Abstract
Substrates of pleiotropic drug resistance (PDR) transporters can induce the expression of corresponding transporter genes by binding to their transcription factors. Penetrating cations are substrates of PDR transporters and theoretically may also activate the expression of transporter genes. However, the accumulation of penetrating cations inside mitochondria may prevent the sensing of these molecules. Thus, whether penetrating cations induce PDR is unclear. Using Saccharomyces cerevisiae as a model, we studied the effects of penetrating cations on the activation of PDR. We found that the lipophilic cation dodecyltriphenylphosphonium (C12TPP) induced the expression of the plasma membrane PDR transporter genes PDR5, SNQ2 and YOR1. Moreover, a 1-hour incubation with C12TPP increased the concentration of Pdr5p and Snq2p and prevented the accumulation of the PDR transporter substrate Nile red. The transcription factor PDR1 was required to mediate these effects, while PDR3 was dispensable. The deletion of the YAP1 or RTG2 genes encoding components of the mitochondria-to-nucleus signalling pathway did not prevent the C12TPP-induced increase in Pdr5-GFP. Taken together, our data suggest (i) that the sequestration of lipophilic cations inside mitochondria does not significantly inhibit sensing by PDR activators and (ii) that the activation mechanisms do not require mitochondria as a signalling module.
Similar content being viewed by others
Introduction
Living cells are capable of resisting toxic compounds due to the activity of plasma membrane pumps with broad substrate specificity. These pumps extrude various chemically unrelated molecules from the cytoplasm to the media (see reviews)1,2. Such a protection requires an energy investment because such pumps consume ATP. Accordingly, Saccharomyces cerevisiae cells with inactivated pleiotropic drug resistance (PDR) ABC transporters show increased ATP concentrations and reach stationary phase cell densities that are higher than those of the wild-type cells3. Moreover, under normal conditions, the overexpression of a major pleiotropic drug resistance ABC transporter PDR5 decreases the fitness of the strain4. For this reason, cells benefit from repressing PDR pump genes under normal conditions and activating them only upon exposure to xenobiotics. PDR pump substrates have been shown to induce the expression of the corresponding genes in yeast5,6,7,8 and animals9. For instance, treating rat liver cells with the MDR (multiple drug resistance) transporter substrate doxorubicin upregulates the expression of the corresponding P-glycoprotein gene9. 2-methyl-4-chlorophenoxyacetic has also been shown to be a substrate of yeast pleiotropic pumps (Pdr5p and Tpo1p), inducing the expression of the corresponding genes5. The induction of the ABC transporters genes PDR5 and YOR1 was observed in response to atorvastatin treatment8. The most striking example is cycloheximide D, an inhibitor of protein synthesis and a Pdr5p substrate10. At low concentrations, cycloheximide D can induce the expression of PDR5 in yeast by binding directly to the upstream transcription factor Pdr1p11.
Another mechanism of pleiotropic drug resistance pumps activation that relies on mitochondria was revealed for some yeast species12,13,14. In Saccharomyces cerevisiae, the loss of mitochondrial DNA or the deletion of nuclear encoded genes essential for mitochondrial functioning (e.g., FZO1) upregulates unspecific drug resistance15,16,17. This effect requires the Pdr3p transcription factor and the components of the mitochondria-to-nucleus retrograde signalling pathway RTG1 and RTG215. However, as MDR pumps consume ATP, energy deprivation caused by inhibiting mitochondria can prevent the extrusion of xenobiotic agents from the cytoplasm. Indeed, the assay for MDR pump activity includes a step of energy deprivation to ensure the maximum absorption of the MDR substrates by the cells18.
Lipophilic cations are substrates of MDR pumps18,19,20 and a potentially promising class of molecules for pharmacological applications. Alkylated modifications of such molecules are used to produce chimeric compounds to deliver drugs to the mitochondrial matrix (e.g., antioxidants)21,22. Dodecyltriphenylphosphonium (C12TPP) has been found to act as an uncoupler, being co-transported with a free fatty acid23. The uncoupling activity allows to use this chemical as an anti-obesity drug in murine models24. Moreover, lipophilic cations including C12TPP appear to be competitive inhibitors of MDR25,26,27. However, whether such compounds can induce the expression of MDR genes is unclear. On the one hand, as substrates of major ABC transporters, these compounds might bind Pdr1p/Pdr3p transcription factors and activate the transcription of the transporters. Moreover, the accumulation of C12TPP in the mitochondria can induce the inhibition of mitochondrial enzymes28 or induce ROS accumulation29, which in turn can potentially activate the RTG-dependent activation of PDR genes. On the other hand, the sequestration of the lipophilic cations in the mitochondria may prevent their possible interactions with Pdr1p/Pdr3p transcription factors. The latter can inhibit the activation of MDR. In this work, we studied the effects of the alkylated lipophilic cation C12TPP on the activation of pleiotropic drug resistance system in yeast.
We found that C12TPP induces MDR in yeast. This activation requires the Pdr1p transcription factor, whereas the deletion of the PDR3 gene or the genes encoding components of the mitochondria-to-nucleus retrograde signalling pathways did not abolish the induction.
Results
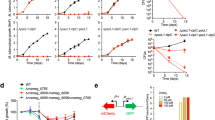
We have shown recently that alkylated rhodamines are substrates of S. cerevisiae ABC transporters27. In a subsequent set of experiments, we observed that upon transfer from a solid medium to a liquid medium containing C12R1 (dodecylrhodamine 19), the cells first accumulated the dye before the amount of accumulated dye began to decrease (Fig. 1). This effect cannot be attributed to a decrease in the C12R1/cell ratio because there was only a modest increase in cell density during the first 180 minutes after the inoculation of yeast cells in the liquid medium (Figure S1). A lag interval preceding a decrease suggests that the observed decrease in the C12R1 signal could be due to the transcriptional or translational activation of pleiotropic efflux pumps. To test this possibility, we measured the dynamics of C12R1 accumulation in a strain with deleted PDR1 and PDR3 transcription factors. We did not detect any decrease in the C12R1 levels in this strain (Fig. 1).
The dynamics of C12R1 accumulation. Yeast cells were resuspended at 2*104 cells/ml in fresh YPD containing dodecylrhodamine (C12R1) at 200 nM. The C12R1 concentration was assessed using FACS (see Materials and Methods). The graphs show averages of four (WT, wild-type) or five (Δpdr1Δpdr3) biological replicates with standard deviations. *P = 0.017 for comparisons with WT according to the unpaired Mann-Whitney test.
As C12R1 fluorescence has partly overlapping emission spectra with GFP, we used the non-fluorescent lipophilic cation dodecyltriphenylphosphonium (C12TPP) for further experiments. We found that C12TPP induces an accumulation of two major pleiotropic drug resistance ABC transporters: Pdr5p and Snq2p (Fig. 2A). The signal of Yor1-GFP was relatively low (Figure S2); therefore, we could not draw any conclusions about this transporter at this stage. The increase in Pdr5-GFP levels was quantified by means of flow cytometry (Fig. 2B). In line with the C12R1 accumulation experiments, the Pdr5-GFP levels started to increase after approximately 30 minutes of incubation with C12TPP (Fig. 2C). Importantly, the C12TPP-induced increase in the Pdr5-GFP levels was not due to an increase in the cell volume or cell surface area, as the cell size was identical in the C12TPP-treated and the control yeast cells (Figure S3). Next, we measured the relative mRNA levels of three transporter genes — PDR5, SNQ2 and YOR1— using quantitative reverse transcription PCR (RT-qPCR). We found that C12TPP increased the mRNA levels of these genes in the wild-type strain, while the effect in the Δpdr1Δpdr3 strain was much less pronounced (Fig. 3). This result suggests that the C12TPP-induced accumulation of ABC transporters is a result of the transcriptional activation of the corresponding genes.
C12TPP induces the accumulation of the major MDR proteins Pdr5 and Snq2. (A) Accumulation of Pdr5-GFP and Snq2-GFP induced by C12TPP. The incubation time was 1 hour. All fluorescence microphotographs were taken with the same exposure time and contrast adjustments, with the exception of the ‘enhanced contrast’ panels. Green is a pseudocolour. (B) Representative histograms of flow cytometry experiments with yeast cells expressing Pdr5-GFP. The numbers above the arrows indicate the concentrations (µM) of supplemented C12TPP. The incubation time was 1 hour. (C) Quantification by flow cytometry of Pdr5-GFP accumulation induced by C12TPP. The average integral intensities of Pdr5-GFP fluorescence with the deduction of autofluorescence of GFP-negative cells were measured.
C12TPP (2.5 µM, 1 hours of incubation) increases the mRNA levels of PDR genes PDR5, SNQ2 and YOR1. All mRNA levels were normalized to ACT1 levels, and the value in untreated wild-type cells (WT) was set at 100%. The bars show the mean ± standard errors, n = 4. *P = 0.028 for comparisons to an untreated control, according to the unpaired Mann-Whitney test.
Does the accumulation of ABC proteins induced by C12TPP indeed lead to the increased efflux of the transporter substrates? To answer this question, we studied the effect of preconditioning with C12TPP on the accumulation of the ABC transporter non-charged fluorescent substrate Nile red. This compound is a substrate of pleiotropic transporters of Candida albicans30. We found that SNQ2 gene overexpression significantly decreased the accumulation of Nile red in S. cerevisiae cells, while its repression had the opposite effect (Figure S4). The repression of the ABC transporter gene PDR5 also increased the accumulation of Nile red in the cells (Figure S4). We found that the preincubation of yeast cells with C12TPP inhibited Nile red accumulation in cells (Fig. 4A–D). At the same time, in line with our previous study26, we noticed that the simultaneous supplementation of Nile red and C12TPP to yeast incubation media facilitated the uptake of Nile red by cells (Fig. 4B–D). Thus, C12TPP plays both inhibitory and activating roles in MDR regulation, and the contribution of each role changes with the duration of cellular exposure to the chemical.
Preincubation with C12TPP prevents the accumulation of the MDR substrate Nile red. (A) Preincubation with 2.5 µM C12TPP for 1 hour decreases the accumulation of Nile red in yeast cells. Red is a pseudocolour. (B) The design of the experiment. (C) Representative histograms of flow cytometry experiments of Nile red accumulation in yeast cells. C12TPP (2.5 µM) was added either before or simultaneously with Nile red (3.5 µM). (D) Quantification of flow cytometry results (average ± standard deviation). Here, 100% corresponds to the average fluorescence (level) of Nile red in the control samples. The numbers of separate experiments are indicated below the bars. P values were calculated for comparison with the untreated control (WT) according to the unpaired Mann-Whitney test with Bonferroni adjustments.
What is the mechanism of C12TPP-dependent MDR upregulation? The lipophilic cations can inhibit respiration28 and facilitate proton leakage23,31. At the same time, dysfunctional mitochondria upregulate mitochondria-to-nucleus signalling mediated by Rtg-proteins, and such signalling has been shown to upregulate PDR515. Thus, we tested the role of the retrograde pathway in C12TPP-induced resistance using the non-charged lipophilic fluorescent dye Nile red. The inhibition of the pathway by RTG2 knockdown did not prevent the C12TPP-induced increase in Pdr5-GFP levels in yeast cells (Fig. 5A). The release of mitochondrially produced hydrogen peroxide into the cytoplasm is another possible signalling pathway from dysfunctional mitochondria to the nucleus (for review, see32). This pathway relies on cytoplasmic Yap1p33, the transcription factor activated under oxidative stress34. While the primary targets of Yap1p are antioxidant genes35, Yap1p may also contribute to the activation of PDR genes36. To test the role of Yap1p in the C12TPP activation of MDR, we produced the Δyap1 Pdr5-GFP strain. Taking into account the possible crosstalk between the Yap1p and Rtg pathways,33 we tested the activation in the double Δyap1Δrtg2 mutant. We found that C12TPP was capable of upregulating the Pdr5-GFP levels in all tested strains with disrupted mitochondria-to-nucleus signalling (Fig. 5A). Thus, we concluded that YAP1 does not play a role in C12TPP-induced PDR. Next, we tested the contribution of the transcription factors PDR1 and PDR3 in the C12TPP-dependent upregulation of Pdr5-GFP levels. We found that the deletion of PDR1 but not PDR3 inhibited this effect (Fig. 5A). The accumulation of Snq2-GFP was also abolished in the Δpdr1 and Δpdr1Δpdr3 strains (Figure S5). Likewise, C12TPP was able to induce a decrease in Nile red levels in the Δyap1, Δrtg1, Δrtg3, Δrtg2, Δyap1Δrtg2, and Δpdr3 strains but displayed no effect if the PDR1 gene was deleted (Fig. 5B). We also showed that repressing the SNQ2 gene prevented the C12TPP-induced decrease in Nile red accumulation (Fig. 5C). This result suggests that the Snq2p ABC transporter is involved in the C12TPP-mediated induction of MDR. However, the repression of PDR genes affects the accumulation of both C12TPP (inducer) and Nile red (sensor). Therefore, the repression of a transporter gene can increase C12TPP concentrations in the cells and, in this way, facilitate a stronger compensatory response by other genes. Thus, the negative result with the PDR5 and YOR1 genes does not exclude the possibility that these genes contribute to C12TPP-mediated MDR activation. Notably, the overexpression of PDR1 targets SNQ2 and PDR5 abolished the effect of C12TPP on Nile red accumulation (Figure S6). This observation suggests that the lipophilic cation needs to accumulate inside the cells to activate MDR. Alternatively, ABC transporter overexpression can be sufficient to drive the maximal efflux rate. However, we believe the latter explanation is unlikely because the addition of C12TPP to the wild-type cells lowered the Nile red levels much more than in the PGAL strains.
The role of retrograde signalling and PDR genes in the C12TPP-induced activation of PDR. (A) Pdr5-GFP levels in knockout yeast strains grown in YPD and treated with C12TPP (2.5 µM, 1 hour). (B) C12TPP-induced (2.5 µM, 1 hour) decreases in Nile red levels in the knockout yeast strains. (C) C12TPP-induced (2.5 µM, 1 hour) decreases in Nile red levels in yeast strains with repressed ABC transporter genes SNQ2, PDR5 or YOR1. (A–C) Cells were grown in YPD medium (the condition of PGAL repression). Data are shown as the mean ± standard error. Numbers of biological replicates are indicated below the bars. P values were calculated for comparisons with the untreated control (WT) according to unpaired Mann-Whitney test with Bonferroni adjustments for multiple comparisons.
Lipophilic cations, as positively charged molecules, accumulate in negatively charged cellular organelles — mitochondria37. We expected that the accumulation of such compounds in the mitochondria would mitigate the interaction of the cations with their sensors in the cytoplasm. To test this, we compared the activation of MDR in different growth media. In the presence of high glucose concentrations, yeast cells suppress mitochondrial functions, while in the presence of a poorly fermentable carbon source (galactose) or non-fermentable carbon source (glycerol), yeast cells rely on mitochondrial energetics38. We tested the effect of C12TPP on Pdr5-GFP levels in yeast cells in rich medium supplemented with either galactose or glycerol. We found that the inoculation of yeast cells in YPGal medium supplemented with C12TPP triggered the accumulation of Pdr5-GFP. At the same time, there was no increase in Pdr5-GFP levels in the glycerol (Fig. 6A). The uncoupling23,31 or inhibitory28 effects of C12TPP in the mitochondria may deplete cellular ATP and prevent Pdr5-GFP protein synthesis. In agreement with this hypothesis, myxothiazol (an inhibitor of the mitochondrial coenzyme Q - cytochrome c reductase) abolished the induction of Pdr5p-GFP upregulation induced by azole antifungal clotrimazole in glycerol-based medium (Figure S7).
Mitochondrial depolarization does not facilitate C12TPP-induced Pdr5-GFP accumulation. (A) The effect of a carbon source on the C12TPP-induced (2.5 µM, 1 hour) change in the Pdr5-GFP level. (B) Myxothiazol at 7 µM and oligomycin A at 5 µg/ml admixture increases the cytoplasmic (diffuse) signal from the lipophilic cation C12R1 (200 nM) in wild-type yeast cells. Red is a pseudocolour. (C,D) Quantification of Pdr5-GFP level in cells (mean ± standard deviation) from flow cytometry experiments with myxothiazol and/or oligomycin A for yeast cells grown in YPD (C, n = 5) or YPGal (D, n = 6). Clotrimazol was added to a final concentration of 20 µM, with a 1-hour incubation time. P values were calculated according to a Wilcoxon signed-rank test with Bonferroni adjustments.
Can mitochondrial depolarization increase the C12TPP-dependent activation of PDR in our experimental model? This would be possible if the equilibration in the lipophilic cation concentrations between the cell and incubation medium was relatively slow (Fig. 1). Thus, the intracellular distribution of the cation is likely to affect the detection of the molecule. If so, an inhibition of the respiratory chain can, due to mitochondrial depolarization, prevent accumulation of the lipophilic cation in the mitochondrial matrix, subsequently increasing the activation of MDR. First, we tested whether the inhibition of the respiratory chain by myxothiazol increases the cytoplasmic levels of the fluorescent lipophilic cation C12R1. For glucose-supplemented rich medium, we used a mixture of myxothiazol and an inhibitor of ATP-synthase, oligomycin A, to prevent the energization of mitochondria by ATP hydrolysis driven by mitochondrial ATP-synthase. We used a concentration of the inhibitors sufficient to inhibit the growth rate of the control laboratory strain in the presence of a non-fermentable carbon source39. We found that the mixture of the inhibitors (“depolarization cocktail”) increased the cytoplasmic signal of C12R1 in yeast cells (Fig. 6B). However, the mitochondrial depolarization did not upregulate the C12TPP-induced increase in Pdr5-GFP levels in either glucose-based (Fig. 6C) or galactose-based medium (Fig. 6D). We suggested that the addition of 2.5 µM of C12TPP causes the maximum possible activation of Pdr5-GFP expression; therefore, we tested lower concentrations of the chemical. However, there was no significant increase in Pdr5-GFP levels in the presence of 1 µM C12TPP. As oligomycin inhibits Pdr5p40 and, therefore, may interfere with the compensatory activation of PDR genes, we tested the C12TPP-induced Pdr5-GFP accumulation in the cells treated with sodium azide. Sodium azide inhibits respiration as well as ATPase activity. In line with our previous results, C12TPP did not activate MDR in the presence of NaN3 (Figure S8). Together, these data suggest that mitochondrial depolarization does not facilitate lipophilic cation-dependent MDR activation or that this activation takes place at a relatively narrow range of the conditions.
Discussion
The exposure of yeast cells to xenobiotics favours the selection of cells with upregulated PDR gene expression41. Accordingly, the activation of pleiotropic drug resistance transporter genes decreases the efficiency of azole antifungal treatments42. Therefore, the inhibition of the PDR transporters appears to be a promising strategy to increase the treatment efficiency of emerging multiple drug resistant pathogenic fungal strains. Some cationic amphiphilic compounds, the substrates of PDR transporters, were shown to sensitize yeast cells to azole antifungals26,27,43,44,45. However, the accumulation of an MDR substrate in the cell upregulates the expression of PDR transporters genes5,11, which can significantly limit the application of this approach. At the same time, the mechanism of PDR upregulation is not absolutely clear.
In our study we tested the ability of the lipophilic cation C12TPP to activate multiple drug resistance in Saccharomyces cerevisiae. We expected that as a positively charged molecule, C12TPP could accumulate in the mitochondria and thus evade sensing by the cytoplasmic/nuclear transcription factors PDR1 and PDR3. However, C12TPP induced an increase in the levels of both PDR5 mRNA and Pdr5-protein and, at the same time, prevented the accumulation of Nile red, the neutral fluorescent substrate of PDR transporters (Figs 4 and 5). Moreover, the depolarization of the mitochondria, which inhibits sequestration of C12TPP inside the mitochondria, did not upregulate C12TPP-induced MDR activation (Fig. 6). At the same time, the presence of the functional transcription factor gene PDR1 was necessary for the effects of C12TPP (Fig. 5). We speculate that there is a concentration window for each toxic chemical within which the chemical upregulates cellular protection mechanisms. This window is limited from above by the energetic capacity of the cell, i.e., the excess amount of a metabolic inhibitor can prevent the protective response at the transcriptional or translational level. From below, the window is limited by the specificity of the sensing factors. Thus, the accumulation in the mitochondria may prevent the activation of MDR by C12TPP in a relatively narrow concentration range. Although this possibility cannot be excluded with our experiments, we have shown that mitochondrial depolarization does not induce the accumulation of Pdr5-GFP in yeast cells in the presence of relatively low (1 µM) concentrations of C12TPP (Fig. 6C,D). Importantly, we proved that under the same conditions (the combination of respiratory chain inhibitor myxothiazol and inhibitor of ATP-synthase oligomycin A), the azole antifungal clotrimazole still induced Pdr5-GFP accumulation (Fig. 6C,D).
How can a cell sense the xenobiotics that accumulate in the mitochondria? One of the known mechanisms of PDR activation relies on direct interaction with the transcription factor Pdr1p11. Therefore, the binding to the transcription factor competes with the mitochondrial uptake of the lipophilic cation. If the affinity of Pdr1p to C12TPP is sufficiently high, the Pdr1p targets will be activated even in the presence of highly energized mitochondria. However, we think that this explanation is unlikely because C12TPP in low concentrations is not able to induce MDR (Fig. 6C,D). Alternatively, the substrates of MDR can indirectly activate PDR1 by inhibiting the efflux of endogenous substrates of PDR transporters. Such substrates might activate the expression of the transporters by the aforementioned mechanism. There are data suggesting that Pdr5p and Snq2p can export a metabolite that acts as a quorum-sensing factor in yeast suspension cultures46,47. In this case, the cellular localization of C12TPP could play a minor role because the insufficient activity of MDR will cause accumulation of the endogenous metabolites, thus activating MDR.
Another possible way of sensing the xenobiotics is to measure some vital cellular metabolites or the energy level. For instance, Rtg2p binding to downstream components of the retrograde signalling cascade Mks1p is regulated by the ATP level48. Thus, the retrograde signalling activation of PDR genes may integrate different pathways via the ATP level. However, in our experiments, the deletion of RTG2 did not prevent the C12TPP-induced activation of Pdr5-GFP accumulation.
To summarize, we have shown that the penetrating lipophilic cation C12TPP upregulates pleiotropic drug resistant ABC transporters in yeast Saccharomyces cerevisiae. The activation mechanism includes the transcription factor PDR1 but not PDR3, RTG2 or YAP1. Mitochondrial depolarization induced by the uncouplers did not facilitate C12TPP-induced multiple drug resistance activation. Taken together, our data suggest that the sensing of pleiotropic drug resistance in yeast cells is robust to variations in xenobiotic localization.
Materials and Methods
Strains and growth conditions
In this study, we used strains with W303 or BY4741 genetic backgrounds. Their derivatives are listed in Table 1. In the mutant strains, the genomic copies of ABC transporter genes were under the control of PGAL. The PGAL promoter was induced by the supplementation of galactose and repressed by glucose. To delete the complete open reading frames, we amplified the deletion cassettes from Invitrogen yeast deletion collection strains with the gene-specific primers. Double and triple mutants were produced by crossing the corresponding single mutants and subsequent tetrad dissection. All strains were verified by PCR with primers that were able to yield product only with the correct insertions. PDR5-GFP, SNQ2-GFP and YOR1-GFP strains were from Invitrogen GFP collection49. Rich growth medium YPD (yeast peptone D-glucose), YPRafGal (yeast peptone raffinose galactose), YPGal (yeast peptone galactose) and YPGly (yeast peptone glycerol) were prepared according to Sherman50. Yeast extract was obtained from BD; bactoagar, bactopeptone and D-glucose were from Amresco. Dodecyltriphenylphosphonium (C12R1) and dodecylrhodamine 19 (C12R1) were previously synthesized in our institute as bromide salts51,52, and the other chemicals were obtained from Sigma-Aldrich.
Fluorescent microscopy
To study the accumulation of GFP proteins, we used the fluorescence microscope Olympus BX41 with the U-MNIBA3 (excitation wavelength 470–495 nm; beamsplitter filter 505 nm; emission 510–550 nm) filter set. The accumulation of C12R1 and Nile red was visualized with the U-MNG2 (excitation 530–550 nm, beamsplitter filter 570 nm; emission >590 nm) filter set. All results were reproduced in at least three biological replicates.
Flow cytometry
Fluorescence of GFP was assessed with a CytoFlex (Beckman) flow cytometer using an excitation wavelength of 488 nm on the emission filter (525/40 nm). The accumulation of Nile red was measured with an emission filter (585/42 nm). Bar plots represent the average of averages of cell populations from separate biological experiments. At least 10 000 events were analysed in each experiment.
Quantitative reverse transcription PCR (RT-qPCR) analysis
RNA was isolated from yeast cells using the hot formamide extraction method described in53. RNA samples were isolated independently four times on separate days. RNA quality and quantity were assessed by electrophoresis and spectrophotometry. cDNA was synthesized by annealing 2 μg of RNA with 0.1 μg of random hexamers and 0.1 μg of Oligo-dT using Superscript III reverse-transcriptase (Thermo Fisher Scientific) for 1 hour at 44 °C. RT-qPCR was carried out using the CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA). We used primer sequences for PDR genes54. For the detection of the target genes, the Eva Green master mix (Syntol, Russia) was used according to the manufacturer’s instructions. The thermal profile for EVA Green RT-qPCR included an initial heat-denaturing step at 95 °C for 3 minutes, 40 cycles at 95 °C for 15 sec, an annealing step for 30 sec and 72 °C for 30 sec, coupled with fluorescence measurements. Following amplification, the melting curves of PCR products were monitored to determine the specificity of the amplification. Each sample was analysed in triplicate, and a non-template control was added to each run. The PCR efficiency (E) was calculated according to the equation E = 10(−1/slope) by performing the standard curves. Target mRNA levels were normalized to the reference gene ACT1.
GFP accumulation assay
Cells with GFP expression were grown overnight in solid medium (YPD, YPGal or YPGly) and then resuspended to a density of 2*104 cells/ml in the same medium. Fluorescence was assessed with a flow cytometer after 1 hour of preincubation with the inhibitors (or solvent) at 30 °C.
Nile red accumulation assay
To assess Nile red accumulation, we grew yeast cells overnight in the corresponding rich growth medium and resuspended the cells to a density of 2*104 cells/ml in the same medium. After 1 hour of preincubation with the inhibitors (or solvent) at 30 °C, Nile red was added to a final concentration of 3.5 μM. After a 10-minute incubation at 30 °C, the amount of accumulated Nile red in the yeast cells was measured by flow cytometry.
Data analysis
Data were analysed with a non-parametric Wilcoxon-Mann-Whitney test using R. The Bonferroni adjustment for multiple comparisons was applied where appropriate.
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Prasad, R. & Goffeau, A. Yeast ATP-binding cassette transporters conferring multidrug resistance. Annu. Rev. Microbiol. 66, 39–63 (2012).
Jungwirth, H. & Kuchler, K. Yeast ABC transporters–a tale of sex, stress, drugs and aging. FEBS Lett. 580, 1131–1138 (2006).
Krasowska, A., Łukaszewicz, M., Bartosiewicz, D. & Sigler, K. Cell ATP level of Saccharomyces cerevisiae sensitively responds to culture growth and drug-inflicted variations in membrane integrity and PDR pump activity. Biochem. Biophys. Res. Commun. 395, 51–55 (2010).
Hull, R. M., Cruz, C., Jack, C. V. & Houseley, J. Environmental change drives accelerated adaptation through stimulated copy number variation. PLoS Biol. 15, e2001333 (2017).
Teixeira, M. C. & Sá-Correia, I. Saccharomyces cerevisiae resistance to chlorinated phenoxyacetic acid herbicides involves Pdr1p-mediated transcriptional activation of TPO1 and PDR5 genes. Biochem. Biophys. Res. Commun. 292, 530–537 (2002).
Lucau-Danila, A. et al. Early expression of yeast genes affected by chemical stress. Mol. Cell. Biol. 25, 1860–1868 (2005).
Alenquer, M., Tenreiro, S. & Sá-Correia, I. Adaptive response to the antimalarial drug artesunate in yeast involves Pdr1p/Pdr3p-mediated transcriptional activation of the resistance determinants TPO1 and PDR5. FEMS Yeast Res. 6, 1130–1139 (2006).
Yibmantasiri, P., Bircham, P. W., Maass, D. R., Bellows, D. S. & Atkinson, P. H. Networks of genes modulating the pleiotropic drug response in Saccharomyces cerevisiae. Mol. Biosyst. 10, 128–137 (2014).
Fardel, O., Lecureur, V., Daval, S., Corlu, A. & Guillouzo, A. Up-regulation of P-glycoprotein expression in rat liver cells by acute doxorubicin treatment. Eur. J. Biochem. 246, 186–192 (1997).
Leppert, G. et al. Cloning by gene amplification of two loci conferring multiple drug resistance in Saccharomyces. Genetics 125, 13–20 (1990).
Thakur, J. K. et al. A nuclear receptor-like pathway regulating multidrug resistance in fungi. Nature 452, 604–609 (2008).
Sanglard, D., Ischer, F. & Bille, J. Role of ATP-binding-cassette transporter genes in high-frequency acquisition of resistance to azole antifungals in Candida glabrata. Antimicrob. Agents Chemother. 45, 1174–1183 (2001).
Bouchara, J. P. et al. In-vivo selection of an azole-resistant petite mutant of Candida glabrata. J. Med. Microbiol. 49, 977–984 (2000).
Devaux, F., Carvajal, E., Moye-Rowley, S. & Jacq, C. Genome-wide studies on the nuclear PDR3-controlled response to mitochondrial dysfunction in yeast. FEBS Lett. 515, 25–28 (2002).
Hallstrom, T. C. & Moye-Rowley, W. S. Multiple signals from dysfunctional mitochondria activate the pleiotropic drug resistance pathway in Saccharomyces cerevisiae. J. Biol. Chem. 275, 37347–37356 (2000).
Traven, A., Wong, J. M., Xu, D., Sopta, M. & Ingles, C. J. Interorganellar communication. Altered nuclear gene expression profiles in a yeast mitochondrial dna mutant. J. Biol. Chem. 276, 4020–4027 (2001).
Panwar, S. L. & Moye-Rowley, W. S. Long chain base tolerance in Saccharomyces cerevisiae is induced by retrograde signals from the mitochondria. J. Biol. Chem. 281, 6376–6384 (2006).
Kolaczkowski, M. et al. Anticancer drugs, ionophoric peptides, and steroids as substrates of the yeast multidrug transporter Pdr5p. J. Biol. Chem. 271, 31543–31548 (1996).
Gros, P., Talbot, F., Tang-Wai, D., Bibi, E. & Kaback, H. R. Lipophilic cations: a group of model substrates for the multidrug-resistance transporter. Biochemistry 31, 1992–1998 (1992).
Fetisova, E. K. et al. Mitochondria-targeted antioxidant SkQR1 selectively protects MDR (Pgp 170)-negative cells against oxidative stress. FEBS Lett. 584, 562–566 (2010).
Kelso, G. F. et al. Selective targeting of a redox-active ubiquinone to mitochondria within cells: antioxidant and antiapoptotic properties. J. Biol. Chem. 276, 4588–4596 (2001).
Skulachev, V. P. Cationic antioxidants as a powerful tool against mitochondrial oxidative stress. Biochem. Biophys. Res. Commun. 441, 275–279 (2013).
Severin, F. F. et al. Penetrating cation/fatty acid anion pair as a mitochondria-targeted protonophore. Proc. Natl. Acad. Sci. USA 107, 663–668 (2010).
Kalinovich, A. V. et al. Mitochondria-targeted dodecyltriphenylphosphonium (C12TPP) combats high-fat-diet-induced obesity in mice. Int. J. Obes. 40, 1864–1874 (2016).
Stermitz, F. R., Lorenz, P., Tawara, J. N., Zenewicz, L. A. & Lewis, K. Synergy in a medicinal plant: Antimicrobial action of berberine potentiated by 5′-methoxyhydnocarpin, a multidrug pump inhibitor. Proceedings of the National Academy of Sciences 97, 1433–1437 (2000).
Knorre, D. A. et al. Dodecyltriphenylphosphonium inhibits multiple drug resistance in the yeast Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 450, 1481–1484 (2014).
Knorre, D. A. et al. Alkylrhodamines enhance the toxicity of clotrimazole and benzalkonium chloride by interfering with yeast pleiotropic ABC-transporters. FEMS Yeast Res. 16, (2016).
Trnka, J., Elkalaf, M. & Anděl, M. Lipophilic triphenylphosphonium cations inhibit mitochondrial electron transport chain and induce mitochondrial proton leak. PLoS One 10, e0121837 (2015).
Ojovan, S. M. et al. Accumulation of dodecyltriphenylphosphonium in mitochondria induces their swelling and ROS-dependent growth inhibition in yeast. J. Bioenerg. Biomembr. 43, 175–180 (2011).
Ivnitski-Steele, I. et al. Identification of Nile red as a fluorescent substrate of the Candida albicans ATP-binding cassette transporters Cdr1p and Cdr2p and the major facilitator superfamily transporter Mdr1p. Anal. Biochem. 394, 87–91 (2009).
Knorre, D. A., Krivonosova, T. N., Markova, O. V. & Severin, F. F. Amiodarone inhibits multiple drug resistance in yeast Saccharomyces cerevisiae. Arch. Microbiol. 191, 675–679 (2009).
Knorre, D. A., Sokolov, S. S., Zyrina, A. N. & Severin, F. F. How do yeast sense mitochondrial dysfunction? Microb. Cell 3, 532–539 (2016).
Zyrina, A. N., Smirnova, E. A., Markova, O. V., Severin, F. F. & Knorre, D. A. Mitochondrial Superoxide Dismutase and Yap1p Act as a Signaling Module Contributing to Ethanol Tolerance of the Yeast Saccharomyces cerevisiae. Appl. Environ. Microbiol. 83, (2017).
Schnell, N., Krems, B. & Entian, K. D. The PAR1 (YAP1/SNQ3) gene of Saccharomyces cerevisiae, a c-jun homologue, is involved in oxygen metabolism. Curr. Genet. 21, 269–273 (1992).
Lee, J. et al. Yap1 and Skn7 control two specialized oxidative stress response regulons in yeast. J. Biol. Chem. 274, 16040–16046 (1999).
Miyahara, K., Hirata, D. & Miyakawa, T. yAP-1- and yAP-2-mediated, heat shock-induced transcriptional activation of the multidrug resistance ABC transporter genes in Saccharomyces cerevisiae. Curr. Genet. 29, 103–105 (1996).
Zielonka, J. et al. Mitochondria-Targeted Triphenylphosphonium-Based Compounds: Syntheses, Mechanisms of Action, and Therapeutic and Diagnostic Applications. Chem. Rev. 117, 10043–10120 (2017).
de Deken, R. H. The Crabtree Effect: A Regulatory System in Yeast. Microbiology 44, 149–156 (1966).
Starovoytova, A. N., Sorokin, M. I., Sokolov, S. S., Severin, F. F. & Knorre, D. A. Mitochondrial signaling in Saccharomyces cerevisiae pseudohyphae formation induced by butanol. FEMS Yeast Res. 13, 367–374 (2013).
Ernst, R. et al. A mutation of the H-loop selectively affects rhodamine transport by the yeast multidrug ABC transporter Pdr5. Proc. Natl. Acad. Sci. USA 105, 5069–5074 (2008).
Hill, J. A., Ammar, R., Torti, D., Nislow, C. & Cowen, L. E. Genetic and genomic architecture of the evolution of resistance to antifungal drug combinations. PLoS Genet. 9, e1003390 (2013).
Shapiro, R. S., Robbins, N. & Cowen, L. E. Regulatory circuitry governing fungal development, drug resistance, and disease. Microbiol. Mol. Biol. Rev. 75, 213–267 (2011).
Quan, H. et al. Potent in vitro synergism of fluconazole and berberine chloride against clinical isolates of Candida albicans resistant to fluconazole. Antimicrob. Agents Chemother. 50, 1096–1099 (2006).
Spitzer, M. et al. Cross-species discovery of syncretic drug combinations that potentiate the antifungal fluconazole. Mol. Syst. Biol. 7, 499 (2011).
Obłąk, E., Piecuch, A., Krasowska, A. & Luczyński, J. Antifungal activity of gemini quaternary ammonium salts. Microbiol. Res. 168, 630–638 (2013).
Hlavácek, O., Kucerová, H., Harant, K., Palková, Z. & Váchová, L. Putative role for ABC multidrug exporters in yeast quorum sensing. FEBS Lett. 583, 1107–1113 (2009).
Prunuske, A. J., Waltner, J. K., Kuhn, P., Gu, B. & Craig, E. A. Role for the molecular chaperones Zuo1 and Ssz1 in quorum sensing via activation of the transcription factor Pdr1. Proc. Natl. Acad. Sci. USA 109, 472–477 (2012).
Zhang, F., Pracheil, T., Thornton, J. & Liu, Z. Adenosine Triphosphate (ATP) Is a Candidate Signaling Molecule in the Mitochondria-to-Nucleus Retrograde Response Pathway. Genes 4, 86–100 (2013).
Huh, W.-K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Sherman, F. Getting started with yeast. Methods Enzymol. 350, 3–41 (2002).
Rokitskaya, T. I. et al. Mitochondria-targeted penetrating cations as carriers of hydrophobic anions through lipid membranes. Biochim. Biophys. Acta 1798, 1698–1706 (2010).
Antonenko, Y. N. et al. Derivatives of rhodamine 19 as mild mitochondria-targeted cationic uncouplers. J. Biol. Chem. 286, 17831–17840 (2011).
Shedlovskiy, D., Shcherbik, N. & Pestov, D. G. One-step hot formamide extraction of RNA from Saccharomyces cerevisiae. RNA Biol. 14, 1722–1726 (2017).
Hu, F. et al. Key cytomembrane ABC transporters of Saccharomyces cerevisiae fail to improve the tolerance to D-limonene. Biotechnol. Lett. 34, 1505–1509 (2012).
Acknowledgements
We are grateful to Aglaya Azbarova for useful comments on our manuscript and to Irina Savchenko for assistance with quantitative RT PCR experiments. The study was supported in part by the Russian Science Foundation grant 14-50-00029.
Author information
Authors and Affiliations
Contributions
K.G., R.Z., F.S. and D.K. designed the experiments. K.G. (major contribution), E.B. and D.K. performed the experiments. K.G. and D.K. analysed the data. K.G. and D.K. prepared the figures. D.K. drafted the manuscript. All authors contributed to manuscript editing and reviewed the final manuscript. D.K. and F.S. supervised the project and acquired the funding.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Galkina, K.V., Besedina, E.G., Zinovkin, R.A. et al. Penetrating cations induce pleiotropic drug resistance in yeast. Sci Rep 8, 8131 (2018). https://doi.org/10.1038/s41598-018-26435-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26435-z
This article is cited by
-
Protonophore FCCP provides fitness advantage to PDR-deficient yeast cells
Journal of Bioenergetics and Biomembranes (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.