Abstract
Alzheimer’s Disease (AD) is the most common cause of dementia in the elderly. Centenarians – reaching the age of >100 years while maintaining good cognitive skills – seemingly have unique biological features allowing healthy aging and protection from dementia. Here, we studied the expression of SIRT1 along with miR-132 and miR-212, two microRNAs known to regulate SIRT1, in lymphoblastoid cell lines (LCLs) from 45 healthy donors aged 21 to 105 years and 24 AD patients, and in postmortem olfactory bulb and hippocampus tissues from 14 AD patients and 20 age-matched non-demented individuals. We observed 4.0-fold (P = 0.001) lower expression of SIRT1, and correspondingly higher expression of miR-132 (1.7-fold; P = 0.014) and miR-212 (2.1-fold; P = 0.036), in LCLs from AD patients compared with age-matched healthy controls. Additionally, SIRT1 expression was 2.2-fold (P = 0.001) higher in centenarian LCLs compared with LCLs from individuals aged 56–82 years; while centenarian LCLs miR-132 and miR-212 indicated 7.6-fold and 4.1-fold lower expression, respectively. Correlations of SIRT1, miR-132 and miR-212 expression with cognitive scores were observed for AD patient-derived LCLs and postmortem AD olfactory bulb and hippocampus tissues, suggesting that higher SIRT1 expression, possibly mediated by lower miR-132 and miR-212, may protect aged individuals from dementia and is reflected in their peripheral tissues.
Similar content being viewed by others
Introduction
With increased longevity in recent decades, humankind is faced with a continued increase in the prevalence of late-onset neurodegenerative disorders, most notably, sporadic Alzheimer’s Disease (AD). In 2012 The World Health Organization has estimated that by 2050, the number of AD patients globally will triple from 35 million to 115 million. Such scenario, supported by recent data1, will dramatically affect healthcare-related costs and the economy, in particular in developed countries where fertility rates are declining2. At the same time, higher numbers of individuals are expected to reach the age of 100 years without dementia3. While extensive research efforts have been allocated to studying AD complex etiology and pathophysiology, including large epidemiological studies4, genome-wide association studies5, and transcriptomic studies6, little efforts, in comparison, have been directed toward studying the genomics and epigenomics of healthy aging, which may provide clues for understanding the molecular basis for neurodegeneration and its protective factors7. Understanding the biology of healthy human aging and dementia-free longevity is key for deciphering the biology of neurodegenerative diseases8. Indeed, centenarians, individuals reaching the age of over 100 years while retaining good cognitive skills, have been extensively studied9,10,11. For example, centenarian LCLs were shown to possess enhanced DNA repair capacity following H2O2 exposure12.
AD frequently results, already in its early phases, in impaired olfactory sensing capacity and neuropathological hallmark lesions of neurodegenerative diseases are present in the olfactory bulb, often at early, subclinical stages13 Olfactory deficits can be noticed in the ability to detect, recognize, and remember odorants14. Olfactory impairment is associated with incident amnestic MCI and progression from amnestic MCI to AD dementia15. Relative proteome measurements in postmortem olfactory bulb tissue from AD have discovered protein networks interaction during the progression of sporadic AD16. Neurogenesis during adult life is crucial for maintaining correct synaptic connectivity and involves the generation of new axons and dendrites. Adult neurogenesis in humans takes place exclusively in the dentate gyrus of the hippocampus and in the subventricular zone of the olfactory bulb, while newborn neurons migrate to other brain regions. Impaired adult neurogenesis, implicated in AD, is underlined by reduced ability for neural stem cell renewal in these brain tissues17.
In our recent work18, we reported reduced expression of SIRT1 (Sirtuin 1) and RGS2 (regulator of G-protein signaling 2) in lymphoblastoid cell lines (LCLs) from AD patients compared with healthy controls. SIRT1 (silent information regulator 1) is one of seven human sirtuins that possess mono-ADP-ribosyltransferase or deacylase activity toward target proteins, including histones. SIRT1 regulates endocrine, mitochondrial, circadian rhythm and hypothalamic functions. In the brain, SIRT1 takes part in memory formation by modulating synaptic plasticity, promoting axonal elongation and dendritic branching19. Knock-in induced higher expression of the Caenorhabditis elegans and yeast homolog of human SIRT1, Sir2, was found to increase their lifespan20,21. Transgenic mice overexpressing brain Sirt1 exhibit a delayed aging phenotype and extension of lifespan22. Injection of SIRT1 lentivirus in the hippocampus of p25 transgenic mice, a model of AD and tauopathies, protected the mice against neurodegeneration23. SIRT1 had lower concentration in serum of AD and MCI patients compared with healthy individuals24. SIRT1 expression in Tg2576 mice and in derived primary neuronal cultures promotes α-secretase activity with reduction of Aβ generation, suggesting that SIRT1 prevents amyloid neuropathology25. During embryogenesis of mice, Sirt1 interacts with Bcl6 to promote proper neurogenesis of the mouse cerebral cortex26. Sirt1 was upregulated in mouse neural progenitor cells by oxidative stress and directed these cells to differentiate to astrocytes instead of neurons27. Interestingly, there is an association between SIRT1 activation and inhibition of Aβ generation: inhibiting SIRT1 through virally mediated expression of a dominant negative SIRT1 construct increased the accumulation of Aβ peptides by Tg2576 neuron cultures28. Exploring the blood gene expression profiles of centenarians along with neurodegenerative disease patients may yield important clues into longevity and protection from neurodegeneration. Among genes associated with healthy aging, those coding for sirtuins, NAD+-dependent protein deacylases that cleave off acetyl or other acyl groups from the ε-amino group of lysine residues in histones and other proteins, have received extra attention, as their high expression levels were demonstrated to contribute to longevity and to the lifespan-prolonging effects of caloric restriction in diverse organisms including yeast and mammals29,30. Among the seven mammalian sirtuins, sirtuin 1, encoded in humans by SIRT1, is best recognized as associated with longevity and neuroprotection31,32.
miRNAs, short noncoding RNA regulators of gene expression, are expressed in the brain and participate in regulating neuronal plasticity, function and development. Dysfunction in miRNA transcription may lead to neurodegenerative disease and neurodevelopment disorders33. For example, miR-132 contributes to dendritic outgrowth of newborn neurons in the adult mouse hippocampus34. miR-132, miR-212 and miR-22 were shown to target SIRT135,36,37. Deletion of miR-132 and miR-212 was shown to induce tau aggregation in mice expressing endogenous or human mutant tau38, and impair mouse cognitive skills39. Additionally, these miRNAs were upregulated in postmortem frontal cortex tissues of AD and mild cognitive impairment (MCI) patients40.
In the current study, we therefore studied SIRT1, RGS2, miR-132, miR-212 and miR-22 expression levels in LCLs from healthy donors of various age groups, including centenarians, and in LCLs from AD patients. Next, we compared the differential expression levels of the above genes in postmortem olfactory bulb and hippocampal tissues from AD patients and non-demented age matched controls. Our findings provide insights into AD etiology, neurodegeneration pathways, and healthy aging.
Results
Expression levels of SIRT1 and miRNAs in centenarian and AD LCLs
Human lymphoblastoid cell lines (LCLs) from healthy adults of various ages including non-demented centenarians and from AD patients were grown under optimal conditions in serum-containing medium. RNA samples were prepared from the 69 selected LCLs (see Methods).
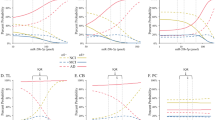
The expression levels of RGS2 and SIRT1, both reported by us to be lower in AD vs. healthy LCLs18 were compared across the different age groups by real-time PCR. The healthy control LCLs were divided into three age groups: 21–35 years (mean 28 ± 1.4 years; N = 12), 56–82 years (mean 71 ± 2.6 years; N = 11), and non-demented centenarians aged 100–105 years (mean 101 ± 0.3 years; N = 16). Figure 1a depicts the individual and mean LCL expression levels of SIRT1 across the non-demented age groups (N = 39) as well as the AD patients (N = 28). As shown, when compared to LCLs from individuals aged 56–82 years, the expression levels of SIRT1 were upregulated (FD = 2.2; P = 0.001) in the centenarian LCLs. By contrast, SIRT1 levels were similar in LCLs from centenarian donors compared with LCLs from individuals aged 21–35 years P = 0.45). In sharp contrast, the SIRT1 expression levels in AD LCLs were dramatically down-regulated compared with centenarian LCLs (FD = −8.4; P = 1.2E-07) as well as compared with the LCLs from healthy donors aged 56–82 years (FD = −4.0; P = 0.001) or 21–35 years (FD = −6.8; P = 0.001; Fig. 1a).
Expression levels and correlations of SIRT1, miR-132 and miR-212 in LCLs from female Alzheimer’s disease patients, female healthy controls (in two age groups) and female centenarians. Expression levels (2−Δct) are shown for the different LCL groups for: (a) SIRT1 (b) miR-132 (c) miR-212. (d,e) Pearson correlation (R) plots for the expression levels of SIRT1 with miR-132 and with miR-212 (combined for all cohorts; N = 69). The LCLs were from Alzheimer’s disease patients (N = 24), healthy controls aged 21–35 years (N = 12), healthy controls aged 56–82 years (N = 11) and centenarians (N = 16). (f) Pearson correlation between age of AD onset and miR-132 expression. Expression levels were determined by real-time PCR (see Methods). Bars represent the expression averages. Fold-difference (FD) values are shown between cohorts along with P values (Kruskal–Wallis test, Mann-Whitney U test). Average ages were similar for the AD patients (76 ± 1.4 years) and the 56–82 years control cohort (71 ± 2.7 years). Data for SIRT1 expression levels in Alzheimer’s disease and control LCLs are from our previous study (Hadar et al. 2016), while new data were obtained for centenarian LCLs; miRNA expression levels were determined in the same RNA preparations (see Methods).
The expression levels of both miR-132 and miR-212 showed a mirror-image of the SIRT1 expression levels in the same LCL cohorts: levels were upregulated in AD LCLs compared with healthy age-matched control LCLs (FD = 1.7; P = 0.014, FD = 2.1; P = 0.036) and, more dramatically, were extremely low in centenarian compared with AD LCLs (FD = 12.9; P = 2.1E-07, FD = 8.6; P = 7.7E-07; Fig. 1b,c). These findings agree with the differential expression levels observed for SIRT1 in the same RNA preparations. In addition, the expression levels of both miR-132 and miR-212 negatively correlated with those of SIRT1 (R = −0.60 P = 4.5E-08; R = −0.49 P = 0.00002; Fig. 1d,e; data were combined for all cohorts, N = 69). Notably, miR-132 LCL expression levels, which negatively correlated with the respective SIRT1 levels (Fig. 1d), negatively correlated also with AD age of onset (R = −0.45, P = 0.029) (Fig. 1f).
As shown in Supplementary Fig. 1a, miR-22 expression was down-regulated (FD = −2.5; P = 0.001) in the centenarian LCLs compared with controls aged 56–82 years. Negative Pearson correlation was observed between miR-22 and SIRT1 expression levels (R = −0.629 P = 1.17E-09; Supplementary Fig. 1b). In an earlier study18 we reported lower RGS2 expression in LCLs from AD patients compared with age-matched healthy controls. Here, we observed similar RGS2 expression in the different healthy control age groups, including the centenarian LCLs (Supplementary Fig. 3b).
AD LCLs expression levels of SIRT1, miR-132 and miR-212 correlate with patient cognitive scores
The expression levels of SIRT1, miR-132 and miR-212 in LCLs from female AD patients (N = 22) were examined for correlations with their Mini Mental State Examination (MMSE) scores and AD Assessment Scale (ADAS) scores (Supplementary Table 1). Higher ADAS scores and lower MMSE scores both indicate more severe cognitive deficits. SIRT1 expression levels in AD LCLs showed negative correlation with patient MMSE scores (R = −0.47, P = 0.033) (Fig. 2a). Corresponding positive correlations were found when comparing MMSE scores and miR-132 (R = 0.47; P = 0.026) and miR-212 LCL expression (R = 0.43; P = 0.046; Fig. 2b,c). Patients ADAS score exhibited lack of correlation with SIRT1 expression (Fig. 2d), albeit negative correlations were observed with expression levels of miR-132 (R = −0.67; P = 0.0006) or miR-212 (R = −0.72; P = 0.00014) (Fig. 2e,f). Of note, the LCL expression levels of miR-212 and miR-132 exhibited a robust positive Pearson correlation (R = 0.893; P = 6.12E-25; Supplementary Fig. 1c; combined cohorts, N = 69). This finding is not surprising given the proximal location of miR-212 and miR-132 on human chromosome 17p13.3 (separated by only 263 nucleotides) and was previously reported for chronic lymphocytic leukemia cells41.
Pearson correlation plots for female Alzheimer’s disease MMSE and ADAS cognitive scores and expression levels (2−Δct) of SIRT1, miR-132 and miR-212 in their LCLs. (a–c) Pearson correlation plot for MMSE scores and expression levels of (a) SIRT1 (N = 21*), (b) miR-132 (N = 22) and (c) miR-212 (N = 22). (d) Lack of correlation for ADAS scores and SIRT1 expression levels (N = 21*). (e,f) Negative Pearson correlation plots for ADAS scores and expression levels of (e) miR -132 (N = 22) and (f) miR -212 (N = 22). ADAS and MMSE scores were recorded at time of blood withdrawal for LCL generation. Note the negative correlation of SIRT1 expression with MMSE but not with ADAS; the negative MMSE correlation agrees with the positive MMSE correlations for miR-132 and miR-212, two miRNAs known to target SIRT1 (see Discussion). ADAS, Alzheimer’s Disease Assessment Scale; MMSE, Mini-Mental State Examination. *A single outlier LCL was removed from the correlation analysis of ADAS and MMSE scores with SIRT1.
Expression levels of miR-132 and miR-212 are downregulated in AD postmortem olfactory bulb and hippocampus
We applied real-time PCR reactions for measuring the expression levels of SIRT1, RGS2, miR-22, miR-132 and miR-212 in postmortem olfactory bulb and hippocampus tissues from AD patients and non-demented age-matched controls. The expression levels of both miR-132 and miR-212 were downregulated in postmortem olfactory bulb (FD = −1.32; P = 0.037 and FD = −1.46; P = 0.025, respectively) and hippocampus tissues (FD = −1.8; P = 0.029 and FD = −2.1; P = 0.004, respectively) from late-onset AD patients (N = 14) compared to non-demented controls (N = 20) (Fig. 3a–d). However, similar SIRT1 expression levels were observed in the same postmortem tissues from AD and non-demented controls. The expression levels of miR-212 and miR-132 exhibited negative Pearson correlations with SIRT1 in the olfactory bulb tissues (Supplementary Fig. 4a,b). Additionally, SIRT1 expression correlated with RGS2 expression in the olfactory bulb tissues (Supplementary Fig. 4c). As observed in human LCLs, miR-132 expression levels correlated with those of miR-212 also in the olfactory bulb and hippocampus postmortem tissues (Supplementary Fig. 5a,b).
Expression levels of miR-132 (a,c) and miR-212 (b,d) in postmortem olfactory bulb (a,b) and hippocampus (c,d) tissues from sporadic Alzheimer’s disease patients and age-matched non-demented controls and their correlations with MMSE scores. Expression levels (2−Δct) were determined in postmortem tissue RNA preparations from Alzheimer’s disease patients (N = 14) and age matched non-demented controls (N = 20). See Methods for tissue collection and storage, RNA extractions, and real-time PCR experiments. Bars represent the miR expression averages. Fold-difference (FD) values are shown between cohorts along with P values (Mann-Whitney U test). (e–g) Pearson correlation between AD patients MMSE scores and their postmortem expression levels of (e) olfactory bulb SIRT1, (f) hippocampal miR-132 and (g) hippocampal miR-212.
SIRT1, miR-132 and miR-212 expression levels in the olfactory bulb and hippocampus are correlated with MMSE scores
The AD postmortem brain tissue MMSE cognitive scores were explored for detecting correlations with the measured expression levels of the above genes and miRNAs. A negative correlation was observed for the AD postmortem olfactory bulb expression levels of SIRT1 with the patients MMSE cognitive scores (R = −0.54; P = 0.015); while in the hippocampus, both miR-132 and miR-212 showed positive correlations with MMSE scores (R = 0.41; P = 0.037, R = 0.44; P = 0.028; Fig. 3e–g). In addition, both miR-132 and miR-212 expression levels were negatively correlated with Braak stage scores and BrainNet Europe (BNE) Aβ phase score in AD olfactory bulb tissues (Supplementary Fig. 6).
Discussion
In the current study, we compared the expression levels of SIRT1 between LCLs derived from healthy controls of different age groups, including centenarians and AD patients. We observed upregulation of SIRT1 expression in LCLs of centenarians compared with healthy controls aged 56–82 years (FD = 2.2, P = 0.01) and with AD patients (FD = 8.4, P = 1.2-E-07). Interestingly, SIRT1 expression levels in LCLs of centenarians were similar to those of younger donors aged 21–35 (Fig. 1a).
Higher SIRT1 levels afford axonal protection by increasing nicotinamide mononucleotide adenylyltransferase activity42. Mice overexpressing Sirt1 exhibited repressed autoimmune encephalomyelitis clinical symptoms and reduced axonal injury43. SIRT1 protein was recently shown to deacetylate tau lysine residues in a mouse model of tauopathy, thereby reducing pathogenic tau generation and spread and suggesting that higher SIRT1 levels are neuroprotective44. SIRT1 protein levels were higher in blood of older healthy controls (56–92 years) compared with younger individuals (32–55 years)45. A study of SIRT1 expression levels in PBMC reported lower expression in individuals aged 60–73 years or in nonagenarians and centenarians compared with younger individuals (19–42 years)46. However, the above studies included only samples from healthy controls. The purpose of our current study was to examine the expression levels of SIRT1, as well as miRNAs known to regulate it, in human LCLs and postmortem brain tissues from both AD patients and controls. One advantage of performing such studies in LCLs is that cell lines from all studied individuals are grown and analyzed under strictly similar conditions, thus minimizing variations arising from different methodological factors. A substantial part of the human genome is transcribed in both brain and LCLs, and indeed, these human cell lines have often proved instrumental for studying CNS disorder biomarkers47.
To further understand SIRT1 regulation we tested (in the same LCL RNA preparations) the expression levels of miR-21236, miR-13235 and miR-2237 reported to regulate SIRT1, and observed opposite expression patterns compared with SIRT1 (Fig. 1, Supplementary Fig. 1). We observed that miR-212 and miR-132 were upregulated in LCLs from AD patients compared to controls, and were extremely low in centenarian LCLs. These findings match the opposite expression patterns detected for SIRT1 in the same cohorts suggesting that all three miRNAs take part in regulating SIRT1 mRNA levels. In AD patients, lower SIRT1 expression levels and higher miR-212 and miR-132 LCL expression levels correlated with better cognitive scores (higher MMSE and lower ADAS scores; Fig. 2), suggestive of earlier AD stage. In addition, higher miR-132 expression levels correlated with lower age of AD onset (Fig. 1f): the higher expression levels of miR-132 possibly allow improved defense against the ongoing neurodegeneration. The hypothesis that higher miR-212 and miR-132 levels are protective in AD shares common features with our recent suggestion that low peripheral RGS2 expression may serve as an early AD biomarker, while lower RGS2 expression levels were associated with better cognitive scores in AD patients18.
Indeed, higher serum miR-132 levels were recently suggested as biomarkers for mild cognitive impairment, a stage often preceding AD48. AD patients whose LCLs exhibited higher Aβ1–42 sensitivity had older age of disease onset (Supplementary Fig. 2c), while no age/Aβ1–42 sensitivity correlations were observed in healthy control LCLs (Supplementary Fig. 2a). Yet, no correlations were observed for SIRT1 expression levels in LCLs with AD age of onset (Supplementary Fig. 2d). The expression levels of miR-212 and miR-132 in the hippocampus and olfactory bulb, two brain regions where neurogenesis takes place throughout human life, were downregulated in AD patient postmortem tissues compared with controls (Fig. 3a-d) and were correlated with worse cognitive MMSE scores (Fig. 3e–g). Our results confirm previous findings that miR-212 and miR-132 were downregulated in postmortem hippocampus brain tissues from AD patients49. Additionally, miR-132 and miR-212 were reduced in medial frontal gyrus49, temporal cortex, and gray matter from prefrontal cortex AD postmortem samples50. Nevertheless, our findings indicated similar SIRT1 expression levels in postmortem olfactory bulb and hippocampus from AD patients compared with controls (Supplementary Fig. 7). SIRT1 expression levels were decreased in miR-132 transfected human umbilical vein endothelial cells. These observations, which differ from those we observed in AD and control LCLs (Fig. 1 and Supplementary Fig. 3), may imply that the regulation of SIRT1 expression by these miRNAs is tissue specific51,52.
miR-132 is required for normal dendrite maturation in newborn mouse hippocampal neurons53. miR-132 expression (measured by in situ hybridization) increases at the onset of synaptic integration in the mouse olfactory bulb. Furthermore sequestration of miR-132 in newborn neurons led to reduced dendritic complexity, spine density and increased survival of newborn neurons54. Additionally, mouse hippocampus miR-132 improves the integration of newborn neurons into synaptic circuitry34. miR-132 was involved in olfactory memory formation, as such, miR-132 expression was upregulated by odor exposure. Interestingly, a local infusion of miR-132 antisense into the olfactory bulb prior to training impaired olfactory learning in C. sphinx55.
Hypertension is recognized as an AD risk factor56. We recently proposed that the reduced RGS2 expression we have identified during early AD progression18 increases angiotensin II receptor (AGTR2) signaling and thus contributes to hypertension, which may in part explain the comorbidity of hypertension and AD57. Interestingly, AGTR2 activation increases the expression of miR-212 and miR-132 in cardiovascular tissues58. Our current observations of higher miR-212 and miR-132 in AD LCLs may thus represent yet another facet of the hypertension/AD link.
Our studies of miR-212 and miR-132 expression levels in AD and control LCLs yielded different findings compared with those obtained in postmortem brain tissues; this may reflect cross-talk between the immune system and the brain. The brain may reduce peripheral inflammation, possibly by parasympathetic cholinergic signaling which decreases cytokine production59. Transfection with miR-132 in normal B lymphocytes led to enhanced inflammation, as evidenced by increased production of lymphotoxin and tumor necrosis factor α60.Thus, the higher miR-132 expression we observed in AD LCLs (Fig. 1) is suggestive of an inflammatory-like state. Lipopolysaccharide-induced inflammation causes microglia activation near newborn neurons which impairs basal hippocampal neurogenesis in rats61. Additionally miR-132 was upregulated in U251 human astrocytoma cells exposed to the inflammatory protein myeloid related protein-862. We propose that high levels of inflammation, known to take part in AD pathogenesis, may thus lead to higher levels of miR-132 in immune cells, as also reflected by AD LCLs in this study, allowing better cognitive status in AD patients through a pathway potentially involving SIRT1 transcriptional down-regulation, as observed here (Fig. 4). Further studies are required for testing this hypothesis.
Our current findings highlight the key role of SIRT1 in healthy aging and in neurodegenerative disorders. SIRT1 expression levels were much higher in centenarian compared with AD LCLs, while the opposite was observed for miR-132 and miR-212 in the same cells. Lower LCL miR-132 expression levels were associated with later AD age of onset. Furthermore, higher miR-132 and miR-212 levels were associated with improved AD patient cognitive scores. Thus, our observations in AD and centenarian LCLs further demonstrate their utility for studying CNS disorder biomarkers47. We suggest that miR-132 and miR-212 may take part in protection from neurodegeneration during early AD progression, a possibility that should be considered by further studies. In addition, our study highlights the value of including biosamples from centenarians in research on neurodegenerative diseases.
Patients, Cell Lines, Materials and Methods
Human LCLs
All LCLs were generated from peripheral blood B lymphocytes donated by consenting individuals. Of note, a written informed consent was obtained from all donors. LCLs from healthy adult donors aged 21 to 81 years were obtained from the National Laboratory for the Genetics of Israeli Populations (NLGIP; http://yoran.tau.ac.il/nlgip/) at Tel-Aviv University, Israel (N = 16) and from the University of Cagliari, Italy (N = 16). LCLs from AD patients were obtained from Prof. Alessio Squassina at the University of Cagliari (N = 28). LCLs from centenarians were obtained from Prof. Jacek Kuźnicki, the International Institute of Molecular and Cell Biology in Warsaw, Poland (N = 16; http://www.iimcb.gov.pl). Detailed demographic data of controls and cognitive scores of the AD patients are presented in Supplementary Table 1. Cells were maintained in similar optimal growth conditions as described63. Tissue-culture reagents were purchased from Biological Industries (Beit-Haemek, Israel).
RNA extraction
RNA extractions were performed from cells incubated in upright T-25 flasks under optimal growth conditions in serum-containing media at a cell density of 0.5 × 106-1 × 106 cells/ml. Cells were centrifuged and then lysed using Tri-reagent (T9424, Sigma-Aldrich), followed by RNA separation using chloroform and precipitation using isopropanol and washed with 80% of cold ethanol. RNA was extracted using the phenol-chloroform method63. RNA was quantified using a NanoDrop spectrophotometer (NanoDrop, Wilmington, DE, USA), with both 260/280 nm and 260/230 nm parameters > 2.0.
Postmortem AD and non-demented controls olfactory bulb and hippocampus
Forty postmortem olfactory bulb brain tissues and a similar number of hippocampus tissues (14 AD patients and 20 age matched non-demented controls) were obtained from Newcastle Brain Tissue Resource (NBTR) in accordance with the approval of the joint Ethics Committee of Newcastle and North Tyneside Health Authority and following NBTR brain banking procedures which includes full written informed consent from tissue donors or relatives. All AD cases fulfilled the criteria for high AD neuropathologic change according to the National Institute on Ageing-Alzheimer’s Association (NIA-AA) guidelines64. Demographic data for the AD patients and age matched non-demented controls are presented in Supplementary Table 2. RNAs were extracted from these postmortem tissues following homogenization using Bullet Blender (Next Advance, Inc., NY, USA), and extraction with ZR-Duet DNA-RNA MiniPrep Plus kit (Zymo Research, Irvine, CA, USA). Next, RNAs were quantified as described above for LCLs. Supplementary Fig. 8 describes our study design and includes N values for each cohort.
Real-time PCR
Real-time quantitative PCR (qPCR) reactions were performed with cDNA samples prepared from 1 μg RNA samples using High Capacity cDNA Reverse Transcription kit (Applied Biosystems, Waltham, MA, USA) containing 10X RT buffer, 10X RT random primers, 25X dNTP mix, RNAse inhibitor and MultiScribe™ Reverse transcriptase. Reverse transcription was performed using a thermal cycler over three steps (25 °C for 10 min, followed by 37 °C for 120 min and 85 °C for 5 min). Real-time PCR reactions were done with 20 μl mixtures containing 20 ng of cDNA, Absolute Blue qPCR ROX mix (Thermo Scientific, MA, USA) and Primers (TaqMan® Gene Expression Assay (Applied Biosystems). GUSB (glucuronidase, beta) was used as reference gene as recommended for transcriptomic analysis of LCLs and for postmortem brain tissues65. TaqMan® Gene Expression Assay IDs are listed below:
Gene Symbol | Assay ID |
|---|---|
GUSB | Hs00939627_m1 |
SIRT1 | Hs01009006_m1 |
RGS2 | Hs01009070_g1 |
PCR reactions were performed using ABI Step One (Applied Biosystems) and the cycle protocol was as follows: 50 °C for 2 min, 95 °C for 15 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min.
For measuring miRNA expression levels, cDNA samples were prepared from 350 ng RNA samples using High Capacity cDNA Reverse Transcription kit (Applied Biosystems, Waltham, MA, USA) in accordance with the protocol for Creating Custom RT Pools using TaqMan ® MicroRNA Assays from Applied Biosystems. Real-time PCR reactions were done with 10 μl mixtures containing 10 ng of cDNA, TaqMan® Fast Advanced Master Mix (Applied Biosystems, Waltham, MA, USA) and Primers (TaqMan™ MicroRNA Assay; Applied Biosystems). U6 snRNA was used as reference gene as recommended for transcriptomic analysis of LCLs66. TaqMan® MicroRNA Assay IDs are listed below:
MicroRNA Symbol | TaqMan™ MicroRNA Assay ID |
|---|---|
U6 snRNA (Control miRNA Assay) | 001973 |
hsa-miR-22-3p | 000398 |
hsa-miR-132-3p | 000457 |
hsa-miR-212-3p | 000515 |
Comparative critical threshold (Ct) values were determined in duplicates for analyzing relative gene and miRNA expression in selected sample groups according to 2−ΔCт (ΔCт = Ct target Gene − Ct reference gene).
Statistical analyses
Statistical analysis for gene and miRNA mean expression in LCLs and postmortem brain tissues was performed by Kruskal-Wallis test (for more than two groups) followed by non-parametric Mann-Whitney U test. Multiple testing corrections were performed by the Benjamini-Hochberg test67; adjusted P-values with a false discovery rate (FDR) < 0.05 were considered significant (Supplementary Table 3). Correlation analysis was performed by either Pearson correlation or Spearman correlation (as indicated in each relevant figure legend). P-values < 0.05 were considered significant. SPSS 23 software (SPSS Inc., Chicago, IL, USA) was used for all statistical analyses.
References
Chibnik, L. B. et al. Trends in the incidence of dementia: design and methods in the Alzheimer Cohorts Consortium. European journal of epidemiology 32, 931–938, https://doi.org/10.1007/s10654-017-0320-5 (2017).
Reed, C. et al. What Drives Country Differences in Cost of Alzheimer’s Disease? An Explanation from Resource Use in the GERAS Study. Journal of Alzheimer’s disease: JAD 57, 797–812, https://doi.org/10.3233/jad-160449 (2017).
Modig, K., Andersson, T., Vaupel, J., Rau, R. & Ahlbom, A. How long do centenarians survive? Life expectancy and maximum lifespan. Journal of internal medicine 282, 156–163, https://doi.org/10.1111/joim.12627 (2017).
Association, A. s. Alzheimer’s disease facts and figures. Alzheimer’s & Dementia 12, 459–509 (2016).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nature genetics 45, 1452–1458 (2013).
Magistri, M., Velmeshev, D., Makhmutova, M. & Faghihi, M. A. Transcriptomics profiling of Alzheimer’s disease reveal neurovascular defects, altered amyloid-β homeostasis, and deregulated expression of long noncoding RNAs. Journal of Alzheimer’s Disease 48, 647–665 (2015).
Sood, S. et al. A novel multi-tissue RNA diagnostic of healthy ageing relates to cognitive health status. Genome biology 16, 185 (2015).
Puca, A. A., Spinelli, C., Accardi, G., Villa, F. & Caruso, C. Centenarians as a model to discover genetic and epigenetic signatures of healthy ageing. Mechanisms of ageing and development (2017).
Hitt, R., Young-Xu, Y., Silver, M. & Perls, T. Centenarians: the older you get, the healthier you have been. The Lancet 354, 652 (1999).
Balistreri, C. R. et al. Genetics of longevity. Data from the studies on Sicilian centenarians. Immunity & Ageing 9, 8 (2012).
Giuliani, C. et al. Centenarians as extreme phenotypes: an ecological perspective to get insight into the relationship between the genetics of longevity and age-associated diseases. Mechanisms of Ageing and Development (2017).
Chevanne, M. et al. Oxidative DNA damage repair and parp 1 and parp 2 expression in Epstein-Barr virus-immortalized B lymphocyte cells from young subjects, old subjects, and centenarians. Rejuvenation research 10, 191–204, https://doi.org/10.1089/rej.2006.0514 (2007).
Attems, J., Walker, L. & Jellinger, K. A. Olfactory bulb involvement in neurodegenerative diseases. Acta neuropathologica 127, 459–475 (2014).
Godoy, M. D., Voegels, R. L., Pinna Fde, R., Imamura, R. & Farfel, J. M. Olfaction in neurologic and neurodegenerative diseases: a literature review. International archives of otorhinolaryngology 19, 176–179, https://doi.org/10.1055/s-0034-1390136 (2015).
Roberts, R. O. et al. Association Between Olfactory Dysfunction and Amnestic Mild Cognitive Impairment and Alzheimer Disease Dementia. JAMA neurology 73, 93–101, https://doi.org/10.1001/jamaneurol.2015.2952 (2016).
Zelaya, M. V. et al. Olfactory bulb proteome dynamics during the progression of sporadic Alzheimer’s disease: identification of common and distinct olfactory targets across Alzheimer-related co-pathologies. Oncotarget 6, 39437–39456, https://doi.org/10.18632/oncotarget.6254 (2015).
Horgusluoglu, E., Nudelman, K., Nho, K. & Saykin, A. J. Adult neurogenesis and neurodegenerative diseases: A systems biology perspective. American journal of medical genetics. Part B, Neuropsychiatric genetics: the official publication of the International Society of Psychiatric Genetics 174, 93–112, https://doi.org/10.1002/ajmg.b.32429 (2017).
Hadar, A. et al. RGS2 expression predicts amyloid-β sensitivity, MCI and Alzheimer’s disease: genome-wide transcriptomic profiling and bioinformatics data mining. Translational psychiatry 6, e909 (2016).
Herskovits, A. Z. & Guarente, L. SIRT1 in neurodevelopment and brain senescence. Neuron 81, 471–483 (2014).
Imai, S.-I., Armstrong, C. M., Kaeberlein, M. & Guarente, L. Transcriptional silencing and longevity protein Sir2 is an NAD-dependent histone deacetylase. Nature 403, 795–800 (2000).
Tissenbaum, H. A. & Guarente, L. Increased dosage of a sir-2 gene extends lifespan in Caenorhabditis elegans. Nature 410, 227–230 (2001).
Satoh, A. et al. Sirt1 extends life span and delays aging in mice through the regulation of Nk2 homeobox 1 in the DMH and LH. Cell metabolism 18, 416–430 (2013).
Kim, D. et al. SIRT1 deacetylase protects against neurodegeneration in models for Alzheimer’s disease and amyotrophic lateral sclerosis. The EMBO journal 26, 3169–3179 (2007).
Kumar, R. et al. Sirtuin1: a promising serum protein marker for early detection of Alzheimer’s disease. PLoS One 8, e61560 (2013).
Qin, W. et al. Neuronal SIRT1 activation as a novel mechanism underlying the prevention of Alzheimer disease amyloid neuropathology by calorie restriction. Journal of Biological Chemistry 281, 21745–21754 (2006).
Tiberi, L. et al. BCL6 controls neurogenesis through Sirt1-dependent epigenetic repression of selective Notch targets. Nature neuroscience 15, 1627–1635 (2012).
Prozorovski, T. et al. Sirt1 contributes critically to the redox-dependent fate of neural progenitors. Nature cell biology 10, 385–394 (2008).
Wang, J. et al. The role of Sirt1: at the crossroad between promotion of longevity and protection against Alzheimer’s disease neuropathology. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics 1804, 1690–1694 (2010).
Howitz, K. T. et al. Small molecule activators of sirtuins extend Saccharomyces cerevisiae lifespan. Nature 425, 191–196, https://doi.org/10.1038/nature01960 (2003).
North, B. J. & Sinclair, D. A. Sirtuins: a conserved key unlocking AceCS activity. Trends in biochemical sciences 32, 1–4, https://doi.org/10.1016/j.tibs.2006.11.002 (2007).
Jeong, H. et al. Sirt1 mediates neuroprotection from mutant huntingtin by activation of the TORC1 and CREB transcriptional pathway. Nature medicine 18, 159–165, https://doi.org/10.1038/nm.2559 (2011).
Zhang, C. et al. Hormetic effect of panaxatriol saponins confers neuroprotection in PC12 cells and zebrafish through PI3K/AKT/mTOR and AMPK/SIRT1/FOXO3pathways. Scientific reports 7, 41082, https://doi.org/10.1038/srep41082 (2017).
Im, H.-I. & Kenny, P. J. MicroRNAs in neuronal function and dysfunction. Trends in neurosciences 35, 325–334 (2012).
Luikart, B. W. et al. miR-132 mediates the integration of newborn neurons into the adult dentate gyrus. PLoS One 6, e19077, https://doi.org/10.1371/journal.pone.0019077 (2011).
Strum, J. C. et al. MicroRNA 132 regulates nutritional stress-induced chemokine production through repression of SirT1. Molecular endocrinology 23, 1876–1884 (2009).
Ramalinga, M. et al. MicroRNA-212 negatively regulates starvation induced autophagy in prostate cancer cells by inhibiting SIRT1 and is a modulator of angiogenesis and cellular senescence. Oncotarget 6, 34446 (2015).
Chen, H. et al. miR-22 inhibits the proliferation, motility, and invasion of human glioblastoma cells by directly targeting SIRT1. Tumor Biology 37, 6761–6768 (2016).
Smith, P. Y. et al. miR-132/212 deficiency impairs tau metabolism and promotes pathological aggregation in vivo. Human molecular genetics 24, 6721–6735, https://doi.org/10.1093/hmg/ddv377 (2015).
Hansen, K. F. et al. Targeted deletion of miR-132/-212 impairs memory and alters the hippocampal transcriptome. Learning & memory (Cold Spring Harbor, N.Y.) 23, 61–71, https://doi.org/10.1101/lm.039578.115 (2016).
Weinberg, R. B., Mufson, E. J. & Counts, S. E. Evidence for a neuroprotective microRNA pathway in amnestic mild cognitive impairment. Frontiers in neuroscience 9, 430, https://doi.org/10.3389/fnins.2015.00430 (2015).
Tavolaro, S. et al. Increased chronic lymphocytic leukemia proliferation upon IgM stimulation is sustained by the upregulation of miR‐132 and miR‐212. Genes, Chromosomes and Cancer 54, 222–234 (2015).
Araki, T., Sasaki, Y. & Milbrandt, J. Increased nuclear NAD biosynthesis and SIRT1 activation prevent axonal degeneration. Science 305, 1010–1013 (2004).
Nimmagadda, V. K. et al. Overexpression of SIRT1 protein in neurons protects against experimental autoimmune encephalomyelitis through activation of multiple SIRT1 targets. The Journal of Immunology 190, 4595–4607 (2013).
Min, S.-W. et al. SIRT1 Deacetylates Tau and Reduces Pathogenic Tau Spread in a Mouse Model of Tauopathy. Journal of Neuroscience 38, 3680–3688 (2018).
Kilic, U. et al. A remarkable age-related increase in SIRT1 protein expression against oxidative stress in elderly: SIRT1 gene variants and longevity in human. PloS one 10, e0117954, https://doi.org/10.1371/journal.pone.0117954 (2015).
Owczarz, M. et al. miR-34a and miR-9 are overexpressed and SIRT genes are downregulated in peripheral blood mononuclear cells of aging humans. Experimental Biology and Medicine 242, 1453–1461 (2017).
Gurwitz, D. Human iPSC-derived neurons and lymphoblastoid cells for personalized medicine research in neuropsychiatric disorders. Dialogues Clin Neurosci 18, 267–276 (2016).
Xie, B. et al. Serum miR-206 and miR-132 as Potential Circulating Biomarkers for Mild Cognitive Impairment. Journal of Alzheimer’s disease: JAD 45, 721–731, https://doi.org/10.3233/jad-142847 (2015).
Cogswell, J. P. et al. Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. Journal of Alzheimer’s disease 14, 27–41 (2008).
Pichler, S. et al. The miRNome of Alzheimer’s disease: consistent downregulation of the miR-132/212 cluster. Neurobiology of aging 50, 167. e161–167. e110 (2017).
Amirkhah, R., Meshkin, H. N., Farazmand, A., Rasko, J. E. & Schmitz, U. Computational and Experimental Identification of Tissue-Specific MicroRNA Targets. MicroRNA Detection and Target Identification: Methods and Protocols, 127–147 (2017).
Blazie, S. M. et al. Alternative Polyadenylation Directs Tissue-Specific miRNA Targeting in Caenorhabditis elegans Somatic Tissues. Genetics 206, 757–774 (2017).
Magill, S. T. et al. microRNA-132 regulates dendritic growth and arborization of newborn neurons in the adult hippocampus. Proceedings of the National Academy of Sciences 107, 20382–20387 (2010).
Pathania, M. et al. miR-132 enhances dendritic morphogenesis, spine density, synaptic integration, and survival of newborn olfactory bulb neurons. PloS one 7, e38174 (2012).
Mukilan, M., Varman, D. R., Sudhakar, S. & Rajan, K. E. Activity-dependent expression of miR-132 regulates immediate-early gene induction during olfactory learning in the greater short-nosed fruit bat, Cynopterus sphinx. Neurobiology of learning and memory 120, 41–51 (2015).
Skoog, I. et al. 15-year longitudinal study of blood pressure and dementia. The Lancet 347, 1141–1145 (1996).
Hadar, A., Gozes, I. & Gurwitz, D. in Neuroprotection in Alzheimer’s Disease (ed I. Gozes) 239-251 (Academic Press, 2017).
Eskildsen, T. V. et al. Angiotensin II regulates microRNA-132/-212 in hypertensive rats and humans. International journal of molecular sciences 14, 11190–11207, https://doi.org/10.3390/ijms140611190 (2013).
Pavlov, V. A. & Tracey, K. J. The cholinergic anti-inflammatory pathway. Brain, behavior, and immunity 19, 493–499 (2005).
Miyazaki, Y. et al. A novel microRNA-132-surtuin-1 axis underlies aberrant B-cell cytokine regulation in patients with relapsing-remitting multiple sclerosis. Plos one 9, e105421 (2014).
Ekdahl, C. T., Claasen, J.-H., Bonde, S., Kokaia, Z. & Lindvall, O. Inflammation is detrimental for neurogenesis in adult brain. Proceedings of the National Academy of Sciences 100, 13632–13637 (2003).
Kong, H. et al. The effect of miR-132, miR-146a, and miR-155 on MRP8/TLR4-induced astrocyte-related inflammation. Journal of Molecular Neuroscience 57, 28–37 (2015).
Oved, K. et al. Genome-wide expression profiling of human lymphoblastoid cell lines implicates integrin beta-3 in the mode of action of antidepressants. Transl Psychiatry 3, e313, https://doi.org/10.1038/tp.2013.86 (2013).
Montine, T. J. et al. National Institute on Aging–Alzheimer’s Association guidelines for the neuropathologic assessment of Alzheimer’s disease: a practical approach. Acta neuropathologica 123, 1–11 (2012).
de Brouwer, A. P., van Bokhoven, H. & Kremer, H. Comparison of 12 reference genes for normalization of gene expression levels in Epstein-Barr virus-transformed lymphoblastoid cell lines and fibroblasts. Molecular diagnosis & therapy 10, 197–204 (2006).
Oved, K. et al. Genome-wide miRNA expression profiling of human lymphoblastoid cell lines identifies tentative SSRI antidepressant response biomarkers. Pharmacogenomics 13, 1129–1139, https://doi.org/10.2217/pgs.12.93 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. Journal of the royal statistical society. Series B (Methodological), 289–300 (1995).
Acknowledgements
This study was supported by the Israel Science Foundation to IG and DG (Grant #1424/14). IG was also supported by the AMN Foundation, Gildor Chair, and the Elton Laboratory. EM is recipient of a contract of the European Regional Development Fund, Competitiveness Operational Program 2014–2020, through the grant P_37_732/2016 REDBRAIN. Tissue for this study was provided by the Newcastle Brain Tissue Resource, which is funded in part by a grant from the UK Medical Research Council (G0400074), by Brains for Dementia research, a joint venture between Alzheimer’s Society and Alzheimer’s Research UK and by the NIHR Newcastle Biomedical Research Centre awarded to the Newcastle upon Tyne Hospitals NHS Foundation Trust and Newcastle University. We thank the anonymous donors of the NLGIP Biobank at Tel Aviv University, Israel, whose altruism and trust in biomedical research have made this study possible. This study is in partial fulfillment for the graduate study requirements for AH at the Dr. Miriam and Sheldon G. Adelson Graduate School of Medicine, Sackler Faculty of Medicine, Tel Aviv University.
Author information
Authors and Affiliations
Contributions
D.G. and I.G. conceived the study. A.H. conducted the real-time PCR experiments, analysed the results, and prepared the figures and tables. E.M. and M.W. assisted with RNA extractions. M.P.K., J.K., A.S., P.N., and C.C. provided human lymphoblastoid cell lines and associated patient data. J.A. provided postmortem tissue samples and associated patient data. A.H., D.G., and I.G. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hadar, A., Milanesi, E., Walczak, M. et al. SIRT1, miR-132 and miR-212 link human longevity to Alzheimer’s Disease. Sci Rep 8, 8465 (2018). https://doi.org/10.1038/s41598-018-26547-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26547-6
This article is cited by
-
Knockout of the longevity gene Klotho perturbs aging and Alzheimer’s disease-linked brain microRNAs and tRNA fragments
Communications Biology (2024)
-
Therapeutic Targeting of Krüppel-Like Factor 4 and Its Pharmacological Potential in Parkinson’s Disease: a Comprehensive Review
Molecular Neurobiology (2024)
-
The expression profile of brain-derived exosomal miRNAs reveals the key molecules responsible for spontaneous motor function recovery in a rat model with permanent middle cerebral artery occlusion
Mammalian Genome (2024)
-
Mind the Gap: Unraveling the Intricate Dance Between Alzheimer’s Disease and Related Dementias and Bone Health
Current Osteoporosis Reports (2024)
-
Chromatin remodeler Activity-Dependent Neuroprotective Protein (ADNP) contributes to syndromic autism
Clinical Epigenetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.