Abstract
Transfer of nitrogen fixation ability to plants, especially crops, is a promising approach to mitigate dependence on chemical nitrogen fertilizer and alleviate environmental pollution caused by nitrogen fertilizer run-off. However, the need to transfer a large number of nitrogen fixation (nif) genes and the extreme vulnerability of nitrogenase to oxygen constitute major obstacles for transfer of nitrogen-fixing ability to plants. Here we demonstrate functional expression of a cyanobacterial nitrogenase in the non-diazotrophic cyanobacterium Synechocystis sp. PCC 6803 (Synechocystis 6803). A 20.8-kb chromosomal fragment containing 25 nif and nif-related genes of the diazotrophic cyanobacterium Leptolyngbya boryana was integrated into a neutral genome site of Synechocystis 6803 by five-step homologous recombination together with the cnfR gene encoding the transcriptional activator of the nif genes to isolate CN1. In addition, two other transformants CN2 and CN3 carrying additional one and four genes, respectively, were isolated from CN1. Low but significant nitrogenase activity was detected in all transformants. This is the first example of nitrogenase activity detected in non-diazotrophic photosynthetic organisms. These strains provide valuable platforms to investigate unknown factors that enable nitrogen-fixing growth of non-diazotrophic photosynthetic organisms, including plants.
Similar content being viewed by others
Introduction
Nitrogen fixation is the conversion of molecular nitrogen (N2) to ammonia, a more accessible nitrogen source for most organisms. Biological nitrogen fixation is catalyzed by nitrogenase, which can be classified into three isoforms according to metal contents in the catalytic components: MoFe-nitrogenase1, VFe-nitrogenase2,3, and FeFe-nitrogenase4,5. MoFe-nitrogenase has been most extensively studied6,7,8, and the reaction catalyzed by MoFe-nitrogenase is as follows:
MoFe-nitrogenase (hereafter called just nitrogenase) consists of two components, Fe protein and MoFe protein. The Fe protein, a homodimer of NifH, acts as an ATP-dependent reductase for the catalytic component MoFe protein by accepting electrons from ferredoxin or flavodoxin (dithionite for in vitro systems) and transferring them to MoFe protein in a process coupled to ATP hydrolysis. MoFe protein, a heterotetramer of NifD and NifK, serves as the catalytic component. The MoFe protein contains two metallocenters, P-cluster and FeMo-cofactor (FeMo-co). Electrons from the Fe protein are transferred to FeMo-co via P-cluster and dinitrogen bound to the FeMo-co is reduced to ammonia. The three metallocenters required for nitrogen fixation, the [4Fe-4S] cluster of Fe protein, the P-cluster, and the FeMo-co, are extremely vulnerable to oxygen with half-lives of seconds to minutes upon exposure to the air9. In addition, biosynthesis of P-cluster and FeMo-co require many nif gene products (at least eight; nifHBSUENVZ)10, and like the metallocenters of nitrogenase, precursors of FeMo-co such as NifB-cofactor are very sensitive to oxygen. Thus, an anaerobic environment is necessary for biosynthesis and operation of nitrogenase.
Nitrogen supply is one of the main factors determining crop yield. Industrial nitrogen fixation, invented at the beginning of the 20th century, has allowed for the large-scale production of chemical nitrogen fertilizer from the atmosphere, which in turn has markedly increased crop yields during the past century11. However, industrial nitrogen fixation consumes massive amounts of fossil fuel, contributing to the increase in atmospheric CO2. Furthermore, significant amounts of nitrogen fertilizer leak into the environment due to excess application and inefficient assimilation into crops, resulting in serious environmental pollution from reactive nitrogen species12,13. Conferring crops with nitrogen fixing ability is one promising technology for alleviating these deleterious environmental consequences. One such approach is the transfer of genes encoding nitrogenase directly to the crop genome14,15. Transgenic crops expressing active nitrogenase could then convert atmospheric nitrogen to ammonia as an in situ nitrogen source.
However, there are two major obstacles to this “new agriculture using the air as nitrogen fertilizer”. One obstacle is the oxygen vulnerability of nitrogenase and enzymes involved in biosynthesis of nitrogenase metallocenters. Crops are photosynthetic organisms that produce oxygen by photosynthesis. Thus, nitrogenase must be protected not only from atmospheric oxygen but also from endogenously produced oxygen in plants. The other obstacle is the required co-expression of at least 9 nif genes for nitrogenase and for biosynthesis of the metalloclusters at appropriate levels16. In addition, abundant reducing power (reduced ferredoxin) and ATP are required for nitrogenase (see reaction formula above). Mitochondria and chloroplasts (plastids) have been proposed as cellular compartments to accommodate active nitrogenase in plant cells14,17. The mitochondrial matrix provides a low oxygen environment due to active respiration at the inner membranes. However, the mitochondrial ferredoxin (adorenodoxin) does not serve as an electron donor to nitrogenase18. Alternatively, chloroplasts produce enough reduced ferredoxin and ATP by active photosynthetic electron transfer in thylakoid membranes under light conditions. Moreover, endogenous ferredoxins are able to serve as electron donors to nitrogenase18. On the other hand, oxygen production by photosystem II may be incompatible with oxygen-sensitive nitrogenase. Chloroplast genomes of many algae and plants such as gymnosperms (e.g., Pinus thunbergii)19 and mosses (e.g., Physcomitrella patens and Marchantia polymorpha)20,21 encode three subunits of a nitrogenase-like enzyme, dark-operative protochlorophyllide oxidoreductase (DPOR), that shows similar oxygen sensitivity as nitrogenase20. This suggests that chloroplasts have the potential ability to express and accommodate active nitrogenase14.
Cyanobacteria are prokaryotes that perform oxygenic photosynthesis similar to plants and share a common evolutionary ancestor with chloroplasts. About half of cyanobacterial species have the ability of nitrogen fixation22. In some of these species, the coexistence of oxygenic photosynthesis and nitrogen fixation is achieved by spatial separation in differentiated cells specialized for nitrogen fixation termed heterocysts (heterocystous cyanobacteria)23. For non-heterocystous cyanobacteria, it has been proposed that photosynthesis and nitrogen fixation are separated temporally under control of the circadian clocks24,25. However, some non-heterocystous cyanobacteria show nitrogenase activity only under light conditions26,27,28, implying the existence of some molecular mechanisms to solve the “oxygen paradox” between oxygen-sensitive nitrogenase and oxygen-producing photosynthesis.
Cyanobacteria without nitrogen fixing ability could serve as a suitable model acceptor for genes involved in nitrogen fixation. In this study, we isolated a transformant CN1 of the non-diazotrophic cyanobacterium Synechocystis sp. PCC 6803 (Synechocystis 6803) harboring a long nif gene cluster (20.8 kb) from the diazotrophic cyanobacterium Leptolyngbya boryana as a model experiment for transplantation of nif genes in non-diazotrophic photosynthetic organisms. In addition, two other transformants CN2 and CN3 were also isolated from CN1. All three transformants, CN1, CN2 and CN3 carrying 26, 27 and 30 nif and nif-related genes, respectively, exhibited low but significant nitrogenase activity. This is the first example of functional nitrogenase expression in an oxygenic photosynthetic organism. These transformants provide promising platforms to identify factors allowing nitrogen fixation in oxygenic photosynthetic organisms and to improve nitrogen fixing growth in non-diazotrophic photosynthetic organisms.
Results
Step-wise integration of the nif gene cluster into the chromosome of Synechocystis 6803
In a previous study, we identified the cnfR gene encoding the transcriptional activator for nif genes in the 50-kb gene cluster of the L. boryana chromosome29. All nif and nif-related genes activated by CnfR are concentrated in the central 20-kb region of the gene cluster. In this region, there are 25 genes containing the minimal gene repertory to produce active nitrogenase components Fe protein and MoFe protein16 (Fig. 1A): 14 nif genes (nifBSUHDKVZT in the right part and nifPENXW in the left part), 5 nif-related genes (fdxB, fdxH, hesAB and fdxN), and 6 genes with unknown functions (feoA, orf70, orf155, orf90, dpsA, and orf84). These genes are divergently transcribed from the intergenic region between nifB and nifP upon activation by CnfR29,30. We also confirmed that this activation system operates in Synechocystis 680330. We then examined whether nitrogen fixation can be conferred to Synechocystis 6803 by introduction of the 20-kb nif gene cluster together with the cnfR gene into its chromosome.
(A) The 50-kb nif gene cluster in L. boryana dg5. Genes that were transferred to the genome of Synechocystis 6803 to isolate CN1 are shown in pink. Genes in this central region (shown in two arrows of right and left parts) are divergently transcribed from the nifB (P nifB ) and nifP (P nifP ) promoters (shown by small pink triangles). Expression of all these genes except cnfR is dependent on CnfR29. Genes denoted in yellow show the highest transcript levels under nitrogen fixing conditions and their transcripts are still detected in the absence of cnfR (∆cnfR), albeit lower than wild type29. Subfragments I–V introduced sequentially into the genome neutral site III are shown by red lines. (B) Gene arrangements of the transferred nif gene cluster in Synechocystis 6803 transformants CN1, CN2, and CN3. Subfragments VI, VII, and VIII (red lines) were used for isolation of CN1, CN2, and CN3, respectively.
Synechocystis 6803 is naturally transformable, and exogenous DNA fragments are incorporated and integrated into the chromosome via homologous recombination. However, the >20-kb fragment needed for transfer of nitrogen fixation ability would be difficult to integrate stably in a single step. Thus, the 20-kb DNA fragment was divided into five subfragments (Fragments I to V; Fig. 1) with some overlap and integrated into a neutral site of the acceptor chromosome in 5 steps (Supplementary Figs 1 and 2).
The first fragment (Fragment I) was introduced into neutral site III (between slr2030-slr2031) along with the kanamycin resistance gene (KmR) as a selection marker to isolate the first transformant SN1 (Supplementary Fig. 2). In SN1, 8 genes and a 3′ region of nifX were integrated. The second fragment (Fragment II) was introduced between Fragment I and slr2031 of SN1 with the spectinomycin resistance gene (SpR) as an alternative selection marker to isolate SN2. The third fragment (Fragment III) was integrated between Fragment II and slr2031 by KmR selection to isolate SN3. The fourth fragment (Fragment IV) was integrated between Fragment III and slr2031 by SpR selection to isolate SN4. Finally the fifth fragment (Fragment V) was integrated between Fragment IV and slr2031 by KmR selection to isolate SN5. In SN5, the 20.8-kb (20,763 bp) fragment containing 25 genes (from fdxB to nifT) and the KmR gene (2,230 bp) was inserted between slr2030 and slr2031.
Isolation of CN1, CN2, and CN3
A chimeric fragment (Fragment VI) in which the cnfR gene was connected to the trc promoter (derived from pQE-70) and the selective marker SpR was integrated into neutral site I, the coding region of psbA2 (slr1311) in SN5, to isolate CN1.
In SN5 and CN1, a 5′ region of the mop gene was introduced downstream of the leftmost nif gene cluster gene fdxB. Another transformant with intact mop gene, CN2, was isolated by integration of the seventh fragment (Fragment VII) between slr2030 and fdxB using the chloramphenicol resistance gene (CmR) as the selection marker (Fig. 1B, Supplementary Figs 1 and 2). In CN2, the 20.9-kb (20,909-bp) fragment containing the 26 genes from mop to nifT, the CmR gene (1,599 bp), and the KmR gene (2,230 bp) was inserted between slr2030 and slr2031.
The cox genes (coxB2, coxA2, and coxC2) located just downstream of cnfR in the nif gene cluster in L. boryana encode subunits of cytochrome c oxidase (COX), and COX is probably involved in oxygen protection and ATP generation for nitrogenase. A chimeric fragment (Fragment VIII) of the truncated nifB promoter (484 bp), the coxB2-coxA2-coxC2 gene cluster, and the intact mop gene was integrated between slr2030 and fdxB in CN1 to yield CN3 (Fig. 1B, Supplementary Figs 1 and 2). In CN3, the 24.9-kb (24,863-bp) fragment containing 29 genes (from mop to nifT, and coxB2 to coxC2), the KmR gene (2,230 bp), and the CmR gene (1,599 bp) was inserted between slr2030 and slr2031. We expected that the cox genes would be expressed by activation of CnfR and that the resultant COX complex would stimulate nitrogenase activity by supplying of ATP and removing residual oxygen.
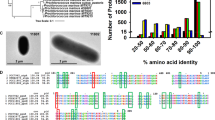
Genome resequencing of CN1, CN2, and CN3 confirmed that all gene manipulations resulted as intended except for a single SNP, a T-to-C conversion causing one amino acid residue substitution, Phe150 to Leu, in the NifE protein. Alignment of various cyanobacterial NifE proteins from 30 species including L. boryana suggested that the residue at this site is not conserved (namely Ile, Val, or Phe), while the NifE protein from Tripothrix campylonemoides has Leu at this position. We concluded that this SNP would not strongly affect the activity of NifE for active nitrogenase production in these transformants.
Nitrogenase activity in CN1, CN2, and CN3
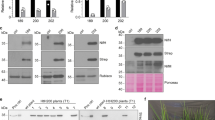
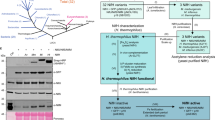
We first examined whether CN1 expresses nitrogenase activity using the ethylene formation assay. Cells grown under aerobic and nitrate-replete conditions were shifted to micro-oxic and nitrogen-deficient conditions to induce expression of nif genes by CnfR, and ethylene formation from acetylene was measured under anaerobic and light conditions. To normalize cell density between two species, L. boryana and Synechocystis 6803, we estimated acetylene reduction activity per cell dry weight (CDW, Supplementary Fig. 3). Dithionite was added to remove oxygen produced endogenously by photosynthesis, thereby enhancing nitrogenase activity. Wild type (WT) L. boryana (dg5) cells showed ethylene production of about 500 (517 ± 226) and 1,000 (1,084 ± 190) nmol ml−1 h−1 mgcdw−1 in the absence and presence of dithionite, respectively (Fig. 2A), confirming that the addition of dithionite has a significant positive effect on in vivo nitrogenase activity as reported31. While no activity was detected in WT Synechocystis 6803, CN1 showed very low but significant ethylene formation in the presence of dithionite (2.8 ± 0.6 nmol ml−1 h−1 mgcdw−1, Fig. 2A and Supplementary Figs 4 and 5). The activity of CN1 was about 0.26% of that of L. boryana WT (in the presence of dithionite). Then, we compared activities of the three transformants, CN1, CN2 and CN3 (Fig. 2B). Activity of CN2 was significantly lower than that of CN1 (p < 0.05). CN3 showed comparable or even slightly lower nitrogenase activity to CN1 (p = 0.38). These results clearly indicate that a small but significant amount of active nitrogenase can be produced in all transformants.
(A) Nitrogenase activity of Synechocystis 6803 WT, CN1, and L. boryana dg5 as estimated by ethylene (C2H4) formation in the presence (blue) and absence (gray) of dithionite. Nitrogenase activity was estimated as ethylene formation per CDW. Incubation time was 2 h. (B) Nitrogenase activity of the three transformants CN1, CN2, and CN3. Incubation time was 5 h. Error bars (green) represent standard deviations (n = 3). n.d. denotes “not detected”.
We examined whether the three transformants CN1, CN2, and CN3 are able to grow under nitrogen fixing conditions. However, none of the Synechocystis 6803 transformants showed significant growth even under micro-oxic conditions for 23 days (Supplementary Fig. 6). This result suggests that nitrogen fixing ability was not conferred in these transformants despite integration of 25, 26, and 29 nif and nif-related genes.
Estimation of nitrogenase proteins in CN1
To examine the expression levels of nif genes in CN1 following CnfR induction, we conducted Western blot analysis of cell extracts using antisera against NifH, NifD, and NifK (Fig. 3). The direct parental transformant for CN1, SN5, does not harbor the cnfR gene and so was examined as a control. After treatment for nitrogenase induction as above, WT, SN5, and CN1 cells were subject to nitrogenase assay (Fig. 3A), and total cell extracts were prepared for Western blot analysis (Fig. 3B). Neither SN5 nor WT Synechocystis 6803 showed detectable activity, while CN1 showed significant activity as in Fig. 2. In Western blots of CN1 extracts demonstrated clear bands for NifH, NifD, and NifK, while no bands for any of these proteins were detected from SN5 extracts. This result supports our initial notion that nifB and nifP promoters can be activated by CnfR to produce nifHDK transcripts and NifHDK proteins in non-diazotrophic cyanobacterium.
(A) Nitrogenase activity as measured by ethylene formation for WT, SN5, and CN1 Synechocystis 6803 in the presence (blue) and absence (gray) of dithionite. (B) Western blot analysis of crude extracts from WT (lane 1), SN5 (lane 2) and CN1 (lane 3) Synechocystis 6803 using antisera against NifH, NifD, and NifK. Five hundred nanograms of protein was loaded onto each lane. (C) Estimated expression levels of NifH, NifD, and NifK by CN1 (0.5 µg, lane 2) compared to whole crude extract of L. boryana dg5 (0.5 µg, lane 3) and serial dilutions (10-fold dilution, lane 7; 20-fold dilution, lane 6; 50-fold dilution, lane 5; and 100-fold dilution, lane 4). Numbers below the signals are relative band intensities estimated by densitometry normalized to 0.1 for the band in 10-fold dilution (ImageJ 1.50i).
Furthermore, we estimated NifHDK protein expression levels in CN1 compared to L. boryana dg5 by Western blot analysis (Fig. 3C). The NifHDK band intensity from CN1 extract was similar to that produced by a 1/10 diluted extract of L. boryana dg5 (compare lanes 2 and 7 in Fig. 3C). The intensities of the signals of NifH, NifD, and NifK bands from CN1 were 0.17, 0.064, and 0.23, respectively (lane 2, Fig. 3C). Considering that the ethylene formation activity of CN1 was about 0.26% (CDW-basis) of that of L. boryana, it is estimated that only 1.5% of total NifH, 4.1% of total NifD, and 1.1% of total NifK in CN1 are assembled into active nitrogenase constituents.
Discussion
In this study we isolated three transformants of the non-diazotrophic cyanobacterium Synechocystis 6803, harboring 25 (strain CN1), 26 (CN2), and 29 (CN3) genes of the diazotrophic cyanobacterium L. boryana. All three CN strains showed a low but significant nitrogenase activity under micro-oxic and nitrogen-deficient conditions, indicating that CnfR activates the divergent transcription of nifB and nifP promoters in the non-diazotrophic heterologous host Synechocystis 6803. This result is consistent with the previous reporter experiments of the nifB and nifP promoters in Synechocystis 680330. Although these transformants did not grow diazotrophically, this is the first successful example of heterologous active nitrogenase expression in an oxygenic photosynthetic organism. This result indicates that the 25 genes within the 20.8-kb fragment integrated into CN1 are sufficient for expression of active nitrogenase in oxygenic photosynthetic cells.
Although the expression levels of NifHDK proteins in CN1 were about 6.4%–23% of those in L. boryana (Fig. 3C), nitrogenase activity was only 0.26% that of L. boryana (Fig. 2A), suggesting that only small proportions (1.1%–4.1%) of expressed NifHDK proteins contribute to formation of active nitrogenase. This simple calculation may underestimate the proportions of active nitrogenase proteins in Synechocystis 6803. Even if the nitrogenase assembles normally in Synechocystis 6803 cells, its operating conditions may be suboptimal due to heterogeneous expression. Large amounts of reducing power and ATP should be supplied to support nitrogenase activity. The most efficient electron donor for cyanobacterial nitrogenase is a ferredoxin FdxH. The fdxH gene is contained in the nif gene cluster that was introduced into the CN strains. Thus, FdxH would donate electrons to nitrogenase in CN1. However, the endogenous ferredoxin PetF of Synechocystis 6803, which could be less efficient as an electron donor to nitrogenase32, may compete with FdxH in binding to nitrogenase during transferring electrons, resulting in lowering nitrogenase activity33. It is also likely that ATP production is not enough to support nitrogenase activity. In any case, it is inferred that the inability for diazotrophic growth by CN strains is due to the extremely low nitrogenase activity.
Furthermore, the low efficiency of NifHDK assembly is probably caused by the oxygen vulnerability of nitrogenase and assembly proteins, such as NifB, NifU and NifEN, involved in the biosynthesis of nitrogenase metal centers. Synechocystis 6803 has the ability to grow under micro-oxic conditions34, but the capacity to remove or consume oxygen may not be high enough for nitrogenase to assemble and operate under micro-oxic conditions compared to L. boryana. This assumption is supported by the dithionite dependence of nitrogenase activity in CN strains but not L. boryana (Fig. 2). Nitrogenase activity of CN strains was observed only in the presence of dithionite, in contrast to the relatively high nitrogenase activity of L. boryana even in the absence of dithionite (Fig. 2A). In Anabaena sp. PCC 7120 (Anabaena 7120), the flavodiiron protein (encoded by flv3B) catalyzing oxygen reduction to water is essential for nitrogenase activity in heterocysts under aerobic conditions35. While Synechocystis 6803 has four Flv isoforms, they appear to be specialized for oxygen protection of PSII (Flv2 and Flv4) and PSI (Flv1 and Flv3). Very low nitrogenase activity in the CN strains suggests that the basal activities of the four Flv isoforms are insufficient for protection of nitrogenase from oxygen. It is possible, however, that overexpression of diazotrophic cyanobacterial flv genes such as flv3B may protect nitrogenase from oxygen, resulting in higher nitrogenase activity.
The mop gene encodes a small 7-kDa protein referred to as molbindin that is implicated in cellular storage and homeostasis of Mo36,37. Semi-quantification of nif gene cluster transcription levels29 indicated that mop is expressed in nitrogen-deficient conditions independent of oxygen level and its transcript is detected even in the absence of CnfR (∆cnfR mutant), in contrast to nif genes, which are expressed exclusively under nitrogen-deficient and micro-oxic (nitrogen fixation) conditions. Thus, the mop gene was not included in the first transformant CN1. However, as a constituent of FeMo-co, molybdenum is required for active nitrogenase. The mop gene is adjacent to the modABC genes encoding the Mo transporter in the nif gene cluster of L. boryana. No mop ortholog is found in the Synechocystis 6803 genome, while there are modABC orthologs (sll0738 as modA and sll0739 as modBC). Given that the mop gene may play an important role in nitrogenase assembly, we isolated a second transformant, CN2, including the mop gene in addition to the gene repertory of CN1. However, the acetylene reduction activity of CN2 was even lower than that of CN1 (Fig. 2B, p < 0.05), suggesting that mop does not enhance nitrogenase activity under the conditions examined in the heterologous host Synechocystis 6803.
The heterocystous cyanobacterium Anabaena 7120 has three cox gene clusters (cox1, cox2, and cox3), each encoding three subunits for COX. In contrast to constitutive expression of cox1 genes in vegetative cells, cox2 and cox3 clusters are expressed specifically in heterocysts in response to nitrogen-deficient conditions38. A targeted mutant in which both cox2 and cox3 gene clusters were disrupted lost diazotrophic growth ability and demonstrated severely reduced nitrogenase activity (8%), indicating a critical contribution of heterocyst-specific cox genes in nitrogen fixation. L. boryana also harbors three cox gene clusters, one of which, coxB2-coxA2-coxC2, is contained within the nif gene cluster29. We isolated the third transformant CN3, in which the coxB2-coxA2-coxC2 gene cluster was artificially connected to the truncated nifB promoter. We expected that these introduced cox genes would contribute to the protection of nitrogenase against oxygen by enhancement of COX activity. However, CN3 exhibited comparable or even slightly lower nitrogenase activity to that of CN1 (Fig. 2B, p = 0.38). To estimate the effect of the cox genes in CN3, we measured respiratory (oxygen consumption) activity (Supplementary Fig. 7). However, the respiratory activity of CN3 was about the same levels as WT, CN1, and CN2, and the expected effect of the cox genes on nitrogenase activity was not observed. While the truncated nifB promoter covers all conserved motifs (motifs I to IX) for recognition by CnfR30, the expression level might not be high enough for overexpression of the cox genes to show a significantly high COX activity. Thus, further improvement is required for higher expression of the cox genes in CN3.
Various strategies have been proposed for transplantation of nitrogen fixing ability to crops. Introduction of nif genes directly into the crop genome to express active nitrogenase is one promising approach. Current basic researches on this approach use E. coli, yeast, and tobacco as models. Based on the functional expression of Klebsiella pneumonia nif genes in E. coli39,40, Dixon and Wang specified a minimal set of 9 nif genes from Paenibacillus required for expression of active nitrogenase in E. coli16. They also identified 10 genes from Azotobacter vinelandii that are sufficient for active FeFe-type nitrogenase expression in E. coli5. Recently, they demonstrated that plant ferredoxins and their reductase systems are competent to donate electrons to MoFe- and FeFe-nitrogenases in the E. coli system18. Mitochondria and chloroplasts have been proposed as intracellular compartments to accommodate nitrogenase in the plant cell14,17. To this end, Rubio’s group reported successful expression of active Fe protein in yeast mitochondria41, but expression of active MoFe protein in yeast mitochondria remains difficult even when using extensive synthetic approaches42. In tobacco, 16 nif genes from K. pneumoniae were expressed as fusion proteins with mitochondrial targeting signals under the control of the 35 S promoter, and most were successfully localized in mitochondria. However, some gene products, including NifD, NifE, and NifQ, were difficult to detect immunologically in tobacco cells43. As an attempt to express nitrogenase in chloroplasts, Researchers at Monsanto Company isolated transgenic tobacco plants harboring the nifH and nifM genes in the chloroplast genome, and a very tiny but significant Fe protein activity was detected44.
In this study, we present the first successful functional expression of nitrogenase in an oxygenic photosynthetic organism, Synechocystis 6803. Synechocystis 6803 has several potential advantages as a model. First, nifM and nifQ are not included in the CN strains because orthologs are not found in the genomes of any diazotrophic cyanobacteria, including L. boryana29,45. This nifM- and nifQ-independent nitrogen fixation is a feature of cyanobacteria and is not found in heterotrophic diazotrophs such as A. vinelandii and K. pneumonia. Second, cyanobacteria share a common origin with plant chloroplasts, so endogenous ferredoxins and their reducing systems can be used as the electron donors for nitrogenase18. Third, nitrogenase coexists with oxygenic photosynthesis in non-heterocystous nitrogen-fixing cyanobacteria such as L. boryana and Cyanothece. Identifying the molecular mechanisms allowing nitrogen fixation and photosynthesis (i.e., solving the “oxygen paradox”) may enable functional expression of nitrogenase in the chloroplasts by simply coexpressing the underlying genes with nif genes. Transcriptional activation of nif genes by CnfR in response to micro-oxic conditions may be a critical step of this process.
The CN strains isolated in this study provide valuable platforms for investigating environmental factors and unknown genetic mechanisms necessary for functional expression of nitrogenase in chloroplasts.
Materials and Methods
Cyanobacterial strains and cultivation conditions
Synechocystis sp. PCC 6803 strain YF46 was used as the recipient of the nif genes from Leptolyngbya boryana strain dg5. We used BG-11 media for all strains. Instead of ferric ammonium citrate (6 mg L−1), NaNO3 (1.5 g L−1), and Co(NO3)2-6H2O (0.0494 g L−1 in Trace metal mix A5 + Co), a ferric citrate mixture (citric acid 1.2 mg, ferric citrate 1.2 mg and Na2EDTA 0.2 mg), NaCl (1.03 g L−1), and CoCl2 (0.022 g L−1 in a modified Trace metal mix A5 + Co) were added to base media that contained no combined nitrogen (BG-110; for nitrogen-depleted medium)47. For nitrogen-replete medium (BG-11), KNO3 (final, 15 mM) was added. All media contained 20 mM HEPES-KOH; pH 8.2 for Synechocystis 680334 or pH 7.4 for L. boryana48. Bacto Agar (Becton, Dickinson and Company, Franklin Lakes, NJ) was used for solidification of media (1.5% (w/v)). For selection of transformants, kanamycin (15 µg ml−1), spectinomycin (15 µg ml−1), and chloramphenicol (10 µg ml−1) were added. Agar plates were cultivated in the light (50 µmol m−2 s−1) at 30 °C. For micro-oxic conditions, agar plates were incubated in an anaerobic jar (BBL GasPak anaerobic systems, BD Biosciences) with a sachet for producing anaerobic conditions (Gas Generating Kit Anaerobic System; Oxoid, Basingstoke, Hants, UK)46. The “micro-oxic” conditions in this work means that the gas phase in a jar is kept anaerobic while the cyanobacterial cells in the jar still evolve oxygen by photosynthesis34. The evolved oxygen is quickly consumed by reaction with hydrogen generated from the sachet in the presence of an active catalyst in the jar. For check of anaerobic conditions in the jar, dry anaerobic indicator strips (Dry Anaerobic Indicator Strips, BD Biosciences) were used. Strips (containing methylene blue) showing blue under aerobic conditions turned white within one hour when the jar was closed for anaerobic incubation. We confirmed that the color of strips was kept white during incubation. In addition, using a small portable oxygen monitor (OXY-2, Jikco, Tokyo, Japan) we have confirmed that the oxygen level in the jar became 0% during induction.
Plasmid construction
Plasmid vectors, PCR templates, PCR primers, and restriction enzymes used for plasmid constructions are summarized in Supplementary Figures 1 and 2, and Supplementary Tables 1 and 2. PCR fragments were amplified by KOD FX Neo DNA polymerase (Toyobo, Osaka, Japan), and purified from agarose gel using the Wizard SV Gel and PCR Clean-up System (Promega, Madison, WI). After digestion of PCR fragments and plasmids with restriction enzymes, DNA fragments were separated by agarose gel electrophoresis and purified by Wizard SV Gel and PCR Clean-up System. Ligation reactions were performed with the DNA Ligation Kit <Mighty Mix> (Takara, Shiga, Japan). E. coli JM109 strain competent cells were used for transformation. DNA sequences of the inserted fragments were confirmed by an ABI3100 sequencer (Applied Biosystems, Foster City, CA).
Construction of pNSsacB2
The kanamycin resistance (KmR) cassette and the sacB gene were amplified from pRL271 (Accession number L05081; C. P. Wolk49,) and pYFC1050, and fused to a single 4.0-kb fragment by overlap PCR (Supplementary Table 1). This fusion fragment was cloned into the BamHI sites of pUC19 to form pUCK19sacB2. The upstream (sll2030) and downstream (slr2031-ssr1880) fragments of chromosomal neutral site III were amplified from genomic DNA of Synechocystis 6803 and cloned sequentially into the XbaI-XhoI and SacI sites of pUCK19sacB2, respectively, to form pNSsacB2. It should be noted that the sacB gene was used just temporarily in these initial plasmids (pUCK19sacB2 and pNSsacB2) and never in the later plasmids.
Construction of pSN1KF
The single KmR gene (amplified from pNSsacB2) was introduced into the XhoI and KpnI sites of pNSsacB2 to form pNSKmF, with the KmR gene in the same direction as slr2030-slr2031. During this process, the sacB-KmR gene was replaced with the KmR gene. A 4.17-kb chromosomal fragment containing fdxN, feoA, fdxH, hesB, hesA, nifW, orf70, orf115, and partial nifX amplified from genomic DNA of L. boryana dg5 was cloned into the XhoI site of pNSKmF to form pSN1KF (Supplementary Figs 1 and 2).
Construction of pSN2SF
pNSsacB2 was digested with XhoI and KpnI, and the spectinomycin resistance (SpR) fragment amplified from p6803NS2S130 was ligated to form pNSspeF, with the SpR gene in the same direction as slr2030-slr2031. A 5.53-kb chromosomal fragment containing nifW, orf70, orf115, nifX, nifN, nifE, orf99, dpsA, orf84, and partial nifP amplified from the L. boryana dg5 genome was introduced into the SalI-XhoI sites to form pSN2SF.
Construction of pSN3KR
pNSsacB2 was digested with KpnI and self-ligated to form pNSKmR, with the KmR gene in the direction opposite to slr2031-slr2031. A 5.41-kb chromosomal fragment containing partial nifE, orf90, dpsA, orf84, nifP, nifB, and partial fdxN was cloned into the SalI-XhoI sites to form pNS3KR.
Construction of pSN4SFc
The fourth plasmid pSN4SF was constructed from pNSspeF by the introduction of a 5.53-kb fragment containing partial nifB, fdxN, nifS, nifU, nifH and partial nifD. However, no colonies carrying pNS4SF was obtained. We suspected that the partial nifB (the leftmost gene of the fragment) was unexpectedly expressed by read-through from the lac promoter of the parental plasmid pUC19 even in the absence of IPTG, resulting in a severe negative effect on growth of E. coli. To alleviate this probable negative effect, a 663-bp fragment (partial modB and modC genes) was introduced into the SalI-XhoI sites of pNSspeF to form pNSspeFc. The 5.53-kb fragment was successfully cloned into the XhoI site of pNSspeFc to form pSN4SFc.
Construction of pSN5KR
A 5.80-kb chromosomal fragment containing partial nifH, nifD, nifK, nifV, nifZ, and nifT was introduced into the SalI-XhoI sites to form pSN5KR.
Construction of pExCnfR5
The KmR gene of pExCnfR430 was removed by SacI and SalI digestion and the SacI-XhoI SpR gene from p6803NS2S130 was ligated to form pExCnfR5.
Construction of pSN6mop
A CmR cartridge (EcoRV-HincII) derived from pBR325 (Accession, L08855;51) was cloned into pBluescript II SK+ (Accession, X52328;52) to form pBSC9. The CmR cartridge was excised as the BamHI-BclI fragment, and inserted into pUC19 to form pUC19Cm2. The slr2030 fragment was inserted into the SalI site of pUC19Cm2 to form pUCC6803L. A 1.77-kb chromosomal fragment containing mop, fdxB, feoA, fdxH, hesB, and partial hesA was inserted into the SacI-BamHI sites of pUCC6803L to form pSN6mop.
Construction of pSN6cox
A nifB promoter fragment (484-bp) was inserted into the BamHI site to form pSN6Emp. Then, a 3.83-kb chromosomal fragment containing coxB2, coxA2, coxC2, and partial orf159 was inserted into the BamHI site of pSN6Emp to form pSN6cox.
Transformation of Synechocystis 6803
Transformation of Synechocystis 6803 was performed as previously described30. The 20-kb nif gene fragment, the central part of the 50-kb nif gene cluster of L. boryana, was divided into 5 subfragments (each 4–5 kb) and sequentially inserted into the genome neutral site III by homologous recombination. The first plasmid pSN1KF carried a 4.17-kb nif subfragment (Fragment I) flanked by two homologous fragments of slr2030 and slr2031, and the KmR gene was used as the selection marker. The 1st transformant SN1 was isolated from WT by transformation with pSN1KF. The second plasmid pNS2SF carried a 5.53-kb nif subfragment (Fragment II) in which the left 1-kb was used as the left homologous sequence. The SpR gene was included as another selection marker. The 2nd transformant SN2 was isolated from SN1 by transformation with pNS2SF. Thus, by utilizing the selection markers (KmR and SpR) alternately, the 5th transformant SN5 was isolated (Supplementary Figs 1 and 2). The cnfR gene under the control of the trc promoter was introduced into another genome neutral site I of SN5 to isolate transformant CN1 carrying the 20.8-kb (20,763 bp) nif gene cluster and the trc-driven cnfR gene with the selective markers KmR and SpR. Using CN1 as the host strain, the other transformants CN2 and CN3 were isolated by the introduction of pSN6mop and pSN6cox, respectively. Completion of the segregation was confirmed in all mutants by genomic PCR.
Growth under nitrogen fixing conditions
CN strains grown under aerobic and nitrogen-replete conditions were suspended in distilled water. After adjustment of cell density (OD730), aliquots (5.0 µl) of the suspensions were spotted onto BG-110 agar plates. The plates were incubated under aerobic or micro-oxic conditions in the light (50 µmol m−2 s−1) at 30 °C.
Assay of nitrogenase activity
Induction of the nif genes and nitrogenase assay were essentially the same as described previously29. Cells were grown on BG-11 agar plates under light conditions (50 µmol m−2 s−1) at 30 °C for 3 days as a pre-culture. The cells were collected in distilled water adjusted to OD730 9.2, and 250 µl of the cell suspension was spread uniformly to form a 4-cm diameter circle on a BG-110 agar plate. The agar plate was incubated in an anaerobic jar under light conditions (50 µmol m−2 s−1) at 30 °C for 14 h. The cells were harvested with 1.5 ml of BG-110 liquid medium and an aliquot (1.0 ml) of the suspension was transferred into a 5-ml glass vial (V-5A, Nichiden-rika glass, Kobe, Japan) in the anaerobic chamber. To stimulate in vivo nitrogenase activity dithionite was added to the suspension (final concentration: 5 mM31). The glass vials were sparged with a gas mixture (10% (vol/vol) acetylene in argon as standard gas; Japan Fine Products, Kawasaki, Japan) for 45 sec. The glass vials were incubated for 2-5 h under illumination (50 µmol m−2 s−1) at 30 °C with stirring. The upper gas phase (500 µl) was analyzed by a gas chromatograph (GC-2014AF, Shimazdu, Kyoto, Japan) equipped with a Porapak N column (0.3 m × 3 mm, Shinwa Chemical Industrials, Kyoto, Japan) under isothermal conditions (40 °C). Elusion of ethylene (at 0.64 min in this condition) was detected by an FID detector. After the assay of ethylene formation assay, the cells were collected for measurement of optical density at 730 nm (OD730) with a spectrophotometer (model UV1700 or UV1600; Shimadzu). Chlorophyll content of the suspension (500 µl) was also determined as described53.
Determination of the relationship between OD730 and CDW
CDWs of cell suspensions of Synechocystis 6803 and L. boryana with various OD730 values (0 to 50) were measured as described in Kato et al.54, and a linear relationship between CDW and OD730 was determined for each species (Supplementary Fig. 3). CDW was estimated from OD730 based on the linear relationship between them.
Determination of respiratory and photosynthesis activities with an oxygen electrode
Cells of WT (strain YF), CN1, CN2 and CN3 were subjected to induction of the nif genes as described above. The cells harvested with 1.5 ml of BG-110 liquid medium were diluted three-fold with BG-110, an aliquot (0.99 ml) of the cell suspension was used for measurement of oxygen consumption in the dark and oxygen evolution in the light with a Clark-type oxygen electrode (DW1, Hansatech, Norfolk, UK). Sodium bicarbonate (10 µl of 1 M, final 10 mM) was added just before measurement.
Western blot analysis
Cells were grown on BG-11 agar plates under light conditions (50 µmol m−2 s−1) at 30 °C for 3 days as a pre-culture. The cells were collected in distilled water and adjusted to OD730 15.6. Aliquots of cell suspension (500 µl) were spread uniformly to form 4-cm diameter circles on BG-110 agar plates. Plates were then incubated in an anaerobic jar under light conditions (50 µmol m−2 s−1) at 30 °C for 14 h. Induced cells in the anaerobic chamber were harvested in 700 µl of protein extraction buffer (50 mM HEPES-KOH; pH 7.5, 10 mM MgCl2) and an aliquot (500 µl) was transferred into a 1.5-ml micro-centrifuge tube containing glass beads (100 mg glass beads; 150–212 microns, Sigma, St. Louis, MO). The cells were disrupted by vigorous shaking (Setting LA; “BugCrasher” GM01, Taitec, Saitama, Japan) at 4 °C for 1 h. The resultant homogenates were centrifuged at 13,600 × g for 3 min at 4 °C to prepare the supernatant fraction. Protein concentration was determined by Bradford assay (Protein Assay, Bio-Rad, Hercules, CA) with bovine serum albumin as the standard. Western blot analysis was carried out as described previously55. The specific protein bands were visualized by a lumino-image analyzer (LAS-3000mini, Fujifilm, Tokyo, Japan). The obtained images were converted to tiff images by Multi Gauge software (Fujifilm). The signal intensities of the NifH, NifD, and NifK proteins on the tiff images were quantified by ImageJ ver 1.50i.
The NifH, NifD, and NifK proteins were detected by specific antisera prepared against L. boryana NifH, NifD, and NifK proteins as Strep-fusion proteins (Scrum, Tokyo, Japan). These proteins were expressed in E. coli and purified by Strep-Tactin affinity column.
References
Hoffman, B. M., Lukoyanov, D., Yang, Z. Y., Dean, D. R. & Seefeldt, L. C. Mechanism of nitrogen fixation by nitrogenase: the next stage. Chem. Rev. 114, 4041–4062 (2014).
Robson, R. L. et al. The alternative nitrogenase of Azotobacter chroococcum is a vanadium enzyme. Nature 322, 388–390 (1986).
Hu, Y., Lee, C. C. & Ribbe, M. W. Vanadium nitrogenase: a two-hit wonder? Dalton Trans. 41, 1118–1127 (2012).
Müller, A., Schneider, K., Knüttel, K. & Hagen, W. R. EPR spectroscopic characterization of an ‘iron only’ nitrogenase. S = 3/2 spectrum of component 1 isolated from Rhodobacter capsulatus. FEBS Lett. 303, 36–40 (1992).
Yang, J., Xie, X., Wang, X., Dixon, R. & Wang, Y. P. Reconstruction and minimal gene requirements for the alternative iron-only nitrogenase in Escherichia coli. Proc. Natl. Acad. Sci. USA 111, E3718–E3725 (2014).
Seefeldt, L. C., Hoffman, B. M. & Dean, D. R. Mechanism of Mo-dependent nitrogenase. Annu. Rev. Biochem. 78, 701–722 (2009).
Hoffman, B. M., Dean, D. R. & Seefeldt, L. C. Climbing nitrogenase: toward a mechanism of enzymatic nitrogen fixation. Acc. Chem. Res. 42, 609–619 (2009).
Seefeldt, L. C., Hoffman, B. M. & Dean, D. R. Electron transfer in nitrogenase catalysis. Curr. Opin. Chem. Biol. 16, 19–25 (2012).
Yates, M. & Planqué, K. Nitrogenase from Azotobacter chroococcum. Purification and properties of the component proteins. Eur. J. Biochem. 60, 467–476 (1975).
Hu, Y. & Ribbe, M. W. Biosynthesis of the metalloclusters of nitrogenases. Annu. Rev. Biochem. 85, 455–483 (2016).
Erisman, J. W., Sutton, M. A., Galloway, J., Klimont, Z. & Winiwarter, W. How a century of ammonia synthesis changed the world. Nat. Geosci. 1, 636–639 (2008).
Galloway, J. N. et al. Transformation of the nitrogen cycle: recent trends, questions, and potential solutions. Science 320, 889–892 (2008).
Sutton, M. A. et al. Too much of a good thing. Nature 472, 159–161 (2011).
Beatty, P. H. & Good, A. G. Plant science. Future prospects for cereals that fix nitrogen. Science 333, 416–417 (2011).
Curatti, L. & Rubio, L. M. Challenges to develop nitrogen-fixing cereals by direct nif-gene transfer. Plant Sci. 225, 130–137 (2014).
Wang, L. et al. A minimal nitrogen fixation gene cluster from Paenibacillus sp. WLY78 enables expression of active nitrogenase in Escherichia coli. PLoS Genet. 9, e1003865, https://doi.org/10.1371/journal.pgen.1003865 (2013).
Stokstad, E. The nitrogen fix. Science 353, 1225–1227 (2016).
Yang, J., Xie, X., Yang, M., Dixon, R. & Wang, Y. P. Modular electron-transport chains from eukaryotic organelles function to support nitrogenase activity. Proc. Natl. Acad. Sci. USA 114, E2460–E2465 (2017).
Yamamoto, H., Kusumi, J., Yamakawa, H. & Fujita, Y. The effect of two amino acid residue substitutions via RNA editing on dark-operative protochlorophyllide oxidoreductase in the black pine chloroplasts. Sci. Rep. 7, 2377, https://doi.org/10.1038/s41598-017-02630-2 (2017).
Yamamoto, H., Kurumiya, S., Ohashi, R. & Fujita, Y. Functional evaluation of a nitrogenase-like protochlorophyllide reductase encoded by the chloroplast DNA of Physcomitrella patens in the cyanobacterium Leptolyngbya boryana. Plant Cell Physiol. 52, 1983–1993 (2011).
Ueda, M., Tanaka, A., Sugimoto, K., Shikanai, T. & Nishimura, Y. chlB requirement for chlorophyll biosynthesis under short photoperiod in Marchantia polymorpha L. Genome Biol. Evol. 6, 620–628 (2014).
Stal, L. J. & Zehr, J. P. Cyanobacterial nitrogen fixation in the ocean: Diversity, regulation, and ecology in The Cyanobacteria: Molecular Biology, Genomics and Evolution, (eds Herrero, A. & Flores, E.) 423–446 (Caister Academic Press, 2008).
Kumar, K., Mella-Herrera, R. A. & Golden, J. W. Cyanobacterial heterocysts. Cold Spring Harb. Perspect. Biol. 2, a000315, https://doi.org/10.1101/cshperspect.a000315 (2010).
Grobbelaar, N., Huang, T. C., Lin, H. Y. & Chow, T. J. Dinitrogen-fixing endogenous rhythm in Synechococcus RF-1. FEMS Microbiol. Lett. 37, 173–177 (1986).
Mitsui, A. et al. Strategy by which nitrogen-fixing unicellular cyanobacteria grow photosynthetically. Nature 323, 720–722 (1986).
Stal, L. J. & Krumbein, W. E. Isolation and characterization of cyanobacteria from a marine microbial mat. Bot. Mar. 28, 351–365 (1985).
Capone, D. G., O’neil, J. M., Zehr, J. & Carpenter, E. J. Basis for diel variation in nitrogenase activity in the marine planktonic cyanobacterium Trichodesmium thiebautii. Appl. Environ. Microbiol. 56, 3532–3536 (1990).
Bergman, B., Gallon, J. R., Rai, A. N. & Stal, L. J. N2 fixation by non-heterocystous cyanobacteria. FEMS Microbiol. Rev. 19, 139–185 (1997).
Tsujimoto, R., Kamiya, N. & Fujita, Y. Transcriptional regulators ChlR and CnfR are essential for diazotrophic growth in nonheterocystous cyanobacteria. Proc. Natl. Acad. Sci. USA 111, 6762–6767 (2014).
Tsujimoto, R., Kamiya, N. & Fujita, Y. Identification of a cis-acting element in nitrogen fixation genes recognized by CnfR in the nonheterocystous nitrogen-fixing cyanobacterium Leptolyngbya boryana. Mol. Microbiol. 101, 411–424 (2016).
Nagatani, H. H. & Haselkorn, R. Molybdenum independence of nitrogenase component synthesis in the non-heterocystous cyanobacterium Plectonema. J. Bacteriol. 134, 597–605 (1978).
Schrautemeier, B., Cassing, A. & Böhme, H. Characterization of the genome region encoding an fdxH-type ferredoxin and a new 2[4Fe-4S] ferredoxin from the nonheterocystous, nitrogen-fixing cyanobacterium Plectonema boryanum PCC 73110. J. Bacteriol. 176, 1037–1046 (1994).
Razquin, P. et al. Differential activities of heterocyst ferredoxin, vegetative cell ferredoxin, and flavodoxin as electron carriers in nitrogen fixation and photosynthesis in Anabaena sp. Photosynth. Res. 43, 35–40 (1995).
Minamizaki, K., Mizoguchi, T., Goto, T., Tamiaki, H. & Fujita, Y. Identification of two homologous genes, chlA I and chlA II , that are differentially involved in isocyclic ring formation of chlorophyll a in the cyanobacterium Synechocystis sp. PCC 6803. J. Biol. Chem. 283, 2684–2692 (2008).
Ermakova, M. et al. Heterocyst-specific flavodiiron protein Flv3B enables oxic diazotrophic growth of the filamentous cyanobacterium Anabaena sp. PCC 7120. Proc. Natl. Acad. Sci. USA 111, 11205–11210 (2014).
Mouncey, N. J., Mitchenall, L. A. & Pau, R. N. Mutational analysis of genes of the mod locus involved in molybdenum transport, homeostasis, and processing in Azotobacter vinelandii. J. Bacteriol. 177, 5294–5302 (1995).
Wagner, U. G., Stupperich, E. & Kratky, C. Structure of the molybdate/tungstate binding protein Mop from Sporomusa ovata. Structure 8, 1127–1136 (2000).
Valladares, A., Herrero, A., Pils, D., Schmetterer, G. & Flores, E. Cytochrome c oxidase genes required for nitrogenase activity and diazotrophic growth in Anabaena sp. PCC 7120. Mol. Microbiol. 47, 1239–1249 (2003).
Dixon, R. A. & Postgate, J. R. Transfer of nitrogen-fixation genes by conjugation in Klebsiella pneumoniae. Nature 234, 47–48 (1971).
Dixon, R., Cannon, F. & Kondorosi, A. Construction of a P plasmid carrying nitrogen fixation genes from Klebsiella pneumoniae. Nature 260, 268–271 (1976).
López-Torrejón, G. et al. Expression of a functional oxygen-labile nitrogenase component in the mitochondrial matrix of aerobically grown yeast. Nature Communications 7(11426) (2016).
Burén, S. et al. Formation of nitrogenase NifDK tetramers in the mitochondria of Saccharomyces cerevisiae. ACS Synth. Biol. 6, 1043–1055 (2017).
Allen, R. S. et al. Expression of 16 nitrogenase proteins within the plant mitochondrial matrix. Front. Plant Sci. 8, 287, https://doi.org/10.3389/fpls.2017.00287 (2017).
Ivleva, N. B., Groat, J., Staub, J. M. & Stephens, M. Expression of active subunit of nitrogenase via integration into plant organelle genome. PLoS One 11, e0160951, https://doi.org/10.1371/journal.pone.0160951 (2016).
Hiraide, Y. et al. Loss of cytochrome c M stimulates cyanobacterial heterotrophic growth in the dark. Plant Cell Physiol. 56, 334–345 (2015).
Aoki, R., Takeda, T., Omata, T., Ihara, K. & Fujita, Y. MarR-type transcriptional regulator ChlR activates expression of tetrapyrrole biosynthesis genes in response to low-oxygen conditions in cyanobacteria. J. Biol. Chem. 287, 13500–13507 (2012).
Suzuki, I., Sugiyami, T. & Omata, T. Regulation by cyanate of the genes involved in carbon and nitrogen assimilation in the cyanobacterium Synechococcus sp. strain PCC 7942. J. Bacteriol. 178, 2688–2694 (1996).
Yamazaki, S., Nomata, J. & Fujita, Y. Differential operation of dual protochlorophyllide reductases for chlorophyll biosynthesis in response to environmental oxygen levels in the cyanobacterium Leptolyngbya boryana. Plant Physiol. 142, 911–922 (2006).
Black, T. A., Cai, Y. & Wolk, C. P. Spatial expression and autoregulation of hetR, a gene involved in the control of heterocyst development in Anabaena. Mol. Microbiol. 9, 77–84 (1993).
Fujita, Y., Takahashi, Y., Chuganji, M. & Matsubara, H. The nifH-like (frxC) gene is involved in the biosynthesis of chlorophyll in the filamentous cyanobacterium Plectonema boryanum. Plant and Cell Physiol. 33, 81–92 (1992).
Prentki, P., Karch, F., Iida, S. & Meyer, J. The plasmid cloning vector pBR325 contains a 482 base-pair-long inverted duplication. Gene 14, 289–299 (1981).
Alting-Mees, M. A. & Short, J. M. pBluescript II: gene mapping vectors. Nucleic Acids Res. 17, 9494 (1989).
Aoki, R., Goto, T. & Fujita, Y. A heme oxygenase isoform is essential for aerobic growth in the cyanobacterium Synechocystis sp. PCC 6803: Modes of differential operation of two isoforms/enzymes to adapt to low oxygen environments in cyanobacteria. Plant Cell Physiol. 52, 1744–1756 (2011).
Kato, A., Takatani, N., Ikeda, K., Maeda, S. & Omata, T. Removal of the product from the culture medium strongly enhances free fatty acid production by genetically engineered. Biotechnol. Biofuels 10, 141, https://doi.org/10.1186/s13068-017-0831-z (2017).
Aoki, R., Hiraide, Y., Yamakawa, H. & Fujita, Y. A novel “oxygen-induced” greening process in a cyanobacterial mutant lacking the transcriptional activator ChlR involved in low-oxygen adaptation of tetrapyrrole biosynthesis. J. Biol. Chem. 289, 1841–1851 (2014).
Acknowledgements
We thank Haruki Yamamoto for preliminary work on this project and for preparing the antisera against NifD and NifK from L. boryana. We thank Peter Wolk for the generous gift of pRL271. We thank Kazuma Uesaka and Kunio Ihara for genome resequencing of CN strains. We also thank Nobuyuki Takatani, Akihiro Kato, and Tatsuo Omata for their technical help in measuring oxygen evolution and consumption with an oxygen electrode and in determining CDW. We also thank Takafumi Yamashino, Tatsuo Omata and Kazuki Terauchi for valuable discussion. This work was supported by the Japan Society for the Promotion of Science (Grants-in-Aid for Scientific Research Nos 26660084, 15H04387, 15H01397, and 17H05525), the Japan Science and Technology Agency (Advanced Low Carbon Technology Research and Development Program, Precursory Research for Embryonic Science and Technology and JST-Mirai Program).
Author information
Authors and Affiliations
Contributions
Y.F. conceived and supervised the study; Y.F. and R.T. designed experiments; R.T. and H.K. performed most experiments; H.Y. and A.N. constructed some plasmids; K.Y. and H.Y. measured respiratory activity of the CN strains; H.Y. analyzed image data and determined CDW; Y.F. and R.T. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsujimoto, R., Kotani, H., Yokomizo, K. et al. Functional expression of an oxygen-labile nitrogenase in an oxygenic photosynthetic organism. Sci Rep 8, 7380 (2018). https://doi.org/10.1038/s41598-018-25396-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-25396-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.