Abstract
The ryanodine receptor 1 is a large calcium ion channel found in mammalian skeletal muscle. The ion channel gained a lot of attention recently, after multiple independent authors published near-atomic cryo electron microscopy data. Taking advantage of the unprecedented quality of structural data, we performed molecular dynamics simulations on the entire ion channel as well as on a reduced model. We calculated potentials of mean force for Ba2+, Ca2+, Mg2+, K+, Na+ and Cl− ions using umbrella sampling to identify the key residues involved in ion permeation. We found two main binding sites for the cations, whereas the channel is strongly repulsive for chloride ions. Furthermore, the data is consistent with the model that the receptor achieves its ion selectivity by over-affinity for divalent cations in a calcium-block-like fashion. We reproduced the experimental conductance for potassium ions in permeation simulations with applied voltage. The analysis of the permeation paths shows that ions exit the pore via multiple pathways, which we suggest to be related to the experimental observation of different subconducting states.
Similar content being viewed by others
Introduction
Ryanodine receptors (RyRs) are large ion channels that play a pivotal role in the regulation of the free calcium concentration in living cells1. Their main purpose is gating calcium releases from internal storages, such as the sarcoplasmic and endoplasmic reticula2,3,4,5. RyRs are tetramers consisting of four identical protomers. The full receptor has a molecular mass of 2.2 megadaltons and its just over 5000 residues per protomer make it the largest known ion channel6. Three mammalian isoforms of RyR are known (RyR1 to RyR3): RyR1 is abundant in skeletal muscle, where it is responsible for the rapid release of calcium ions from the sarcoplasmic reticulum (SR)6,7,8. Subsequent to their release into the cytoplasm, the calcium ions bind to troponin-C and trigger muscular contraction9,10. Hence, RyR1 plays a crucial role in the excitation-contraction coupling of skeletal muscle (e.g.11,12,13). Mutations in the RyR1 isoform have been linked to malignant hyperthermia and central core disease2,3. RyR2 is responsible for calcium induced calcium release in cardiac muscle14,15, whereas RyR3 is found in smooth muscle and brain tissue16.
Ryanodine receptors gained a lot of attention recently17,18, after multiple authors published cryo electron microscopy (cryo-EM) data of the receptor in unprecedented resolution, reaching down to 3.8 Å in the closed19,20,21,22,23 and 4.1 Å in the open conformation21,22,23,24. Although Molecular Dynamics (MD) simulations on homology models of RyR1 have been carried out before25,26,27,28, the availability of these high resolution structural data sets allows for direct simulation of RyR1.
In this manuscript, we assessed the conduction and selectivity mechanisms of RyR1 by using MD simulations. Therefore, we first constructed a full model of RyR1, based on the most recent cryo-EM data. To address the conduction mechanism of RyR1 and its energetics, we calculated potentials of mean force (PMFs) along the central pore for different ion types and identified key residues, directly responsible for the conduction of ions in RyR1. Furthermore, we investigated the conduction pathways by performing ion permeation simulations, in which ions are driven across the channel by an externally applied electric field, closely mimicking electrophysiological measurements, and found new ion exit pathways in the channel.
Methods
Molecular dynamics simulations
All MD simulations were carried out using the software package Gromacs 5.129,30,31,32,33 and the CHARMM36 force field34,35,36,37. The system was propagated in time using the leap frog integrator with a time step of 2 fs, with the only exception being the first equilibration run, in which a reduced time step of 1 fs was used for stability reasons. During the simulations, H-bonds were restrained using the LINCS algorithm38, long range electrostatic interactions were treated using the Particle-Mesh Ewald (PME) method39 and van-der-Waals and Coulomb cutoffs were set to 1.2 nm. The TIP3P water model40 was used and all molecular visualizations were done with the VMD software41.
Receptor modelling
Although cryo-EM datasets of the open-state receptor are available, the quality of the closed state data is significantly better compared to the open state. Therefore, we first constructed an atomistic model of the closed receptor, which was equilibrated and later transitioned into the conductive state by adding biasing forces to the simulation in a steered MD-like fashion.
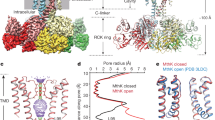
A full structural model of the closed RyR1 was constructed utilizing data obtained by Yan et al.19, Rouslan et al.21 and Zalk et al.20 (see Fig. 1). Loops, unresolved in all of the referenced data sets, were modeled using the Modeller software with the DOPE scorer42,43,44,45. In addition to loops, the structures contain five large unresolved segments, which are partially disordered19,20. For two of those, located in the SPRY3 and between the handle and central domains, Modeller was used to construct homology models separately, which were later added to the model. The further unresolved parts are contained in the handle domain, between the central and channel domains as well as in the channel domain and are likely disordered19,20. In the first and last case, predicted models were used for the simulations46. The second disordered part, almost 300 residues in size, needed to be excluded from the model. Therefore, the residues A4269 and L4544 were directly connected.
Side view of the RyR1-model. The left side is color-coded according to the domains of the receptor. Major domains are the N-terminal domain ( ), the handle domains (
), the handle domains ( ), the helical domains (
), the helical domains ( ), the central domain (
), the central domain ( ) and the channel domain (
) and the channel domain ( ). The right side highlights the reduced model, consisting of the central and channel domains, in red. The orange arrow indicates the direction of the calcium current from the SR into the cytoplasm. A top-view is shown in the inset.
). The right side highlights the reduced model, consisting of the central and channel domains, in red. The orange arrow indicates the direction of the calcium current from the SR into the cytoplasm. A top-view is shown in the inset.
The obtained model of RyR1 was relaxed using Gromacs in a multistep fashion. First, the system was subject to 20000 steps of in vacuo energy minimization. After that, the model was solvated using approximately a million water molecules and charge neutrality was achieved by adding 360 Ca2+ ions, to create a realistic environment for the luminal pore. Further energy minimization was conducted on the full system, containing the protein, solvent, and ions, until the maximum force (on any atom) consistently reached values below 1000 kJ/mol/nm. For additional relaxation, the solvated receptor was simulated for 100 ps, while the temperature was kept constant at 300 K, using a velocity rescaling thermostat47. During this simulation, harmonic restraints were placed on all protein atoms. To allow relaxation of the side chains, an additional simulation run, lasting 200 ps, during which only Cα atoms were restrained, was conducted. The remaining restraints were reduced in a stepwise fashion to every 4th Cα atom and later to every 16th Cα atom in simulation runs of 100 ps length. To stabilize the well resolved pore (consisting of the central and channel domains), harmonic restraints were imposed on every 16th Cα atom of the pore during an additional relaxation run, lasting 2.5 ns. In total, the system could relax for 3 ns as a NVT ensemble. To obtain a relaxed system at atmospheric pressure, further 7 ns were simulated using a Berendsen barostat48 (approximate NPT ensemble).
Membrane embedding
A small POPC patch, which had been equilibrated for 35 ns using the CHARMM36 force field49, was used to form a suitable membrane. It was enlarged to match the size of the receptor by concatenating multiple copies of it and the enlarged patch was subject to further 2 ns of NPT equilibration. The protein was embedded into the membrane, using the program g_membed50,51. With this algorithm, the receptor was scaled to 0.1 of its size along the dimensions parallel to the lipid bilayer. Lipids, intersecting with the shrunken protein, were removed and the receptor was gradually scaled back to its original size within 2000 molecular dynamics steps. After embedding, the complete system was subject to additional relaxation, lasting 1 ns (NVT) and 1.5 ns (NPT).
Adding ions
The calcium ions, used to neutralize the system during equilibration, were removed. New ions were added to the compartments of either side of the membrane. In particular, 3.2 mmol/L of Mg2+ and 10 mmol/L of Ca2+ ions were added to the luminal side of the system. On the cytosolic side, 140 mmol/L of K+ were added and the entire system was neutralized using Cl− ions. Whereas a physiological amount of potassium ions was used, the luminal calcium concentration is typically reported to be 1 mmol/L2. Due to the small system sizes accessible by MD simulation, divalent ions were added at the very high end of experimentally observed concentrations. In total, 37 Ca2+, 12 Mg2+, 1274 K+ and 652 Cl− ions were included. To maintain the asymmetric ion concentrations in the single-membrane system during simulation runs under periodic boundary conditions, an energy step method similar to the one developed by Khalili-Araghi et al.52 was implemented. Here, spatial restraints were imposed upon all cations using a step-shaped potential. They were kept in their respective compartment by a repulsive force, that acted on the ions within a narrow transition region around the periodic boundary. A force of 10000 kJ/mol/nm turned out to allow for stable simulation runs, while simultaneously being strong enough to reliably restrain all ions in their compartments. Further details on the usage and implementation of the energy step method are available in the supplementary material.
Equilibrating and obtaining a reduced system
The entire simulated system was contained in a box of 36.6 × 36.6 × 21.5 nm3 in size and consisted of three million particles. The full system was equilibrated for 100 ns, using a Parrinello-Rahman barostat53,54,55 for semiisotropic pressure coupling. Although fully equilibrating the entire receptor in 100 ns was impossible, the simulation run was sufficient to converge the root-mean-square deviation (RMSD) of the relevant subsystem, which would later make up the reduced model (see Supplementary Material Fig. 2).
For all further simulation runs, the equilibrated model was reduced to the central and channel domains, which form the pore of the receptor. Due to the large size of RyR1, a reduced model is more suitable to study potentials of mean force along the central axis and the ion conduction mechanisms. Therefore, all residues up to A3680 were deleted. Because of the mushroom-like shape of the receptor (see Fig. 1), this procedure allowed for a dramatic reduction of the membrane patch size and as a consequence, a drastically smaller system and thus a reduced computational load.
The reduced system was resolvated and ions were added again, as described above. To avoid conformational changes due to the missing parts of the receptor, harmonic restraints were placed upon the interface of the reduced model to its removed parts. For this purpose, a force constant of 1000 kJ/mol/nm2 was used for all Cα atoms of residues A3680 to L4170. The reduced system consisted of 700000 particles.
Obtaining a model of open state RyR1
The model was steered into a conductive state using the recent structural data by des Georges et al.22 of open-state RyR1 (PDB: 5TAL). All Cα atoms of the pore region (defined by V4820 to T4970), including the pore-forming helices (S1–S6) and the selectivity filter, were dragged into their new positions as dictated by the cryo-EM data within 5 ns. This was done using harmonic potentials with force constants of 500 kJ/mol/nm, applied through the flooding algorithm56,57. To equilibrate the structure of the open-state receptor, constraints were maintained on every forth Cα atom for a simulation run of 30 ns. After restrained equilibration, we observed a converged Cα RMSD of 0.75 nm for the entire reduced system and 0.35 nm for the pore region, compared to the cryo-EM structure. The value decreased to 0.3 nm if the extremely flexible luminal loops were excluded from the calculation.
Further 30 ns were simulated without restraining the structure in its conductive conformation, to see if the channel would spontaneously close again. No significant closure movement was observed (see Supplementary Material Fig. 3). Additionally, longer unrestrained equilibration yielded an increase of RMS distances by only 0.1 nm compared to the restrained run, indicating that the open structure is stable within the accessible timescales (see Supplementary Material Fig. 4). Therefore, no open-state restraints were used during the 0.6 ns-short runs during umbrella sampling. However, to guarantee the channel remains fully open during the longer 100 ns-long ion permeation runs, harmonic open-state restraints were placed on every forth Cα atom of the pore region during the applied voltage runs, as described above.
Calculating potentials of mean force
Potentials of mean force (PMFs) were calculated for barium, calcium, magnesium, sodium, potassium and chloride ions across the central axis of the ryanodine receptor in its open state. Each PMF was computed using umbrella sampling and the weighted histogram analysis method (WHAM)58,59,60, based on 209 umbrella windows. Each window was prepared by exchanging the ion of interest with a water molecule. Although the typical distance between water molecules is larger, a window spacing of 0.7 Å was used. This was done by taking four snapshots from the last 80 ps of the proceeding equilibration run. For each umbrella window, the snapshot that allowed for optimal spacing was used. During the umbrella sampling simulations, the ion was held in place by a harmonic biasing potential with a spring constant of 500 kJ/mol/nm2 in axial direction. Each window was simulated for 0.6 ns without open-state restraints, of which the first 0.1 ns were excluded from analysis. Because the channel is typically occupied by many ions simultaneously, other ions were not restrained from entering the pore during the umbrella sampling. However, no further ions were present in the narrow hydrophobic core, ranging between −2 nm and 1 nm, during umbrella sampling. We calculated cyclic PMFs with g_wham60 and obtained error estimates by using 200 bootstraps.
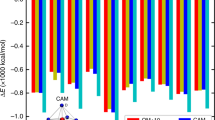
From the PMF we obtained standard free energies of binding (ΔG0) using the method derived by Doudou et al.61 without restraints perpendicular to the reaction coordinate. The calculation was carried out as described by Pongprayoon et al.62 in the supporting information of the cited paper, who used the formalism in a similar context previously. Ions were considered in bound state between −6.0 nm and 7.5 nm along the pore axis and considered unbound (in bulk phase) outside the interval (see Fig. 2a). We estimated the effectively sampled area perpendicular to the reaction coordinate in analogy to equation (8) in61:
where \({\xi }_{1}^{2}\) and \({\xi }_{2}^{2}\) are the eigenvalues of the covariance matrix of the bulk phase windows in the plane perpendicular to the reaction coordinate. The effectively sampled area amounted to 3.09, 3.04, 5.15, 9.09, and 3.30 nm2 for Ba2+, Ca2+, Mg2+, K+, and Na+ respectively.
(a) PMFs across the open pore. Data shown for Ba2+ ( ), Ca2+ (
), Ca2+ ( ), Mg2+ (
), Mg2+ ( ), Na+ (
), Na+ ( ), K+ (
), K+ ( ) and Cl− (
) and Cl− ( ) ions. The main PMF basins are labeled as ①, ② and ③. (b) The pore of RyR1, aligned with the pore axis of the panel above. Key residues are highlighted in color, depending on which PMF feature they contribute to; the orange arrow indicates the direction of ion permeation. The semitransparent bar indicates the position of the membrane. Panel (c) shows the number of water molecules in the second hydration shell of each ion type across the pore. The inset shows the pore radius and its standard error, based on the structures of all umbrella windows, calculated using HOLE101. The panels (d) to (i) show a calcium ion interacting with key residues, as labeled in panel (b). Note that 4 copies of each residue are shown, since RyR1 is a tetramer.
) ions. The main PMF basins are labeled as ①, ② and ③. (b) The pore of RyR1, aligned with the pore axis of the panel above. Key residues are highlighted in color, depending on which PMF feature they contribute to; the orange arrow indicates the direction of ion permeation. The semitransparent bar indicates the position of the membrane. Panel (c) shows the number of water molecules in the second hydration shell of each ion type across the pore. The inset shows the pore radius and its standard error, based on the structures of all umbrella windows, calculated using HOLE101. The panels (d) to (i) show a calcium ion interacting with key residues, as labeled in panel (b). Note that 4 copies of each residue are shown, since RyR1 is a tetramer.
To obtain a better handle on the electrostatic contributions to the PMF, we calculated the electrostatic potential across the channel by solving the Poisson equation. The calculation was done using the APBS program63, using dielectric constants of 2 and 78.5 for protein/membrane and solvent respectively.
Conducting ion-permeation simulations
Our initial attempts to perform ion permeation simulations with an applied voltage using calcium ions were unsuccessful, as the ions tended to get stuck in the pore. Unfortunately, the very high affinity of RyR1 for calcium ions renders direct permeation simulations impossible. This is due to the well known problem of over-affinity for divalent ions in classical, additive force fields, which will be addressed further in the Discussion. Therefore, we decided to use monovalent potassium ions instead of divalent calcium ions for the ion permeation simulations in the conductive state of RyR1. All ions were removed from the system and new potassium ions were added in symmetric concentrations of 140 mmol/L to either side of the membrane. An external electric field was added by specifying it in the Gromacs parameter file. The strength of the electric field E, specified in units V/nm, was chosen such that the product E · L with the box dimension in axial direction (L = 14.626 nm) resulted in voltages of −0.2, 0.0, 0.2, 0.3 and 0.5 V64. Although high in physiological terms, voltages of this strength are commonly used in MD simulations and have proven to yield good results in the past65,66,67. Of each run, lasting 100 ns, the first 10 ns were excluded from analysis. Ionic currents were determined by counting the net number of ions that crossed the periodic boundary parallel to the membrane. Additionally, a further run was conducted using a voltage of 1.0 V, which lasted 30 ns. Again, the first 10 ns were excluded from analysis.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Results
Umbrella sampling
All potentials of mean force of the open-state receptor are plotted in Fig. 2a. Additionally, the maximum energetic wells or barriers as well as the standard free energies of binding for all cations are summarized in Table 1.
Clearly, the ion affinity depends heavily on the ionic charge, which is due to the negative net charge of the receptor pore. Therefore, the receptor shows greater affinity to divalent cations than for monovalent cations and it is repulsive for the chloride anion. The PMFs for all cations show similar features: There are two main basins around −2 nm and +1 nm, respectively, of which the first one is split into two basins by a small peak in the profile for Ba2+, Ca2+ and Mg2+. The two main basins are separated by a free energy barrier at 0 nm in the center of the pore. Furthermore, the divalent ions show affinity for a minor third binding site, located at 6 nm in Fig. 2a.
To explain those features, we examined the umbrella windows at relevant points and highlighted as well as labeled key residues of the pore in Fig. 2b. Additionally, close-up renderings of a calcium ion interacting with some of the residues are shown in Fig. 2d–i.
Starting on the luminal side of the SR membrane, the pore creates an attractive environment for cations through the abundance of negatively charged residues in the flexible, acidic luminal loops. In particular, the presence of residues E4867, D4868, E4869, D4870, D4877, D4878 as well as E4902, D4903, D4907 and E4908 directly in front of the pore contribute to a local increase of cationic concentrations.
At the entrance of the pore around 2.5 nm, its radius decreases quickly towards the selectivity filter (SF), a GGGIGDE motif, as shown in Fig. 2c. Two rings of glutamate and aspartate residues (E4900 and D4899) mark the beginning of the filter and contribute to the first PMF basin ①. As shown in Fig. 2d and e, the ion makes contact with the negatively charged carboxylate groups of either of the four copies of the residues in the tetrameric protein.
Slightly further down in the SF, the pore reaches its constriction site of 3 Å at G4894. Further towards the cytoplasmic side and inside the hydrophobic transmembrane part, two rings, formed by Q4933 and I4937 residues, create a secondary constriction site. Comparison with the equilibrated closed structure shows that these residues act as a gate, as it was also seen by others68. In this case, I4937 rotates into the pore and blocks it completely. In the open state, side chains only slightly extend into the pore, as depicted in Fig. 2f. Therefore, the free energy barrier at around 0 nm is likely caused by a combination of a small steric barrier due to Q4933 and I4937 and the free energy penalty for the hydrated ion in the hydrophobic transmembrane section. This also suggests that the PMF minimum ① is co-formed by the restrictions by G4894 and the hydrophobic core to either side and the consequentially increased entropy of the ion inside the cavity of increased radius between the two constriction points.
The ring of isoleucine residues (I4937) is directly followed by a negatively charged ring of aspartate residues (D4938). Since the channel has a radius of only 3.5 Å at this point, all four acidic residues strongly interact with the permeating cation. As shown in Fig. 2g, the ion makes contact with at least one of the four copies of the residue, while the remaining three residues are still close enough to interact with the ion. However, it seems that D4938 is not single handedly responsible for the second main PMF basin. Instead, further acidic residues, such as D4945, E4948, E4952 and D4699, extend into the channel and contribute to basin ②, as the pore is getting wider. The ion makes contact with each of those rings, as shown in Fig. 2h exemplarily, while the preceding and succeeding rings are still close enough for electrostatic interaction.
PMF basin ③ is caused by E5033 and to a lesser extent by D5026. At this point, the E5033-residues of adjacent protomers are already more than 2 nm apart, so that the PMF is dominated by the protomer that the ion binds to, whereas the remaining three are too far away for strong interactions.
To check if the PMF basin ② is indeed not caused by a single amino acid, but rather by the electrostatics of many residues combined, we quantified the influence of the acidic residues on the electrostatic environment by numerically solving the Poisson equation for the open-state RyR1. Indeed, the results in Fig. 3 show a strong and attractive potential for cations. In particular, basin ② coincides with the minimum of the electrostatic potential across the channel axis.
(a) One-dimensional electrostatic potential along the channel axis (white dotted line in panel (b)). The PMF for calcium is shown in red to aid orientation. (b) The two-dimensional electrostatic potential on a slice across RyR1. Dashed lines mark the outline of the channel to aid orientation. The locations of the two main PMF basis are marked by white crosses. The orange arrow indicates the direction of the calcium current. (c) Two red spheres were placed in the main PMF basins.
Concerning the different ion types, Ba2+, Ca2+ and Mg2+ show the highest affinities. Barium and calcium show similar profiles: both with an approximate PMF depth of 30 kJ/mol. The PMF for magnesium is only 21.5 kJ/mol in depth. All three divalent ions bind similarly strongly to D4899 and E4900 in basin ①. Barium has a more pronounced basin ① compared to the other divalent ions, which is plausible as it is the largest of the three and thus the steric penalties due to the constriction sites to either side are larger. The Mg2+ ion does not bind as strongly to the second binding site as the other divalent cations, but all three show distinct dips in the profile at the positions of D4938, D4945, E4948, E4952, and D4966, while those dips are especially prominent for magnesium. The behavior of all three divalent ion types is almost identical at the third basin. The standard free energies of binding show an identical trend, as Ba2+ and Ca2+ bind strongly (−30.9 and −30.1 kJ/mol respectively) and Mg2+ follows with a standard free energy of −23.6 kJ/mol.
Monovalent cations like Na+ and K+ are also attracted by the receptor, although the affinity is considerably smaller compared to divalent cations. Potassium ions show the weakest PMF well of only 10.6 kJ/mol and the least attractive standard free energy of binding of −14.1 kJ/mol, which indicates that with no further cations present, RyR1 is a good conductor for potassium ions. Sodium ions have a more attractive standard binding free energy of −20.5 kJ/mol. Furthermore, Na+ ions seem to be almost unaffected by Q4933 and I4937, as the PMF of sodium does not show a free energy well in the center. The PMF for chloride reveals that RyR1 is strictly a cation channel.
To explain the differences in the PMFs of ions with identical valency, it is worth looking at the behavior of their hydration shells. We calculated the number of water molecules in the first and second hydration shell for all ion types across the pore, but only the profile of the second hydration number is shown in Fig. 2c. The first hydration number remained almost constant for all ions, as they kept more than 85% of their first hydration shell across the entire pore axis. With respect to the 2nd hydration number, we find that the profiles closely follow the pore radius. The ions are the most dehydrated at the constriction site caused by the SF, at the secondary constriction site as well as at D4938, where the ions bind closely to the residue and the pore is still narrow. Unsurprisingly, the size of the hydration shell as well as the amount of dehydration depend on the ionic size. In particular, large ions such as chloride, potassium and barium are dehydrated to a much greater extent than the small magnesium ion. Mg2+ is the smallest of all tested ions and as visible in Fig. 2c, it hardly loses water molecules from its second hydration shell. Therefore, the ion remains well shielded against the electrostatic influence of acidic residues, which could explain its small interaction free energy with the pore.
Although sodium ions are considerably smaller than potassium ions, Fig. 2c shows that Na+ is dehydrated to a similar extent as K+. Its smaller size could explain why sodium is almost unaffected by the steric and hydrophobic barrier of Q4933 and I4937. As previously seen with sodium channels69, its higher charge density yields stronger interactions with acidic residues, such as D4938, resulting in a larger binding affinity compared to potassium ions.
Based on the much larger affinity for divalent ions at basin ①, we expect divalent ions to substitute monovalent ions at this point and subsequently block them from permeating. We tested this hypothesis in permeation simulations with applied voltage.
Ion permeation simulations
We have already shown that K+ ions yield a reduced standard binding free energy of only −14.1 kJ/mol, while at the same time showing a similarly shaped PMF compared to Ca2+ ions. We therefore expected to see a conduction behavior similar to calcium ions, in time scales that are currently accessible for simulations. Fortunately, RyR1 is well examined experimentally for both Ca2+ and K+, as many experiments were in fact done using potassium ions70,71,72,73. Hence, the comparison to experiments is possible and could shed light on the general ion permeation mechanism.
The ion permeation simulations allowed for a direct measurement of the current-voltage (IV) dependence of potassium ions through the receptor.
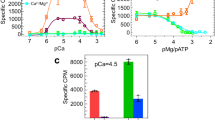
Figure 4 shows a linear IV relationship over the entire tested voltage range, even for the unphysiological value of 1 V. A polynomial of first order was fitted to the data to determine conductances of (668 ± 26) pS at a symmetric physiological concentration of 140 mmol/L. The result is in good agreement with the experimental values between 700 and 800 pS at concentrations of 21074 to 250 mmol/L71,72,73 and is larger than reported values for calcium ions of roughly 100 pS75.
Current-voltage relationship as determined for potassium ions of the open-state RyR1. A linear fit to all data points from −0.2 to 1.0 V reveals a conductance of (668 ± 26) pS at 140 mmol/L. The inset shows a rendering of the simulated system; the arrow symbolizes the direction of the electric field added to the simulation and thus also indicates the direction of permeation.
To determine the role of residues D4899 and E4900 in ion selectivity, we performed a second simulation run of 60 ns at 0.3 V, but saturated each of those eight acidic residues with a calcium ion at its carboxylate group. Indeed, the current decreased from (194 ± 19) pA in the calcium-free case to only (38 ± 11) pA with calcium present. The result indicates that the presence of calcium ions at these high-affinity binding sites imposes a rate limiting energetic barrier for potassium ions. The respective residues therefore serve as a filter for divalent ions over monovalent ions, which was also seen experimentally71,72.
Furthermore, we analyzed the ion trajectories in more detail as depicted in Fig. 5. While all ions passed through the central transmembrane pore, the channel splits into multiple exits at the location of the second main PMF basin. Three modes of exiting the channel into the cytoplasm were observed. Most ions continued drifting in axial direction and exited trough the central pore (Ⓐ). Others moved into one of the water-filled clefts between the four protomers and exited either by diffusing in axial direction inside the cleft (Ⓑ) or by moving laterally (Ⓒ). In the latter case, ions left the receptor on the cytoplasmic side parallel to the membrane. Most, but not all of the ions that left the central pore sideways did so through a window, spanned by R4944, E4948, K4951, E4952 of one monomer and D4945, E4946, and E4949 of the monomer on the other side of the cleft. We propose these residues as targets for a direct experimental validation through mutagenesis studies.
Ion trajectories of the simulation at 0.3 V are visualized by plotting axial coordinate against the distance to the pore axis (radius). Each trajectory is classified and color coded based on the path taken: Ⓐ  , Ⓑ
, Ⓑ  , Ⓒ
, Ⓒ  . The outline of the membrane and the receptor are shown by dashed gray lines for orientation. The inset shows a schematic sketch of the 3 modes of exiting the receptor with their respective labels.
. The outline of the membrane and the receptor are shown by dashed gray lines for orientation. The inset shows a schematic sketch of the 3 modes of exiting the receptor with their respective labels.
To quantify this observation, the path of each pore-passing ion was analyzed and classified into one of the three pathways using a heuristic approach described in the supporting material.
A classified and color coded set of trajectories is shown in Fig. 5.
Taking the values of simulated external voltages up to 0.5 V into account, 53% of trajectories exited via pathway Ⓐ, whereas 25% and 17% exited through Ⓑ and Ⓒ, respectively. The remaining 5% of trajectories could not be classified uniquely according to the criteria described in the supporting material. We could not observe a clear correlation between the external voltage and the number of times a specific exit was chosen in the voltage range from −0.2 V to +0.5 V. However, in the additional simulation conducted at an external voltage of 1 V, 80% of trajectories exited via pathway Ⓐ, as the strong external electric field biases the ionic movement in axial direction.
Discussion
We presented MD simulations on the RyR1 to shed light on the atomistic processes involved in ionic permeation. To address the conduction mechanism and its energetics, we performed umbrella sampling simulations for different ion types and obtained potentials of mean force and standard binding free energies. Major binding sites as well as the responsible key residues were identified. Furthermore, we conducted permeation simulations with an applied voltage and observed previously unknown conduction pathways, in which the ions exit the channel into the cytoplasm through the side of the receptor.
Our study is based on the latest cryo-EM data, which enabled us to perform atomistic simulations of ion conduction in RyR1 for the first time. The used cryo-EM structures, that represent the closed conformation of the receptor, have the highest resolution of all available data sets, especially as compared to these representing open conformations. Therefore, a model of closed RyR1 was built based on these structures and subsequently equilibrated, and later reduced to the significant domains and transitioned into the open state. With the final RMSD of 0.35 nm in the relevant pore region compared to the published open structure by des Georges et al.22, we believe we have built a realistic model of the receptor. Comparison of the radius profile of our model, as shown in Fig. 2c, with the profile published by des Georges et al.22, yields a qualitative agreement. Both profiles show a main constriction site of 3 Å, formed by G4894 in the SF, but we note that after equilibration, the sidechains of I4937 and D4966 extend further into the pore than in the cryo-EM structure. However, the small Cα RMSD of the pore region to the cryo-EM structure (0.35 nm and 0.3 nm without luminal loops) show that the most important part of the channel is well modeled and very stable. As indicated by the overall RMSD of 0.75 nm, larger deviations exist in the peripheral channel regions, where fewer residues were fully resolved by cryo-EM. Furthermore, the reduced model introduces a general caveat to our simulation results; however, the PMFs are typically dominated by local, short range interactions76. Also, the deleted part of the pore is already multiple nanometers in width and thus a direct influence on the conductance and the conduction mechanism seems unlikely.
Our simulations included a pure POPC membrane, which is commonly used for simulations and has been shown to be a good model of the SR membrane77,78. Although phosphatidylcholines (PC) lipids are found in the SR membrane in large quantities, pure POPC does not fully reflect the natural diversity in terms of lipid composition79. Brady et al.80 determined the thickness of the SR membrane as 4.1 Å, which is matched by our model membrane (average P-P distance).
By obtaining PMFs for different ion types, major binding sites for both monovalent and divalent cations were identified. The PMFs for cations, shown in Fig. 2a, have similar features and local minima ①, ②, and, for divalent ions, ③. The PMF shapes, depths, and the free energies of binding are very similar for calcium and barium ions, which is supported by the experimental finding that RyR conducts both ions at similar rates75. The less affine binding free energy of magnesium ions in our simulation suggests a higher conductance for those ions compared to barium and calcium. This deviates from previous work, where conductances followed the sequence Ba2+ > Ca2+ > Mg2+ 75. The disagreement might be due to non-optimal simulation parameters for divalent ions. For monovalent ions, the PMF for K+ is significantly shallower that for Na+, indicating a better potassium conductance, which is also seen in experiments74.
Our findings, summarized in Fig. 2, reveal residues involved in ion permeation, of which some were already identified by mutagenesis studies as vital for the conduction properties of the receptor. Our results offer an atomistic explanation of their functional importance. Starting at the luminal side, acidic luminal loops create an increasingly attractive PMF for monovalent cations, and to an even larger extent, for divalent cations towards the pore entrance. The consequential local increase of cationic concentrations (e.g. visible in the inset of Fig. 4) has been studied previously26. Indeed, Mead-Savery et al.81 showed that the double neutralization ED4832AA on RyR2, which corresponds to ED4902AA on RyR1, has a significant impact on cation selectivity.
Gao et al.71 and Wang et al.72 demonstrated the importance of D4899 and E4900 for the conductance and selectivity of the receptor. Neutralizing the charges of the residues by specific mutations led to decreased currents and a loss of calcium selectivity. The charge-conserving mutations D4899E and E4900D did not have such an effect, which supports our observations that the ion interacts mainly with the charged groups of these residues and that ion conduction is indeed governed by strong electrostatic interactions.
In our simulations, the primary constriction site of 3 Å in the SF is formed by G4894. This explains previous mutation studies where an increase in the side chain volume at this point, by G4894A mutation, dramatically reduces both potassium and calcium currents70,71. Out of the residues forming the second PMF basin, the charge-neutralizing mutations D4938N, D4945N, and E4952Q were carried out successfully in experiments73. In line with our observation of strong electrostatic interactions governing ion permeation in RyR1, D4938N and D4945N showed a decrease in both current and selectivity73. E4952Q did not show a significant effect73, however it is located farther away from the permeation pathway. Additionally, E4955Q yielded unchanged calcium and potassium currents in experiments73, which is also explained by our simulations: although the residue is part of a pore forming helix, its side chain is facing outwards and hence cannot directly interact with permeating ions.
The PMFs, summarized in Fig. 2, reveal that RyR1 has a high affinity for divalent ions, whereas the monovalent cations K+ and Na+, abundant in the SR and the cytoplasm, face a significantly shallower free energy well. This observation suggests a large conductance for those ions, as seen during the permeation simulations with potassium ions (see Fig. 4). Indeed, monovalent ions were found to be conducted at much higher rates than Ca2+ through calcium channels if no calcium was present71,72,73,82,83,84,85,86,87. Due to the high calcium affinity, the pore was occupied and blocked at basin ① by at least one divalent ion during all unbiased equilibration simulations with both monovalent and divalent ions present in a simulation box. Therefore, the shallow free energy wells for monovalent cations do not necessarily translate to an increased conductance for those ions under physiological conditions (i.e. with both K+ and Ca2+ ions available in the solution), which is shown best by the reduced potassium current in our permeation simulations when calcium ions are present. Even when accounting for a probable over-estimation of the calcium affinity in our simulations, which will be discussed in the next paragraph, this observation is consistent with the idea that the RyR achieves its calcium selectivity by expressing over-affinity for calcium ions in the same fashion as previously proposed for both RyR88,89,90 and the L-type calcium channel91. In this model, a calcium ion binds tightly to the pore and blocks the permeation of monovalent cations, since their coulombic repulsion is too weak to release the calcium ion from its binding site. Only with the aid of the stronger electrostatic interaction (e.g. a second calcium ion), the first ion can be released from the binding site and permeate the channel. This idea is strongly supported by the finding that the charge of D4899 and E4900 is vital for the calcium selectivity71,72. The high calcium affinity also suggests a cooperative calcium conduction mechanism, as observed in both Na+ and K+ channels66,92,93 which could be investigated using multi-ion PMFs in future studies. Indeed, we observe multiple ions occupying binding site ② during the applied voltage simulations with potassium ions.
Ion permeation simulations with applied voltage were conducted with potassium, as attempts with calcium were unsuccessful due to the ions getting stuck in the pore. Indeed, overestimated binding affinities of divalent ions, and of calcium ions in particular, are a known problem in classical force fields, including CHARMM36, which we used in this study. In this classical representation, partial charges are fixed during the simulation, which makes the model unable to cover electrostatic polarization effects94. Those effects are particularly important for divalent ions due to their strong electrostatic field. It has been shown that the lack of polarization effects leads to an overestimation of ion-protein affinity in the order of 20% for calcium ions95.
The conductance of (668 ± 26) pS, as obtained using potassium ions, for which the polarization problem is much less critical95, is in very good agreement with the experimental values, ranging between 700 and 800 pS71,72,73,74, which demonstrates the validity of our simulation approach. More careful analysis of the ion permeation simulations revealed possible additional modes of exiting the RyR into the cytoplasm. Although the observation is made on a reduced structure, visual inspection of the full model as well as of the cryo-EM density by des Georges et al.22 indicates that the additional pathways are not blocked by the remaining parts of the receptor (see Supplementary Material Fig. 12). It is an intriguing idea to link those additional pathways to the three commonly observed subconducting states at 1/4, 2/4 and 3/4 of the maximal conductance of RyR196 that were observed for different ion species, including potassium97. In the past, other groups have explained the subconducting states by four individual contributions of the four protomers8,97,98,99,100. Although today’s high resolution 3-D reconstructions clearly show a single, central pore, our study suggests that the ionic current could indeed split up, forming multiple conduction pathways.
The presented MD simulations of RyR1, which are based on the latest structural data, provide new insights into the conduction mechanism of the receptor and its energetics. The PMFs, calculated for different ion types, showed that the residues D4899, E4900, G4894, Q4933, I4937, D4938, E4945, E4952, D4966, D5026 and E5033 are the most important for efficient ion permeation, in a broad agreement with available mutagenesis data. The stronger affinity of RyR1 for divalent ions compared to monovalent ions indicates that the receptor achieves calcium selectivity by over-affinity. We observed this effect directly in ion permeation simulations with applied voltage, using potassium ions. The calculated conductance for K+ ions is in excellent agreement with experimental estimates. The analysis of the conduction pathways revealed that some ions leave the receptor via previously unknown exits pathways, which we suggest as a possible explanation for the commonly observed subconductance states of RyR1.
Change history
02 July 2018
The PDF version of this Article previously published contained an incorrectly sized Figure 3. In addition, the font sizes for the coloured words in the legends of Figures 1, 2 and 5 were incorrect in the HTML version. This has now been corrected in the PDF and HTML versions of the Article.
References
Laver, D. Regulation of ryanodine receptors from skeletal and cardiac muscle during rest and excitation. Clin. experimental pharmacology physiology 33, 1107–1113 (2006).
Wehrens, X. H. & Marks, A. R. Ryanodine Receptors: Structure, function and dysfunction in clinical disease, vol. 254 (Springer Science & Business Media 2004).
Zalk, R., Lehnart, S. E. & Marks, A. R. Modulation of the ryanodine receptor and intracellular calcium. Annu. Rev. Biochem. 76, 367–385 (2007).
Lanner, J. T., Georgiou, D. K., Joshi, A. D. & Hamilton, S. L. Ryanodine receptors: structure, expression, molecular details, and function in calcium release. Cold Spring Harb. perspectives biology 2, a003996 (2010).
Van Petegem, F. Ryanodine receptors: structure and function. J. Biol. Chem. 287, 31624–31632 (2012).
Takeshima, H. et al. Primary structure and expression from complementary dna of skeletal muscle ryanodine receptor. Nat. 339, 439–445 (1989).
Imagawa, T., Smith, J., Coronado, R. & Campbell, K. Purified ryanodine receptor from skeletal muscle sarcoplasmic reticulum is the Ca2+ -permeable pore of the calcium release channel. J. Biol. Chem. 262, 16636–16643 (1987).
Lai, F. A., Erickson, H. P., Rousseau, E., Liu, Q.-Y. & Meissner, G. Purification and reconstitution of the calcium release channel from skeletal muscle. Nat. (1988).
Berchtold, M. W., Brinkmeier, H. & Müntener, M. Calcium ion in skeletal muscle: its crucial role for muscle function, plasticity, and disease. Physiol. Reviews 80, 1215–1265 (2000).
Batters, C., Veigel, C., Homsher, E. & Sellers, J. R. To understand muscle you must take it apart. Front. Physiology 5, 90 (2014).
Rebbeck, R. T. et al. Skeletal muscle excitation–contraction coupling: who are the dancing partners? The international journal of biochemistry & cell biology 48, 28–38 (2014).
Bannister, R. A. Bridging the myoplasmic gap ii: more recent advances in skeletal muscle excitation–contraction coupling. J. Exp. Biol. 219, 175–182 (2016).
Dulhunty, A. F., Wei-LaPierre, L., Casarotto, M. G. & Beard, N. A. Core skeletal muscle ryanodine receptor calcium release complex. Clin. Exp. Pharmacol. Physiol. 44, 3–12 (2017).
Inui, M., Saito, A. & Fleischer, S. Isolation of the ryanodine receptor from cardiac sarcoplasmic reticulum and identity with the feet structures. J. Biol. Chem. 262, 15637–15642 (1987).
Nakai, J. et al. Primary structure and functional expression from cdn a of the cardiac ryanodine receptor/calcium release channel. FEBS letters 271, 169–177 (1990).
Hakamata, Y., Nakai, J., Takeshima, H. & Imoto, K. Primary structure and distribution of a novel ryanodine receptor/calcium release channel from rabbit brain. FEBS letters 312, 229–235 (1992).
Clarke, O. B. & Hendrickson, W. A. Structures of the colossal RyR1 calcium release channel. Curr. Opin. Struct. Biol. 39, 144–152 (2016).
Van Petegem, F. How to open a ryanodine receptor. Cell Res. (2016).
Yan, Z. et al. Structure of the rabbit ryanodine receptor RyR1 at near-atomic resolution. Nat. 517, 50–55 (2015).
Zalk, R. et al. Structure of a mammalian ryanodine receptor. Nature 517, 44–49 (2015).
Efremov, R. G., Leitner, A., Aebersold, R. & Raunser, S. Architecture and conformational switch mechanism of the ryanodine receptor. Nat. 517, 39–43 (2015).
des Georges, A. et al. Structural basis for gating and activation of RyR1. Cell 167, 145–157 (2016).
Bai, X.-C., Yan, Z., Wu, J., Li, Z. & Yan, N. The central domain of RyR1 is the transducer for long-range allosteric gating of channel opening. Cell Research (2016).
Wei, R. et al. Structural insights into Ca2+-activated long-range allosteric channel gating of RyR1. Cell Research 26, 977–994 (2016).
Schilling, R. Modellierung und Molekulardynamik, Simulation der Porenregion des Ryanodin-Rezeptors 1. Diploma Thesis, Heidelberg University (2011).
Schilling, R., Fink, R. H. & Fischer, W. B. MD simulations of the central pore of ryanodine receptors and sequence comparison with 2B protein from coxsackie virus. Biochimica et Biophys. Acta (BBA)-Biomembranes 1838, 1122–1131 (2014).
Shirvanyants, D. et al. Pore dynamics and conductance of RyR1 transmembrane domain. Biophys. Journal 106, 2375–2384 (2014).
Schilling, R., Fink, R. H. & Fischer, W. B. Interaction of ions with the luminal sides of wild-type and mutated skeletal muscle ryanodine receptors. J. Molecular Modeling 22, 1–9 (2016).
Berendsen, H. J., van der Spoel, D. & van Drunen, R. Gromacs: a message-passing parallel molecular dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
Van Der Spoel, D. et al. Gromacs: fast, flexible, and free. J. Computational Chemistry 26, 1701–1718 (2005).
Hess, B., Kutzner, C., Van Der Spoel, D. & Lindahl, E. Gromacs 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chemical Theory Computation 4, 435–447 (2008).
Pronk, S. et al. Gromacs 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinforma. btt055 (2013).
Abraham, M., Hess, B., van der Spoel, D. & Lindahl, E. Gromacs manual 5.1, version 2016-rc1. Accessed: 03.01.2017, http://manual.gromacs.org/documentation/ (2016).
Brooks, B. R. et al. Charmm: a program for macromolecular energy, minimization, and dynamics calculations. J. Computational Chemistry 4, 187–217 (1983).
MacKerell, A. D. Jr. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins? The journal physical chemistry B 102, 3586–3616 (1998).
Brooks, B. R. et al. Charmm: the biomolecular simulation program. J. Computational Chemistry 30, 1545–1614 (2009).
Huang, J. & MacKerell, A. D. Charmm36 all-atom additive protein force field: Validation based on comparison to nmr data. J. Computational Chemistry 34, 2135–2145 (2013).
Hess, B. et al. Lincs: a linear constraint solver for molecular simulations. J. Computational Chemistry 18, 1463–1472 (1997).
Darden, T., York, D. & Pedersen, L. Particle mesh ewald: An n log (n) method for ewald sums in large systems. The J. Chemical Physics 98, 10089–10092 (1993).
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. The J. Chemical Physics 79, 926–935 (1983).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Molecular Graphics 14, 33–38 (1996).
Sali, A. & Blundell, T. Comparative protein modelling by satisfaction of spatial restraints. Protein structure by distance analysis 64, C86 (1993).
Marti-Renom, M. A. et al. Comparative protein structure modeling of genes and genomes. Annu. review biophysics biomolecular structure 29, 291–325 (2000).
Fiser, A., Do, R. K. G. & Šali, A. Modeling of loops in protein structures. Protein science 9, 1753–1773 (2000).
Webb, B. & Sali, A. Comparative protein structure modeling using modeller. Curr. Protocols Bioinformatics 5–6 (2014).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. The phyre2 web portal for protein modeling, prediction and analysis. Nat. Protocols 10, 845–858 (2015).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. The J. chemical physics 126, 014101 (2007).
Berendsen, H. J., Postma, Jv, van Gunsteren, W. F., DiNola, A. & Haak, J. Molecular dynamics with coupling to an external bath. The J. chemical physics 81, 3684–3690 (1984).
Klauda, J. B. Molecular coordinates: Charmm36 FF POPC. Accessed: 13.12.2016, https://terpconnect.umd.edu/~jbklauda/research/download.html (2010).
Yesylevskyy, S. O. Protsqueeze: simple and effective automated tool for setting up membrane protein simulations. J. chemical information modeling 47, 1986–1994 (2007).
Wolf, M. G., Hoefling, M., Aponte-Santamara, C., Grubmüller, H. & Groenhof, G. g_membed: Efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation. J. computational chemistry 31, 2169–2174 (2010).
Khalili-Araghi, F., Ziervogel, B., Gumbart, J. C. & Roux, B. Molecular dynamics simulations of membrane proteins under asymmetric ionic concentrations. The J. general physiology 142, 465–475 (2013).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Physics 52, 7182–7190 (1981).
Nosé, S. & Klein, M. Constant pressure molecular dynamics for molecular systems. Mol. Phys. 50, 1055–1076 (1983).
Martoňák, R., Laio, A. & Parrinello, M. Predicting crystal structures: The parrinello-rahman method revisited. Phys. Review Letters 90, 075503 (2003).
Grubmüller, H. Predicting slow structural transitions in macromolecular systems: Conformational flooding. Phys. Rev. E 52, 2893 (1995).
Lange, O. F., Schäfer, L. V. & Grubmüller, H. Flooding in GROMACS: Accelerated barrier crossings in molecular dynamics. J. Computational Chemistry 27, 1693–1702 (2006).
Kumar, S., Rosenberg, J. M., Bouzida, D., Swendsen, R. H. & Kollman, P. A. The weighted histogram analysis method for free-energy calculations on biomolecules. i. the method. J. Computational Chemistry 13, 1011–1021 (1992).
Roux, B. The calculation of the potential of mean force using computer simulations. Comput. Phys. Commun. 91, 275–282 (1995).
Hub, J. S., De Groot, B. L. & Van Der Spoel, D. g-wham a free weighted histogram analysis implementation including robust error and autocorrelation estimates. J. Chem. Theory Comput. 6, 3713–3720 (2010).
Doudou, S., Burton, N. A. & Henchman, R. H. Standard free energy of binding from a one-dimensional potential of mean force. J. Chemical Theory Computation 5, 909–918 (2009).
Pongprayoon, P., Beckstein, O., Wee, C. L. & Sansom, M. S. Simulations of anion transport through oprp reveal the molecular basis for high affinity and selectivity for phosphate. Proceedings of the National Academy of Sciences 106, 21614–21618 (2009).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. 98, 10037–10041 (2001).
Gumbart, J., Khalili-Araghi, F., Sotomayor, M. & Roux, B. Constant electric field simulations of the membrane potential illustrated with simple systems. Biochimica et Biophys. Acta (BBA)-Biomembranes 1818, 294–302 (2012).
Jensen, M. Ø., Jogini, V., Eastwood, M. P. & Shaw, D. E. Atomic-level simulation of current–voltage relationships in single-file ion channels. The J. General Physiology 141, 619–632 (2013).
Köpfer, D. A. et al. Ion permeation in K+ channels occurs by direct coulomb knock-on. Sci. 346, 352–355 (2014).
Dhakshnamoorthy, B., Rohaim, A., Rui, H., Blachowicz, L. & Roux, B. Structural and functional characterization of a calcium-activated cation channel from tsukamurella paurometabola. Nat. Communications 7, 12753 (2016).
Mowrey, D. D. et al. Ion-pulling simulations provide insights into the mechanisms of channel opening of the skeletal muscle ryanodine receptor. J. Biol. Chem. jbc–M116 (2017).
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nat. 475, 353 (2011).
Zhao, M. et al. Molecular identification of the ryanodine receptor pore-forming segment. J. Biol. Chem. 274, 25971–25974 (1999).
Gao, L. et al. Evidence for a role of the lumenal m3-m4 loop in skeletal muscle ca 2+ release channel (ryanodine receptor) activity and conductance. Biophys. J. 79, 828–840 (2000).
Wang, Y., Xu, L., Pasek, D. A., Gillespie, D. & Meissner, G. Probing the role of negatively charged amino acid residues in ion permeation of skeletal muscle ryanodine receptor. Biophys. J. 89, 256–265 (2005).
Xu, L., Wang, Y., Gillespie, D. & Meissner, G. Two rings of negative charges in the cytosolic vestibule of type-1 ryanodine receptor modulate ion fluxes. Biophys. Journal 90, 443–453 (2006).
Lindsay, A., Manning, S. D. & Williams, A. J. Monovalent cation conductance in the ryanodine receptor-channel of sheep cardiac muscle sarcoplasmic reticulum. The J. Physiology 439, 463–480 (1991).
Tinker, A. & Williams, A. J. Divalent cation conduction in the ryanodine receptor channel of sheep cardiac muscle sarcoplasmic reticulum. The J. general physiology 100, 479–493 (1992).
Allen, T. W., Andersen, O. S. & Roux, B. Energetics of ion conduction through the gramicidin channel. Proc. Natl. Acad. Sci. 101, 117–122 (2004).
Espinoza-Fonseca, L. M. & Thomas, D. D. Atomic-level characterization of the activation mechanism of serca by calcium. Plos One 6, e26936 (2011).
Sonntag, Y. et al. Mutual adaptation of a membrane protein and its lipid bilayer during conformational changes. Nat. Communications 2, 304 (2011).
Fajardo, V. A. et al. Sarcoplasmic reticulum phospholipid fatty acid composition and sarcolipin content in rat skeletal muscle. The J. membrane biology 248, 1089–1096 (2015).
Brady, G., Fein, D., Harder, M., Spehr, R. & Meissner, G. A liquid diffraction analysis of sarcoplasmic reticulum. i. compositional variation. Biophys. Journal 34, 13–34 (1981).
Mead-Savery, F. C. et al. Changes in negative charge at the luminal mouth of the pore alter ion handling and gating in the cardiac ryanodine-receptor. Biophys. Journal 96, 1374–1387 (2009).
Hess, P., Lansman, J. B. & Tsien, R. W. Calcium channel selectivity for divalent and monovalent cations. voltage and concentration dependence of single channel current in ventricular heart cells. The J. general physiology 88, 293–319 (1986).
Kostyuk, P., Mironov, S. & Shuba, Y. M. Two ion-selecting filters in the calcium channel of the somatic membrane of mollusc neurons. The J. Membr. Biol. 76, 83–93 (1983).
Almers, W. & McCleskey, E. Non-selective conductance in calcium channels of frog muscle: calcium selectivity in a single-file pore. The J. Physiol. 353, 585 (1984).
Almers, W., McCleskey, E. & Palade, P. A non-selective cation conductance in frog muscle membrane blocked by micromolar external calcium ions. The J. Physiol. 353, 565 (1984).
Fukushima, Y. & Hagiwara, S. Currents carried by monovalent cations through calcium channels in mouse neoplastic b lymphocytes. The J. Physiology 358, 255 (1985).
Kuo, C.-C. & Hess, P. Ion permeation through the L-type Ca2+ channel in rat phaeochromocytoma cells: two sets of ion binding sites in the pore. The J. Physiology 466, 629 (1993).
Gillespie, D., Xu, L., Wang, Y. & Meissner, G. (de) constructing the ryanodine receptor: modeling ion permeation and selectivity of the calcium release channel. The J. Phys. Chem. B 109, 15598–15610 (2005).
Gillespie, D. Energetics of divalent selectivity in a calcium channel: the ryanodine receptor case study. Biophys. J. 94, 1169–1184 (2008).
Gillespie, D., Giri, J. & Fill, M. Reinterpreting the anomalous mole fraction effect: the ryanodine receptor case study. Biophys. journal 97, 2212–2221 (2009).
Corry, B. & Chung, S.-H. Mechanisms of valence selectivity in biological ion channels. Cell. Mol. Life Sci. CMLS 63, 301–315 (2006).
Chakrabarti, N. et al. Catalysis of Na+ permeation in the bacterial sodium channel navab. Proc. Natl. Acad. Sci. 110, 11331–11336 (2013).
Berneche, S. & Roux, B. Energetics of ion conduction through the K+ channel. Nat. 414, 73–77 (2001).
Harder, E., MacKerell, A. D. Jr. & Roux, B. Many-body polarization effects and the membrane dipole potential. J. Am. Chem. Soc. 131, 2760–2761 (2009).
Li, H. et al. Representation of ion–protein interactions using the drude polarizable force-field. The J. Phys. Chem. B 119, 9401–9416 (2015).
Ahern, G. P., Junankar, P. R. & Dulhunty, A. F. Subconductance states in single-channel activity of skeletal muscle ryanodine receptors after removal of fkbp12. Biophys. J. 72, 146–162 (1997).
Smith, J. S. et al. Purified ryanodine receptor from rabbit skeletal muscle is the calcium-release channel of sarcoplasmic reticulum. The J. Gen. Physiol. 92, 1–26 (1988).
Liu, Q.-Y., Lai, F. A., Rousseau, E., Jones, R. V. & Meissner, G. Multiple conductance states of the purified calcium release channel complex from skeletal sarcoplasmic reticulum. Biophys. Journal 55, 415 (1989).
Brillantes, A.-M. B. et al. Stabilization of calcium release channel (ryanodine receptor) function by fk506-binding protein. Cell 77, 513–523 (1994).
Ondrias, K., Brillantes, A., Scott, A., Ehrlich, B. & Marks, A. Single channel properties and calcium conductance of the cloned expressed ryanodine receptor/calcium-release channel. Soc. Gen. Physiol. Series 51, 29–45 (1995).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Molecular Graphics 14, 354–360 (1996).
Acknowledgements
L.P.H. acknowledges support by the Friedrich Ebert Foundation. R.H.A.F. and L.P.H. acknowledge support by the state of Baden-Württemberg through bwHPC. W.K. and B.L.d.G. are supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) - FOR 2518. The research by R.H.A.F. and L.P.H. was supported financially by the German Excellence Initiative II - Global Networks.
Author information
Authors and Affiliations
Contributions
L.P.H. carried out the simulations, analyzed the data and wrote the manuscript. W.K. and B.L.d.G. supervised the research and revised the manuscript. R.H.A.F. designed and supervised the research and revised the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Heinz, L.P., Kopec, W., de Groot, B.L. et al. In silico assessment of the conduction mechanism of the Ryanodine Receptor 1 reveals previously unknown exit pathways. Sci Rep 8, 6886 (2018). https://doi.org/10.1038/s41598-018-25061-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-25061-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.