Abstract
The enzyme alanine racemase (Alr) has been a new target for the development of antibacterial drugs based on the involvement of D-Ala in bacterial cell wall biosynthesis. Our previous study noted that Alr is essential for the growth and interspecies competitiveness of S. mutans, the major causative organism of dental caries. However, physiological activity and cariogenicity of S. mutans affected by Alr remains unknown. The current study examined the biofilm biomass, biofilm structure, extracellular polysaccharide (EPS) synthesis, glucosyltransferase (gtf) gene expression, acid production and acid tolerance in the alr-mutant strain. We found that biofilm formation, biofilm structure, and EPS synthesis was in a D-Ala dose-dependent manner. Biofilm structure was loose in alr-mutant group and the ratio of EPS/bacteria was also elevated. Additionally, the expression levels of multiple gtfs were up-regulated, and acid tolerance was decreased. We also established in vivo models of dental caries and found that the incidence and severity of the caries were decreased in the alr-mutant group in comparison to the parental S. mutans group. Our in vivo and in vitro experiments demonstrate that Alr is essential for the cariogenicity of S. mutans and that Alr might be a potential target for the prevention and treatment of caries.
Similar content being viewed by others
Introduction
Alanine racemase (Alr) is a bacterial enzyme that catalyses the conversion of L-alanine to D-alanine (D-Ala)1. This function is critical for the growth of bacteria due to their need for D-alanine, an essential component in the biosynthesis of cell wall peptidoglycan in both gram-positive and gram-negative bacteria2. Two kinds of Alr have been identified in bacteria: the alr-encoded racemase, which is constitutive and used for D-Ala biosynthesis, and the dadX-encoded racemase, which is inducible and used for the catabolism of D-Ala1,3,4. Lack of expression of the alr gene is lethal when there is no addition of exogenous D-Ala5. Many studies have focused on alanine racemase, which aim to develop antibacterial drugs for multiple bacterial species, such as Pseudomonas aeruginosa6, Acinetobacter baumannii7, Staphylococcus aureus8, Bacillus anthracis9, Lactobacillus plantarum10, and Escherichia coli11.
Dental caries is one of the most common chronic diseases worldwide and is considered a major health concern for professionals12. It is the main cause of tooth pain, tooth loss and can affect overall health by making food intake more difficult13. Therefore, focusing on the factors affecting caries progression and finding efficient strategies for the management of caries is of great significance. Streptococcus mutans (S. mutans) is the dominant bacterium in dental plaque and has long been implicated as the major causative organism of dental caries14. S. mutans is able to ferment carbohydrates and produce acids that reduce the local pH15. In addition, S. mutans embedded in the dental plaque is resistant to host defences because the extracellular matrix inside the biofilms provides protection from harmful factors16. Exploring factors that influence the cariogenic virulence of S. mutans might aid in the discovery of a more effective target for an anti-caries drug. Considering the essential role of D-alanine for the bacterial cell wall, the alr gene is a potential antibacterial target for S. mutans.
In our previous study, an alr-mutant strain was constructed, and the role of alr in cell growth and cell wall integrity was explored. Our data showed that a minimal concentration of D-Ala (150 µg/ml) was required for the optimal growth of the alr-mutant strain. The depletion of D-Ala in the growth medium led to cell wall perforation and cell lysis in the alr-mutant strain17. Another previous study also demonstrated the important role of D-Ala metabolism for the growth of S. mutans18. Given the importance of alr to the growth of S. mutans, the aim of this study was to explore the role of Alr on the physiological activity, including biofilm formation, extracellular polysaccharide (EPS) synthesis, glucosyltransferases (gtfs) gene expression, acids production and tolerance, and cariogenicity of S. mutans.
Results
A D-Ala dose-dependent manner in alr-mutant biofilm formation
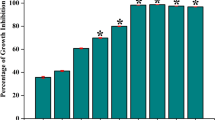
We have studied the growth of WT S. mutans affected by exogenous D-Ala, and the result showed that exogenous D-Ala did not obviously affect the growth of S. mutans (see Supplementary Fig. S1). And we also have determined the biofilm biomass by crystal violet staining of the WT S. mutans affected by different concentrations of exogenous D-Ala, and the result showed no significantly difference either (see Supplementary Fig. S2). Therefore, the addition of D-Ala was a complementation to the alr-mutant. As shown in Fig. 1, Alr is important for alr-mutant strain biofilm formation and biofilm biomass increased in a D-Ala dose-dependent manner. Compared to the parental S. mutans group, biofilm biomass was higher in 200 μg/ml of D-Ala with statistical significance. However, there was no obvious difference among the other two alr-mutant groups and WT S. mutans group.
Biofilm biomass assay by crystal violet staining. The absorbance of the crystal violet-stained S. mutans biofilm at 600 nm is shown with the mean plus standard deviation (SD). The asterisks indicate the significant differences compared to the parental S. mutans strain group. The error bars represent the SD. *P < 0.05.
Loose biofilm structure and short planktonic cells chain of alr-mutant strain
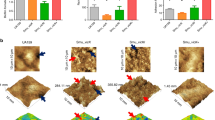
Compared to parental S. mutans group (Fig. 2), the biofilm structure was loose at a concentration of 100 µg/ml of D-Ala, and there were fewer cells but more extracellular matrix within the biofilms. Biofilms became denser as D-Ala concentration increased (150 μg/ml, 200 μg/ml). The morphology of the planktonic cells supplemented with 150 µg/ml of D-Ala was also recorded by a scanning electron microscopy; the alr-mutant cells were more randomly distributed and showed a shorter chain length than the parental S. mutans strain.
Increased EPS synthesis and decreased bacterial cell numbers in the alr-mutant group according to CLSM
As EPS are major components of S. mutans biofilms, we captured images by confocal laser scanning microscopy and performed three-dimensional reconstructions to explore how EPS were affected by Alr. As shown in Fig. 3A, EPS synthesis was markedly enhanced in a D-Ala-dose-dependent manner. At a D-Ala concentration of 100 µg/ml, the EPS around the bacterial cells were rare and became more abundant in the 200 µg/ml group. However, we found that the bacterial cell numbers in three alr-mutant groups were less than cell numbers in the parental S. mutans group (Fig. 3B). The coverage of wild type S. mutans cells was over 20% in most layers while the coverage of alr-mutant strain was less than 20%. The distance between the red line and green line represents the relative ability of bacteria producing EPS. It shows that the alr-mutant strain produced more EPS compared to the parental S. mutans strain. Moreover, the ratio of EPS/bacterial cells formed by the alr-mutant strain was also higher than that of the parental S. mutans strain (Fig. 3C).
Bacterial cell multiplication and EPS synthesis by confocal laser scanning microscopy (CLSM). (A) The three-dimensional reconstruction of biofilms. Reconstruction of the biofilms was performed with IMARIS 7.0. Bacterial cells were labelled with the SYTO 9 green fluorescent dye (left column), and EPS was labelled with the Alexa Fluor 647 red fluorescent dye (middle column). (B) The EPS and bacteria distributions on the reconstructed biofilm. (C) The EPS/bacteria ratio. The asterisks indicate significant differences compared to the parental S. mutans strain group. The error bars represent the standard deviation (SD). *P < 0.05.
Up-regulated expression of EPS synthesis-associated genes in the alr-mutant group according to qRT-PCR analysis
Compared to the parental S. mutans strain, the expression of gtfB, gtfC, and gtfD in the alr-mutant cells was up-regulated in the three different D-Ala concentration groups. However, there was no statistical significance in the 200-µg/ml group, and only gtfC gene expression significantly increased (by 29.3-fold) in the 150 µg/ml group. Nevertheless, the mRNA levels of gtfB, gtfC, and gtfD in the alr-mutant cells of the 100-µg/ml group were significantly up-regulated by 41.7-, 117.9-, and 16.2-fold, respectively (Fig. 4).
Decreased acid tolerance in alr-mutant strain
There was no obvious difference in acid production among alr-mutant groups and wild type S. mutans group (Fig. 5A). However, acid tolerance was significantly decreased in the absence of the alr gene (Fig. 5B). Fewer bacterial colonies were formed by the alr-mutant groups than by the control group. No statistically significant differences were observed for acid tolerance between the three alr-mutant groups that had been supplemented with different D-Ala concentrations.
Decreased cariogenicity of alr-mutant strain in rats
The caries lesions of each rat were scored and recorded according to the Keyes scoring system, which divides lesions into four grades: enamel only (E), slightly dentinal (Ds), moderate dentinal (Dm), and extensive dentinal (Dx). Ds lesions represent the involvement of 1/4 of the dentin between the enamel and the pulp chamber. Dm lesions represent the involvement of 1/4 ~ 3/4 of the dentin region. Dx lesions represent caries progression beyond 3/4 of the dentin region.
Photographs of the caries lesions on the teeth of the rats were obtained using a stereo microscope (Fig. 6A). No caries lesions were observed in the blank group. In the wild type S. mutans group, the range of caries lesions was larger and deeper in depth as most of the sulcal caries had progressed to the slight dentine (Ds); a few moderate dentinal lesions (Dm) were also observed. The 150 µg/ml D-Ala alr-mutant group, however, had fewer Ds lesions.
Caries lesions in rats challenged with the wild type S. mutans or alr-mutant strain. (A) Stereo microscopy images of caries lesions. (B) Statistical analysis of the Keyes scores. The asterisks indicate significant differences compared to the parental S. mutans strain group. The error bars represent the standard deviation (SD). *P < 0.05. (C) The scores for the enamel lesions (E) represent the incidence of caries. The scores for the slightly dentinal lesions (Ds), moderate dentinal lesions (Dm), and extensive dentinal lesions (Dx) represent the severity of the caries.
The statistical analysis is shown in Fig. 6B. More E and Ds lesions were observed in the S. mutans group (P < 0.05) than the alr-mutant group. In addition, no Dx lesions occurred in either group. Figure 6C shows that there was an obvious decrease in the incidence and severity of sulcal caries in the alr-mutant strain group.
Discussion
To explore whether Alr can represent an effective drug target to modulate the cariogenicity of oral biofilm and benefit the management of dental caries, we firstly determined that Alr is essential for the growth and interspecies competitiveness of S. mutans17. A minimal concentration of D-Ala (150 µg/ml) was required for the optimal growth of the alr-mutant strain. D-Ala starvation in the growth medium led to cell wall perforation and cell lysis in the alr-mutant strain. Although Alr has been well demonstrated to be closely associated with the survival and cell wall in S. mutans, its cariogenicity and physiological importance in biofilm is yet to be explored. In the current study, we examined the effect of Alr on S. mutans biofilm formation, biofilm structure, EPS synthesis, the expression of genes involved in sugar production, acid production, and acid tolerance, all of which are responsible for caries development. We also observed that Alr is closely associated with caries development. Similar D-Ala dose-dependent manner was observed in alr-mutant biofilm formation, biomass and EPS synthesis, which suggested alr is important for the mutant biofilm structure.
The loose biofilm structure of the alr-mutant was observed at a concentration of 100 µg/ml of D-Ala. The biofilms became denser as the D-Ala concentration increased. The observed biofilm change may also be attributed to the mutant death and cell lysis we have reported previously. A pronounced short chain length of alr-mutant compared to the wild type under the scanning electron microscopy also supported the possibility.
Interestingly, we also observed that the alr-mutant produced more EPS compared to the parental S. mutans strain with fewer cells in most layers of the biofilm. The EPS/bacteria ratio of alr-mutant was elevated, and the elevated ratio was consistent with the qPCR results of the expression levels of the gtfB, gtfC, and gtfD genes, which might be explained by the regulatory systems inside S. mutans such as two-component signal transduction systems (TCSTS). Some TCSTS e.g. VicRK and CovR are responsible for sucrose-dependent adherence and biofilm formation19,20. The expression of gtfB/C/D is positively regulated by VicR and/or repressed by CovR20. The cell lysis of the alr-mutant was sensed by TCSTS, which subsequently turned on the up-regulation pathway to EPS synthesis for survival as high EPS levels lead to a thick biofilm, causing resistance to immune factors or harmful metabolites16. There was no adequate D-Ala in the 100 μg/ml group, and alr-mutant exhibited strongest EPS-producing ability, indicating stronger survivability in the worst environment. Therefore gtfs expression was highest at 100 μg/ml. However, EPS synthesis was highest in the mutant treated with 200 μg/ml D-Ala. We speculate that the high concentration of D-Ala served as an additional carbon source that the bacteria utilized to produce polysaccharide. As previously mentioned, S. mutans is able to metabolize carbohydrates that leads to acid accumulation and a subsequent pH declination in the dental biofilm21. EPS provide a limiting barrier for acid diffusion by generating acidic microenvironments within the dental plaque. This dual-effect leads to a continuous pH decline to the critical pH, below which tooth hard-tissue demineralization begins and dental caries occurs22,23. Although the alr-mutant produced high EPS levels and limited acid diffusion, the distinctly weak acid tolerance (Fig. 5B) also limited the acid accumulation and continuous pH declination. Alr significantly affected acid tolerance, which we speculate is caused by cell wall instability in the alr-mutant strain, even though there was adequate D-Ala supplied in the culture medium. This hypothesis, however, needs to be verified in a further study. The exact mechanisms are still unknown, but deletion of alr affects the formation as well as structure of the biofilm, indicating that alr plays an important role in the physiological activity of S. mutans.
To better understand the exact role of Alr in the cariogenicity of S. mutans, we developed the caries rat model and found that the caries incidence and severity in the alr-mutant group were lower than in the parental S. mutans. Although EPS synthesis and gtfs expression were increased in the alr-mutant in introductory studies, caries lesions still decreased in the alr-mutant. This might be due to the poor acid tolerance of the alr-mutant strain. There were no sufficient living cells within biofilms and thus no sufficient acid accumulation. These data indicate the compromised cariogenicity of the alr-mutant, further supporting our hypothesis that Alr could be a promising target to control the prevalence of cariogenic of S. mutans.
In conclusion, the current study demonstrated that Alr is important for the physiological activity and cariogenicity of the S. mutans. Alr can represent a promising target for the management of dental caries.
Materials and Methods
Chemicals, bacterial strains and growth conditions
S. mutans UA 159 was obtained from the American Type Culture Collection (Manassas, VA, USA) and was routinely cultured in brain heart infusion broth (BHI; Difco, Sparks, MD, USA) at 37 °C aerobically (95% air/5% CO2). The alr-mutant strain17 was cultured in BHI plus 100, 150, or 200 µg/ml of D-Ala (Sigma). The medium was supplemented with 1% sucrose (called BHIS) when needed. 1 × 107 CFU/ml of S. mutans or the alr-mutant strain was used for both in vitro and in vivo studies.
Crystal violet staining for biofilm biomass analysis
After culturing in 96-well microtiter plates for 24 h, the biofilms were gently washed three times with phosphate buffer saline (PBS), fixed with 95% methanol for 30 min, washed three times with PBS, stained with 0.5% crystal violet for 30 min and washed three times with PBS. The crystal violet was extracted with 200 µl of 100% ethanol. The extract was evaluated at 600 nm using a spectrophotometer24.
Scanning electron microscopy (SEM) analysis of biofilm and planktonic cell morphology
After culturing in 24-well microtiter plates for 24 h, the biofilms (grown on glass slides) were gently washed three times with PBS, fixed with glutaraldehyde (2.5%) overnight at 4 °C, washed three times with PBS, dehydrated using a series of ethanol rinses (30, 50, 70, 80, 85, 90, 95 and 100%), immersed for 10 min in 100% ethanol and dried in a desiccator24,25. For planktonic cells, cells were pelleted by centrifugation at 4000 rpm for 5 min, washed twice with PBS and re-suspended. An aliquot (5.0 µl) of bacterial suspension was deposited on a sterile glass slide and air-dried at 37 °C. Samples were then fixed, dehydrated, immersed and dried26. After coating with gold-palladium, samples were analysed in a scanning electron microscope (Inspect F; FEI, Eindhoven, The Netherlands) at 5000×, 10000×, 20000× magnification.
Confocal laser scanning microscopy (CLSM) for the assessment of extracellular polysaccharide (EPS) synthesis within biofilms
Biofilms cultured on glass slides were grown in BHIS medium and supplemented with 1 µM Alexa 647 red fluorescent dye (Molecular Probes Inc., OR, USA) and protected from light. Alexa 647 are labelled dextran conjugates and were incorporated into EPS. After incubation for 24 h, the biofilms were washed with PBS three times and incubated with 1 µM SYTO 9 green fluorescent dye (Molecular Probes Inc., OR, USA) at room temperature for 20 min. SYTO 9 bound to bacterial nucleic acid and represented bacteria in biofilms. Then the biofilms were washed with PBS three times and dried. ProLong Gold Antifade Reagent was dropped on the biofilms, and images were obtained using a confocal laser scanning microscope (TCS SP2; Leica Microsystems, Wetzlar, Hessen, Germany). The Alexa Fluor 488 and 647 fluorescent channels were selected to detect green and red fluorescence respectively. The whole procedure was performed away from light27,28.
Quantitative real-time PCR (qRT-PCR) analysis of S. mutans EPS synthesis-associated genes
The qRT-PCR protocol has been described in detail in our previous studies29,30,31. Briefly, the biofilms were harvested after incubation in BHIS medium for 24 h, washed three times with PBS, and stabilized using the RNAprotect Bacteria Reagent (1 ml, Qiagen, MD, USA). Then, the biofilms were suspended in 180 µl of lysis buffer (containing 20 mM Tris-HCl, pH 8.0, 1.2% Triton-100, 1 mM EDTA-Na2, and 30 mg/ml of lysozome) with 10 µl of mutanolysin (10 KU/ml, Sigma-Aldrich, MO, USA) and 15 µl of proteinase K (Qiagen), incubated with agitation at 37 °C for 120 min, and sonicated for 5 cycles (10 s/cycle, 52% amplitude, Sonic Dismembrator, Model 500, Fisher Scientific). The biofilm RNA was extracted and purified using the RNeasy Mini kit (Qiagen). The RNA concentration was measured using a NanoDrop 2000 (Thermo Fisher Scientific Inc., USA). A high-capacity cDNA reverse transcription kit with random primers (Applied Biosystems, Life Technologies Corp., CA, USA) was used to synthesize cDNA from 10 µg RNA. PCR reactions comprised two micrograms of cDNA, S. mutans primers (0.375 µM) and the Fast SYBR green master mix (Applied Biosystems) and were run using an ABI Prism 7000 sequence detection system for quantitative PCR. The primers were 16 S, gtfB, gtfC, and gtfD31. The values of 2−∆∆Ct were analysed to calculate the gene expression fold changes21.
Glycolytic pH drop assay for acid production
The role of Alr in S. mutans glycolysis was measured using the method described by Xu et al.32. The bacteria were harvested at the mid-logarithmic growth phase, washed with PBS and resuspended (OD600 nm = 0.5) in 0.5 mM potassium phosphate buffer consisting of 1.25 mM MgCl2 and 37.5 mM KCl (pH = 6.5). Glucose was added to the bacterial suspension, and the final concentration was adjusted to 1% (wt/vol). The glycolytic activity of S. mutans and the alr-mutant strains resulted in a decrease in pH, and the pH levels were monitored at 15-min intervals over a period of 120 min.
Acid tolerance assay
The role of Alr in S. mutans acid tolerance was also measured using the method described by Xu et al.32. The bacteria were harvested at the mid-logarithmic growth phase, collected by centrifugation and resuspended (OD600 nm = 0.2) in TYEG medium (containing 10% tryptone, 5% yeast extract, 3% K2HPO4, and 1% glucose) buffered with 40 mM phosphate-citrate buffer solution (pH = 5.0) and incubated at 37 °C aerobically for 2 h. After incubation at pH 5.0, samples were removed for viable counts.
In vivo models of dental caries
The in vivo study was approved by the ethics committee of West China School of Stomatology, Sichuan University (WCCSIRB-D-2014-072), and all experiments were performed according to the National Institutes of Health Guide for the Care and Use of Laboratory Animals.
Sixteen specific pathogen-free (SPF) male Wistar rats, aged 21 days, were randomly divided into three groups: S. mutans (n = 7), alr-mutant (150 µg/ml, n = 7), and blank (n = 2). Upon arrival, animals were determined to be free of any indigenous oral microorganism by feeding the rats antibiotics, namely, ampicillin, chloramphenicol and carbenicillin (1.0 g/kg), for 3 consecutive days33. After a washout period of 3 days, in which the residual antibiotics in the oral cavity were removed by feeding rats antibiotic-free distilled water, rats were challenged with 1.0 × 107 CFU/ml S. mutans or alr-mutant strain suspensions for 3 consecutive days (twice per day at 30-min intervals with no food or water for 1 h after incubation). All rats were fed the National Institutes of Health cariogenic diet 2000 and 5% sucrose water34. For the alr-mutant group, D-Ala was added to the water (150 µg/ml). The experiment lasted for 27 days, after which the rats were sacrificed. The jaws of the rats were aseptically dissected, and the caries status was scored using the Keyes method35.
Statistical Analysis
Each experiment was independently repeated at least three times. One-way analysis of variance (ANOVA) was used to analyse the crystal violet staining, acid tolerance, and qRT-PCR. Independent t-tests were used for the in vivo study. The data were statistically analysed using the SPSS 21.0 software. P < 0.05 was considered statistically significant.
Data Availability
The datasets generated and analysed during the current study are available from the corresponding authors on reasonable requests.
References
Walsh, C. T. Enzymes in the D-alanine branch of bacterial cell wall peptidoglycan assembly. J Biol Che 264, 2393–2396 (1989).
Strych, U., Huang, H. C., Krause, K. L. & Benedik, M. J. Characterization of the alanine racemases from Pseudomonas aeruginosa PAO1. Curr Microbiol 41, 290–294, https://doi.org/10.1007/s002840010136 (2000).
He, W., Li, C. & Lu, C. D. Regulation and characterization of the dadRAX locus for D-amino acid catabolism in Pseudomonas aeruginosa PAO1. J Bacteriol 193, 2107–2115, https://doi.org/10.1128/jb.00036-11 (2011).
Liu, J. L., Liu, X. Q. & Shi, Y. W. Expression, purification, and characterization of alanine racemase from Pseudomonas putida YZ-26. World J Microbiol Biotechnol 28, 267–274, https://doi.org/10.1007/s11274-011-0816-1 (2012).
Hernandez, S. B. & Cava, F. Environmental roles of microbial amino acid racemases. Environ Microbiol 18, 1673–1685, https://doi.org/10.1111/1462-2920.13072 (2016).
LeMagueres, P. et al. Crystal structure at 1.45 angstrom resolution of alanine racemase from a pathogenic bacterium, Pseudomonas aeruginosa, contains both internal and external aldimine forms. Biochemistry 42, 14752–14761, https://doi.org/10.1021/bi030165v (2003).
Davis, E., Scaletti-Hutchinson, E., Opel-Reading, H., Nakatani, Y. & Krause, K. L. The structure of alanine racemase from Acinetobacter baumannii. Acta Crystallogr F Struct Biol Commun 70, 1199–1205, https://doi.org/10.1107/s2053230x14017725 (2014).
Scaletti, E. R., Luckner, S. R. & Krause, K. L. Structural features and kinetic characterization of alanine racemase from Staphylococcus aureus (Mu50). Acta Crystallogr D Biol Crystallogr 68, 82–92, https://doi.org/10.1107/S0907444911050682 (2012).
Counago, R. M., Davlieva, M., Strych, U., Hill, R. E. & Krause, K. L. Biochemical and structural characterization of alanine racemase from Bacillus anthracis (Ames). BMC Struct Biol 9, 53, https://doi.org/10.1186/1472-6807-9-53 (2009).
Palumbo, E. et al. Knockout of the alanine racemase gene in Lactobacillus plantarum results in septation defects and cell wall perforation. FEMS Microbiol Lett 233, 131–138, https://doi.org/10.1016/j.femsle.2004.02.001 (2004).
Strych, U. & Benedik, M. J. Mutant analysis shows that alanine racemases from Pseudomonas aeruginosa and Escherichia coli are dimeric. J Bacteriol 2002, 184, (15), 4321–4325 https://doi.org/10.1128/jb.184.15.4321-4325 (2002)
Kaste, L. M. et al. Coronal caries in the primary and permanent dentition of children and adolescents 1-17 years of age: United States, 1988–1991. J Dent Res 75, 631–641, https://doi.org/10.1177/002203459607502S03 (1996).
Selwitz, R. H., Ismail, A. I. & Pitts, N. B. Dental caries. Lancet 369, 51–59, https://doi.org/10.1016/s0140-6736(07)60031-2 (2007).
Loesche, W. J. Role of Streptococcus-mutans in human dental decay. Microbiol Rev 50, 353–380 (1986).
de Soet, J. J., Nyvad, B. & Kilian, M. Strain-related acid production by oral streptococci. Caries Res 34, 486–490, https://doi.org/10.1159/000016628 (2000).
Donlan, R. M. & Costerton, J. W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev 15, 167–193, https://doi.org/10.1128/cmr.15.2.167-193.2002 (2002).
Wei, Y. et al. Alanine racemase is essential for the growth and interspecies competitiveness of Streptococcus mutans. Int J Oral Sci 8, 231–238, https://doi.org/10.1038/ijos.2016.34 (2016).
Qiu, W. et al. D-alanine Metabolism is Essential for Growth and Biofilm Formation of Streptococcus mutans. Mol Oral Microbiol 31, 435–444, https://doi.org/10.1111/omi.12146 (2016).
Rasamiravaka, T. & El Jaziri, M. Quorum-Sensing Mechanisms and Bacterial Response to Antibiotics in P. aeruginosa. Curr Microbiol 73, 747–753, https://doi.org/10.1007/s00284-016-1101-1 (2016).
Smith, E. G. & Spatafora, G. A. Gene Regulation in S. mutans: Complex Control in a Complex Environment. J Dent Res 91, 133–141, https://doi.org/10.1177/0022034511415415 (2012).
Decker, E.-M., Klein, C., Schwindt, D. & von Ohle, C. Metabolic activity of Streptococcus mutans biofilms and gene expression during exposure to xylitol and sucrose. Int J Oral Sci 6, 195–204, https://doi.org/10.1038/ijos.2014.38 (2014).
Bowen, W. H. & Koo, H. Biology of Streptococcus mutans-Derived Glucosyltransferases: Role in Extracellular Matrix Formation of Cariogenic Biofilms. Caries Res 45, 69–86, https://doi.org/10.1159/000324598 (2011).
Xiao, J. et al. The Exopolysaccharide Matrix Modulates the Interaction between 3D Architecture and Virulence of a Mixed-Species Oral Biofilm. PLoS Pathog 8, e1002623, https://doi.org/10.1371/journal.ppat.1002623 (2012).
Huang, R., Li, M. & Gregory, R. L. Effect of nicotine on growth and metabolism of Streptococcus mutans. Eur J Oral Sci 120, 319–325, https://doi.org/10.1111/j.1600-0722.2012.00971.x (2012).
Li, M. et al. Effect of nicotine on cariogenic virulence of Streptococcus mutans. Folia Microbiol 61, 505–512, https://doi.org/10.1007/s12223-016-0465-8 (2016).
Tu, H. et al. Activity of Synthetic Antimicrobial Peptide GH12 against Oral Streptococci. Caries Res 50, 48–61, https://doi.org/10.1159/000442898 (2016).
Zhou, H. et al. Evaluation of three-dimensional biofilms on antibacterial bonding agents containing novel quaternary ammonium methacrylates. Int J Oral Sci 6, 77–86, https://doi.org/10.1038/ijos.2014.18 (2014).
Li, M. et al. Effect of nicotine on dual-species biofilms of Streptococcus mutans and Streptococcus sanguinis. FEMS Microbiol Lett 350, 125–132, https://doi.org/10.1111/1574-6968.12317 (2014).
Huang, R. et al. Effects of Nicotine on Streptococcus gordonii Growth, Biofilm Formation, and Cell Aggregation. Appl Environ Microbiol 80, 7212–7218, https://doi.org/10.1128/AEM.02395-14 (2014).
Huang, R., Li, M. & Gregory, R. L. Nicotine promotes Streptococcus mutans extracellular polysaccharide synthesis, cell aggregation and overall lactate dehydrogenase activity. Arch Oral Biol 60, 1083–1090, https://doi.org/10.1016/j.archoralbio.2015.04.011 (2015).
Li, M., Huang, R., Zhou, X. & Gregory, R. L. Role of sortase in Streptococcus mutans under the effect of nicotine. Int J Oral Sci 5, 206–211, https://doi.org/10.1038/ijos.2013.86 (2013).
Xu, X., Zhou, X. D. & Wu, C. D. The Tea Catechin Epigallocatechin Gallate Suppresses Cariogenic Virulence Factors of Streptococcus mutans. Antimicrob Agents Chemother 55, 1229–1236, https://doi.org/10.1128/AAC.01016-10 (2011).
Bao, R. et al. Flagellin-PAc Fusion Protein Inhibits Progression of Established Caries. J Dent Res 94, 955–960, https://doi.org/10.1177/0022034515582224 (2015).
Falsetta, M. L. et al. Symbiotic Relationship between Streptococcus mutans and Candida albicans Synergizes Virulence of Plaque Biofilms In Vivo. Infect Immun 82, 1968–1981, https://doi.org/10.1128/IAI.00087-14 (2014).
Keyes, P. H. ental caries in the molar teeth of rats. II. A method for diagnosing and scoring several types of lesions simultaneously. J Dent Res 37, 1088–1099, https://doi.org/10.1177/00220345580370060901 (1958).
Acknowledgements
We are grateful to Chaoliang Zhang for the technical support of SEM and CLSM. This study was supported by the National Natural Science Foundation of China (81400501 to M.L. and 81430011 to X.Z.), the International Science and Technology Cooperation Program of China 2014DFE30180 (X.Z.), the Special Fund of State Key Laboratory of Oral Diseases, Sichuan University (SKLOD201525 to M.L.).
Author information
Authors and Affiliations
Contributions
Shiyu Liu, Yuan Wei and Mingyun Li contributed to the design, data acquisition, analysis, drafting and revision of the manuscript. Biao Ren and Xian Peng contributed to the design of the study. Keke Zhang contributed to the data acquisition. Xuedong Zhou, Lei Cheng, and Vivian Chen contributed to the revision of the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, S., Wei, Y., Zhou, X. et al. Function of alanine racemase in the physiological activity and cariogenicity of Streptococcus mutans. Sci Rep 8, 5984 (2018). https://doi.org/10.1038/s41598-018-24295-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-24295-1
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.