Abstract
A rare variant (BAFF-var) of the tumor necrosis factor superfamily 13b (TNFSF13B) gene has been recently associated with multiple sclerosis (MS) and systemic lupus erythematosus (SLE). The aim of this study was to investigate the association between TNFSF13B BAFF-var and susceptibility to rheumatoid arthritis (RA) and replicate that association in SLE. 6,218 RA patients, 2,575 SLE patients and 4,403 healthy controls from three different countries were included in the study. TNFSF13B BAFF-var was genotyped using TaqMan allelic discrimination assay. PLINK software was used for statistical analyses. TNFSF13B BAFF-var was significantly associated with RA (p = 0.015, OR = 1.21, 95% CI = 1.03–1.41) in the Spanish cohort. A trend of association was observed in the Dutch (p = 0.115) and German (p = 0.228) RA cohorts. A meta-analysis of the three RA cohorts included in this study revealed a statistically significant association (p = 0.002, OR = 1.24, 95% CI = 1.10–1.38). In addition, TNFSF13B BAFF-var was significantly associated with SLE in the Spanish (p = 0.001, OR = 1.41, 95% CI = 1.14–1.74) and the German cohorts (p = 0.030, OR = 1.86, 95% CI = 1.05–3.28), with a statistically significant p-value obtained in the meta-analysis (p = 0.0002, OR = 1.46, 95% CI = 1.09–2.32). The results obtained confirm the known association of TNFSF13B BAFF-var with SLE and, for the first time, demonstrate that this variant contributes to susceptibility to RA.
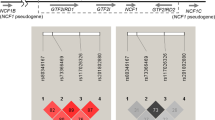
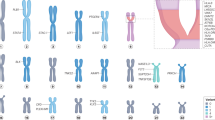
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) and systemic lupus erythematosus (SLE) are complex autoimmune diseases influenced by both genetic and environmental factors1,2. Genome-wide association studies (GWAS) have revealed more than 100 independent signals for RA and 50 for SLE associated with these diseases3,4. In a recent study Steri et al.5 identified an association signal in the TNFSF13B gene which was associated with multiple sclerosis (MS) and SLE. TNFSF13B encodes tumor necrosis factor superfamily member 13b (also known as B-cell activating factor (BAFF)). BAFF is a cytokine that is primarily produced by monocytes and neutrophils and plays a crucial role in B-cell homeostasis and the regulation of B-cell maturation, differentiation and survival6. This cytokine plays a particularly important role in the pathogenesis of SLE, to such an extent that the first targeted therapy approved for SLE was Belimumab, a monoclonal antibody targeting human BAFF7,8,9. The causal variant identified by Steri et al.5 for this association, “BAFF-var”, is an insertion-deletion in which five nucleotides are replaced by one (GCTGT > A), being this A the risk allele. Persons carrying the A allele have an increased risk for the disease. This risk allele results in a shorter transcript that escapes microRNA inhibition thus leading to an increase in the production of soluble BAFF. Besides, it has been observed that TNFSF13B BAFF-var is strongly associated with augmented levels of total IgG and IgM and with reduced monocyte counts5. Thus, TNFSF13B BAFF-var is considered a shared genetic risk variant for autoimmune diseases based on its association with MS and SLE, and it plays an important role in autoimmune processes5. Considering that replication of previous results is of vital importance for the correct development of scientific knowledge, we performed a replication study to corroborate the association of TNFSF13B BAFF-var with susceptibility to SLE.
Otherwise, BAFF system dysregulation is involved in the pathogenesis of RA1,10 and abnormal levels of BAFF have been detected in serum, synovial fluid and saliva from RA patients11. It is noteworthy that more than 10 shared risk loci have been identified for SLE and RA12,13. Taking into considerations above, we aimed to assess for the first time the potential association of TNFSF13B BAFF-var with RA in patients from Spain, the Netherlands and Germany.
Materials and Methods
Study subjects
Overall, 6,218 RA patients and 4,403 healthy controls from Spain, Germany and the Netherlands were enrolled in our study. Regarding the Spanish cohort, DNA was obtained from 4,429 RA patients from the Xeral-Calde University Hospital, Lugo; Marqués de Valdecilla University Hospital, Santander; Bellvitge University Hospital, Barcelona; Virgen del Rocío University Hospital, Seville; Campus de la Salud University Hospital, Granada; and Hospital Clínico San Carlos, La Princesa University Hospital, La Paz University Hospital, 12 de Octubre University Hospital and Gregorio Marañon University Hospital, all located in Madrid. DNA from 3,200 healthy controls was obtained from Banco Nacional de ADN. DNA was obtained from 890 RA patients and 733 controls from the Netherlands at the Radboud University Medical Center (Nijmegen) and the Nijmegen Biomedical Study (NBS)14 respectively. DNA was obtained from 890 RA patients and 470 controls of German descent at the Hannover Medical School (Hannover) and Düsseldorf University Hospital. All the patients included in this study were of European descent and had been diagnosed with RA according to the 1987 classification criteria of the American College of Rheumatology15.
A total of 2,575 SLE patients from Spain and Germany were included in this study. DNA from 1,160 patients from Spain was obtained at Xeral-Calde University Hospital, Lugo; Virgen del Rocío University Hospital, Sevilla; Virgen de las Nieves University Hospital, Granada; Virgen de la Victoria University Hospital, Málaga, and Parc Taulí University Hospital, Sabadell. DNA from 460 German patients was obtained at the Hannover Medical School, Hannover. All SLE patients were of European descent and fulfilled the American College of Rheumatology criteria for the classification of SLE16.
The study was approved by the local ethical committees of the different participating centers (Ethics Committee of the Spanish National Research Council, Xeral-Calde University Hospital, Marqués de Valdecilla University Hospital, Bellvitge University Hospital, Virgen del Rocío University Hospital, Campus de la Salud University Hospital, Hospital Clínico San Carlos, La Princesa University Hospital, La Paz University Hospital, 12 de Octubre University Hospital and Gregorio Marañon University Hospital, Virgen de las Nieves University Hospital, Virgen de la Victoria University Hospital, Parc Taulí University Hospital, Radboud University Medical Center, Hannover Medical School and Düsseldorf University Hospital) in accordance with the tenets of the Declaration of Helsinki. Written informed consent was obtained from all participants prior to enrolment in the study.
Genotyping
Genomic DNA was isolated from peripheral blood samples using standard salting-out techniques. TNFSF13B BAFF-var was genotyped using a TaqMan allelic discrimination custom assay (assay ID: AH0JGPG) (Applied Biosystems, Foster City, California, USA) on an ABI 7900HT Fast Real-Time PCR System (Thermo Fisher). Genotyping call rate was >95% in all sample sets.
Statistical analysis
All the statistical analyses were carried out with PLINK v1.9 (http://pngu.mgh.harvard.edu/purcell/plink/)17. For all groups of individuals, Hardy-Weinberg equilibrium was determined at a significance level of 0.01. Differences between allele and genotype frequencies in individuals were determined by χ2 test. Odds ratios (ORs) and 95% confidence intervals were calculated according to Woolf’s method. Meta-analyses were performed with inverse-variance method under a fixed-effects model. Heterogeneity of the ORs across cohorts was assessed using both Cochran’s Q test and inconsistency index (I2). The statistical power of our study was calculated by using Power Calculator for Genetic Studies 2006 (CaTS) software (http://www.sph.umich.edu/csg/abecasis/CaTS/)18, under an additive model. The statistical power of the study for determining significance at a p value of 0.05 is shown in Supplementary Table S1.
Data availability
The complete datasets generated and analyzed during the current study are available from the corresponding author on reasonable request.
Ethical use of human participants statement
All participants signed a written informed consent prior to their enrolment in the study for identified data to be published. All procedures were performed after signed informed consent was obtained from patients and in accordance with the local (Spanish National Research Council) ethics committee and the Declaration of Helsinki of 1975, as revised in 1983.
Results
First, in order to evaluate the association between the TNFSF13B gene variant (BAFF-var) and susceptibility to RA, a comparative analysis of allelic and genotype frequencies in RA patients vs. healthy individuals was performed in the Spanish, Dutch and German populations. Results are shown in Table 1. The minor allele frequency (A) of TNFSF13B BAFF-var was higher in RA patients in the three populations included in this study. This difference was statistically significant in the Spanish subgroup (p = 0.015, OR = 1.21, 95% CI = 1.03–1.41), and suggestive associations were observed in the Dutch (p = 0.115, OR = 1.47, 95% CI = 0.91–2.36) and German (p = 0.228, OR = 1.39, 95% CI = 0.81–2.36) subgroups. In addition, a statistically significant p value was obtained in the meta-analysis of the three RA populations included in this study (p = 0.002, OR = 1.24, 95% CI = 1.10–1.38) with low heterogeneity levels (heterogeneity q value = 0.64, I2 = 0) (Table 1).
Secondly, to try to replicate the results previously published by Steri et al.5, a comparative analysis of allelic and genotype frequencies in SLE patients vs. healthy individuals from the Spanish and German populations was also performed (Table 2). The minor allele frequency (A) in TNFSF13B BAFF-var was statistically significantly higher in SLE patients than in controls in both, the Spanish (p = 0.0001, OR = 1.41, 95% CI = 1.14–1.74) and German (p = 0.030, OR = 1.86, 95% CI = 1.05–3.28) subgroups. Meta-analysis of the two SLE populations included in this study revealed a statistically significant p value (p = 0.0002, OR = 1.46, 95% CI = 1.09–2.32) with low heterogeneity levels (heterogeneity q value = 0.37, I2 = 0) (Table 2).
Discussion
We confirm the recently reported association of the TNFSF13B BAFF-var with SLE in two independent populations of Spanish and German origin. In addition, this is the first time that evidence is provided of an association between TNFSF13B BAFF-var and RA. These findings, together with the data reported by Steri et al.5, suggest that TNFSF13B BAFF-var is a common genetic risk factor for autoimmunity.
SLE and RA mice models showed increased serum BAFF levels, and BAFF blockade reduced diseases manifestations19,20. Nevertheless, although clinical trials are being performed on BAFF in RA and SLE patients, only moderate results are emerging. An example of these modest results is the BAFF-inhibitor drug belimumab, which has been found to be only moderately effective in a small number of patients with RA and SLE8,21. Because RA and SLE are heterogeneous diseases with different biological subsets, the efficacy of different drugs could be divergent in specific patient subgroups. Therefore, our new findings could help in the advance of RA therapies, as patients stratified by TNFSF13B BAFF-var status may show a differential benefit from anti-BAFF therapies. In this sense, a recent study showed that a TNFSF13B genetic variant influenced response to B-cell targeted therapy (rituximab) in patients with RA22. Furthermore, B-cell-depleting therapies in patients carrying TNFSF13B BAFF-var would have a weaker effect due to a rapid resurgence of memory B cells induced by high soluble BAFF levels, which could increase the risk of inadequate response and/or relapse23.
Genome-wide association studies (GWAS) have identified multiple risk genetic factors associated with human autoimmune disease. Of note, GWAS are designed to detect only common genetic variants (>5%); however, low-frequency and rare variants may represent an important component of autoimmune risk and provide a key insight into both novel and previously implicated immunological pathways that are disrupted in autoimmune diseases24. Thus, large genetic studies have identified low frequency variants with relevant functional significance associated with autoimmune diseases (e.g. TREX1 in SLE25 and TYK2 in several autoimmune diseases26,27). Our findings support the important role of rare variants, such as TNFSF13B BAFF-var, in understanding the unknown mechanisms of autoimmunity.
For the first time, our results showed that allele A of TNFSF13B BAFF-var was associated with the risk of RA in the Spanish population. Trends of association were observed in the Dutch and German RA cohorts. This could be due to a low statistical power as the Dutch and the German cohorts sample sizes are relatively small as compared to the Spanish cohort, as well as the minor allele frequency is less common in northern Europe (2.02% in Germany and 1.91% in the Netherlands) than in southern Europe (4.2% in Spain). This north-south gradient in the minor allele frequency of TNFSF13B BAFF-var has been previously reported, with higher frequencies observed in southern Europe (5.7% in Italy and 4.9% in Spain) than in northern Europe (1.8% in United Kingdom and Sweden)5. However, the three RA cohorts showed higher levels of allele A in cases than in controls and statistically significant association was observed in the meta-analysis. Thus, further studies involving larger cohorts would be necessary to elucidate if this variant is associated with autoimmune risk in northern populations.
Regarding the SLE replication study, our results confirm the association of allele A of TNFSF13B BAFF-var with the risk of SLE in the Spanish and German populations. The association of the variant was found to be stronger in SLE than in RA and, in both cases, the allele A acts as a risk factor.
GWAS have revealed a significant shared genetic component in autoimmunity28,29. The association of TNFSF13B BAFF-var with RA and SLE observed in our study extends the number of overlapping genetic risk factors across different autoimmune diseases. In this regard, performing further studies to elucidate the specific association of TNFSF13B BAFF-var with other autoimmune diseases would be of great interest as new therapy approaches involving BAFF could emerge.
In conclusion, our results suggest that TNFSF13B BAFF-var plays an important role in susceptibility to RA, and confirm the association of this variant with SLE.
References
McInnes, I. B. & Schett, G. The Pathogenesis of Rheumatoid Arthritis. N. Engl. J. Med. 365, 2205–2219 (2011).
Tsokos, G. C. Systemic Lupus Erithematosus. N. Engl. J. Med. 365, 2110–2121 (2011).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature. 506, 376–381 (2013).
Morris, D. L. et al. Genome-wide association meta-analysis in Chinese and European individuals identifies ten new loci associated with systemic lupus erythematosus. Nat. Genet. 48, 940–946 (2016).
Steri, M. et al. Overexpression of the Cytokine BAFF and Autoimmunity Risk. N. Engl. J. Med. 376, 1615–1626 (2017).
Woodland, R. T., Schmidt, M. R. & Thompson, C. B. BLyS and B cell homeostasis. Semin. Immunol. 18, 318–326 (2006).
Vincent, F. B., Morand, E. F., Schneider, P. & Mackay, F. The BAFF/APRIL system in SLE pathogenesis. Nat. Rev. Rheumatol. 10, 365–373 (2014).
Furie, R. et al. A phase III, randomized, placebo-controlled study of belimumab, a monoclonal antibody that inhibits B lymphocyte stimulator, in patients with systemic lupus erythematosus. Arthritis Rheum. 63, 3918–3930 (2011).
Navarra, S. V., Guzman, R. M. & Gallacher, A. E. Efficacy and safety of belimumab in patients with active systemic lupus erythematosus: a randomised, placebo-controlled, phase 3 trial. Lancet. 377, 721–731 (2011).
Wei, F., Chang, Y. & Wei, W. The role of BAFF in the progression of rheumatoid arthritis. Cytokine. 76, 537–544 (2015).
Bosello, S. et al. Concentrations of BAFF Correlate with Autoantibody Levels, Clinical Disease Activity, and Response to Treatment in Early Rheumatoid Arthritis. J. Rheumatol. 35, 1256–1264 (2008).
Márquez, A. et al. A combined large-scale meta-analysis identifies COG6 as a novel shared risk locus for rheumatoid arthritis and systemic lupus erythematosus. Ann. Rheum. Dis. 76, 286–294 (2017).
Orozco, G. et al. Study of the common genetic background for rheumatoid arthritis and systemic lupus erythematosus. Ann. Rheum. Dis. 70, 463–8 (2011).
Wetzels, J. F., Kiemeney, L. A., Swinkels, D. W., Willems, H. L. & den Heijer, M. Age- and gender-specific reference values of estimated GFR in Caucasians: the Nijmegen Biomedical Study. Kidney Int. 72, 632–637 (2007).
Arneti, F. C. et al. The American Rheumatism Association 1987 Revised Criteria for the Classification of Rheumatoid Arthritis. Arthritis Rheum. 31, 316–324 (1988).
Tan, E. M. et al. The 1982 Revised Criteria for the Classification of Systemic Lupus Erythematosus. Arthritis Rheum. 25, 1271–1277 (1982).
Purcell, S. et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Skol, A. D., Scott, L. J., Abecasis, G. R. & Boehnke, M. Joint analysis is more efficient than replication-based analysis for two-stage genome-wide association studies. Nat. Genet. 38, 209–213 (2006).
Ding, H. et al. Blockade of B-cell-activating factor suppresses lupus-like syndrome in autoimmune BXSB mice. J. Cell Mol. Med. 14, 1717–1725 (2009).
Lai Kwan Lam, Q., King Hung Ko, O., Zheng, B. J. & Lu, L. Local BAFF gene silencing suppresses Th17-cell generation and ameliorates autoimmune arthritis. Proc. Natl. Acad. Sci. USA. 105, 14993–14998 (2008).
Stohl, W. et al. Efficacy and Safety of Belimumab in Patients with Rheumatoid Arthritis: A Phase II, Randomized, Double-blind, Placebo-controlled, Dose-ranging Study. J. Rheumatol. 40, 579–589 (2013).
Juge, P. A. et al. Variants of genes implicated in type 1 interferon pathway and B-cell activation modulate the EULAR response to rituximab at 24 weeks in rheumatoid arthritis. RMD Open. 3, e000448, https://doi.org/10.1136/rmdopen-2017-000448 (2017).
Dass, S. et al. Highly sensitive B cell analysis predicts response to rituximab therapy in rheumatoid arthritis. Arthritis Rheum. 58, 2993–2999 (2008).
Massey, J. & Eyre, S. Rare variants and autoimmune disease. Brief. Funct. Genomics. 13, 392–397 (2014).
Lee-Kirsch, M. A. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 are associated with systemic lupus erythematosus. Nat. Genet. 39, 1065–1067 (2007).
Dendrou, C. A. et al. Resolving TYK2 locus genotype-to-phenotype differences in autoimmunity. Sci. Transl. Med. 8, 363–149 (2016).
Diogo, D. et al. TYK2 protein-coding variants protect against rheumatoid arthritis and autoimmunity, with no evidence of major pleiotropic effects on non-autoimmune complex traits. PloS One. 10, e0122271, https://doi.org/10.1371/journal.pone.0122271 (2015).
Cho, J. H. & Gregersen, P. K. Genomics and the multifactorial nature of human autoimmune disease. N. Engl. J. Med. 365, 1612–1623 (2011).
Cho, J. H. & Feldman, M. Heterogeneity of autoimmune diseases: pathophysiologic insights from genetics and implications for new therapies. Nat. Med. 21, 730–738 (2015).
Acknowledgements
We thank Sonia Garcia, and Gema Robledo for their excellent technical assistance, and all the patients and healthy controls for kindly accepting participating in this study. This work was supported by the following grants: P12-BIO-1395 from Consejería de Innovación, Ciencia y Tecnología, Junta de Andalucía (Spain), and the Cooperative Research Thematic Network (RETICS) program, RD16/0012/0004 (RIER), from Instituto de Salud Carlos III (ISCIII, Health Ministry, Madrid, Spain). DGS was supported by the Spanish Ministry of Economy and Competitiveness through the program FPI (SAF2015-66761-P).
Author information
Authors and Affiliations
Contributions
D.G.S., L.O.F. and J.M. were involved in the conception and design of the study as well as in the interpretation of data. D.G.S. performed the statistical analyses and drafted the manuscript. J.M. critically revised the manuscript for important intellectual content. S.V., A.G., E.R., B.F.G., F.J.L.L., A.B., I.G.A., J.N., C.G.V., J.M.S., R.G.P., M.F.G.E., C.T., P.C., B.K., M.J.H.C., T.W., M.S. and M.A.G.G. contributed samples and/or data acquisition and all authors revised and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
González-Serna, D., Ortiz-Fernández, L., Vargas, S. et al. Association of a rare variant of the TNFSF13B gene with susceptibility to Rheumatoid Arthritis and Systemic Lupus Erythematosus. Sci Rep 8, 8195 (2018). https://doi.org/10.1038/s41598-018-26573-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26573-4
This article is cited by
-
Protein and functional isoform levels and genetic variants of the BAFF and APRIL pathway components in systemic lupus erythematosus
Scientific Reports (2022)
-
BAFF, APRIL and BAFFR on the pathogenesis of Immunoglobulin-A vasculitis
Scientific Reports (2021)
-
Susceptibility of BAFF-var allele carriers to severe SLE with occurrence of lupus nephritis
BMC Nephrology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.