Abstract
Programmed death ligand 1 (PD-L1)-mediated induction of immune tolerance has been vigorously investigated in autoimmunity and anti-tumor immunity. However, details of the mechanism by which PD-L1 is induced in CD4+ T cells are unknown. Here, we revealed the potential function of Klf1 and Egr2-mediated induction of PD-L1 in CD4+ T cells. We focused on the molecules specifically expressed in CD4+CD25−LAG3+ regulatory T cells (LAG3+ Tregs) highly express of PD-L1 and transcription factor Egr2. Although ectopic expression of Egr2 induced PD-L1, a deficiency of Egr2 did not affect its expression, indicating the involvement of another PD-L1 induction mechanism. Comprehensive gene expression analysis of LAG3+ Tregs and in silico binding predictions revealed that Krüppel-like factor 1 (Klf1) is a candidate inducer of the PD-L1 gene (Cd274). Klf1 is a transcription factor that promotes β-globin synthesis in erythroid progenitors, and its role in immunological homeostasis is unknown. Ectopic expression of Klf1 induced PD-L1 in CD4+ T cells through activation of the PI3K-mTOR signaling pathway, independent of STATs signaling and Egr2 expression. Our findings indicate that Klf1 and Egr2 are modulators of PD-L1-mediated immune suppression in CD4+ T cells and might provide new insights into therapeutic targets for autoimmune diseases and malignancies.
Similar content being viewed by others
Introduction
The pathological bases of autoimmune diseases such as rheumatoid arthritis (RA) and systemic lupus erythematosus (SLE) have gradually been elucidated1,2. Vigorous analyses of cytokines and inhibitory cell surface molecules of immune cells have led to novel treatment methods. Tumor necrosis factor-α (TNF-α) inhibitors, antibodies against the interleukin-6 (IL-6) receptor and cytotoxic T lymphocyte associated protein-4 (CTLA-4) immunoglobulin (Ig) fusion protein have been widely used in the clinic3. In inflammatory conditions, T cell responses are regulated both positively and negatively by cell surface molecules. CD28/CD80 (B7-1), CD28/CD86 (B7-2), inducible costimulator (ICOS) and its ligand ICOSL stimulate immune responses. In contrast, immune responses are inhibited by binding of CTLA4/CD80, CTLA4/CD86 and lymphocyte activation gene 3 (LAG3)/MHC class II4. Recently, a B7 family co-inhibitory molecule, Programmed death-ligand 1 (PD-L1, encoded by Cd274), belonging to the CD28 receptor family5, has become a subject of active investigation. Engagement of PD-1 by PD-L1 or PD-L2 transduces a signal that inhibits T-cell proliferation, cytokine production, and cytolytic function. PD-L1 and PD-L2 have marked structural similarities, but they display different expression patterns. Although the expression of PD-L2 is restricted to activated dendritic cells and macrophages6, PD-L1 is expressed by activated CD4+ T cells, CD8+ T cells, natural killer cells, activated monocytes, myelocytes and CD4-CD8- (double negative: DN) T cells in the thymus. PD-L1 is also expressed in non-hematopoietic organs such as heart, lung, spleen, thymus and kidney7. High expression of PD-L1 has been observed in various tumors including lung cancer and pancreatic cancer7.
Whereas PD-L1 is induced by stimulation of the T cell receptor (TCR)8, CD4+ regulatory T cells (Tregs) constitutively express PD-L1 in the steady state9,10. Tregs play a major role in maintaining immune tolerance and are divided into two types: those emerging from the thymus and those induced in the periphery. CD4+CD25+ Tregs (CD25+ Tregs) mainly emerge from the thymus and express the transcription factor forkhead protein 3 (Foxp3) as a master regulator gene (Foxp3)11. CD25+ Tregs inhibit effector cell proliferation and cytokine production8,12 and directly suppress B cell activation via PD-L113. Immunological homeostasis is thought to be maintained by CD25+ Tregs in concert with peripherally induced Foxp3-independent Tregs. We reported CD4+CD25− Tregs that characteristically express LAG3 and Early growth response gene 2 (Egr2) and produce immune suppressive cytokine IL-1014,15. Egr2 is a zinc-finger transcription factor that plays an important role in the maintenance of T cell anergy by negatively regulating T cell activation16. We previously reported that forced expression of Egr2 in naïve T cells induces LAG3, IL-10, and transcription factor Blimp-1, indicating that Egr2 confers the phenotype of CD4+CD25−LAG3+ Tregs (LAG3+ Tregs) on CD4+ T cells14,17. LAG3+ Tregs develop in the periphery and exert their suppressive activities in a Foxp3-independent manner14. LAG3+ Tregs also produce large amounts of the inhibitory cytokine transforming growth factor-β3 (TGF-β3), which improves the pathology of lupus in a murine model (MRL-Faslpr/lpr)10. LAG3+ Tregs lacking Egr2 attenuate TGF-β3 production and lose suppressive ability for B cells10. Interestingly, LAG3+ Tregs express high levels of PD-L1. Moreover, TGF-β3-mediated suppression of B cells is dependent on PD-1 expression on B cells10. Those observations indicate that PD-L1 expression on LAG3+ Tregs play a critical role in the maintenance of immune tolerance. Taken together, these findings suggest that elucidating the molecular control of the PD-1/PD-L1 axis in CD4+ T cells is important for clinical applications targeting costimulatory molecules.
In the current study, we show new functions of transcription factor Krüppel like factor 1 (Klf1) and Egr2 in the induction of PD-L1 in CD4+ T cells. Klf1 promotes β-globin synthesis in erythroid progenitor cells and its role in immunological homeostasis has not been investigated. Mechanisms that induce PD-L1 expression have been studied mainly in immune cells and tumor cells, and the induction mechanisms of PD-L1 appear to be dependent on cell type. However, the detailed mechanism by which PD-L1 is induced in CD4+ T cells remains to be determined. Moreover, it is important to elucidate the cell type-specific mechanism of PD-L1 induction in CD4+ T cells. Here, we reveal that Klf1 and Egr2, both of which are characteristically expressed in PD-L1-expressing LAG3+ Tregs, induce PD-L1 in CD4+ T cells independently of each other.
Results
Egr2 induces PD-L1 expression in CD4+ T cells
Constitutive high level expression of PD-L1 in the steady state occurs only in Tregs, such as CD25+ Tregs and LAG3+ Tregs9,10. We first confirmed our previous findings10,14 that among CD4+ T cell subsets, both Egr2 and PD-L1 were most highly expressed in LAG3+ Tregs (Fig. 1a–c). Then, to elucidate whether overexpression of Egr2 induced PD-L1 protein, we utilized the pMIG retroviral vector containing IRES-regulated green fluorescent protein (GFP) as a reporter to generate a pMIG-Egr2 vector (Fig. 1d). As expected, forced expression of Egr2 in naïve CD4+ T cells induced PD-L1 protein expression (Fig. 1e). These findings suggest the important role of Egr2 in the induction of PD-L1 in CD4+ T cells.
Egr2-mediated PD-L1 induction in CD4+ T cells. (a) LAG3 expression in splenocytes from C57BL/6 (B6) mice was analyzed by flow cytometry (FCM). CD25+ Tregs: CD4+CD25+ T cells; naïve T: CD4+CD25−CD45RBhigh T cells; LAG3+ Tregs: CD4+CD25−CD45RBlowLAG3+ T cells. (b) PD-L1 expression in T cell subsets. Histograms are gated on CD4+ T cells (left). Data are representative of three independent experiments. Mean fluorescence intensity (MFI) of the indicated T cell subsets is shown (right). (n = 3 per group). *p < 0.001 (Bonferroni’s multiple comparison test) (c) Quantitative PCR assessing Egr2 mRNA expression in the indicated T cell subsets (n = 3 per group). (d) Retroviral constructs of pMIG vector for the transduction of Egr2. (e) Evaluation of PD-L1 expression in Egr2 gene-transduced CD4+ T cells. MFI ratio represents the PD-L1 MFI signals of GFP negative versus GFP positive. Plots and histogram are the representative of 3 independent experiments.
Egr2 is not essential for the induction of PD-L1 expression in CD4+ T cells
To examine whether expression of PD-L1 in LAG3+ Tregs was dependent on Egr2, we employed T cell-specific Egr2 conditional knockout mice (Egr2fl/flCD4Cre+: Egr2 CKO). Unexpectedly, under physiological conditions, LAG3+ Tregs from Egr2 CKO mice expressed PD-L1 protein to the same extent as the littermate control mice (Fig. 2a). It has been reported that TCR stimulation induces the upregulation of PD-L1 in CD4+ T cells.8 Here, we found that the expression levels of PD-L1 in Egr2-deficient CD4+ T cells stimulated with anti-CD3ε and anti-CD28 antibodies in vitro was not affected (Fig. 2b). These results indicate that Egr2 is not necessary for the induction of PD-L1 in either LAG3+ Tregs or activated CD4+ T cells, suggesting the involvement of another PD-L1 induction mechanism in CD4+ T cells.
PD-L1 expression in CD4+ T cells that were Egr2-deficient. (a) PD-L1 expression in splenocytes from T cell-specific Egr2 conditional knockout (Egr2fl/flCD4Cre+: Egr2 CKO) mice and control mice analyzed by FCM. (b) CD4+ T cells purified from each mouse were analyzed by FCM after 48 h stimulation with the indicated concentration of anti-CD3ε/anti-CD28 antibodies. Histograms are the representative of 3 independent experiments.
Klf1 induces PD-L1 expression in CD4+ T cells independent of Egr2
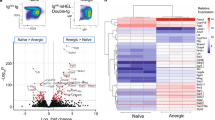
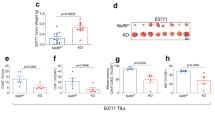
To analyze other transcription factors that induce PD-L1 expression, we selected 1562 LAG3+ Treg-specific genes from microarray data of T cell subsets, including freshly isolated naïve T cells, CD25+ Tregs, LAG3+ Tregs and CD4+CD25−CD45RBlowLAG3− memory T cells registered in the ArrayExpress database (access number E-MEXP-1343)14. Furthermore, we performed a transcription factor binding site enrichment analysis using the JASPAR Database18, an open-access database for eukaryotic transcription factor binding profiles. Then, we combined a transcriptomics and in silico binding prediction approach to identify PD-L1-inducible genes in LAG3+ Tregs. As shown in Figs 3a, 5 genes overlapped in these two data sets: signal transducer and activator of transcription 1 (Stat1), Stat3, Klf1, T-cell acute lymphocytic leukemia protein 1 (Tal1), and Egr2. Stat1 and Stat3 directly bind to the promoter region of the Cd274 gene19,20. Intriguingly, transcription factor genes Klf1 and Tal1 were picked up as novel candidate PD-L1-inducible genes. To validate the expression profiles obtained by microarray data, qRT-PCR was performed on Klf1 and Tal1 genes, and both genes were characteristically upregulated in LAG3+ Tregs (Figs 3b and S1). PD-L1 protein was induced in pMIG-Klf1 transfected cells, but not pMIG-Tal1 transduced cells (Figs 3c,d, and S2). We next focused on the mechanism by which Klf1 induced PD-L1. We examined the dependency of Egr2 expression on Klf1-mediated induction of PD-L1 in CD4+ T cells. Expression of Egr2 was not induced in Klf1-transduced CD4+ T cells (Fig. 3e). On the other hand, Klf1 was not induced in Egr2-transduced CD4+ T cells (Fig. 3f). Moreover, CD4+ T cells from Egr2 CKO mice expressed PD-L1 when Klf1 was overexpressed (Fig. 3g). Although we and others have shown that Egr3 compensates for the function of Egr2 during Egr2 deficiency17,21, PD-L1 expression was induced by Klf1 in CD4+ T cells from Egr3fl/flEgr2fl/flCD4Cre+ (Egr2/3 DKO) mice (Fig. S3). These results clearly indicate that Klf1 induces PD-L1 in CD4+ T cells independently of Egr2/Egr3 expression.
Egr2-independent induction of PD-L1 expression in CD4+ T cells by ectopic expression of Klf1. (a) Venn diagram representing the genomic overlap between LAG3+ Tregs specific transcription factors from microarray analysis of CD4+ T cell subsets from ArrayExpress database (E-MEXP-1343) and PD-L1-inducible candidate genes predicted by JASPAR database. (b) Gene expression levels of Klf1 in CD4+ T cells. Klf1 gene expression in indicated T cell subsets relative to unstimulated naïve CD4+ cells using micro array data set as in (a) (left) and relative to Actb mRNA of each indicated T cell subset confirmed by qRT-PCR (right). *p < 0.001 (Bonferroni’s multiple comparison test) (c) Evaluation of PD-L1 expression in Klf1 gene-transduced CD4+ T cells. Klf1 cDNA was inserted into the retrovirus vector, pMIG with a GFP reporter gene. pMIG-Mock or pMIG-Klf1 was transfected into CD4+ T cells. MFI ratio represents the PD-L1 MFI signals of GFP negative versus GFP positive. (d) GFP+ and GFP- cells in (c) were fractionated and mRNA expression of Cd274 (encoding PD-L1) was evaluated by qRT-PCR (n = 3). (e,f) Egr2 and Klf1 mRNA expression was evaluated by qRT-PCR using the cells transfected in (c) and Fig. 1d. (g) Klf1 and Cd274 mRNA expression in Klf1 gene-transduced CD4+ T cells from Egr2fl/flCD4Cre+ mice was evaluated by qRT-PCR (n = 3). *p < 0.05; n.s.: not significant (unpaired two-tailed Student’s t-test) Plots are representative of 3 independent experiments.
Marked induction of the Klf1 gene in CD4+ T cells modulates the gene expression profile
To clarify the mechanism by which PD-L1 is induced by Klf1 in CD4+ T cells, changes in the gene expression profile in pMIG-Klf1- or pMIG-Mock-transduced cells were divided into three groups according to GFP expression levels and analyzed by RNA sequencing (Fig. 4a). Principal component analysis (PCA) of these cells showed that Klf1-transduced cells (Klf1 GFPhigh) had unique characteristics compared with the other cells (Fig. 4b). Pathway analysis of the differentially expressed genes (DEGs) was conducted using Ingenuity Pathways Analysis software (IPA). IPA analysis showed candidate intracellular signal transduction pathways induced by Klf1, which include mammalian target of rapamycin (mTOR) signaling pathway as the top canonical pathway, indicating that mTOR signaling played a crucial role in Klf1-mediated PD-L1 expression (Fig. 4c).
Comprehensive analysis of gene expression profiles in Klf1-transduced CD4+ T cells. (a) Sorting strategy for next generation sequencing (NGS) of Klf1-gene-transduced CD4+ T cells. Splenocytes from WT mice were transfected with pMIG-Mock or pMIG-Klf1 vector according to their GFP expression. (b) Principal component analysis plots for NGS data of the indicated CD4+ T cell subsets as in (a,c) Differentially expressed genes of NGS data sets in (b) were analyzed by pathway analysis tool Ingenuity Pathway Analysis (IPA) software. The top 10 most significantly enriched canonical pathways in IPA are shown.
Inhibition of the PI3K-mTOR signaling pathway decreases Klf-1-mediated induction of PD-L1 expression in CD4+ T cells
Consistent with our own analysis (Fig. 3a), previous reports showed that Stat1 and Stat3 directly bind to the promoter region of Cd27419,20,22,23, suggesting the important role of STATs signaling in the induction of PD-L1. The induction mechanism of PD-L1 is cell-type dependent (see Discussion). To clarify the nature of the Klf1-mediated PD-L1 induction mechanism, we employed Stat1 KO mice, T cell-specific Stat3 KO mice (Stat3fl/flCD4Cre+: Stat3 CKO), Stat4 KO mice, Stat5a KO mice, Stat6 KO mice and wild-type (WT) mice. Unexpectedly, the expression levels of PD-L1 on Klf1-transduced CD4+ T cells were not affected by Stat1, Stat3, Stat4, Stat5a nor Stat6 deficiency, indicating that these STATs do not regulate Klf1-mediated upregulation of PD-L1 in CD4+ T cells (Fig. 5a).
Klfl-mediated PD-L1 induction requires the activation of PI3K-mTOR signaling pathway. (a) Splenocytes from each Stat-deficient or WT mouse were transfected with a pMIG-Mock or pMIG-Klf1 vector. PD-L1 expression levels were assessed by FCM 48 h after the transfection. In the Stat1 KO mouse, anti-CD3ε/anti-CD28 stimulation was used whereas Concanavalin A (ConA) stimulation was used for the other KO mice. (b) Phosphorylation of Akt and S6K was analyzed by Western blotting. The indicated blots were derived from the same experiments. Full length blots are presented in Supplementary Figure S4. (c) Splenocytes from B6 mice were transfected with pMIG-Mock or pMIG-Klf1 and the indicated inhibitors were added 24 h later. Then, cells were cultured for 48 h and PD-L1 expression was evaluated by FCM. The proportion of cells expressing PD-L1 at high levels was compared between the GFP-positive and GFP-negative groups. (n = 3). Control: without inhibitor; SP 600125: JNK1/2/3 inhibitor (5 μM); SB 203580: MAPK-p38 inhibitor (20 μM); PD 98059: MEK/ERK inhibitor (50 μM); LY 294002: PI3K inhibitor 20 μM); Torin: mTOR inhibitor (200 nM). *p < 0.01 (Bonferroni’s multiple comparison test). Plots are representative of 3 independent experiments.
The PI3K-Akt-mTOR signaling pathway is activated in TCR-stimulated T cells and affects the fate of Th cells and Tregs24,25,26,27. Our IPA analysis of DEGs in Klf1-transduced cells suggested the PI3K-Akt-mTOR signaling pathway is involved in the Klf1-mediated induction of PD-L1. To verify this hypothesis, PI3K-Akt-mTOR cellular signaling pathways were evaluated by Western blot analysis. Unexpectedly, there was no difference in the phosphorylation levels of Akt and its downstream effector (S6K protein) between Klf1-transduced CD4+ T cells and mock-transduced CD4+ T cells (Figs 5b, S4). To examine whether PD-L1 expression was dependent on active PI3K-AKT-mTORsignaling, Klf1-transduced CD4+ T cells were treated with pharmacologic inhibitors, including a JNK1/2/3 inhibitor (SP600125), a p38 inhibitor (SB203580), an MEK inhibitor (PD9805981), a PI3K inhibitor (LY294002) and an mTOR inhibitor (Torin1). Treatment with LY294002 and Torin1 significantly reduced the Klf1-mediated induction of PD-L1 (Fig. 5c). These results indicate that a basal level PI3K-Akt-mTOR signaling is necessary for PD-L1 induction by Klf1 in CD4+ T cells, suggesting that Klf1 and the PI3K-Akt-mTOR signaling pathway might cooperate to induce PD-L1 expression on CD4+ T cells.
Discussion
Previous work has established that PD-L1 has critical functions as an immunosuppressive cell surface molecule. However, the precise mechanisms by which PD-L1 is induced on CD4+ T cell have long remained elusive. Here, we showed that the expression of PD-L1 on CD4+ T cells is induced by Klf1 and Egr2. Klf1 was predicted by in silico analysis from the gene expression profile of Egr2-expressing LAG3+ Tregs that characteristically express PD-L1. Klf1-mediated PD-L1 induction by CD4+ T cells, and it was partially dependent on the PI3K-AKT-mTOR signaling pathway, but independent of Egr2 and STATs signaling. Our findings have important implications for understanding the molecular mechanisms underlying cell-mediated immune tolerance modulated by CD4+ T cells that express PD-L1.
Interaction of PD-1 and its ligand PD-L1 or PD-L2 negatively regulates intracellular signaling by recruiting protein tyrosine phosphatases, SHP-1 and SHP-228. PD-L1 KO mice show enhanced Th1 responses, exacerbation of experimental autoimmune encephalomyelitis and autoimmune hepatitis29,30. In human, the clinical relevance of genetic polymorphisms in the human CD274 gene, encoding PD-L1, has been reported in several autoimmune diseases, such as Addison’s disease and Graves’ disease31,32. On the other hand, deficiency of PD-L2 results in orally administered antigen intolerance33, indicating different roles for PD-L1 and PD-L2 in immunity and pathogenesis. Although the expression of PD-L2 is restricted mainly to activated dendritic cells and macrophages6, PD-L1 is expressed in various immune cells, including activated CD4+ T cells and Treg subsets. PD-1/PD-L1 interaction is also known as an inhibitory mechanism in CD25+ Tregs13 and LAG3+ Tregs. These findings clearly suggest that PD-L1 expression on CD4+ T cells is important for maintenance of immune tolerance.
We previously reported that forced expression of Egr2 in CD4+ T cells provides the functional traits of LAG3+ Treg, such as induced expression of Prdm1, IL-10 and LAG314. However, the role of Egr2 in induction of PD-L1 in CD4+ T cells has not been clarified. We demonstrated that Egr2 is not required for PD-L1 expression on CD4+ T cells. It has been reported that among the 4 Egr family members, Egr3 compensates for the function of Egr2 when the latter is deficient21. We reported that expression of latent TGF-β binding protein (Ltbp)-3 was maintained by both Egr2 and Egr3 and was required for TGF-β3 production from LAG3+ Tregs17. However, T cells that were doubly-deficient for Egr2 and Egr3 were not affected in their Klf-1-mediated induction of PD-L1, indicating that Klf-1 induces PD-L1 in CD4+ T cells via an Egr2/Egr3- independent manner.
These findings indicate a possibility of the involvement of another Egr2-independent PD-L1 induction mechanism in CD4+ T cells, and we focused on Klf1 based on the expression on LAG3+ Tregs. The Klf family is a transcription factor group composed of 17 genes, many of which are important for immune homeostasis. Klf1 is also called erythroid KLF (EKLF) and is predominantly expressed in erythroid progenitor cells. Klf1 has important functions in a variety of mechanisms, including gene activation and repression, regulation of chromatin arrangement, transcription initiation and elongation34. Klf1 binds to the CACCC site in the promoter region of the β-globin gene35,36. In humans, KLF1 also plays a role in transcribing the β-globin gene. A mutation in the CACCC region causes β-thalassemia37,38. Although Klfl is characterized primarily as a transcriptional activator, it can also act as a repressor that interacts with the components of the corepressor Sin3A and the NuRD complex39,40. KLF1 participates in the repression of the fetal γ-globin genes in human adult erythroid progenitors by regulating the expression of BCL11A. However, the role of Klf1 in the immune system has not been clarified since deficiency of Klf1 results in embryonic lethality due to congenital thalassemia41. In the present study, we revealed a new immunological function of Klf1 in PD-L1 induction in CD4+ T cells that was achieved in an Egr2/Egr3-independent manner.
The induction mechanism of PD-L1 appears to depend on the cell type. Extracellular stimuli and intracellular signaling mechanisms that contribute to the expression of PD-L1 have been studied mainly in tumor cells. A cell line derived from lung cancer cells (A549) expresses PD-L1 following interferon (IFN)-γ stimulation, but PD-L1 expression is attenuated by a Jak/Stat inhibitor42. Mouse and human T cell lymphoma cells express PD-L1 in the steady state, and Stat3 binds to the promoter region of the Cd274 gene encoding PD-L119,22,23. In Hodgkin’s lymphoma cells, expression of PD-L1 was reduced through inhibition of STAT3 expression by a MAPK inhibitor20. PD-L1 expression is promoted by Toll-like receptor 4- (TLR4) stimulation in human multiple myeloma-derived plasma cells, and this PD-L1 expression is dependent on the mitogen- activated protein kinase (MAPK) signal pathway43. In human dermal fibroblasts, phosphorylation of ERK1/2 and PI3K induces expression of PD-L144. On the other hand, in dendritic cells, Stat4- or Stat6-deficiency does not affect PD-L1 expression45. Although accumulating evidence has revealed the induction mechanisms of PD-L1 in various cell types, that of PD-L1 in CD4+ T cells has yet to be determined. In the present study, we revealed the unique signaling pathway that cooperates to induce PD-L1 expression in CD4+ T cells. Although our in silico analysis suggests the involvement of Stat1 and Stat3, deficiencies of various STATs, including Stat1 and Stat3, did not affect Klf1-induced PD-L1 expression levels. Thus, it appears that Stat1 and Stat3 are not necessary for the Klf1-mediated PD-L1 induction in CD4+ T cells.
Previous studies revealed that the PI3K-Akt-mTOR pathway plays a critical role in PD-L1 expression. In murine squamous cell carcinoma cells, breast cancer, prostate cancer46 and human glioma cells, inactivation of phosphatase and tensin homolog deleted from chromosome 10 (PTEN) promotes PD-L1 expression through the PI3K-Akt pathway and mTOR activation47,48. In human non-small cell lung cancer, PD-L1 is also induced by mTOR activation49. In T cells, Akt-mTOR signaling is a key driver of activation, differentiation and function. T cells possess a unique metabolic profile and a corresponding set of mTOR signal requirements. In the present study, we revealed that Klf1-mediated PD-L1 induction in CD4+ T cells is dependent on the PI3K-Akt-mTOR signaling pathway.
In recent years, treatments focused on PD-1/PD-L1 signaling have been clinically applied to the field of tumor immunity, but this approach has not been expanded to autoimmune diseases. Targeting PD-L1 and PD-1 is a potentially novel treatment strategy, and it has attracted attention. In the field of tumor immunity, several monoclonal antibodies against PD-1 or PD-L1 have been developed and clinical studies initiated. These include treatments with the anti-PD-1 antibody Nivolumab and the anti-PD-L1 antibody Avelumab for treatment of skin cancer and non-small cell lung cancer50,51,52. Tumor immunity and autoimmunity are inseparable, and autoimmune side effects are observed in about 5% of subjects treated with anti-PD-1 antibody and anti-PD-L1 antibody53,54. Pancreatic islet cells highly express PD-L1 in nonobese diabetic (NOD) mice, and exacerbation of the disease is observed by administration of anti-PD-1 antibody or anti-PD-L1 antibody, whereas no change was observed with anti-PD-L2 antibody55. Similar trends are also seen in a contact hypersensitivity model56 and a coronary artery lesion model following heart transplantation57. These reports indicate that PD-1/PD-L1 interaction is a novel therapeutic target for autoimmune diseases.
Here, it was shown that Klf1 and Egr2, transcription factors specifically expressed in LAG3+ Tregs, induce PD-L1 expression in CD4+ T cells independent of each other. These novel findings shed light on the understanding of PD-L1-mediated immune tolerance and suggest the potential therapeutic targeting of Klf1 and Egr2 in the treatment of autoimmune diseases and malignancies.
Materials and Methods
Mice
C57BL/6J (B6) mice were purchased from Japan SLC. 129S6/SvEv-Stat1tm1Rds (Stat1 KO) mice and CD4Cre+ mice were purchased from Taconic Biosciences (Germantown, NY, USA). Stat3fl/flCD4Cre+ (Stat3 CKO) mice, C129S2-Stat4tm1Gru/J (Stat4 KO) mice, C.129 S (B6) -Stat5aatm1Mam/J (Stat5a KO) mice and C.129S2-Stat6tm1Gru/J (Stat6 KO) mice were purchased from Oriental Yeast Co. (Tokyo, Japan). T cell-specific Egr2 conditional knockout (Egr2fl/flCD4Cre+: Egr2 CKO) mice were produced by crossing Egr2fl/fl mice (provided from Patrick Charnay (INSERM, France)) and CD4Cre+ mice. Egr2/3 DKO mice (Egr2fl/flEgr3fl/flCD4Cre+) were generated by crossing Egr2 CKO mice with Egr3fl/fl mice17. Control mice were littermates of Egr2fl/fl, CD4Cre+ or wild-type (wild type: WT) mice. All mice were housed in a specific pathogen-free environment. For the experiments, mice were at least 6–8 weeks of age. Mice of 24–28 weeks of age were used for analysis of LAG3+ Tregs. All animal experiments were approved by the ethics committee of the University of Tokyo Institutional Animal Care and Use Committee, and all experiments were conducted based on the approved experimental plan according to the guidelines of the University of Tokyo.
Reagents
Anti-CD3ε antibody (145-2C11), anti-CD28 antibody (37.51), Fc blocking antibody (anti-CD16/CD32 antibody), FITC- anti-CD45RB antibody (16A) and PE- anti-LAG3 antibody (C9B7W) were obtained from BD Biosciences (San Jose, CA, USA). Biotinylated anti-CD8a antibody (53-6.7), biotinylated anti-CD11c antibody (HL3), PE-anti-CD25 antibody (PC61), APC- anti-CD25 antibody (3C7), biotinylated anti-PD-L1 antibody (10 F.9G2), avidin (SA)-APC antibody, APC/Cy7-anti-CD4 antibody (RM4-5) and 7-AAD were purchased from BioLegend (San Diego, CA, USA). Biotinylated anti-CD19 antibody (1D3) was purchased from eBioscience (Frankfurt am Main, Germany). Streptavidin microbeads were purchased from Miltenyi Biotec (Bergisch Gladbach, Germany and recombinant murine IL-2 was purchased from R&D Systems (Minneapolis, MN, USA). For cultivation, T cells were incubated in RPMI 1640 supplemented with 10% fetal bovine serum (FBS), 100 μg/mL L-glutamine, 100 U/mL penicillin, 100 μg/mL streptomycin and 50 μM 2-mercaptoethanol (all purchased from Sigma). Packaging Cells Plat-E were prepared by culturing in Dulbecco’s modified Eagle’s medium (DMEM) containing 10% FBS, 100 μg/mL L-glutamine, 100 U/mL penicillin, 100 μg/mL streptomycin, 1 μg/mL puromycin and 10 μg/mL blasticidin.
Flowcytometry
Mouse spleens were treated with type IV collagenase (Sigma, St. Louis, USA), and CD4+ T cells were collected using a magnet-assisted cell sorting (MACS) system (Miltenyi). Cells were magnetically labeled using biotinylated anti-CD8a, CD11c, and CD19 antibodies and streptavidin microbeads, and the negative fraction that passed through the MACS LS column (Miltenyi) was collected as CD4+ T cells. LAG3+ Tregs were collected by negative selection using biotinylated anti-CD45 RB antibody. Collected cells were labeled using FITC-anti-CD45RB antibody, PE-anti-LAG3 antibody, APC-anti-CD25 antibody, APC/Cy7-anti-CD4 antibody after blocking Fc receptors. Cell collection was performed with CellQuest (BD Biosciences) using a FACS Vantage flow cytometer (BD Biosciences), and analysis was performed by FlowJo (Tree Star, USA). The purity of cells concentrated by MACS was 90% or more, and the purity of cells collected by flow cytometry (FCM) was 99%.
Quantitative real-time PCR
RNA was extracted using an RNeasy Micro kit (QIAGEN, Germantown, USA) and reverse transcribed into cDNA using SuperScript III and Random Primers (both Invitrogen). Quantitative RT-PCR (qRT-PCR) was performed with CFX Connect (Bio-Rad) using QuantiTect SYBR Green PCR Kit (QIAGEN). The RNA expression level of the target gene was evaluated relative to β-actin (Actb). Sequences of primers are shown in Supplementary Table S1.
Microarray analysis
Microarray analysis was performed as described elsewhere14. Briefly, the gene expression profile of murine CD4+ T cell subsets was obtained from an ArrayExpress database, access number E-MEXP-1343 data. Background adjustment and standardization were performed using Bioconductor’s Mas5 package in statistical software R. Hierarchical clustering was performed in GeneSpring GX (Agilent Technologies, Santa Clara, CA, USA) for genes whose expression was enhanced 1.5-fold compared to CD4+CD25−CD45RBhigh T cells (naïve CD4+ T cells), or expression decreased at least 0.6-fold. Gene expression level was evaluated as the expression level relative to naïve CD4+ T cells.
T cell isolation
T cells were isolated from mouse splenocytes and then collected in the MACS system using a naïve CD4+ T cell isolation kit (Miltenyi Biotec) as follows. Mouse splenocytes were harvested, passed through a 70 μm cell strainer (BD Bioscience), and hemolyzed using an ACK lysis buffer. A solution containing antibodies against a biotin-antibody cocktail (anti-CD8a, anti-CD11b, anti-CD11c, anti-CD19, anti-CD25, anti-CD45R (B220), anti-CD49b (DX5), antiCD105, anti-MHC Class II, anti-Ter119 and anti-CD16/CD32 (Fcg III/II Receptor) was added and reacted at 4 °C for 10 min. Then, 200 μL of anti-biotin microbeads were added and reacted at 4 °C for 15 min, and T cells were separated by negative selection using an LS column. After separation, staining was carried out using anti-CD44 and anti-CD62L antibodies and analyzed by FCM; the purity was >90%.
Gene transduction using retroviral vectors
DNA encoding Egr2 protein (NM-010118) was isolated from mouse T cell cDNA and subcloned into the retroviral vector pMIG. The full-length mRNA of mouse Klf1 (NM-010635) was synthesized by Takara Bio and subcloned into the retroviral vector pMIG. PMIG-Mock, pMIG-Egr2 and pMIG-Klf1 were transfected into packaging cells (Plat-E) using the Extreme Gene 9 transfection reagent (Roche) and the virus-containing culture supernatant was collected. Mouse splenocytes were cultured for 48 h in the presence of Concanavalin A (ConA) (10 μg/mL) (Sigma) and IL-2 (50 μg/mL) to prepare ConA blasts, and gene transduction was conducted using a retroviral supernatant. Specifically, a viral supernatant was added to a Falcon 24-well plate coated with a recombinant human fibronectin fragment CH296 (RetroNectin, Takara), centrifuged at 1200 × g at room temperature for 3 h, and the same operation was repeated three times. After removal of the viral supernatant, ConA blasts were transferred to a plate and cultured in an incubator for 48 h for viral infection. Cultured cells were collected and used for subsequent experiments. In intracellular signal suppression experiments, JNK1/2/3 inhibitor SP600125 (5 μM, Wako Pure Chemical Industries, Ltd., Osaka, Japan), PI3K inhibitor LY294002 (20 μM, Wako), MAPK (p38) inhibitor SB203580 (20μM, Cayman), the MAPK (MEK) inhibitor PD98059 (40 μM, Cayman) or mTOR inhibitor Torin1 (200nM, Cayman) were added 24 h after initiation of infection, and cultured for further 48 h before FCM analysis.
Cell culture
Naïve CD4+ T cells were isolated from splenocytes from Egr2 CKO mice and Egr2fl/fl, Egr2fl/+, CD4Cre+ (control) mice using the mouse naïve CD4+ T cell isolation kit (Miltenyi Biotech) with a MACS system. The cells were cultured for 72 h on anti-CD3ε antibody (2–10 μg/mL) and anti-CD28 (5 μg/mL) antibody-coated 96-well flat bottom plates, followed by FCM analysis.
RNA-sequencing
CD4+ T cells that were transfected with retroviral vectors were divided into 3 fractions according to GFP fluorescence (strongly positive, weakly positive and negative) using a FACSVantage flow cytometer. Total RNA was then extracted. Sequence libraries were prepared according to standard methods using Smart-seq2 (Illumina), and they were analyzed with a next generation sequencer, a MiSeq system (Illumina). Resultant data were mapped to UCSC mm10 using the STAR program, and the read count was calculated using HTSeq. Principal component analysis (PCA) was performed using Prcomp (R package), and visualization in three dimensions was performed by Plot 3D (R package). Differentially expressed genes were analyzed using Ingenuity Pathway Analysis (QIAGEN).
Western blotting
CD4+ T cells transfected with retroviral vectors were cultured in lysis buffer (50 mM Tris HCL, 0.15 M NaCl, 1% Triton-X, 1 mM EDTA) supplemented with Halt protease inhibitor cocktail kit (1:100; Thermo Fisher), an equal amount of 2 × Laemmli sample buffer (Sigma) was added, and the mixture was reacted at 95 °C for 5 min. Total protein was measured using a BCA Protein Assay kit (Pierce). Each sample was electrophoresed on a 7.5% Mini-PROTEAN TGX precast gel (Bio-Rad) and transferred to an Immobilon-PVDF membrane (Millipore). After blocking with 5% bovine serum albumin, the membrane was reacted with anti-phospho-Akt (Thr308) antibody (1:1000), anti-Akt (Pan) antibody (1:1000), anti phospho-S6K1 (Thr389) antibody (1:1000), anti-S6K1 antibody (1:1000) or anti-GAPDH antibody (1:1000) at 4 °C overnight, and then goat anti rabbit IgG-horseradish peroxidase (HRP) (Invitrogen 1:3000) was added at room temperature for 1 h. Signals were detected using ECL Prime or ECL Select (GE Healthcare).
Statistical analysis
Statistically significant differences were analyzed using GraphPad Prism 7 (GraphPad Software, Inc.). Comparative study of qRT-PCR and mean fluorescence intensity was conducted by Student’s t-test in the case of two groups. Comparative studies between multiple groups were conducted by one-way ANOVA and then a multiple comparison test was performed using the Bonferroni method.
Data availability
Please contact the corresponding author for data requests.
Accession number
The microarray and RNA-sequencing data are available online at in the ArrayExpress database (access number E-MEXP-1343, E-MTAB-6778).
References
Benson, R. A., McInnes, I. B., Brewer, J. M. & Garside, P. Cellular imaging in rheumatic diseases. Nat Rev Rheumatol 11, 357–367, https://doi.org/10.1038/nrrheum.2015.34 (2015).
Tsokos, G. C. Systemic lupus erythematosus. N Engl J Med 365, 2110–2121, https://doi.org/10.1056/NEJMra1100359 (2011).
van Vollenhoven, R. F. Treatment of rheumatoid arthritis: state of the art 2009. Nat Rev Rheumatol 5, 531–541, https://doi.org/10.1038/nrrheum.2009.182 (2009).
Nguyen, L. T. & Ohashi, P. S. Clinical blockade of PD1 and LAG3–potential mechanisms of action. Nat Rev Immunol 15, 45–56, https://doi.org/10.1038/nri3790 (2015).
Sharpe, A. H. & Freeman, G. J. The B7-CD28 superfamily. Nat Rev Immunol 2, 116–126, https://doi.org/10.1038/nri727 (2002).
Yamazaki, T. et al. Expression of Programmed Death 1 Ligands by Murine T Cells and APC. The Journal of Immunology 169, 5538–5545, https://doi.org/10.4049/jimmunol.169.10.5538 (2002).
Flies, D. B. & Chen, L. The new B7s: playing a pivotal role in tumor immunity. J Immunother 30, 251–260, https://doi.org/10.1097/CJI.0b013e31802e085a (2007).
Francisco, L. M., Sage, P. T. & Sharpe, A. H. The PD-1 pathway in tolerance and autoimmunity. Immunol Rev 236, 219–242, https://doi.org/10.1111/j.1600-065X.2010.00923.x (2010).
Baecher-Allan, C., Brown, J. A., Freeman, G. J. & Hafler, D. A. CD4+ CD25+ regulatory cells from human peripheral blood express very high levels of CD25 ex vivo. Novartis Found Symp 252, 67–88, discussion 88–91, 106–114 (2003).
Okamura, T. et al. TGF-beta3-expressing CD4+CD25(−)LAG3+ regulatory T cells control humoral immune responses. Nat Commun 6, 6329, https://doi.org/10.1038/ncomms7329 (2015).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061, https://doi.org/10.1126/science.1079490 (2003).
Sandner, S. E. et al. Role of the programmed death-1 pathway in regulation of alloimmune responses in vivo. J Immunol 174, 3408–3415 (2005).
Gotot, J. et al. Regulatory T cells use programmed death 1 ligands to directly suppress autoreactive B cells in vivo. Proc Natl Acad Sci USA 109, 10468–10473, https://doi.org/10.1073/pnas.1201131109 (2012).
Okamura, T. et al. CD4+ CD25−LAG3+ regulatory T cells controlled by the transcription factor Egr-2. Proc Natl Acad Sci USA 106, 13974–13979, https://doi.org/10.1073/pnas.0906872106 (2009).
Huard, B., Prigent, P., Pagès, F., Bruniquel, D. & Triebel, F. T cell major histocompatibility complex class II molecules down-regulate CD4+ T cell clone responses following LAG-3 binding. Eur J Immunol 26, 1180–1186, https://doi.org/10.1002/eji.1830260533 (1996).
Safford, M. et al. Egr-2 and Egr-3 are negative regulators of T cell activation. Nat Immunol 6, 472–480, https://doi.org/10.1038/ni1193 (2005).
Morita, K. et al. Egr2 and Egr3 in regulatory T cells cooperatively control systemic autoimmunity through Ltbp3-mediated TGF-β3 production. Proc Natl Acad Sci USA 113, E8131–E8140, https://doi.org/10.1073/pnas.1611286114 (2016).
Sandelin, A., Alkema, W., Engström, P., Wasserman, W. W. & Lenhard, B. JASPAR: an open-access database for eukaryotic transcription factor binding profiles. Nucleic Acids Res 32, D91–94, https://doi.org/10.1093/nar/gkh012 (2004).
Marzec, M. et al. Oncogenic kinase NPM/ALK induces through STAT3 expression of immunosuppressive protein CD274 (PD-L1, B7-H1). Proc Natl Acad Sci USA 105, 20852–20857, https://doi.org/10.1073/pnas.0810958105 (2008).
Yamamoto, R. et al. B7-H1 expression is regulated by MEK/ERK signaling pathway in anaplastic large cell lymphoma and Hodgkin lymphoma. Cancer Sci 100, 2093–2100, https://doi.org/10.1111/j.1349-7006.2009.01302.x (2009).
Zhu, B. et al. Early growth response gene 2 (Egr-2) controls the self-tolerance of T cells and prevents the development of lupuslike autoimmune disease. J Exp Med 205, 2295–2307, https://doi.org/10.1084/jem.20080187 (2008).
Wölfle, S. J. et al. PD-L1 expression on tolerogenic APCs is controlled by STAT-3. Eur J Immunol 41, 413–424, https://doi.org/10.1002/eji.201040979 (2011).
Matta, B. M., Raimondi, G., Rosborough, B. R., Sumpter, T. L. & Thomson, A. W. IL-27 production and STAT3-dependent upregulation of B7-H1 mediate immune regulatory functions of liver plasmacytoid dendritic cells. J Immunol 188, 5227–5237, https://doi.org/10.4049/jimmunol.1103382 (2012).
Haxhinasto, S., Mathis, D. & Benoist, C. The AKT-mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J Exp Med 205, 565–574, https://doi.org/10.1084/jem.20071477 (2008).
Sauer, S. et al. T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc Natl Acad Sci USA 105, 7797–7802, https://doi.org/10.1073/pnas.0800928105 (2008).
Turner, M. S., Kane, L. P. & Morel, P. A. Dominant role of antigen dose in CD4+Foxp3+ regulatory T cell induction and expansion. J Immunol 183, 4895–4903, https://doi.org/10.4049/jimmunol.0901459 (2009).
Hawse, W. F., Boggess, W. C. & Morel, P. A. TCR Signal Strength Regulates Akt Substrate Specificity To Induce Alternate Murine Th and T Regulatory Cell Differentiation Programs. J Immunol 199, 589–597, https://doi.org/10.4049/jimmunol.1700369 (2017).
Okazaki, T., Maeda, A., Nishimura, H., Kurosaki, T. & Honjo, T. PD-1 immunoreceptor inhibits B cell receptor-mediated signaling by recruiting src homology 2-domain-containing tyrosine phosphatase 2 to phosphotyrosine. Proc Natl Acad Sci USA 98, 13866–13871, https://doi.org/10.1073/pnas.231486598 (2001).
Latchman, Y. E. et al. PD-L1-deficient mice show that PD-L1 on T cells, antigen-presenting cells, and host tissues negatively regulates T cells. Proc Natl Acad Sci USA 101, 10691–10696, https://doi.org/10.1073/pnas.0307252101 (2004).
Dong, H. et al. B7-H1 determines accumulation and deletion of intrahepatic CD8(+) T lymphocytes. Immunity 20, 327–336 (2004).
Mitchell, A. L. et al. Programmed death ligand 1 (PD-L1) gene variants contribute to autoimmune Addison’s disease and Graves’ disease susceptibility. J Clin Endocrinol Metab 94, 5139–5145, https://doi.org/10.1210/jc.2009-1404 (2009).
Hayashi, M., Kouki, T., Takasu, N., Sunagawa, S. & Komiya, I. Association of an A/C single nucleotide polymorphism in programmed cell death-ligand 1 gene with Graves’ disease in Japanese patients. Eur J Endocrinol 158, 817–822, https://doi.org/10.1530/EJE-07-0649 (2008).
Zhang, Y. et al. Regulation of T cell activation and tolerance by PDL2. Proc Natl Acad Sci USA 103, 11695–11700, https://doi.org/10.1073/pnas.0601347103 (2006).
Yien, Y. Y. & Bieker, J. J. EKLF/KLF1, a tissue-restricted integrator of transcriptional control, chromatin remodeling, and lineage determination. Mol Cell Biol 33, 4–13, https://doi.org/10.1128/MCB.01058-12 (2013).
Miller, I. J. & Bieker, J. J. A novel, erythroid cell-specific murine transcription factor that binds to the CACCC element and is related to the Kruppel family of nuclear proteins. Mol Cell Biol 13, 2776–2786 (1993).
Kulozik, A. E., Bellan-Koch, A., Bail, S., Kohne, E. & Kleihauer, E. Thalassemia intermedia: moderate reduction of beta globin gene transcriptional activity by a novel mutation of the proximal CACCC promoter element. Blood 77, 2054–2058 (1991).
Orkin, S. H., Antonarakis, S. E. & Kazazian, H. H. Jr. Base substitution at position −88 in a beta-thalassemic globin gene. Further evidence for the role of distal promoter element ACACCC. J Biol Chem 259, 8679–8681 (1984).
Orkin, S. H. et al. Linkage of beta-thalassaemia mutations and beta-globin gene polymorphisms with DNA polymorphisms in human beta-globin gene cluster. Nature 296, 627–631 (1982).
Chen, X. & Bieker, J. J. Stage-specific repression by the EKLF transcriptional activator. Mol Cell Biol 24, 10416–10424, https://doi.org/10.1128/MCB.24.23.10416-10424.2004 (2004).
Siatecka, M., Xue, L. & Bieker, J. J. Sumoylation of EKLF promotes transcriptional repression and is involved in inhibition of megakaryopoiesis. Mol Cell Biol 27, 8547–8560, https://doi.org/10.1128/MCB.00589-07 (2007).
Perkins, A. C., Sharpe, A. H. & Orkin, S. H. Lethal beta-thalassaemia in mice lacking the erythroid CACCC-transcription factor EKLF. Nature 375, 318–322, https://doi.org/10.1038/375318a0 (1995).
Lee, S. J. et al. Interferon regulatory factor-1 is prerequisite to the constitutive expression and IFN-gamma-induced upregulation of B7-H1 (CD274). FEBS Lett 580, 755–762, https://doi.org/10.1016/j.febslet.2005.12.093 (2006).
Liu, J. et al. Plasma cells from multiple myeloma patients express B7-H1 (PD-L1) and increase expression after stimulation with IFN-{gamma} and TLR ligands via a MyD88-, TRAF6-, and MEK-dependent pathway. Blood 110, 296–304, https://doi.org/10.1182/blood-2006-10-051482 (2007).
Lee, S. K. et al. IFN-gamma regulates the expression of B7-H1 in dermal fibroblast cells. J Dermatol Sci 40, 95–103, https://doi.org/10.1016/j.jdermsci.2005.06.008 (2005).
Liang, S. C. et al. Regulation of PD-1, PD-L1, and PD-L2 expression during normal and autoimmune responses. Eur J Immunol 33, 2706–2716, https://doi.org/10.1002/eji.200324228 (2003).
Crane, C. A. et al. PI(3) kinase is associated with a mechanism of immunoresistance in breast and prostate cancer. Oncogene 28, 306–312, https://doi.org/10.1038/onc.2008.384 (2009).
Squarize, C. H. et al. PTEN Deficiency Contributes to the Development and Progression of Head and Neck Cancer. Neoplasia 15, 461–471, https://doi.org/10.1593/neo.121024 (2013).
Parsa, A. T. et al. Loss of tumor suppressor PTEN function increases B7-H1 expression and immunoresistance in glioma. Nat Med 13, 84–88, https://doi.org/10.1038/nm1517 (2007).
Lastwika, K. J. et al. Control of PD-L1 Expression by Oncogenic Activation of the AKT-mTOR Pathway in Non-Small Cell Lung Cancer. Cancer Res 76, 227–238, https://doi.org/10.1158/0008-5472.CAN-14-3362 (2016).
Brahmer, J. et al. Nivolumab versus Docetaxel in Advanced Squamous-Cell Non-Small-Cell Lung Cancer. N Engl J Med 373, 123–135, https://doi.org/10.1056/NEJMoa1504627 (2015).
Robert, C. et al. Nivolumab in previously untreated melanoma without BRAF mutation. N Engl J Med 372, 320–330, https://doi.org/10.1056/NEJMoa1412082 (2015).
Kaufman, H. L. et al. Avelumab in patients with chemotherapy-refractory metastatic Merkel cell carcinoma: a multicentre, single-group, open-label, phase 2 trial. The Lancet Oncology 17, 1374–1385, https://doi.org/10.1016/s1470-2045(16)30364-3 (2016).
Topalian, S. L. et al. Survival, durable tumor remission, and long-term safety in patients with advanced melanoma receiving nivolumab. J Clin Oncol 32, 1020–1030, https://doi.org/10.1200/JCO.2013.53.0105 (2014).
Weber, J. S., Yang, J. C., Atkins, M. B. & Disis, M. L. Toxicities of Immunotherapy for the Practitioner. J Clin Oncol 33, 2092–2099, https://doi.org/10.1200/JCO.2014.60.0379 (2015).
Ansari, M. J. et al. The programmed death-1 (PD-1) pathway regulates autoimmune diabetes in nonobese diabetic (NOD) mice. J Exp Med 198, 63–69, https://doi.org/10.1084/jem.20022125 (2003).
Tsushima, F. et al. Preferential contribution of B7-H1 to programmed death-1-mediated regulation of hapten-specific allergic inflammatory responses. Eur J Immunol 33, 2773–2782, https://doi.org/10.1002/eji.200324084 (2003).
Yang, J. et al. Critical role of donor tissue expression of programmed death ligand-1 in regulating cardiac allograft rejection and vasculopathy. Circulation 117, 660–669, https://doi.org/10.1161/CIRCULATIONAHA.107.741025 (2008).
Acknowledgements
We are grateful to K. Watada, K. Sakashita, J. Takezawa, R. Yamagata, and N. Nemoto for their excellent technical assistance. We thank Dr. Patrick Charnay (INSERM, France) for providing Egr2-floxed mice and Dr. Toshio Kitamura (The Institute of Medical Science, The University of Tokyo, Japan) for providing the packaging cells, Plat-E. This work was supported by grants from the Japan Society for the Promotion of Science, and the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Contributions
K.Y. supervised the project. S.T., T.O. and K.F. designed and planned the study. S.T. T.K. and M.I. performed experiments. S.T. and T.O. wrote the manuscript with contributions from Y.I., S.S. and H.S. All authors discussed the results and commented on the article.
Corresponding authors
Ethics declarations
Competing Interests
T.O. received financial support or fees from Chugai and Bristol-Myers Squibb (BMS). K.Y. received financial support or fees from AbbVie, Astellas, BMS, Daiichi-Sankyo, Mitsubishi Tanabe, Pfizer, Sanofi, Santen, Takeda, Teijin, Boehringer Ingelheim, Chugai, Eisai, Ono, Taisho Toyama, UCB, ImmunoFuture, Asahi Kasei, Janssen, and NIPPON KAYAKU. K.F. received financial support or fees from Astellas, BMS, Daiichi-Sankyo, Mitsubishi Tanabe, Pfizer, Santen, Takeda, Chugai, Eisai, Taisho Toyama, UCB, Janssen, and NIPPON KAYAKU. The remaining authors declare no competing financial interests. T.K., T.O., K.Y., and K.F. received patent-licensing arrangements with Chugai.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Teruya, S., Okamura, T., Komai, T. et al. Egr2-independent, Klf1-mediated induction of PD-L1 in CD4+ T cells. Sci Rep 8, 7021 (2018). https://doi.org/10.1038/s41598-018-25302-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-25302-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.