Abstract
Whole genome duplication is now accepted as an important evolutionary force, but the genetic factors and the life history implications affecting the existence and abundance of polyploid lineages within species are still poorly known. Polyploidy has been mainly studied in plant model species in which the sporophyte is the dominant phase in their life history. In this study, we address such questions in a novel system (Porphyra, red algae) where the gametophyte is the dominant phase in the life history. Three Porphyra species (P. dioica, P. umbilicalis, and P. linearis) were used in comparisons of ploidy levels, genome sizes and genetic differentiation using flow cytometry and 11 microsatellite markers among putative polyploid lineages. Multiple ploidy levels and genome sizes were found in Porphyra species, representing different cell lines and comprising several cytotype combinations among the same and different individuals. In P. linearis, genetic differentiation was found among three polyploid lineages: triploid, tetraploid and mixoploids, representing different evolutionary units. We conclude that the gametophytic phase (n) in Porphyra species is not haploid, contradicting previous theories. New hypotheses for the life histories of Porphyra species are discussed.
Similar content being viewed by others
Introduction
Polyploidy, the increase in genome size by the acquisition of more than one set of chromosomes has been a key factor in eukaryote evolution. In fact, most flowering plants and vertebrates descend from polyploid ancestors1. In angiosperms, many species have been suggested to have polyploid ancestry2. While whole genome duplication (WGD) is now accepted as an important evolutionary force, the genetic factors and the life history implications affecting the existence and abundance of polyploid lineages are still poorly understood3,4,5. Recent studies have also demonstrated that polyploid genomes can be highly dynamic and undergo rapid structural and functional changes. These findings have renewed the interest in examining WGD as an evolutionary process6,7, which coupled with molecular genetics and phylogenetic analyses can provide new insights into the evolution of the genome8.
In polyploid research, most of the systems studied are higher plants (e.g. Brassica, Cotton, Maize, Spartina, Wheat, Tragopogon) representing life cycles in which the sporophyte is the dominant phase in a complex cycle. This cycle alternates between sporophytes (spore-producing organisms) and gametophytes (gamete-producing organisms)9. Although the sporophyte generation dominates in most extant lineages of land plants, in the earliest groups of land plants and also in several macroalgal groups, the gametophyte generation was (and it still is) the dominant component of the life cycle. Up to date, little is known about polyploidy in system groups that have the gametophyte as the dominant phase. This study uses red algae, an Archaeplastida group on the evolutionary line leading to plants, to study polyploid processes in a “gametophyte dominant” system. By studying polyploidy in such a life cycle, it may be possible to gain insights into polyploid evolution and to address questions related to the origins of polyploid lineages/individuals, the genetic consequences of genome duplication, and/or the importance of the triploid bridge in the origin of tetraploid populations, a topic still under debate10,11,12.
Porphyra is a polyphyletic genus of red algae with approximately 57 recognized species13 with similar morphology and with a wide variety of life history strategies14. A recent taxonomic revision of the Bangiales created new genera, and many species previously classified as Porphyra species have been transferred to other genera within the Bangiales, such as Pyropia and Wildemania among others15. Although this and other molecular studies (e.g. refs16,17) have revealed more species diversity in Porphyra than was previously understood, little is known about the mechanisms of speciation in the genus Porphyra. Three of the recognized Porphyra species are the subject of this study: Porphyra dioica J Brodie & L. M. Irvine, Porphyra linearis Greville, and Porphyra umbilicalis Kützing. It is worth to note that although the gametophyte is the dominant phase in their life history, each species has a different mating system in Southern Europe: Porphyra dioica has mostly individuals with separate sexes (dioecious), P. umbilicalis can be hermaphroditic with both sexes in the same individual and gametangia separated in different sections of the thallus (monoecious) or also dioecious, and P. linearis has individuals with both systems, dioecious and hermaphroditic, but probably as a consequence of a protandrous system (or sequential hermaphroditism, with males being formed first, later becoming females).
Porphyra species typically have a heteromorphic life cycle consisting of a gametophytic blade phase (haploid) and a filamentous sporophytic phase called the conchocelis (diploid). The conchocelis phase of Porphyra species is difficult to find in nature18 with only a few in situ records (e.g. refs19,20), in contrast with the gametophytic blades, which are common and abundant in nature. The first description of the Porphyra life cycle by21 revolutionized the farming of this edible seaweed. Drew proved that Conchocelis rosea Batters, originally described as a separate species, was the germinated carpospores (sporophyte, phase 2n) from the Porphyra blades (gametophyte, phase n). It has been assumed since then than two ploidy levels are involved in the Porphyra life cycles and several chromosome studies have corroborated the ploidy level of both phases, with the gametophytic blade being haploid and the sporophytic carpospores/conchocelis being diploid (e.g. refs 22,23,24,25). Drew’s study enabled mass cultivation and the development of the aquaculture industry of Porphyra (Nori, laverbread), the single most valuable marine product in the orient with a current retail value over $1.3 billion per year26.
Recently, within the context of the project NORIGENOMICS (PTDC/MAR/099698/2008), microsatellite markers have been developed for the three Porphyra species that are the subject of this study, to describe the population structure and historical biogeography of these species in the North Atlantic. These markers have allowed us to detect gametophytic blades of Porphyra possessing heterozygous genotypes with 1–6 alleles per individual in many samples along European coasts (Varela-Álvarez et al., unpublished). These findings led us to question if different ploidy levels exist on different individuals/strains and or populations and whether these may be the result of hybridization with other Porphyra species/lineages. However, if polyploids exist in our populations/species, how these results can be integrated into the haploid/diploid life history assumed by21 is an enigma to be resolved. The origins of polyploid lineages, strains, or individuals within the species-complex, have yet to be determined.
The main objective of this study was to assess the existence of putative polyploid lineages in species of Porphyra with different mating systems and, if present, to get insights into the mode of origin of polyploids (i.e., allopolyploidy and/or autopolyploidy). A second objective was to use the evidence from ploidy levels together with the estimation of genetic differentiation among the ploidy types found to understand if the species/cytotypes maintain gene flow. Finally, we expected that these results would provide insight into the evolutionary consequences of ploidy variation for the life history of species of Porphyra representing different mating systems; and to contrast hypothetical models for the evolution of polyploidy in organisms where the gametophyte is the dominant phase, relative to those where it is the sporophyte. Besides improving the understanding of the evolutionary dynamics in polyploid complexes in non-specialized plant species, our study provides evidence of a new evolutionary mechanism in the economically important genus Porphyra and that might perhaps be common in other red algae.
Material and Methods
Sampling collection
A total of 122 blades (P. dioica, 40 blades; P. umbilicalis, 35 blades; and P. linearis, 47 blades) were collected in order to assess their ploidy levels, genome sizes and genetic differentiation among putative polyploid lineages. The localities visited were: Buarcos, Figueira da Foz (40°10′N, 8°53′W) (for P. dioica and P. umbilicalis) and Belém, Lisboa (38°41′N, 9°12′W) (for P. linearis), both in Portugal. Each individual was examined in the binocular microscope and up to 4 sections were made per blade in duplicate (one replicate for genetic analyses and a second replicate for flow cytometric analyses) for further analyses: vegetative blade, zygotospores, female gametes and spermatia (male gametes). The terms zygotosporangia and zygotospores are used to replace carposporangia and carpospores as proposed by Guiry27 for the Bangiales.
Flow cytometry
The nuclear DNA content of Porphyra spp. was estimated by flow cytometry using fresh material. Nuclei were released following the procedure of Loureiro et al.28 by chopping approximately 1 cm2 of each blade section with a razor blade together with 50 mg of fresh leaf tissue Solanum lycopersicum (internal reference standard with 2 C = 1.96 pg29) in 1 mL of WPB buffer28 to ensure that nuclei of both species were exposed to identical chemical and mechanical conditions. To double check for the existence of smaller genome sizes we also applied to a small set of samples a standard of smaller genome size, Raphanus sativus (2 C = 1.11 pg)29. The nuclear suspension was then filtered through a 30 μm nylon filter to remove large fragments, and 50 μg/mL of PI - Propidium iodide (Fluka, Buchs, Switzerland) together with 50 μg/mL of RNase (Fluka, Buchs, Switzerland) were subsequently added to respectively stain the nuclei and prevent staining of double-stranded RNA. After incubation for 5 min, the fluorescence intensity of an average of 2,000 nuclei per sample was analysed using a Partec CyFlow Space flow cytometer (Partec GmbH, Görlitz, Germany), equipped with a green solid state laser for PI excitation. The G1 peak of the standard was set to channel 720 and the amplification system settings were then kept at a constant voltage and gain throughout the experiment. For the data analyses, measurements were acquired using FloMax software v2.4d (Partec GmbH) in the form of six graphs: fluorescence pulse integral in linear scale (FL) (Graph 1); FL vs. time (Graph 2); side light scatter (SSC) vs. forward light scatter (FSC), both in logarithmic (log) scale (Graph 3); FL vs. fluorescence pulse height (Graph 4); FL vs. FSC (Graph 5) and FL vs. SSC (Graph 6). A region of interest comprising mostly the isolated nuclei was defined in the FL vs. SSC cytogram and subsequently used to gate all the other graphs. The presence of aggregates (doublets or triplets) was evaluated in the FL vs. fluorescence pulse height cytogram. The genome size (in picograms, pg) of each sample was determined according to the following formula: (mean PI fluorescence of the sample/mean PI fluorescence of the standard nuclei) * nuclear DNA content of the standard nuclei. The reliability of the genome size measurements was verified by evaluating the quality of the flow cytometry histograms based on the coefficient of variation (CV) of the G1 peaks and the background debris, and the CV of the genome size estimation of each isolate based on three independent measurements.
DNA extraction, PCR and genotyping
For genotyping, the same individuals (gametophytic blades) used previously for flow cytometry were scored and DNA extractions were obtained from up to 4 sections of each blade referred above.
Genomic DNA was isolated using the LiCl extraction protocol described by30 as modified by31. Genotyping using 11 selected microsatellite markers32,33 generated 121 multilocus genotypes (see Table S1 for sources, loci and amplification details). Polymerase chain reactions (PCR) for all markers were performed separately in a 20 μl reaction volume containing 5–50 ng genomic DNA. Amplifications using an Applied Biosystems thermal cycler (GeneAmp 2720) were conducted by the PCR programs and conditions described in32,33. Amplified fragments were separated electrophoretically using an ABI PRISM 3130xl (Applied Biosystems) automated capillary sequencer at CCMAR, Portugal, and sized with GeneScan-350ROX size standard.
Genetic analyses
Calculation of genetic distances for all pairwise comparisons and estimation of genetic differentiation were performed on the whole data set and in 19 groups according to the ploidy levels found (see results below). Ploidy levels in genotypes were determined by flow cytometry and also by the maximum number of alleles found in a genotype for comparison. Number of alleles, allele range, and unbiased total heterozygosity were determined per group containing similar genotypes. The indices of heterozygosity are used as a theoretical index of genetic diversity; they do not represent observed heterozygosity. For mixoploids (blades containing more than one ploidy level) and samples with different types of cells mixed (e.g. zygotospores mixed with vegetative cells), the highest ploidy level of the cell lines found in the sample was considered. For multiploid gametes (gametes of several ploidy levels produced in the same blade, see results), the basic ploidy level found in the samples was considered (3x or 4x). Alleles were scored in GENEMAPPER v.4.1 (Applied Biosystems). Binning and allele rounding performed by GENEMAPPER v.4.1 was also checked with TANDEM34. The calculation of genetic diversity among the samples was estimated using GENODIVE, which allows analysis of polyploid data35. Summary statistics of genetic diversity within populations were calculated, including N: Number of alleles (total number of alleles found), Eff_num: Effective number of alleles (the number of equally frequent alleles it would take to achieve a given level of gene diversity), Hs: Heterozygosity within populations (the expected frequency of heterozygotes within populations, assuming Hardy-Weinberg equilibrium), also known as Gene Diversity, and Ht: Total heterozygosity, the expected frequency of heterozygotes over all populations, assuming Hardy-Weinberg equilibrium. Both Hs and Ht have a correction for sampling bias stemming from sampling a limited number of individuals per population. Each parameter was calculated with and without the maximum likelihood-correction for allele dosage in polyploids implemented in GENODIVE35.
Genetic distances and genotypic differentiation
Genetic distance represents the amount of between-population difference in allele frequency summed over all loci. Genetic differentiation between individual loci was measured by pairwise genetic distances (Nei GST/JostD) for all pairs of groups between the three species and among groups within each species (with and without the maximum likelihood-correction for allele dosage for polyploids) and they were also calculated with GENODIVE35.
In order to illustrate the variation in genetic diversity and genotypic differentiation (distribution of genotypes among the populations), a principal component analysis (PCA) based on allelic variation across all genotyped individuals was performed using GENODIVE35. Genetic population structuring among the three species and among the ploidy levels found was evaluated with the software STRUCTURE36. Structure analysis applies a Bayesian clustering approach and it was used to identify the population structure inferred from microsatellites. Structure analyses were performed with all individuals combined into one dataset for a global analysis and also using one data set per species. Analyses were run without any a priori population assignments and admixture was allowed. We assumed a model where the number of clusters (K) was unknown and the population structure was inferred (lower and upper bound for K equal to 1 and 20, respectively). The modal value of ΔK distribution for the posterior probability of the data for a given K was used as an indicator of the strength of the signal detected by Structure and considered as the real number of K clusters37. Each K was replicated 10 times with 10,000 Markov Chain Monte Carlo (MCMC) iterations after a burn-in period of 50,000, without any prior information on the ploidy and/or species group of each sampled individual. The average membership coefficients for the 10 simulation runs of a given K value were generated by CLUMPP v1.1.238 and a graphical presentation of the average membership coefficients for each isolate was generated in Microsoft Excel. An estimate of the true number of populations, K, was calculated using an ad hoc statistic-based approach implemented in STRUCTURE HARVESTER v0.6.139, as described previously. To avoid bias caused by missing values in the PCA and Structure analyses, missing data were replaced with random values randomly picked from the allele pool of each population as in40. Finally, in order to evaluate the degree of association between genotypic variation and ploidy levels, a neighbour-joining (NJ) network was generated from a matrix of pairwise41 genetic distances of individuals (genotypes), using POPULATIONS 1.242.
Data availability
All the data (flow cytometry data set and genotypes) are available in supplementary information online (Appendices 1 and 2).
Results
Results from the cytometric approach
In total, 217 flow cytometric analyses were performed (P. linearis: 71, P. umbilicalis: 69, P. dioica: 77) using 122 blades (Appendix 1). Genome sizes in this study varied from 0.2 to 1.6 pg (Table 1 and Appendix 1). In each individual, 2 to 4 populations of sample nuclei were found (Fig. 1). In most, a minor subpopulation was evident at a fluorescence intensity corresponding to the G2-phase. The minimum ploidy level was set in the nuclei population with the lowest genome size, which was found in the gametes (DNA corresponding to the non-replicated haploid chromosome complement). Considering this reference, the remainder genome size values obtained were assigned a ploidy level. Evidence from genotyping (next section) also indicated that gametophytes and gametes presented 2 or more alleles per locus, so being ruled out as haploid; thus, we assigned the gametes with the smaller genome size (0.2 pg) the first ploidy level as 2x. Eight different genome size values were obtained across the Porphyra samples, whose ratios could only correspond to seven ploidy levels shown in Table 1. The ploidy levels 2x, 3x, 4x, 6x and 8x were the most common among the samples. Two cell lines were associated with these ploidy levels: cell line 1: 2x–4x–8x and cell line 2: 3x–6x. In some samples of male gametes, two further genome sizes were obtained, 0.9 pg and 1.2 pg, which correspond with ploidy levels 9x and 12x. Genome sizes of 1.6 pg/1C corresponded to the G2-phase of ploidy levels 8x (Table 1).
Histograms of relative fluorescence intensities in Porphyra. Examples of histograms of relative fluorescence intensities obtained through simultaneous flow cytometric analysis of propidium iodide-stained nuclei of Porphyra (peaks 1–4) and internal reference standard S. lycopersicum (peak at the right; with 2C = 1.96 pg). X-axis: Fluorescence (channel number). Y-axis: nuclei counts in each sample. (a) Triploid gametes in P. dioica. (b) Octoploid zygotospores from a tetraploid cytotype in P. umbilicalis. (c) Mixoploid cytotype 2x/4x in P. linearis. (d) Mixoploid cytotype 3x/4x in P. linearis. (e) Male gametes 3x and 4x. (f) Multiploid gametes 4x in P. dioica. Insets show cytograms of side scatters in logarithmic scale (sslog, axis x) vs. Propidium iodide fluorescence (PI, axis y) of nuclear suspensions of the same samples.
In total, 7 different cytotype combinations were found in these Porphyra species that can be classified in three groups (Table 2 and Fig. 2). Group 1 in which gametophytes of each polyploid race (triploid or tetraploid) produced gametes of the same ploidy: (a) Triploid blades 3x with 3x gametes, (b) Tetraploid blades 4x with 4x gametes; Group 2 in which gametophytes of each polyploid race (triploid or tetraploid) produced gametes of the same and/or different ploidy, (c) Triploid blades 3x with gametes 2x and/or 3x, (d) Tetraploid blades 4x with gametes of the same or different ploidy (2x, 3x, and/or 4x); and Group 3 in which gametophytes belong to mixoploid races in which cell lines 1 and 2 were present in the same individual and they produce gametes of different ploidies also: (e) Mixoploid blades (2x/3x), (f) Mixoploid blades (3x/4x) and (g) Mixoploid blades (2x/4x). In general, for mixoploid blades or “Mx blades” (containing more than one ploidy level), we indicated each ploidy level separated by a “/” (e.g., 2x/3x). When gametes with the same and lower ploidies were found in the same blade, we separated the different ploidy levels by a “,” (e.g., 2x, 3x). G1 and G2 on the top of each peak represent nuclei at G1-phase and G2-phase.
In addition, in Group 2, in the Tetraploid race, (Cytotype 4, Fig. 2 and Table 2) some blades produced gametes of a higher ploidy level than the gametophytic blade. We named these gametes as multiploid gametes or “Mp gametes”. We found Mp gametes 3x, (3x, 6x, 9x), and Mp gametes 4x (4x, 8x, 12x). The 2x ploidy level was only found in gametes and in vegetative parts of mixoploids; no single blade was detected as exclusively diploid (2x).
The diplophasic phase 2n was found among the samples in zygotospores for all the species and it was always 6x (in 3x blades) and 8x (in 4x blades). In mixoploids, the zygotospores were only the double of the highest ploidy level (e.g., in 2x/3x blades, the zygotospores were 6x). Furthermore, zygostospores (Zp) were found in either the marginal cells of the blade or in the vegetative part of the gametophytic blade for each polyploid race. When zygotospores and blades were sometimes found mixed with gametes (unfertilized); we indicated each of the ploidy levels of each cell separated by a “+” (e.g., 4x + 8x).
Although the overall average genome size did not differ among the three Porphyra species, the incidence of each cytotype combination differed among them. In roughly, P. linearis was the species with the highest number of mixoploids (40%) and blades produced gametes of any ploidy level (2x, 3x, 4x). Also, 17.5% of the blades were 4x with 4x gametes; 37.5% blades were 3x with 3x gametes, and a small proportion (5%) of triploids produced male gametes with lower ploidy. In P. umbilicalis, 40% of the individuals were 4x with 4x gametes, 17% were 3x with 3x gametes and 37% were either 4x or 3x that produced gametes with lower ploidy. Mixoploidy was also present, but in a much lower percentage (6%) than in P. linearis. For P. dioica: 42% were 4x with gametes 4x, 3% were triploid 3x with gametes 3x and 55% were fronds 4x with gametes of different ploidy levels.
Results from the molecular approach
In total, 121 multi-locus genotypes were found (Appendix 2). These where divided in 19 groups according to the species and ploidy level found in the flow cytometric analyses. The number of genotypes in each group varied from 1 to 23, representing “single ploidy levels” (3x, 4x, 6x, 8x), and “combined ploidy levels” (2x/3x; 2x/4x; 3x/4x; 4x + 8x; 3x + 8x, among other combinations) (Table 3). For all the species, we divided the data in as many groups as possible to decipher if there was any genetic differentiation among them; for example, 6x zygotospores coming from triploid blades or from mixoploid blades were separated in two distinct groups. When there was only one replicate for one type of ploidy level, we grouped the samples in a heterogeneous group (e.g. group 10 in P. umbilicalis) containing genotypes with different ploidy levels and different type of cells in the same sample.
Genetic diversity, both measured as mean number of alleles per locus and the standardized effective number of alleles, was similar for the three species (Table 4). When comparing groups, no genetic differences were found among individuals that produced gametes with lower or multiple ploidy levels and gametes of the same ploidy (data not shown) for any of the species. After applying the maximum likelihood correction for the unknown polyploid dosage, genetic variability statistics did not vary greatly relative to the full data set. Gene diversity (expressed by Hs = 0.5 or Ht = 0.5, approximately) was similar for the three species and for the polyploid lineages for each species. Genetic differentiation expressed as Nei GST and Jost D (Table 5) was found among some groups in P linearis between triploids vs. tetraploids (GST/Jost D: 0.27/0.25) and triploids vs. mixoploids 3x/4x (GST/Jost D: 0.17/0.15). The rest of the group comparisons presented values close to zero. In addition, no differentiation was found between any groups in the other two species (data not shown).
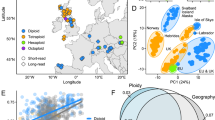
When performing a PCA (principal component analyses) with all the genotypes together, the three species were strongly differentiated (Fig. 3), as were some ploidy groups within one species (P. linearis), indicating reproductive isolation mediated by ploidy. Within P. linearis, genotypes with different ploidy levels (triploids vs. tetraploids vs. mixoploids) were genetically differentiated. Also, the genotypes of the 2n phase, zygotospores 6x and 8x, were closely related to cytotypes 3x and 4x respectively. Within P. umbilicalis and P. dioica, no genetic structuring by ploidy level could be identified in the PCA (Fig. 3), but some introgression between these species was suggested. The NJ network (Fig. 4) highlighted clearly the genetic differentiation by species, and within P. linearis and P. umbilicalis, also according with ploidy level (triploid vs. tetraploids vs. mixoploids or multiple ploidy levels). However, for P. dioica, there was not a clear structure among triploids and tetraploids. The STRUCTURE analyses separated well the three species but also showed genetic lineages within each species (Fig. 5) with best Ks (clusters/groups with similar/related genotypes) found for P. linearis K = 2 (two clusters), P. umbilicalis K = 3 (three clusters), and P. dioica K = 2 (two clusters) (Fig. 5a). With all the data set, the best K was K = 5 (ΔK criterion), results were similar to the analyses of species separately, including distinct clusters for each species, and then distinct genetic lineages for each species. With further subdivision until K = 9 all groups different ploidy levels within each species were revealed (see Fig. 5b). Clusters in each analysis are represented by colours, and individuals are represented as columns. Within each column (individual), the extent of the component colours indicates the magnitude of the membership coefficient (Q) corresponding to each cluster. Up to three genetically distinct groups, coincident with triploid, tetraploid and mixoploid/multiploid individuals were identified within both P. dioica and P. umbilicalis, either in the analyses per single species or in the analyses with all the data combined. These three main ploidy types in P. linearis were not coincident with only three genetic groups. These results also indicate some levels of admixture or cryptic presence of common genotypes in P. dioica and P. umbilicalis.
Spatial representation of genetic differentiation within three Porphyra species. Principal component analyses (PCA) based on allelic variation at 11 loci. On the right, G1, G2, G3 etc. represent the genotypes in Group 1, Group 2 etc. in Table 3.
Relationship between genotypic variation among three Porphyra species. Neighbour-joining (NJ) network generated from a matrix of pairwise41 genetic distances (genotypes), using Populations software. Data from 121 genotypes grouped in 19 groups. The red lines show the well distinct lineages for P. linearis, triploids, tetraploids (in circles) and the transition mixoploids lineages (dotted line).
Genetic subdivision of genotypes found in three Porphyra species inferred by Structure. Bar colours represent the proportions of individual genotypes attributable to K genetic clusters (K). Ploidy groups per species: Zp = zygotospores, Tetra = tetraploid (genotypes and gametes), Mx = Mixoploid, Mp = multiploid gametes, Triploid (genotypes and gametes). Ploidy types are denoted on the top of the plot, and ploidy levels on the bottom of the plot. Each genotype corresponds to either blades, gametes and/or zygotospores. (a) Structure analysis per species separately. (b) Structure analysis with all the data combined. Best Ks for each group are highlighted with the symbol (**).
Discussion
Polyploid races in Porphyra
Our results show that the gametophytes of the three Porphyra species studied are polyploid. Porphyra species showed high variability in genome sizes (eight), ploidy levels (seven), with two different cell lines found and at least three distinct lineages which comprised seven cytotypes. We show that the gametophytic phase (n) in Porphyra is not haploid, in disagreement with the assumptions derived from Drew21 landmark paper. Our results confirm the haplophasic/diplophasic cycle in Porphyra, with replicated and unreplicated nuclei, but not with the ploidy levels previously assumed, nor the simplicity of only two ploidy levels (haploid vs. diploid) assumed. The terminology haplophasic/diplophasic phase is already described for other plant species43.
There was important genetic differentiation and structure among the three species (Figs 3–5). Within each species, genetic differentiation of the polyploid types was different; in P. linearis, genetic differentiation between polyploid types (triploids vs. tetraploids vs. mixoploids) was strong; in P. dioica there was no significant genetic differentiation (most of the genotypes were tetraploid); and in P. umbilicalis, there was some degree of genetic differentiation (weak) which revealed discrete genetic clusters. Gene flow in polyploid lineages in these Porphyra species is likely mediated either with intermediate lineages (mixoploids) or by individuals that release gametes of lower ploidy levels that can fecundate different cytotypes. In P. dioica, the population is dominated by tetraploid individuals with gametes of multiple ploidy levels.
We also support the idea that the Porphyra system is a complex system formed by autopolyploids, mixoploids and allopolyploids that interact among and between them. Triploids and tetraploids are normally originated by the fusion of gametes of different ploidies originated by irregular mitotic processes. Mixoploids are more likely originated by meiotic processes in the germination of the conchospore into four different tetrads (genetic chimeras, see section below) as reported before by previous authors for some Porphyra spp. (e.g.ref44), and consequently different cell lines within the same individual. We have not tested here the hypothesis of whether gametes from one species fecundate gametes from another species (with the same or different ploidy level), but we have observed some shared genotypes between P. dioica and P. umbilicalis (Figs 3–5). Yet, allopolyploids have been already observed by45 for some Japanese species in nature.
Mixoploidy, genetic chimeras and multiploid gametes
Genetic chimeras or organisms with more than one cell line have been reported for several plant species (e.g. ref46), including algal species such as Gracilaria tikvahiae McLachlan47,48 and a Porphyra species: P. yezoensis44,49,50. In Porphyra yezoensis Ueda (actual name: Pyropia yezoensis (Ueda) M.S. Hwang & H.G. Choi) meiosis occurs as the conchospores germinate and the four resultant cells, which all have different genetic compositions, form the mature blade through successive divisions44. However, Niwa et al.51,52, have been reporting both chimeric blades and uniform blades, the last ones associated to asexual reproduction. In the species analysed in the current study, we have detected chimeric blades (mixoploids) and blades with one vegetative cell line (triploids and tetraploids), of which some individuals produced gametes of the same ploidy and others produced gametes of multiple ploidy levels. Three types of mixoploidy have been discovered: 2x/3x, 2x/4x, and 3x/4x. When analysing samples randomly collected in each blade, we always obtained the same result. Thus, it can be concluded that the formation of mixoploids for the samples examined was not sectorial, and the two different cell lines seem to be mixed within the blade. However, further research would be needed to study and confirm this aspect in detail.
We hypothesize that in our polyploid system, mixoploids or blades with two cell lines are a bridge between ploidy types. In polyploid research, triploids are believed to act as bridges between ploidy types; in such a role, they mediate the formation of tetraploids from diploids or facilitate gene flow between diploids and tetraploids4. Triploidy is associated to a long-term strategy, that could produce either asexual progeny or sexual progeny. For example, with a specialized meiosis, triploids can produce haploid sperm and diploid eggs (e.g. in frogs)53, or diploid sperm and haploid eggs (e.g. in plants)54,55. The mixoploids blades found here could reflect a long-term strategy in the Porphyra system of concealing and admixture of different cytotypes in the same species, preventing the exclusion of new cytotypes after emergence (similarly as the minority cytotype exclusion theory56). In the same way, blades that produce gametes of different ploidy levels act as bridges among cytotypes. The generation of gametes of lower ploidy (2x, 3x) in tetraploid blades prevents the elimination of triploids and also ensures the admixture of different polyploid lineages.
We can’t answer how the production of gametes of different ploidy levels is performed. The processes involved in the DNA reduction in Porphyra are yet to be described. Decrease in genome size has been reported to be adaptive. Rapid elimination of DNA sequences from autopolyploid genomes seems to occur in species such as Elymus elongatus, where both newly synthesized autopolyploids as well as natural accessions showed a loss of 10% DNA compared with their diploid progenitors57. However, this reduction is performed in a different way, since genome reduction is done in all the cells in the plant, not only the reproductive cells.
In the case of gametes of higher ploidies (“multiploid gametes”) found in P. dioica (9x, 12x) the only explanation to these sizes is the presence of spontaneous chromosome duplication/triplication phenomena. We discard the hypothesis of the presence of aggregates (doublets or triplets) since no changes in nuclei pulse width were detected among the nuclear populations, as expected if doublets or triplets existed (doublets show larger pulse width when compared with singlets). However, spontaneous chromosome duplication has already been reported in Porphyra species58 in59. However, as no blades of higher ploidy levels (>4x) were found, we presumed that those gametes are unviable. A simple explanation for this would be that since only male gametes presented those ploidy levels, fecundation with female gametes is not possible. Still, further research is needed to clarify this point.
Proposed life histories for Porphyras in the Iberian Peninsula
Based on our results, three new life histories are proposed for Porphyra species in relation to the general assumption of a haploid/diploid life history (Fig. 6). In life history 1 (Fig. 7), two genetically distinct lineages coexist, triploids and tetraploids, but they are isolated from each other. In life history 2 (Fig. 8), both lineages start to release gametes of different ploidy level. In this way, they can potentially cross with each other. However, since zygotospores 6x are only found in triploids, it is believed that gene flow goes from the tetraploids to the triploids. In life history 3 (Fig. 9), three lineages coexist, triploids, tetraploids and mixoploids, with the last lineage the acting as bridge between the other two lineages. Life history 1 and 2 are present in the three Porphyra species (P. linearis, P. umbilicalis and P. dioica), while life history 3 is typical of P. linearis, and present in P. umbilicalis.
The alternation of generations in algae and land plants typically describe the gametophytic generation as being haploid, and the sporophyte generation as diploid. However, there are many examples, which are not correlated with this haploid/diploid model. For example, the nuclei of sporophytes and gametophytes of the brown seaweed Haplospora globosa (Tilopteridales) possess the same number of chromosomes. However, the DNA level of sporophytic nuclei is twice that of gametophytic nuclei60. Another example is the gametophyte of Mediterranean Caulerpas, which can be diploid or triploid61. In Porphyra, incongruences in chromosome numbers have also been reported. For example, for some species of Porphyra, there is no change in chromosome number between the gametophyte vs. the sporophyte (e.g. refs62,63,64,65). However, if those species are exhibiting a parthenogenic life history, there is no explanation for the production and release of male gametes unless they were released to fecundate other polyploid races. We wonder now if the incongruences of the life histories reported in the literature for Porphyra spp. and other algal/plant taxa may be related to the recurrent presence of polyploid events and/or to the occurrence of mixoploidy undiscovered. And in that case, we believe that many algal groups have to be revisited and the life histories re-described according to the ploidy levels. Knowing that polyploids form at relatively high frequency (1 per 100,000) in flowering plants10 and also that formation of polyploids is possible in animals like fish, reptiles, amphibians and even mammals12,66, provides support to the hypothesis that many other polyploid algal species might remain undiscovered.
Our results suggest the hypothesis that the debate created on the cryptic diversity in Porphyra species may be associated with the diversity of ploidy levels that we found. Since cryptic diversity in Porphyra has never been explored in this way, and also considering the evidence for genetic differentiation and isolation for the two polyploid races in Porphyra linearis, triploids and tetraploids, we believe they represent distinct evolutionary lineages with incipient speciation in P. linearis from the Iberian Peninsula, and therefore they may be considered distinct species. Future studies regarding the formal nomenclatural change in P. linearis are required.
Porphyra as an interesting system to study polyploid evolution
The results of this study highlight the interest in using the genus Porphyra to study polyploid evolution. Since the gametophyte is the dominant phase and different cytotypes (triploids, tetraploids and mixoploids and may be other cytotypes) coexist in the same population, the genus Porphyra can add novel insight into the evolutionary origins of polyploids. Interactions among these cytotypes can be studied in the same geographical region and hypotheses about intermediate cytotypes that may act as bridges can be formulated and assessed. The sister genus Bangia being the oldest eukaryotic multicellular taxon with sexual reproduction recorded67, also supports the interest of using the Bangiales to study other evolutionary questions unresolved such as, in an evolutionary scale, what was the first to occur: the gametophytes or sporophyte, and also may help to contrast hypotheses on the origin of the sporophyte and of the alternating generations in land plants68. Polyploidy is often associated to asexual reproduction, but in this case all the species studied are sexual, which is also interesting. Besides, many Porphyra/Pyropia species present asexual reproduction.
Genome doubling (polyploidy) has frequently been associated with evolution and diversification in plants and has been a major factor in the evolution of other lineages of eukaryotes, including yeast69 and many groups of vertebrates and invertebrates reviewed in refs3,70,71,72,73. Here we have reported a polyploid system with a high variability in ploidy levels, cytotypes and lineages with different life history strategies. Porphyra is a very interesting genus to continue the study of polyploid evolution, and towards the elaboration of evolutionary models for other red algae, and may be for other eukaryotes.
Concluding remarks
In our study, many distinct ploidy levels (seven) and genome sizes (eight) were found in Porphyra species. These represent two cell lines and comprise seven different cytotype combinations (with single and compound ploidy levels) among the same and different individuals. It is concluded that the gametophytic phase (n) in Porphyra is not haploid and more than one ploidy level is involved in this phase of the life history. In addition, three main distinct lineages for each Porphyra species are found: Triploids, Tetraploids and Mixoploids. In P. linearis, genetic differentiation was found among the three lineages: triploid and tetraploid, and mixoploid, representing different evolutionary units. Overall, the results of the present study reveal that Porphyra spp. constitute a complex polyploid system, composed by autopolyploids, mixoploids, multiploid gametes and probably spontaneous chromosome doubling, where three different types of life histories coexist but in different proportions depending on the species.
References
Otto, S. P. The evolutionary consequences of polyploidy. Cell 131, 452–462, https://doi.org/10.1016/j.cell.2007.10.022 (2007).
Wood, T. E. et al. The frequency of polyploid speciation in vascular plants. Proc Natl. Acad. Sci. USA 106, 13875–13879, https://doi.org/10.1073/pnas.0811575106 (2009).
Levin, D. The role of chromosomal change in plant evolution (ed. Harvey, P. H. & May, R. M.) 240 pp. (Oxford University Press, New York, 2002).
Comai, L. The advantages and disadvantages of being polyploid. Nat. Rev. Genet. 6, 836–846, https://doi.org/10.1038/nrg1711 (2005).
Soltis, P. S. & Soltis, D. E. The role of hybridization in plant speciation. Annu. Rev. Plant. Biol. 60, 561–588, https://doi.org/10.1146/annurev.arplant.043008.092039 (2009).
Doyle, J. J. et al. Evolutionary genetics of genome merger and doubling in plants. Ann. Rev. Genet. 42, 443–461, https://doi.org/10.1146/annurev.genet.42.110807.091524 (2008).
Leitch, A. R. & Leitch, I. J. Genomic plasticity and the diversity of polyploidy plants. Science 320, 481–483 (2008).
LaJeunesse, T. C., Lambert, G., Anderson, R. A., Coffroth, M. A. & Galbraith, D. W. Symbiodinium (Pyrrhophyta) genome sizes (DNA content) are smallest among dinoflagellates. J. Phycol. 41, 880–886, https://doi.org/10.1111/j.1529-8817.2005.00111.x (2005).
Friedman, W. E. One Genome, Two Ontogenies. Science 339, 1045–1046, https://doi.org/10.1126/science.1234992 (2013).
Ramsey, J. & Schemske, D. W. Pathways, mechanisms, and rates of polyploidy formation in flowering plants. Annu. Rev. Ecol. Syst. 29, 467–501, https://doi.org/10.1146/annurev.ecolsys.29.1.467 (1998).
Ramsey, J. & Schemske, D. W. Neopolyploidy in flowering plants. Annu. Rev. Ecol. Syst. 33, 589–639, https://doi.org/10.1146/annurev.ecolsys.33.010802.150437 (2002).
Otto, S. P. & Whitton, J. Polyploidy: incidence and evolution. Annu. Rev. Genet. 34, 401–437 (2000).
Guiry, M. D. & Guiry, G. M. AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. [WWW document], http://www.algaebase.org. [Accessed 16 April 2018] (2018).
Varela Álvarez, E. Phenology, life history and genetics of Porphyra linearis Greville, a candidate for Nori mariculture in Europe. PhD thesis (unpublished dissertation). National University of Ireland, Galway, Ireland. 219 pp (2002).
Sutherland, J. E. et al. A new look at an ancient order: Generic revision of the Bangiales (Rhodophyta). J. Phycol. 47(5), 1131–1151, https://doi.org/10.1111/j.1529-8817.2011.01052.x (2011).
Lindstrom, S. C. & Fredericq, S. RbcL gene sequences reveal relationships among north-east Pacific species of Porphyra (Bangiales, Rhodophyta) and a new species, P. aestivalis. Phycol. Res. 51, 211–24, https://doi.org/10.1046/j.1440-1835.2003.00312.x (2003).
Kucera, H. & Saunders, G. W. A survey of Bangiales (Rhodophyta) based on multiple molecular markers reveals cryptic diversity. J. Phycol. 48(4), 869–882, https://doi.org/10.1111/j.1529-8817.2012.01193.x (2012).
Conway, E. & Cole, K. Studies in the Bangiaceae: structure and reproduction of the conchocelis of Porphyra and Bangia in culture (Bangiales, Rhodophyceae). Phycologia 16, 205–216, https://doi.org/10.2216/i0031-8884-16-2-205.1 (1977).
Miura, A. A new species of Porphyra and its conchocelis-phase in nature. J. Tokyo Univ. Fish. 47, 305–311 (1961).
Martínez, E. The Conchocelis-phase of Porphyra (Rhodophyta) in the intertidal of San-Juan Island, Washington, USA. Phycologia 29, 391–395, https://doi.org/10.2216/i0031-8884-29-4-391.1 (1990).
Drew, K. M. Conchocelis-phase in the life-history of Porphyra umbilicalis (L.) Kutz. Nature 164, 748–749, https://doi.org/10.1038/164748a0 (1949).
Magne, F. La structure du noyau et le cycle nucleaire chez Ie Porphyra lineraris Greville. C. R. Acad. Sci. 234, 986 (1952).
Yabu, H. & Tokida, J. Mitosis in Porphyra. Bull. Fac. Fish. Hokkaido. Univ. 14(3), 133–136 (1963).
Kito, H. Cytological studies of several species of Porphyra I. Morphological and cytological observations on a species of Porphyra epiphytic on Grateloupia filicina var. porracea (Mert.) Howe. Bull. Fac. Fish. Hokkaido. Univ. 16 (4), 206–208 (in Japanese) (1966).
Kito, H., Yabu, H. & Tokida, J. The number of chromosome in some species of Porphyra. Bull. Fac. Fish. Hokkaido. Univ. 18 (2) 59–60, http://hdl.handle.net/2115/23302 (1967).
Blouin, N. A., Brodie, J. A., Grossman, A. C., Xu, P. & Brawley, S. H. Porphyra: a marine crop shaped by stress. Trends. Plant. Sci. 16(1), 29–37, https://doi.org/10.1016/j.tplants.2010.10.004 (2011).
Guiry, M. D. Sporangia and spores. In Biology of the Red Algae (ed. Cole, K. M. & Sheath, R. G.) 347–376 pp. (Cambridge University Press, Cambridge) (1990).
Loureiro, J., Kopecký, D., Castro, S., Santos, C. & Silveira, P. Flow cytometric and cytogenetic analyses of Iberian Peninsula Festuca spp. Plant. Syst. Evol. 269, 89–105, https://doi.org/10.1007/s00606-007-0564-8 (2007).
Doležel, J. et al. Plant genome size estimation by flow cytometry: Inter-laboratory comparison. Ann. Botany. 82(Supplement A), 17–26 (1998).
Hong, Y. K., Coury, D. A., Polne-Fuller, M. & Bibor, A. Lithium chloride extraction of DNA from the seaweed Porphyra perforata (Rhodophyta). J. Phycol. 28, 217–20, https://doi.org/10.1111/j.0022-3646.1992.00717.x (1992).
van Oppen, M. J. H., Draisma, S. G. A., Olsen, J. L. & Stam, W. T. Multiple trans-Arctic passages in the red alga Phycodrys rubens: evidence from nuclear rDNA ITS sequences. Mar. Biol. 123, 179–188, https://doi.org/10.1007/BF00350338 (1995).
Varela-Álvarez, E., Paulino, C. & Serrão, E. A. Development and characterization of twelve microsatellite markers for Porphyra linearis Greville. Genetica 145, 127–130, https://doi.org/10.1007/s10709-016-9941-y (2017).
Varela-Álvarez, E., et al Isolation and characterization of microsatellite markers for the red alga Porphyra umbilicalis. Plant Genetic Resources, 1–4, https://doi.org/10.1017/S147926211700034X (2017).
Matschiner, M. & Salzburger, W. TANDEM: integrating automated allele binning into genetics and genomics workflows. Bioinformatics 25, 1982–1983, https://doi.org/10.1093/bioinformatics/btp303 (2009).
Meirmans, P. G. & Van Tienderen, P. H. Genotype and Genodive: two pro- grams for the analysis of genetic diversity of asexual organisms. Mol. Ecol. Not. 4, 792–794, https://doi.org/10.1111/j.1471-8286.2004.00770.x (2004).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software Structure: a simulation study. Mol. Ecol. 14, 2611–2620, https://doi.org/10.1111/j.1365-294X.2005.02553.x (2005).
Jakobsson, M. & Rosenberg, N. A. CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23(4), 1801–1806, https://doi.org/10.1093/bioinformatics/btm233 (2007).
Earl, D. A. & von Holdt, B. M. Structure Harvester: a website and program for visualizing Structure output and implementing the Evanno method. Conserv. Genet. Resour. 4(2), 359–361, https://doi.org/10.1007/s12686-011-9548-7 (2012).
Emanuelli, F. et al. Genetic diversity and population structure assessed by SSR and SNP markers in a large germplasm collection of grape. BMC Plant. Biol. 13, 39, https://doi.org/10.1186/1471-2229-13-39 (2013).
Cavalli-Sforza, L. L. & Edwards, A. W. F. Phylogenetic analysis: models and estimation procedures. Am. J. Hum. Genet. 19, 233–257 (1967).
Langella, O. Populations 1.2.28 (12/5/2002): a population genetic software. CNRS UPR9034. Available at, http://bioinformatics.org/~tryphon/populations/ [last accessed 24.06.2010] (1999).
Greilhuber, J., Doležel, J., Martin, A., Lysa, K. & Bennett, M. D. The Origin, evolution and proposed stabilization of the Terms ‘Genome Size’ and ‘C-Value’ to describe nuclear DNA contents. Ann. Botany 95, 255–260 (2005).
Ohme, M. & Miura, A. Tetrad analysis in conchospore germlings of Porphyra yezoensis (Rhodophyta, Bangiales). Plant. Sci. 57, 135–140, https://doi.org/10.1016/0168-9452(88)90079-9 (1988).
Niwa, K., Kobiyama, A. & Sakamoto, T. Interspecific hybridization in the haploid blade- forming marine crop Porphyra (Bangiales, Rhodophyta): occurrence of allopolyploidy in surviving F1 gametophytic blades. J. Phycol. 46, 693–702, https://doi.org/10.1111/j.1529-8817.2010.00853.x (2010).
Szymkowiak, E. J. & Sussex, I. M. What chimeras can tell us about plant development. Annu. Rev. Plant. Physiol. Plant. Mol. Biol. 47, 351–376, https://doi.org/10.1146/annurev.arplant.47.1.351 (1996).
van der Meer, J. P. Genetics of Gracilaria sp. (Rhodophyceae, Gigartinales). II. The life history and genetic implications of cytokinetic failure during tetraspore formation. Phycologia 16, 367–371, https://doi.org/10.2216/i0031-8884-16-4-367.1 (1977).
van der Meer, J. P. & Zhang, X. Similar unstable mutations in three species of Gracilaria (Rhodophyta). J. Phycol. 24, 198–202, https://doi.org/10.1111/j.1529-8817.1988.tb04234.x (1988).
Miura, A. Porphyra cultivation in Japan. In: Tokida, J. & Hirose, H., eds Advance of Phycology in Japan. Dr. W. Junk Publishers, The Hague, 273–304 (1975).
Kobara, T., Miura, A. & Aruga, Y. In vitro studies on the green type mutant of Porphyra yezoensis Ueda. La Mer 14, 8–13 (1976).
Niwa, K., Hayashi, Y., Abe, T. & Aruga, Y. Induction and isolation of pigmentation mutants of Porphyra yezoensis (Bangiales, Rhodophyta) by heavy-ion beam irradiation. Phycol. Res. 57, 194–202 (2009).
Niwa, K. & Abe, T. Chimeras with mosaic patterns in archeospore germlings of Pyropia yezoensis Ueda (Bangiales, Rhodophyta). J. Phycol. 48, 706–709, https://doi.org/10.1111/j.1529-8817.2012.01143 (2012).
Stöck, M. et al. A bisexually reproducing all-triploid vertebrate. Nat. Genet. 30, 325–328, https://doi.org/10.1038/ng839 (2002).
Norrmann, G. A. & Quarin, C. L. Permanent odd polyploidy in a grass (Andropogon ternatus). Genome 29, 340–344 (1987).
Smith-White, S. Polarized segregation in the pollen mother cells of a stable triploid. Heredity 2, 119–129, https://doi.org/10.1038/hdy.1948.7 (1948).
Husband, B. C. Constraints on polyploid evolution: a test of the minority cytotype exclusion principle. Proc. R. Soc. Lond. [Biol] 267, 217–223, https://doi.org/10.1098/rspb.2000.0990) (2000).
Eilam, T., Anikster, Y., Millet, E., Manisterski, J. & Feldman, M. Genome size in natural and synthetic autopolyploids and in a natural segmental allopolyploid of several Triticeae species. Genome 52(3), 275–285, https://doi.org/10.1139/G09-004 (2009).
Zhong, C. H. Study on parthenogenesis and spontaneous chromosome doubling in Porphyra haitanensis Chang et Zheng (Bangiales, Rhodophyta). Masters thesis, Shanghai Ocean University 74 pp., China (2011).
Zhang, Y., Yan, X.-H. & Aruga, Y. The Sex and Sex Determination in Pyropia haitanensis (Bangiales, Rhodophyta). Plos One 8(8), e73414, https://doi.org/10.1371/journal.pone.0073414 (2013).
Kuhlenkamp, R., Müller, D. G. & Whittick, A. Genotypic variation and alternating DNA levels at constant chromosome numbers in the life history of the brown alga Haplospora globosa (Tilopteridales). J. Phycol. 29, 377–380, https://doi.org/10.1111/j.0022-3646.1993.00377.x (1993).
Varela-Álvarez, E. et al. Mediterranean species of Caulerpa are polyploid with smaller genomes in the invasive ones. Plos One 7(10), e47728, https://doi.org/10.1371/journal.pone.0047728 (2012).
Conway, E. & Cole, K. Observations on an unusual form of reproduction in Porphyra (Rhodophyceae, Bangiales). Phycologia 12 3,4, 213–225, https://doi.org/10.2216/i0031-8884-12-3-213.1 (1973).
Mumford, F. T. & Cole, K. Chromosome numbers for fifteen species in the genus Porphyra (Bangiales, Rhodophyta) from the West coast of North America. Phycologia 16, 373–377, https://doi.org/10.2216/i0031-8884-16-4-373.1 (1977).
Krishnamurthy, V. Chromosome number in Porphyra C. Agardh. Phykos 23, 185–190 (1984).
Kapraun, D. F. & Freshwater, D. W. Karyological studies of five species of Porphyra (Bangiales, Rhodophyta) from the North Atlantic and Mediterranean. Phycologia 26, 82–87, https://doi.org/10.2216/i0031-8884-26-1-82.1 (1987).
Mable, B. K. Breaking down taxonomic barriers in polyploidy research. Trends. Plant. Sci. 8(12), 582–590, https://doi.org/10.1016/j.tplants.2003.10.006 (2003).
Butterfield, N. J. Bangiomorpha pubescens n. gen., n. sp.: implications on the evolution of sex, multicellularity, and the Mesoproterozoic/Neoproterozoic radiation of eukaryotes. Paleobiology 26, 386–404, https://doi.org/10.1666/0094-8373(2000)0260386:BPNGNS2.0.CO;2 (2000).
Graham, L. K. & Wilcox, L. W. The origin of alternation of generations in land plants: a focus on matrotrophy and hexose transport. Proc. R. Soc. Lond. [Biol] 355, 757–767, https://doi.org/10.1098/rstb.2000.0614 (2000).
Kellis, M., Birren, B. W. & Lander, E. S. Proof and evolutionary analysis of ancient genome duplication in the yeast Saccharomyces cerevisiae. Nature 428, 617–624, https://doi.org/10.1038/nature02424 (2004).
Wendel, J. F. & Doyle, J. J. Polyploidy and evolution in plants. In: Plant diversity and evolution in plants: genotypic and phenotypic variation in higher plants. (ed. Henry, R.) 97–117 pp, https://doi.org/10.1079/9780851999043.0097 (CABI Publishing, Oxon, UK 2005).
Adams, K. L. & Wendel, J. F. Polyploidy and genome evolution in plants. Curr. Opi. Plant. Biol. 8, 135–141, https://doi.org/10.1016/j.pbi.2005.01.001 (2005).
Gregory, T. R. & Mable, B. K. Polyploidy in animals. In: The Evolution of the Genome. (ed. Gregory, T. R.) 428–501 pp, https://doi.org/10.1016/B978-012301463-4/50010-3 (Elsevier, San Diego, 2005).
Tate, J. A., Soltis, D. E. & Soltis, P. S. Polyploid in Plants. In The Evolution of the Genome. (ed. Gregory, T. R.) 372–414 pp, https://doi.org/10.1016/B978-012301463-4/50009-7 (Elsevier, San Diego, 2005).
Nei, M. Molecular Evolutionary Genetics. Columbia University Press, New York (1987).
Jost, L. GST and its relatives do not measure differentiation. Mol. Ecol. 17, 4015–4026 (2008).
Acknowledgements
We thank Liam Cronin who helped with the sampling in the field. Funded by a FCT (Fundação para a Ciência e a Tecnologia, Portugal) through post-doctoral fellowship SFRH/BPD/109452/2015 and project NORIGENOMICS - PTDC/MAR/099698/2008 both attributed to EVÁ, and UID/Multi/04326/2013 to CCMAR; and also BIODIVERSA/004/2015-MARFOR and Pew Marine Fellows attributed to EAS.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: E.V.Á. Performed the experiments: E.V.Á., C.P., J.L. Analysed the data: E.V.Á., J.L., E.A.S. Contributed reagents/materials/analysis tools: E.V.Á., J.L., E.A.S. Wrote the paper: E.V.Á., E.A.S.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Varela-Álvarez, E., Loureiro, J., Paulino, C. et al. Polyploid lineages in the genus Porphyra. Sci Rep 8, 8696 (2018). https://doi.org/10.1038/s41598-018-26796-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26796-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.