insight into the nature of skin microbiome in obese women undergoing CD. To our knowledge there have been no prior studies of the skin microbiota in pregnant women or as this relates to BMI. Our study provides first necessary step before prospective clinical studies of surgical outcome could be effectively designed to underscore the importance of incorporating targeted SSI prophylaxis and practices to disrupt skin biofilms that are not routinely disrupted using the current alcohol-based chlorahexidine solutions.
Methods
Patients and Study Design
Using a prospective observational study design, we enrolled 58 consecutive women pregnant with singletons who were scheduled for CD at term (≥37weeks). Indications for CD were: prior CD (n = 51), suspected fetal macrosomia (n = 3), malpresentation (n = 2), history of prior shoulder dystocia (n = 1) and patient preference (n = 1). Clinical signs of labor, membrane rupture, diabetes, positive HIV or hepatitis status, known active infections or antibiotics use during the past 7 days were considered exclusion criteria. All patients received routine prophylactic antibiotics in accordance with the current recommendations10 .All women were recruited at The Ohio State University Wexner Medical Center and provided written informed consent. Experimental protocol, data collection, and consent forms were approved by The Ohio State University Human Research Protection Program and all methods were performed in accordance with the relevant guidelines and regulations.
Women were enrolled at admission for surgery. We took a pragmatic approach with respect to pre- and intra-operative skin antisepsis and to the surgical technique for CD. Details about data collection prior to skin preparation before surgery are provided in Supplementary Appendix S1. A schematic of the sampled sites relative to antisepsis and surgical times is provided in Fig. 1. Time 1 (pre-antisepsis), involved swabs from retro-auricular crease (behind the ear), volar forearm, mid-abdomen, anticipated site of Pfannenstiel incision, ChloraPrepTM applicator, vagina, and surgeon’s glove.
At Time 2 (pre-operative), after skin prep and before skin incision we obtained swabs from the same mid-abdomen and Pfannenstiel sites, and ChloraPrepTM applicator after skin prep. Time 3 (intra-operative), after skin incision and prior to opening of the peritoneal cavity involved excision of a small sample of skin from the lower edge of the incision, and collection of samples from lower uterine segment and surgeon’s glove after delivery of the neonate. Time 4 (post-operative), after skin closure, and prior to application of the sterile dressing, involved collection of a swab directly from the surface of the incision. To minimize variability, the skin prep and collection of the swabs was performed by a single investigator (KMR). Technical details about sites and sample collection are provided in Supplementary Appendix S1.
DNA Extraction, Measurement of Bacterial Load and Identification of Bacterial Biofilm
Following collection, all swabs were immediately transported in their sterile containers and stored at −20 °C until processing (2–3 months). DNA was isolated and cleaned up with QIAamp DNA mini-kit (QIAGEN). 16 S rRNA gene was amplified by RT-PCR using specific primers and probe sets. Skin samples from the Pfannenstiel site were fixed in formalin and cryo-embedded. Details on laboratory techniques are provided in Supplementary Appendix S1.
Sequencing of 16 s RNA Genes and Microbial Community Analysis
DNA samples were submitted for metagenomics analysis. The amplified gene segments where sequenced using the ABI Prism 3100xl genetic analyzer (Applied Biosystems), a 16-capillary instrument using fluorescence based electrophoresis technology for DNA sequencing and fragment analysis at The Ohio State University DNA core facility (Wooster, OH). 16 S metagenomics analysis was carried out as per Schloss et al.45 and Kozich et al.46 using the most recently available version of mothur (http://www.mothur.org, ver1.40.0).
Data Processing and Statistical Analyses
Details about processing of the sequence reads, analysis of the genera as described by Grice and Segre18, and additional statistical analyses are provided in Supplementary Appendix S1.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Fitzsimons, K., Modder, J. & Greer, I. A. Obesity in pregnancy: risks and management. Obstet. Med. 2, 52–62 (2009).
Kawakita, T. et al. Indications for primary cesarean delivery relative to body mass index. Am. J. Obstet. Gynecol. . 215(515), e1–e9, https://doi.org/10.1016/j.ajog.2016.05.023 (2016).
Myles, T. D., Gooch, J. & Santolaya, J. Obesity as an independent risk factor for infectious morbidity in patients who undergo cesarean delivery. Obstet. Gynecol. 100, 959–964 (2002).
Machado, L. Cesarean section in morbidly obese parturients: practical implications and complications. N. Am. J. Med. Sci. 4, 13–18 (2012).
Wloch, C. et al. Risk factors for surgical site infection following caesarean section in England: results from a multicenter cohort study. BJOG. 119, 1324–1333 (2012).
Perlow, J. H. & Morgan, M. A. Massive maternal obesity and perioperative cesarean morbidity. Am. J. Obstet. Gynecol. 170, 560–565 (1994).
Rajkumar, R., Press, M. J. & Conway, P. H. The CMS Innovation Center–a five-year self-assessmen. t. N. Engl. J. Med. 372, 1981–1983 (2015).
Bakkum-Gamez, J. N. & Dowdy, S. C. Improving Surgical Site Infection Rates Through Continuous Quality Improvement. Ann. Surg. Oncol. 24, 305–307 (2017).
Tita, A. T. et al. Adjunctive Azithromycin Prophylaxis for Cesarean Delivery. N. Engl. J. Med. 375, 1231–1241 (2016).
ACOG Practice Bulletin No. 120: Use of prophylactic antibiotics in labor and delivery. American College of Obstetricians and Gynecologists. Obstet. Gynecol. 117, 1472–1483 (2011).
Centers for Disease Control and Prevention. National and state healthcare-associated infections standardized infection ratio report (January-December 2010). http://www.cdc.gov/hai/pdfs/sir/national-SIR-Report_03_29_2012.pdf (2012).
Altemeier, W. A., Culbertson, W. R. & Hummel, R. P. Surgical considerations of endogenous infections–sources, types, and methods of contro. l. Surg. Clin. North. Am. 48, 227–240 (1968).
Cogan, N. G., Cortez, R. & Fauci, L. Modeling physiological resistance in bacterial biofilms. Bull. Math. Biol. 67, 831–853 (2005).
Costerton, J. W., Stewart, P. S. & Greenberg, E. P. Bacterial biofilms: a common cause ofpersistent infections. Science. 284, 1318–1322 (1999).
Bonez, P. C. et al. Chlorhexidine activity against bacterial biofilms. Am J Infect Control. 41, e119–e122, https://doi.org/10.1016/j.ajic.2013.05.002 (2013).
Knobloch, J. K., Horstkotte, M. A., Rohde, H., Kaulfers, P. M. & Mack, D. Alcoholic ingredients in skin disinfectants increase biofilm expression of Staphylococcus epidermidis. J. Antimicrob. Chemother. 49, 683–687 (2002).
Tuuli, M. G. et al. A Randomized Trial Comparing Skin Antiseptic Agents at Cesarean Delivery. N. Engl. J. Med. 374, 647–655 (2016).
Grice, E. A. & Segre, J. A. The skin microbiome. Nat. Rev. Microbiol. 9, 244–253 (2011).
Goodman, S. D. et al. Biofilms can be dispersed by focusing the immune system on a common family of bacterial nucleoid-associated proteins. Mucosal Immunology. 4, 625–637 (2011).
Brockson, M. E. et al. Evaluation of the kinetics and mechanism of action of anti-integration host factor-mediated disruption of bacterial biofilms. Mol. Micro. 93, 1246–1258 (2014).
Gustave, J. E., Jurcisek, J. A., McCoy, K. S., Goodman, S. D. & Bakaletz, L. O. Targeting bacterial integration host factor to disrupt biofilms associated with cystic fibrosis. J. Cys. Fibrosis. 12, 384–389 (2013).
Sen, R. K., Murthy, N., Gill, S. S. & Nagi, O. N. Bacterial load in tissues and its predictive value for infection in open fractures. J. Orthop. Surg. (Hong Kong). 8, 1–5 (2000).
Turtiainen, J., Hakala, T., Hakkarainen, T. & Karhukorpi, J. The impact of surgical wound bacterial colonization on the incidence of surgical site infection after lower limb vascular surgery: a prospective observational study. Eur. J. Vasc. Endovasc. Surg. 47, 411–417 (2014).
Whyte, W., Hambraeus, A., Laurell, G. & Hoborn, J. The relative importance of routes and sources of wound contamination during general surgery. I. Non-airborne. J. Hosp. Infect. 22, 41–54 (1991).
Saleh, K., Sonesson, A., Persson, B., Riesbeck, K. & Schmidtchen, A. A descriptive study of bacterial load of full-thickness surgical wounds in dermatologic surgery. Dermatol. Surg. 37, 1014–1022 (2011).
Cronquist, A. B., Jakob, K., Lai, L., Della, L. P. & Larson, E. L. Relationship between skin microbial counts and surgical site infection after neurosurgery. Clin. Infect. Dis. 33, 1302–1308 (2001).
Prescott, S. L. et al. The skin microbiome: impact of modern environments on skin ecology, barrier integrity, and systemic immune programming. World Allergy Organ. J. 10, 29, https://doi.org/10.1186/s40413-017-0160-5 (2017).
Sanford, J. A. & Gallo, R. L. Functions of the skin microbiota in health and disease. Semin. Immunol. 25, 370–377 (2013).
Bouffard, G. G., Blakesley, R. W., Wolfsberg, T. G., Turner, M. L. & Segre, J. A. A diversity profile of the human skin microbiota. Genome Res. 18, 1043–1050 (2008).
Roth, R. R. & James, W. D. Microbial ecology of the skin. Annu. Rev. Microbiol. 42, 441–464 (1988).
Leeming, J. P., Holland, K. T. & Cunliffe, W. J. The microbial ecology of pilosebaceous units isolated from human skin. J. Gen. Microbiol. 130, 803–807 (1984).
Alexiou, K. et al. A prospective randomised trial of isolated pathogens of surgical site infections (SSI). Ann. Med. Surg. (Lond). 21, 25–29 (2017).
Downes, J. et al. Pyramidobacter piscolens gen. nov., sp. nov., a member of the phylum ‘Synergistetes’ isolated from the human oral cavity. Int. J. Syst. Evol. Microbiol. 59, 972–980 (2009).
Xiao, L. et al. High-fat feeding rather than obesity drives taxonomical and functional changes in the gut microbiota in mice. Microbiome 5, 43, https://doi.org/10.1186/s40168-017-0258-6 (2017).
Sarsam, S. E., Elliott, J. P. & Lam, G. K. Management of wound complications from cesarean delivery. Obstet. Gynecol. Surv. 60, 462–473 (2005).
Martens, M. G., Kolrud, B. L., Faro, S., Maccato, M. & Hammill, H. Development of wound infection or separation after cesarean delivery; prospective evaluation of 2,431 cases. J. Reprod. Med. 40, 171–175 (1995).
Roberts, S., Maccato, M., Faro, S. & Pinell, P. The microbiology of post-cesarean wound morbidity. Obstet. Gynecol. 81, 383–386 (1993).
Committee Opinion No. 725: Vaginal Seeding. Committee on Obstetric Practice. Obstet Gynecol. 130, 274-278 (2017).
Haahr, T. et al. Vaginal seeding or vaginal microbial transfer from the mother to the caesarean-born neonate: a commentary regarding clinical management. BJOG. 125, 533–536 (2018).
Starr, R. V., Zurawski, J. & Ismail, M. Preoperative vaginal preparation with povidone-iodine and the risk of postcesarean endometritis. Obstet. Gynecol. 105, 1024–1029 (2005).
Haas, D. M. et al. Vaginal cleansing before cesarean delivery to reduce postoperative infectious morbidity: a randomized controlled trial. Am. J. Obstet. Gynecol. 202, 310.e1–e6, https://doi.org/10.1016/j.ajog.2010.01.005 (2010).
Epstein, S. S. Microbial awakenings. Nature. 457, 1083 (2009).
Oliver, J. D. Recent findings on the viable but nonculturable state in pathogenic bacteria. FEMS Microbiol. Rev. 34, 415–425 (2010).
Donlan, R. M. & Costerton, J. W. Biofilms: survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 15, 167–193 (2002).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–7541 (2009).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a dual index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 79, 5112–5120 (2013).
Acknowledgements
We are indebted to the nurses, fellows and residents at The Ohio State Wexner Medical Center, Department of Obstetrics and Gynecology, and to all patients who participated in the study. This project was partially supported through a grant awarded by the American College of Obstetrics and Gynecology (ACOG): Duchesnay USA Research Fellowships in Quality Improvement in Maternity Care. The funding source had no involvement in study design, interpretation of data, writing of the report or decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
(K.M.R.)- hypothesis, designing study, specimen collection, conducting experiments, acquiring data, analyzing data, and writing the manuscript; (I.A.B.)- designing study, specimen collection, conducting experiments, acquiring data, analyzing data, and writing the manuscript; (J.J.)- specimen processing, biofilm identification and writing the manuscript; (T.S.)- specimen processing, managing budget and writing manuscript; (WW)- specimen processing and acquiring data; (G.Z.)- specimen processing, acquiring data and writing manuscript; (W.E.A.)- analyzing data and writing manuscript; (R.W.R.)- analyzing data and writing manuscript; (L.O.B.)- specimen processing, biofilm identification and writing manuscript; (S.F.T.)- acquiring data and writing manuscript; (C.S.B.)- designing study, specimen collection, conducting experiments, acquiring data, analyzing data, and writing the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rood, K.M., Buhimschi, I.A., Jurcisek, J.A. et al. Skin Microbiota in Obese Women at Risk for Surgical Site Infection After Cesarean Delivery. Sci Rep 8, 8756 (2018). https://doi.org/10.1038/s41598-018-27134-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-27134-5
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.