Abstract
Micro RNAs (miRNA) are small non-coding RNAs that post-transcriptionally regulate gene expression. Dysregulation of miRNA cluster 143/145 has been reported in several malignancies, but their role in non-small cell lung cancer (NSCLC) remains elusive. This study investigates the prognostic impact of miR-143 and miR-145 in primary tumors and metastatic lymph nodes in NSCLC tissue. Tissue from 553 primary tumors and 143 matched metastatic lymph nodes were collected and tissue microarrays were constructed. In situ hybridization was used to evaluate miR-143 and miR-145 expression in tumor epithelial cells and stromal cells in the primary tumors and lymph nodes. In vivo data was supplemented with functional studies of cell lines in vitro to evaluate the role of miR-143 and miR-145 in NSCLC tumorigenesis. In our cohort, stromal miR-143 (S-miR-143) and miR-145 (S-miR-145) expression in primary tumor tissue were independent prognosticators of improved disease-specific survival (DSS) in female (S-miR-143, HR: 0.53, p = 0.019) and male patients (S-miR-145, HR: 0.58, p = 0.021), respectively. Interesting correlations between the miR cluster 143/145 and previously investigated steroid hormone receptors from the same cohort were identified, substantiating their gender dependent significance.
Similar content being viewed by others
Introduction
Lung cancer remains the leading cancer killer in the world with more than 1.6 million estimated annual deaths, worldwide1. The predominant histological subtype, non-small cell lung cancer (NSCLC), accounts for 85% of cases and can be further divided into subgroups according to the recent WHO classification; the most frequent being adenocarcinoma and squamous cell carcinoma2. Surgical resection is the main curative treatment modality for NSCLC, but unfortunately, the majority of patients are diagnosed in advanced stages and thus not eligible for surgery. Despite development in surgical techniques, diagnostic technologies and the implementation of biologic treatment including immunotherapy, the 5-year survival remains depressing at only 18%3. To optimize therapy and improve the overall survival, it is pivotal to uncover better prognostic and predictive molecular markers.
microRNAs (miRNAs) are small non-coding RNA elements important in various biological processes, including tumorigenesis4. They negatively regulate protein translation by binding to the 3′UTR of target messenger RNAs (mRNAs) leading to mRNA degradation or suppression of translation5. miRNA expression correlates with biological and clinical characteristics of tumors; differentiation, aggression, tissue type and therapy response6. Further, “miRNA replacement therapy” provides a novel treatment opportunity by reintroducing downregulated miRNA into cancer cells7. A phase I clinical trial of miRNA replacement therapy in thoracic cancers, based on the miR-15/107 group of miRNAs, was recently completed with promising results8.
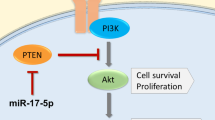
miR cluster 143/145 consists of two miRNAs, miR-143 and miR-145, transcribed from a gene cluster on chromosome 5. It regulates multiple genes involved in cancer cell growth, including well-established cancer related hormone receptors such as ERα, and is generally regarded as a tumor suppressor9,10,11. Reports have indicated a possible prognostic role in non-small cell lung cancer12,13. The presented study investigates the prevalence and prognostic significance of miR-143 and miR-145 in NSCLC. The utilization of in situ hybridization allow both localization of expression according to cell-type and sub-cellular compartment. Further, correlations with steroid hormone receptors progesterone receptor (PR), estrogen receptor alpha (ERα), estrogen receptor beta (ERβ) and aromatase enzyme (AR), previously investigated by our group, were explored. The clinicopathological findings were supplied with data from functional in vitro studies.
Materials and Methods
Patients
NSCLC patients who underwent radical resection at the Nordland Central Hospital and the University Hospital of North Norway from 1990 to 2011, were retrospectively included in this study. Six-hundred-and-thirty-three patients were identified from the hospital records. Of these, 80 patients were excluded due to (1) inadequate fixation of paraffin-embedded tissue blocks (n = 26), (2) radiotherapy or chemotherapy prior to surgery (n = 15), (3) other malignancy within 5 years ahead of an NSCLC diagnosis (n = 39), leaving 553 patients eligible for inclusion. One-hundred-and-seventy-two of the included patients had confirmed metastatic lymph node tissue disease (LN+). Of these, 143 patients had lymph node specimens available for analysis. The eight edition of the International Union Against Cancer TNM classification was used to re-stage all patients, and the tumors were histologically re-classified according to the 2015 World Health Organization Classification of Lung Tumors2,14. Follow-up data as of October 1st 2013.
Tissue microarray construction
All specimens were embedded in paraffin blocks and examined by two experienced pathologists. Detailed methodology regarding TMA construction has previously been published15. Briefly, (1) representative areas of stromal and tumor tissue in primary tumors and tumor tissue from lymph nodes were identified and sampled with a 0.6 mm stylet, (2) transferred to the recipient TMA block and (3) cut into 4μm sections with a Micron microtome (HM355S) prior to in situ hybridization. Normal lung tissue far from the site of the tumor, and lung tissue samples from 20 emphysema patients without any history of neoplastic disease, were used as controls and for comparing biomarker expression level in malignant vs non-malignant tissue.
In situ hybridization (ISH)
miR-143 and miR-145 expression was analyzed by in situ hybridization (ISH) using the Ventana Discovery Ultra (Ventana Medical Inc, Arizona, USA). Optimization of biomarker detection included: RNA degradation prevention, testing of reagent concentration for the tissue of interest and detection method, and testing of hybridization temperatures for each probe with RNA Tm (melting temperature) as guideline. Digoxigenin (DIG) labeled lock nucleic acid (LNA) probes for miR-145-5p (hsa-miR-145, Prod. No. 88068-15, concentration: 2.5 nM), miR-143-3p (hsa-miR-143, Prod. No. 38515-15, concentration: 10 nM), negative control (Scramble miR, Prod. No. 99004-15, concentration: 10 nM) and positive control (U6 has/mmu/rno, Prod. No. 99002-15, concentration: 0.5 nM) were used in this study. Exiqon validated the LNATM miR probes by CE (Capillary Electrophoresis) or HPLC (High-Performance Liquid Chromotography) and confirmed identity of compound by MS (Mass Spectrometry). A TMA multi organ block was used as positive and negative tissue controls.
4 µm TMA sections were incubated overnight at 60 °C to attach tissue to Super Frost Plus slides. To ensure good distribution of reagents and protect sections from desiccation, LCS (Liquid Coverslip oil, Roche, 5264839001) was added. Deparaffinization was performed in EZ Prep buffer (Roche 5279755001) at 68 °C (3 × 12 min). Demasking was done at 95 °C with CC1 buffer (Roche, 6414575001) for 40 minutes. Subsequently, sections were rinsed with Reaction Buffer (Roche 5353955001) and RiboWash, SSPE buffer (Roche 5266262001).
All slides were denaturated for 8 min. at 90 °C. Hybridization with probes was performed for 60 min at 54 °C for miR-145, 55 °C for miR-143, 57 °C for scramble miR and 55 °C for U6. Stringent wash procedures were done at 2 × 8 min with 2.0X RiboWash, SSPE buffer with the same temperatures as used under hybridization for each probe. Blocking against unspecific bindings followed, with blocking solution (Roche, 5268869001) for 16 min. at 37 °C. Alkaline phosphatase (AP)-conjugated anti DIG (Anti-DIG-AP Multimer, Roche 07256302001) was incubated for 20 min. at 37 °C for immunologic detection. After rinsing, substrate enzymatic reactions were carried out with NBT/BCIP (CromoMap Blue kit, Roche 526661001) for 60 min at 37 °C, to give a blue precipitate to detect the microRNA. Sections were again rinsed and counterstained in 4 min with Red Stain II (Roche 5272017001). Increasing gradients of ethanol solutions was used for dehydration. Finally, all sections were mounted using the Histokitt mounting medium (Assistant-Histokitt, 1025/250 Sondheim/Rhoen Germany).
Scoring of ISH
All tissue samples were independently and semi-quantitatively scored by an experienced pathologist (SAS) and a trained medical doctor (KS). Biomarkers were evaluated by intensity in neoplastic epithelial cells and stromal cells; 0 (no staining), 1 (weak), 2 (intermediate) and 3 (strong) and density in stromal cells; 0 = absent, 1 = 1–5%, 2 = 6–50%, 3 = >50%. Due to homogenous staining in neoplastic epithelial cells, scoring of biomarker density was not deemed necessary. For stromal biomarker expression (S-miR) the mean value of intensity and density combined, was calculated. Staining of fibroblasts, fibrocytes, lymphocytes, smooth muscle cells (SMC) and endothelial cells in blood and lymph vessels were included while scoring tumor stroma. Striking positivity was noted in endothelial cells lining the blood vessels and SMCs, including the smallest capillaries. Each variable was dichotomized for survival analyses based on a minimal p-value approach. A high score was defined as a score ≥ mean value for stromal-miR-143 (S-miR-143, mean value: 1.87) and tumor-miR-143 (T-miR-143, mean value: 1.98) and >0 for S-miR-145 and T-miR-145. The same scoring approach was used in PT, LN+, positive and negative tissue controls. For LN+ however, the stromal compartment was not scored due to large numbers of excessively stained lymphocytes. In normal lung tissue from NSCLC patients, collected far from the site of the tumor, miR-143 was prominently expressed in type 2 pneumocytes and macrophages. Collagen and endothelial cells lining the alveolar wall were mostly negative. miR-145 expression was observed in a few pneumocytes type I, while most were negative. Staining in macrophages was predominantly negative.
Functional studies
Cell cultures
Four lung cancer cell lines were used: the adenocarcinoma cell line A549 (ATCC® CCL-185™), the squamous cell carcinoma cell line H520 (ATCC® HTB182™), and the two large cell carcinoma cell lines H460 (ATCC ® HTB-177™) and H661 (ATCC® HTB183™). All cells were cultured in RPMI-1640 media (# R8758, Sigma-Aldrich, St. Louis, USA) supplemented with 10% fetal bovine serum (# S0415, Biochrom, Berlin, Germany) and 1x penicillin-streptomycin antibiotic mixture (# P0781, Sigma-Aldrich, St. Louis, USA) and incubated at 37 °C in 5% CO2 humidified atmosphere.
Cell transfection
Cells were transiently transfected with either 100 nM has-miR-143-3p Pre-miR™ miRNA Precursor (catalog# PM10883, Thermo Fisher Scientific, USA) and/or 100 nM has-miR-145-5p Pre-miR™ miRNA Precursor (catalog# PM11480, Thermo Fisher Scientific, USA), alongside the Cy3™ Dye-Labeled Pre-miR Negative Control #1 (catalog# AM17120, Thermo Fisher Scientific, USA) using the transfection reagent Lipofectamine® 2000 (catalog#11668-019, Life Technologies, Waltham, USA). Transfected Cy3™ Dye-Labeled Pre-miR Negative Control emits fluorescent light when exposed to UV-light, and using a fluorescence microscope, the transfection efficiency was evaluated to be 80–95%.
Total RNA isolation
Total RNA from the cells were isolated using the miRNeasy Mini Kit (cat.# 217004, Qiagen, Hilden, Germany). First, 700 μl QIAzol lysis reagent was used to lyse the cells before homogenization and a 5 minute incubation at room temperature. Second, 140 μl chloroform was added, samples shaken, and then incubated for 3 minutes at room temperature. Third, samples were centrifuged at 12000 G for 15 minutes at 4 °C before the upper aqueous phase was transferred and mixed with 100% ethanol. Finally, the samples were transferred to the RNeasy® Mini column and washed in several steps before elution with 50 μl ddH2O. Samples were stored at −70 °C.
cDNA synthesis
For the first strand cDNA synthesis, the miScript II RT Kit (cat.# 218160, Qiagen, Hilden, Germany) was used. First, 100 ng total RNA was mixed with 4 μl 5X miScript HiSpec buffer, 2 μl 10X Nucleics mix, 2 μl miScript reverse transcriptase mix, and RNase-free water to a final volume of 20 μl. Second, samples were incubated for 1 hour at 37 °C, and then incubated at 95 °C for 5 minutes. Finally, all samples were diluted to a total volume of 200 μl using RNase-free water, and stored at −70 °C.
RT-PCR
Endogenous levels of miR-143 and miR-145 in the cancer cells were quantified relative to the non-cancerous lung cell line NL20 (ATCC® CRL-2503™), and normalized to the stably expressed reference snRNA RNU6 using real-time PCR and the miScript SYBR® Green PCR Kit (catalog# 218073, Qiagen, Hilden, Germany). Primers were miScript Primer Assays Hs_miR-143_1 miScript Primer Assay (catalog# MS00003514, Qiagen, Hilden, Germany), Hs_miR-145_1 miScript Primer Assay (catalog# MS00003528, Qiagen, Hilden, Germany) and Hs_RNU6-2_11 miScript Primer Assay (catalog# MS00033740, Qiagen, Hilden, Germany), according to the manufacturers protocol. In short, a total volume of 25 µl/well in a 96-well plate included 1 µl cDNA mixed with 12.5 µl 2x QuantiTect SYBR Green PCR Master Mix, 2.5 µl 10x miScript Universal Primer, 2.5 µl 10x miScript Primer Assay, and 6.5 µl RNase-free Water. The plate was sealed and centrifuged for 1 minute at 1000 G before it was placed in the 7300 Real-Time PCR System (Thermo Fisher Scientific, Waltham, Massachusetts, USA). Each sample was analyzed in quadruplicates, and two independent experiments were performed.
Proliferation assay
The ability of cancer cells to proliferate was evaluated using the real-time cell analyzer xCELLigence, RTCA DP (catalog#05469759001, ACEA Biosciences, San Diego, USA) fitted with the E-plate 16 (catalog#05469830001, ACEA Biosciences, San Diego, USA). Prior to seeding, cells were trypsinized until detached, resuspended in complete growth media, and counted. In accordance with the manufacturer protocol, cells were seeded in quadruplicates into an E-plate after baseline measurements. The E-plate containing cells was positioned in the RTCA DP instrument, located in an incubator preserving the same conditions as used for routine cultivation of cell lines. The cell index was automatically measured every 30 minutes throughout the experiment duration. Growth curves were calculated with the RTCA software version 1.2.1. A minimum of three independent experiments were performed for each cell line.
Migration assay
The ability of cancer cells to migrate was assessed using ibidiTM culture inserts (ibidi GmbH, Planegg, Germany). The inserts consist of two 0.22 cm2 silicone chambers separated by a 0.5 mm divider. The inserts were positioned into a 12-well tissue culture dish, one insert per well. Roughly 70 µl pre-transfected cell-suspension containing 4–6 × 105 cells/ml were added to each chamber. The cells were left to adhere for 24 hours before the insert was removed and images acquired across the cell-free zone at time points 0 hours and 20 hours. The migration potential into the 0.5 mm gap was calculated using the free online software TScratch, version 1.0 (CSElab, Computational Science and Engineering Laboratory, Switzerland). Initially, the functional experiments for this study were designed using three cell lines; the large cell carcinoma cell line H460, the squamous cell carcinoma cell line H520, and the adenocarcinoma cell line A549. In our experiments, however, the cell lines H460 and H520 did not exhibit migrational properties, leaving only the A549 cell line representing the migration experiment. To strengthen our results, we therefor included the large cell carcinoma cell line H661 in the migration study.
Statistical methods
The statistical package IBM SPSS (version 24 IBM Corp., Armonk, NY USA) was used to perform all statistical analyses.
Interobserver reliability between scorers was assessed by a two-way random effects model with absolute agreement definition. Associations between marker expression, and marker expression and clinicopathological parameters, were examined by Spearman’s rank correlation and χ2 test or Fisher’s exact. Wilcoxon non-parametrical test was used to assess the difference in biomarker expression between lung tumor tissue and non-malignant lung tissue. Statistical significance between proliferation curves was assessed by one-way ANOVA. The Kaplan-Meier method was used to visualize association between marker expression and disease-specific survival (DSS) and the statistical significance between survival curves was tested using the log-rank test. DSS was defined as the time from surgery to lung cancer death. Variables with significant p-values from the univariate analyses were entered into Cox proportional Hazard models. The final models were derived from a backward conditional method with probability for stepwise entry and removal at 0,05 and 0,10.
Ethics
The Regional Committee for Medical and Health Research Ethics (REK Nord), alongside the Norwegian Data Protection have approved this study (protocol ID: 2011/2503). Due to the retrospective study design, the majority of patients were diseased and the tissue specimens over 10 years old. Thus, written patient consent was not deemed necessary by REK Nord. All patients were anonymously included in the database. A trial number for each patient was used when pairing clinical information with the respective patients. Clinical information was reported according to the REMARK guidelines16. The authors confirm that all experiments were performed in accordance with relevant guidelines and regulations. The database buildup was approved by The Data Protection Official for Research (NSD).
Results
Patient characteristics
Clinical, histopathological and demographic variables and their impact on DSS are presented in Table 1. The median age was 67 years (range, 28–85), 373 patients (68%) were male, and the majority, 532 patients (96%), were current or previous smokers. The median follow-up time of survivors was 86 months (range, 34–267). Postoperative radiotherapy was administered to 76 (14%) patients due to non-radical surgical margins or nodal metastasis. Adjuvant chemotherapy was introduced in Norway in 2005, 43 (8%) patients received this treatment.
Scoring agreement
Scoring agreement between the scorers (SAS and KS) was excellent; ICC were 0.80 (p < 0.001) and 0.97 (p < 0.001) for miR-143 and miR-145, respectively.
miR-143 and miR-145 expression in NSCLC cells
ISH expression of miR-143 and miR-145 in NSCLC cells and metastatic lymph nodes
miR-143 was primarily observed in the cytoplasm of tumor epithelial and stromal cells, while miR-145 was mainly observed in the epithelial and stromal cell nuclei (Fig. 1). Table 2 reports miR-143 and miR-145 expression according to tissue compartment and gender. Neoplastic epithelial and stromal cells had significantly increased levels of miR-143 and miR-145 compared to non-malignant lung tissue (T-miR-143: p < 0.001, S-miR-143: p < 0.001, T-miR-145: p = 0.005, S-miR-145: p = 0.020). T-miR-143 expression in PT and LN+ was significantly correlated (0.220, p < 0.001). There was a significant correlation between miR-145 expression in neoplastic epithelial cells and stromal cells (0,362, p < 0.001). Similarly, miR-145 expression in PT and LN+ was significantly correlated (0,366, p < 0.001)
In situ hybridization staining of miR-143 and miR-145 in NSCLC. High miR-143 expression: Panel (A) stromal cells, Panel (C) tumor cells. Low miR-143 expression: Panel (B) stromal cells, Panel (D) cancer cells. High miR-145 expression: Panel (E) stromal cells, Panel (G) cancer cells. Low miR-145 expression: Panel (F) stromal cells, Panel (H) cancer cells. 400× magnification.
Relative expression of miR-143 and miR-145 in NSCLC cell lines
Endogenous levels of miR-143 and miR-145 in the studied NSCLC cell lines were quantified by qPCR, relative to the non-cancerous lung cell line NL20. Both miR-143 and miR-145 were downregulated in all the selected cell lines, compared to NL20 (Supplementary Fig. 1).
Functional studies on miR-143 and miR-145 in vitro
To investigate the potential function of miR-143 and miR-145 in NSCLC tumorigenesis, we performed a series of in vitro experiments. By transfecting various NSCLC cell lines with miR-143 mimic, miR-145 mimic and miR-143+miR145 mimic, we observed the biomarkers effect on cell migration and proliferation.
miR-143 and miR-145 inhibit NSCLC migration
Transfection with miR-143 and miR-145 inhibited migration in both the A549 and H661 cell line when compared with cells transfected with the negative control miRNA (Fig. 2). The inhibition was strongest for miR-145 in both cell lines.
Inhibition of proliferation by miR-143 and miR-145
Both miR-143 and miR-145 inhibited proliferation in the cell lines H460 and A549, and the inhibition was more evident for cells transfected with miR-145 (Fig. 3A,B). Transfection of miR-143 promoted proliferation in the H520 cell line, whereas miR-145 had an inhibitory effect on proliferation in the same cell line (Fig. 3C). In the cell lines A549 and H460, the inhibitory effects of co-transfection with miR-143 and miR-145 in equal concentrations, were equivalent to that of the miR-145 transfection alone. When co-transfecting the H520 cell line with equal concentrations of miR-143 and miR-145, the inhibitory effects displayed by transfecting miR-145 alone were reduced to a degree where the proliferation-rate was not significantly different to the negative control. Simultaneously, the increase in proliferation caused by the miR-143 transfection alone, was greatly reduced when the H520 cell line was co-transfected with both miR-143 and miR-145 in equal concentrations.
Functional studies on NSCLC cell lines: Proliferation. Panels show proliferation rate in cell lines H460, A549 and H520. Cell Index represents proliferation as a function of time. Panel A-H460: Both miR-143 and miR-145, alone or in combination, inhibits proliferation in H460 cells. The inhibitory effect is strongest in cells receiving the miR-145. Panel B-A549: Same inhibition pattern as panel (A). Panel C-H520: Introducing miR-143 promotes proliferation in this cell line, while transfection with miR-145 inhibits proliferation. *** indicates p < 0.001.
Correlation with clinical variables and other molecular markers
There were no significant associations between miR-143 and miR-145 expression in PT or LN+ and clinicopathological prognosticators listed in Table 1.
Between marker correlations with likely biological significance were as follows: LN+T-miR-143 was positively correlated with PT stromal AR expression (r = 0.494: p < 0.001), and inversely correlated with PT tumor epithelial PGR expression (−r = 0.453: p < 0.001). T-miR-143 in PT was correlated with cytoplasmic ERβ in PT (r = 0.215: p < 0.001) and T-miR-145 in PT was correlated with nuclear ERβ expression in tumor cells (r = 0.212: p < 0.001). Other significant correlations were also observed (Supplementary Table 1).
Univariate survival analyses
Clinicopathological variables and their impact on DSS are presented in Table 1. The impacts of biomarkers on DSS in PT are presented in Table 2 and Fig. 4. Neither epithelial nor stromal expression of miR-143 or miR-145 showed significant impact on DSS in the overall cohort. Following gender stratification, however, high S-miR-143 was a positive prognosticator in female patients (p = 0.011), while high S-miR-145 was a positive prognosticator in male patients (p = 0.013). Further, the combination of low S-miR-143 and low S-miR-145 was associated with an unfavorable prognosis in the overall cohort (p = 0.007, Fig. 5). In LN+, neither miR-143 nor miR-145 showed impact on DSS in the overall cohort or stratified by gender.
Survival curves. Kaplan-Meier curves showing disease-specific survival (DSS) in relation to stromal miR-143 and miR-145 expression in primary tumor. By dichotomizing biomarker expression level into high vs low, we present an association between improved DSS and high stromal biomarker expression. Overall cohort (OC): panel (A) (miR-143) and panel (D) (miR-145), female patients: panel (B) (miR-143) and panel (E) (miR-145), male patients: panel (C) (miR-143) and panel (F) (miR-145).
Multivariate analysis
Significant clinicopathological and biomarker variables from univariate analyses were entered into the multivariate analyses. Results are presented in Table 3. In primary tumors (PT), high S-miR-143 (HR: 0.53, 95% CI: 0.31–0.90, p = 0.019) and high S-miR-145 (HR: 0.58, 95% CI: 0.37–0.92, p = 0.021) were independent, positive prognosticators in female and male patients, respectively. The combination low S-miR-143/low S-miR-145 (overall cohort: HR: 0.57, 95% CI: 0.35–0.94, p = 0.027) was independently associated with an unfavorable DSS.
Discussion
In this large retrospective study of 553 NSCLC patients, S-miR-143 and S-miR-145 expression in PT were positive prognosticators in female and male patients, respectively. Further, the combination of low stromal expression of both miR-143 and miR-145 predicted poor DSS in the overall cohort. Cell line studies confirm the tumor suppressive role of miR-143 and miR-145 in NSCLC, further substantiating their importance in lung cancer pathogenesis. We also observe significant correlations with our previously investigated steroid hormone receptors, suggesting that a biologic rationale may cause, or contribute to, the gender related survival impact observed.
To our knowledge, this is the first study investigating the prognostic impact of miR-143 and miR-145 in neoplastic epithelial cells, tumor associated stromal cells and matched metastatic lymph nodes in the same NSCLC cohort.
Associations between miR cluster 143/145 and cancer survival have been reported for different malignancies, results are, however, conflicting. In prostate cancer (PCa), Avgeris et al.17 demonstrated a shorter disease-free survival in PCa patients with low miR-145 expression levels. Campayo et al.18 reported similar results for miR-145 in NSCLC patients. These reports confirm our suggestion of high miR-145 expression as a positive prognosticator, herein in NSCLC patients. Contradicting our results, Al Feber et al.19 and Avgeris et al.20, both reported associations between high levels of miR-143 and miR-145 and poor overall survival in esophageal and bladder cancer, respectively. Importantly, none of the aforementioned studies have evaluated survival impact according to cellular compartment, as was performed in our study.
Our findings suggest miR-143 and miR-145 to play protective roles when expressed in stromal cells. This is in concordance with the biomarkers being regarded as tumor suppressors21. miR-143 and miR-145 target several important genes involved in tumorigenesis including KRAS22 and ERα11. However, their biological function in NSCLC remains largely unknown. Functional studies have proposed different roles; Chen et al., 2010 demonstrated that miR-145 inhibited NSCLC proliferation by targeting the transcription factor regulatory gene c-Myc13, thus confirming the tumor suppressive qualities of this particular miR. In a recent report Zhang et al.12 suggest epidermal growth factor receptor (EGFR) as a downstream target of miR-143, contributing to tumor suppression. Consistent with our findings, Zhang et al.12 demonstrated an inhibition of migration and proliferation of NSCLC cells following transfection with miR-143. Surprisingly, when we transfected the squamous cell carcinoma cell line H520 with miR-143, proliferation was dramatically increased (Fig. 3C). This is in contrast to most studies reporting on the effects of miR-143 on proliferation23,24,25,26,27. However, there are studies depicting alternative roles for the miR cluster 143/14528, and members of our research group have reported similar findings using breast cancer cell lines29. Interestingly, when co-transfecting miR-143 and miR-145 in equal concentrations, the proliferative capacity was markedly reduced (Fig. 3A,B), meaning the net effect of co-transfecting is tumor suppressive.
The use of verified laboratory techniques with meticulously prepared protocols for biomarker handling is a strength with regards to reliability and reproducibility of our results. Our patient cohort is large with an extensive follow-up time, which further substantiate our results. The retrospective study design may represent a weakness with inaccurate clinical patient data.
In line with previous reports, we detect downregulation of both miR-143 and miR-145 in four independent cancer cell lines relative to levels in a non-cancerous cell line (Supplementary Fig. 1)30,31. Thus, extending the general sense of miR-143/miR-145 downregulation in cancer, including NSCLC21. Interestingly, this is in contrast with our ISH-results, reporting significantly increased expression of miR-143/miR-145 in tumor cells and adjoining stromal cells in comparison to non-malignant tissue. To our knowledge, this is the first large-scale miRNA in situ NSCLC tissue hybridization analysis reporting an upregulated miR-143/miR-145 expression. These findings are conflicting with the smaller study (n = 48) by Shen et al.31, reporting a downregulation of miR-145 expression in NSCLC, by the use of ISH technique. We present a thorough and comprehensive study of miR-143 and miR-145 expression in appropriate cell types by the use of several acknowledged techniques, giving an optimal account of miR-expression in the tumor environment. Due to contributions from stromal cells in tumor growth, it is pivotal to consider the stromal compartment when elucidating biological mechanisms in epithelial cancers32. We found that miR-143 was primarily observed in the cell cytoplasm, while miR-145 was mainly observed in the nuclei of epithelial and stromal cells. Further, we report an abundant positivity of miR-143 and miR-145 in fibroblasts and SMCs lining the blood and lymph vessels, consistent with previous reports9,33. miRs are traditionally considered to act within the cell cytoplasm, regulating gene expression post-transcriptionally34. However, a number of miRNAs have been localized in the nuclei, although their nuclear functions remain elusive35.
In an extensive meta-analysis, Kent et al.9 highlighted the crucial importance of cell-type localization of miRNAs, and how a lack of consideration of specific cellular expression of miRNAs may lead to a general misconception that miRNAs are downregulated in neoplastic tissue. Chivukula et al.36, Dimitrova et al.28 and Akao et al.37 all published results indicating that neither miR-143 nor miR-145 are expressed in tumor epithelial cells, causing the latter group to conclude that these miRNAs are downregulated in malignant tissue. Similar results have been published by other groups investigating a variety of malignancies21,38,39. However, only one28 of the previous studies has focused on the histological cell-type localization of miR-143/miR-145. These factors, combined with the lung cancer cell lines lack of stroma, inflammatory cells and vascularization, may contribute to the discrepancy observed between miR-143/miR-145 expression in cell lines and tissue samples.
In this study, we present interesting correlations between miR-143/miR-145 and steroid hormone receptors expressed in the NSCLC tissue. The finding of gender specific survival significance of miR-143 and miR-145, forces us to consider sex hormones as a relevant factor. Delfino et al.40 reported a gender-specific miRNA targeting of molecules related to glioblastoma survival. Further, Duttagupta et al.41 reported differential miRNA expression levels in a gender specific manner. Mounting evidence confirms activation of hormone receptors to be of outmost importance in lung cancer pathogenesis and several interesting cross-talk pathways between steroid hormones and miR-143/miR-145 have been found42,43,44,45,46. Both miRNAs play a critical role in ovarian functioning, and a recent report presents miR-143 affecting estradiol production in granulosa cells by targeting KRAS47,48. Further, Spizzo et al., 2011 reported that miR-145 downregulates ERα expression in breast cancer11. We found correlations between miR-143/miR-145 and AR, the rate limiting enzyme in estradiol production, suggesting the miRs may interact with regional estradiol production and ER signaling in the lung, as observed in breast tissue. In 2012, Paris et al. presented a study on estrogen effects in breast cancer, showing a direct regulation of miRNA expression and ERβ signaling49. Herein, ERβ expression correlated with miR-143/miR-145 expression, suggesting a similar link may exist in NSCLC. The aforementioned reports, assembled with our findings, provide a compelling rationale for a biological cross-talk between miR-143/miR-145 and hormone receptors. If validated in larger, confirmatory studies, this may in fact represent new possibilities for targeted therapy for NSCLC patients, using gender, miRNA and hormone receptor expression as therapy selection criteria.
Conclusion
We present high stromal expressions of miR-143 and miR-145 as positive prognosticators in a gender specific manner in early stage NSCLC patients. Our findings indicate that miR-143/miR-145 acts as tumor suppressor molecules in lung cancer, suggesting that these miRNAs may be useful in miRNA based therapy in NSCLC. Further, we highlight the complexity of miR expression, and stress the importance of cell-type specific expression profiling. By accentuating the correlation between miRNA expression and hormone receptor expression, we emphasize the importance of exploring multi-targeted therapies in the treatment of NSCLC patients, as anti-hormonal therapy is highly accessible.
References
Ferlay, J. et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. International journal of cancer 136 (2015).
Travis, W. D., Brambilla, E., Burke, A., Marx, A. & Nicholson, A. G. WHO classification of tumours of the lung, pleura, thymus and heart. (International Agency for Research on Cancer, 2015).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2016. CA: a cancer journal for clinicians 66, 7–30 (2016).
Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004).
Doench, J. G. & Sharp, P. A. Specificity of microRNA target selection in translational repression. Genes & development 18, 504–511 (2004).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. 6, 857, https://doi.org/10.1038/nrc1997 (2006).
Wiggins, J. F. et al. Development of a lung cancer therapeutic based on the tumor suppressor microRNA-34. Cancer research 70, 5923–5930 (2010).
Reid, G. et al. Clinical development of TargomiRs, a miRNA mimic-based treatment for patients with recurrent thoracic cancer. Epigenomics 8, 1079–1085 (2016).
Kent, O. A., McCall, M. N., Cornish, T. C. & Halushka, M. K. Lessons from miR-143/145: the importance of cell-type localization of miRNAs. Nucleic acids research 42, 7528–7538 (2014).
Zhang, J. et al. Loss of microRNA-143/145 disturbs cellular growth and apoptosis of human epithelial cancers by impairing the MDM2-p53 feedback loop. Oncogene 32, 61 (2013).
Spizzo, R. et al. miR-145 participates with TP53 in a death-promoting regulatory loop and targets estrogen receptor-α in human breast cancer cells. Cell death and differentiation 17, 246 (2010).
Zhang, H. B., Sun, L. C., Ling, L., Cong, L. H. & Lian, R. miR-143 suppresses the proliferation of NSCLC cells by inhibiting the epidermal growth factor receptor. Experimental and Therapeutic Medicine 12, 1795–1802 (2016).
Chen, Z. et al. miRNA-145 inhibits non-small cell lung cancer cell proliferation by targeting c-Myc. Journal of Experimental & Clinical Cancer Research 29, 1 (2010).
Goldstraw, P. et al. The IASLC Lung Cancer Staging Project: proposals for revision of the TNM stage groupings in the forthcoming (eighth) edition of the TNM classification for lung cancer. Journal of Thoracic Oncology 11, 39–51 (2016).
Bremnes, R. et al. High-throughput tissue microarray analysis used to evaluate biology and prognostic significance of the E-cadherin pathway in non–small-cell lung cancer. Journal of Clinical Oncology 20, 2417–2428 (2002).
McShane, L. M. et al. REporting recommendations for tumour MARKer prognostic studies (REMARK). British journal of cancer 93, 387–391 (2005).
Avgeris, M., Stravodimos, K., Fragoulis, E. & Scorilas, A. The loss of the tumour-suppressor miR-145 results in the shorter disease-free survival of prostate cancer patients. British journal of cancer 108, 2573–2581 (2013).
Campayo, M. et al. Low miR-145 and high miR-367 are associated with unfavourable prognosis in resected nonsmall cell lung cancer. European Respiratory Journal 41, 1172–1178 (2013).
Feber, A. et al. MicroRNA prognostic signature for nodal metastases and survival in esophageal adenocarcinoma. The Annals of thoracic surgery 91, 1523–1530 (2011).
Avgeris, M. et al. Uncovering the clinical utility of miR-143, miR-145 and miR-224 for predicting the survival of bladder cancer patients following treatment. Carcinogenesis 36, 528–537 (2015).
Das, A. V. & Pillai, R. M. Implications of miR cluster 143/145 as universal anti-oncomiRs and their dysregulation during tumorigenesis. Cancer cell international 15, 92 (2015).
Kent, O. A. et al. Repression of the miR-143/145 cluster by oncogenic Ras initiates a tumor-promoting feed-forward pathway. Genes & development 24, 2754–2759 (2010).
Wang, Q., Cai, J., Wang, J., Xiong, C. & Zhao, J. MiR-143 inhibits EGFR-signaling-dependent osteosarcoma invasion. Tumour Biol 35, 12743–12748, https://doi.org/10.1007/s13277-014-2600-y (2014).
Xia, H. et al. miR-143 inhibits NSCLC cell growth and metastasis by targeting Limk1. Int J Mol Sci 15, 11973–11983, https://doi.org/10.3390/ijms150711973 (2014).
Liu, J. et al. MiR-143 inhibits tumor cell proliferation and invasion by targeting STAT3 in esophageal squamous cell carcinoma. Cancer letters 373, 97–108, https://doi.org/10.1016/j.canlet.2016.01.023 (2016).
Xu, Y. F. et al. Identification of miR-143 as a tumour suppressor in nasopharyngeal carcinoma based on microRNA expression profiling. Int J Biochem Cell Biol 61, 120–128, https://doi.org/10.1016/j.biocel.2015.02.006 (2015).
Zhang, W., Wang, Q., Yu, M., Wu, N. & Wang, H. MicroRNA-145 function as a cell growth repressor by directly targeting c-Myc in human ovarian cancer. Technology in cancer research & treatment 13, 161–168, https://doi.org/10.7785/tcrt.2012.500367 (2014).
Dimitrova, N. et al. Stromal Expression of miR-143/145 Promotes Neoangiogenesis in Lung Cancer Development. Cancer Discov 6, 188–201, https://doi.org/10.1158/2159-8290.CD-15-0854 (2016).
Johannessen, C. et al. Expression and function of the miR-143/145 cluster in vitro and in vivo in human breast cancer. PLoS One 12, e0186658, https://doi.org/10.1371/journal.pone.0186658 (2017).
Takagi, T. et al. Decreased expression of microRNA-143 and-145 in human gastric cancers. Oncology 77, 12–21 (2009).
Shen, H. et al. Low miR-145 expression level is associated with poor pathological differentiation and poor prognosis in non-small cell lung cancer. Biomedicine & Pharmacotherapy 69, 301–305 (2015).
Almeida, M. I. & Calin, G. A. The miR-143/miR-145 cluster and the tumor microenvironment: unexpected roles. Genome medicine 8, 29 (2016).
Cordes, K. R. et al. miR-145 and miR-143 regulate smooth muscle cell fate decisions. Nature 460, 705 (2009).
Roberts, T. C. The MicroRNA Biology of the Mammalian Nucleus. Molecular therapy. Nucleic acids 3, e188, https://doi.org/10.1038/mtna.2014.40 (2014).
Liao, J. Y. et al. Deep sequencing of human nuclear and cytoplasmic small RNAs reveals an unexpectedly complex subcellular distribution of miRNAs and tRNA 3′ trailers. PloS one 5, e10563, https://doi.org/10.1371/journal.pone.0010563 (2010).
Chivukula, R. R. et al. An essential mesenchymal function for miR-143/145 in intestinal epithelial regeneration. Cell 157, 1104–1116 (2014).
Akao, Y., Nakagawa, Y. & Naoe, T. MicroRNAs 143 and 145 are possible common onco-microRNAs in human cancers. Oncology reports 16, 845–850 (2006).
Lui, W.-O., Pourmand, N., Patterson, B. K. & Fire, A. Patterns of known and novel small RNAs in human cervical cancer. Cancer research 67, 6031–6043 (2007).
Szczyrba, J. et al. The microRNA profile of prostate carcinoma obtained by deep sequencing. Molecular cancer research 8, 529–538 (2010).
Delfino, K., Serão, N., Southey, B. & Rodriguez-Zas, S. Therapy-, gender-and race-specific microRNA markers, target genes and networks related to glioblastoma recurrence and survival. Cancer Genomics-Proteomics 8, 173–183 (2011).
Duttagupta, R., Jiang, R., Gollub, J., Getts, R. C. & Jones, K. W. Impact of cellular miRNAs on circulating miRNA biomarker signatures. PloS one 6, e20769 (2011).
Weinberg, O. K. et al. Aromatase inhibitors in human lung cancer therapy. Cancer research 65, 11287–11291 (2005).
Mah, V. et al. Aromatase expression predicts survival in women with early-stage non–small cell lung cancer. Cancer research 67, 10484–10490 (2007).
Márquez-Garbán, D. C., Chen, H.-W., Fishbein, M. C., Goodglick, L. & Pietras, R. J. Estrogen receptor signaling pathways in human non-small cell lung cancer. Steroids 72, 135–143 (2007).
Skjefstad, K. et al. Prognostic relevance of estrogen receptor α, β and aromatase expression in non-small cell lung cancer. Steroids 113, 5–13, https://doi.org/10.1016/j.steroids.2016.05.008 (2016).
Skjefstad, K. et al. The prognostic role of progesterone receptor expression in non-small cell lung cancer patients: Gender-related impacts and correlation with disease-specific survival. Steroids 98, 29–36 (2015).
Zhang, L. et al. MiRNA-143 mediates the proliferative signaling pathway of FSH and regulates estradiol production. Journal of Endocrinology 234, 1–14 (2017).
Hossain, M., Sohel, M., Schellander, K. & Tesfaye, D. Characterization and importance of microRNAs in mammalian gonadal functions. Cell and tissue research 349, 679–690 (2012).
Paris, O. et al. Direct regulation of microRNA biogenesis and expression by estrogen receptor beta in hormone-responsive breast cancer. Oncogene 31, 4196 (2012).
Acknowledgements
This study was funded by the Northern Norway Health Region Authority (Helse Nord RHF) and The Norwegian Cancer Society. The authors would like to thank them for supporting their work. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. The publication charges for this article have been funded by a grant from the publication fund of UiT The Arctic University of Norway.
Author information
Authors and Affiliations
Contributions
Everyone included in the author list have been involved in the development and design of this study and manuscript. K.S. prepared the first manuscript draft, revisions and finalization was a collaboration between all co-authors. Statistical analysis and interpretation of results was performed by K.S. and C.J., with special contributions from T.G., S.A.S., and L.T.B. E.E.P., T.D., S.A.S., T.K., M.P. and K.S. have contributed to acquisition of data, enabling conduction of this study. C.J. has prepared Figures 2 and 3, T.K. has prepared Figures 4 and 5.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Skjefstad, K., Johannessen, C., Grindstad, T. et al. A gender specific improved survival related to stromal miR-143 and miR-145 expression in non-small cell lung cancer. Sci Rep 8, 8549 (2018). https://doi.org/10.1038/s41598-018-26864-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26864-w
This article is cited by
-
Overexpression of ST7-AS1 Enhances Apoptosis and Inhibits Proliferation of Papillary Thyroid Carcinoma Cells Via microRNA-181b-5p-Dependent Inhibition Tripartite Motif Containing 3
Molecular Biotechnology (2023)
-
LncRNA ST7-AS1, by regulating miR-181b-5p/KPNA4 axis, promotes the malignancy of lung adenocarcinoma
Cancer Cell International (2020)
-
RETRACTED ARTICLE: N6-methyladenosine induced miR-143-3p promotes the brain metastasis of lung cancer via regulation of VASH1
Molecular Cancer (2019)
-
Low Expression of miR-424-3p is Highly Correlated with Clinical Failure in Prostate Cancer
Scientific Reports (2019)
-
Therapeutic microRNAs in human cancer
Cytotechnology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.