Abstract
In the apolipoprotein E–deficient mouse, the gut microbiota has an impact on the development of atherosclerosis, but whether such correlations are also present in rats requires investigation. Therefore, we studied female SD-Apoetm1sage (Apoe−/−) rats fed either a Western diet or a low-fat control diet with or without gluten, which is known to promote gut microbiota changes, until 20 weeks of age. We hypothesized that the manifestation of atherosclerosis would be more severe in Apoe−/− rats fed the Western high-fat diet, as compared with rats fed the low-fat diet, and that atherosclerosis would be accelerated by gluten. Both Western diet-feeding and gluten resulted in significant changes in gut microbiota, but the microbiota impact of gluten was transient. Compared with Apoe−/− rats fed a low-fat diet, Western diet-fed Apoe−/− rats were heavier and became glucose intolerant with increased levels of oxidative stress. They developed early fatty streak lesions in their aortic sinus, while there was no evidence of atherosclerosis in the thoracic aorta. No conclusions could be made on the impact of gluten on atherosclerosis. Although Western diet-fed Apoe−/− rats exhibited a more human-like LDL dominated blood lipid profile, signs of obesity, type 2 diabetes and cardiovascular disease were modest.
Similar content being viewed by others
Introduction
In the past 25 years, genetically altered mouse models, such as the apolipoprotein E (ApoE) deficient (Apoe−/−) mouse, have been studied to understand the underlying pathophysiological mechanisms of cardiovascular disease and the development of therapies1,2. The Apoe gene is expressed in numerous tissues including brain, liver and adipose tissue1,3 and ApoE mediates the binding of chylomicrons and LDL to the LDL-receptor, preventing the accumulation of cholesterol rich particles in the plasma1,4,5,6,7,8. Mouse studies have demonstrated that loss of ApoE results in accumulation of triglyceride-rich lipoprotein (TRL) remnants, elevated cholesterol in the blood and the development of atherosclerotic lesions; a condition, which can be accelerated by diets rich in fat and cholesterol9,10. Aortic lesions progress to advanced human-like stages, characterized by the presence of necrotic cores, fibrous caps, extracellular matrix components, and cholesterol clefts.
Within recent years the gut microbiota has been accepted as an important player in development of a series of conditions – including cardiovascular disease and insulin resistance11,12,13. Genetic factors and equally important environmental factors such as diet, antibiotic or prebiotic treatments play central roles in establishing and also maintaining an individual’s gut microbiota composition14,15,16,17,18. Mice fed diets containing gluten or its active component gliadin differ significantly in their gut microbiota composition from mice fed a gluten free diet19. A gluten free diet has a dramatic impact on reducing the incidence of type 1 diabetes20 in NOD mice, a less dramatic impact on the development of type 2 diabetes21,22, and as we have recently demonstrated, it does not impact development of atherosclerosis in Apoe−/− mice23. In these mice, the impact of the gut microbiota is implicated by results demonstrating that ampicillin treatment reduces LDL and VLDL cholesterol levels and protects from aortic lesion development23. Also the gluten effect on gut microbiota is strong at the age of five weeks, but it seems to disappear at the age of 16 weeks23. The size of rats is advantageous, as this facilitates plaque lesion analysis, tissue and blood collection and more extensive plasma analyses. Rodents generally have rapid LDL-clearance, and HDL-cholesterol is the largest of the lipoprotein fractions1,24,25, which might explain why rodents do not spontaneously develop atherosclerosis.
Due to limited possibilities of cultivating embryonic stem cells from other species, gene knockout has mostly been performed on mice until a few years ago. Today, the introduction of the nuclease techniques, such as Zinc Finger (ZF)26, Transcription Activator-Like Effector Nuclease (TALEN)27 and recently Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR/Cas9)28 have enabled easy gene deletion in species other than mice. In 2011, scientists at Sigma’s Advanced Genetic Engineering (SAGE) Labs created ApoE gene knockout (Apoe−/−) rats by the use of ZF technology29. Only limited data on Apoe−/− rats exist29,30. In two previous studies describing Apoe−/− rat models29,30, very early fatty streak lesions were demonstrated. However, in one of the studies, in which the rats were fed a high fat diet for only 12 weeks, no vascular lesions were detected unless the carotid artery was partially ligated30. So, in general there is an obvious need to further validate Apoe−/− rats as a model of dyslipidemia, glucose intolerance, changes in gut microbiota, and atherosclerosis and more specifically, to investigate the translational aspects of our previous observations on the relationship between gut microbiota and atherosclerosis in Apoe−/− mice in a larger model such as a rat.
The aim of the present study, therefore, was to evaluate the development of atherosclerosis, weight gain, insulin resistance and early signs of liver damage, in Apoe−/− rats under two different gut microbiota compositions, i.e. with or without gluten, when fed a Western high-fat diet (WD) compared to a control low-fat diet (LF) for a longer period of time than previous studies (Fig. 1). We hypothesized that gluten would change the gut microbiota composition, that the manifestation of atherosclerosis would be more severe in Western high-fat diet fed compared to low-fat diet fed rats, and that the manifestation of atherosclerosis would be accelerated in gluten-fed rats as compared with rats fed a gluten-free diet.
Timeline and Study design. (A) Timeline of the study showing time points for Oral glucose tolerance test (OGTT), blood and faeces sampling. (B) Animals were divided into four dietary groups; Group LF received a low fat control diet (n = 16 at all time points), group LF + G received a low fat control diet containing 3.5% gluten (n = 16 at week 8, n = 15 at week 14 and n = 14 at week 20; 2 rats were euthanized at week 11 and 19, respectively), group WD received Western Diet (n = 14 at week 8, n = 13 at week 14 and n = 13 at week 20; 3 rats were euthanized at week 6, 7 and 8, respectively), and group WD + G received Western Diet containing 3.5% gluten (n = 8 at all time points (4 rats were euthanized on or before week 7).
Results
Dietary fat and gluten change gut microbiota composition in Apoe −/− rats
Female Apoe−/− rats were divided into four groups, and each group was fed one of four different diets; a gluten-free low-fat diet (LF), a gluten-containing LF diet (LF + G), a gluten-free Western diet (WD) or a gluten-containing WD (WD + G), as shown in Fig. 1A,B. To avoid maternal milk gluten transfer, the experimental diets were fed ad libitum to the mothers from approximately 10 days pre-partum, and subsequently to the offspring until 20 weeks of age. Qualitatively, compositional gut microbiota changes in eight week-old rats were demonstrated as a response to both dietary fat (WD vs. LF; P = 0.001 and WD + G vs. LF + G; P = 0.001, un-weighed data, Fig. 2A) and dietary gluten content (WD vs. WD + G; P = 0.001 and LF vs. LF + G; P = 0.038, un-weighed data, Fig. 2A). At 20 weeks of age, clustering could only be demonstrated with respect to the dietary fat content (WD vs. LF; P = 0.001 and WD + G vs. LF + G; P = 0.027, un-weighed data, Fig. 2C). Quantitatively, significant differences as a response to dietary fat content were observed in the eight week-old animals (WD vs. LF; P = 0.001, weighed data, Fig. 2B), and after 20 weeks (WD + G vs. LF + G; P = 0.042, weighed data, Fig. 2D). WD differed significantly from LF + G at both eight and 20 weeks and both quantitatively (P = 0.001 and P = 0.014, respectively) and qualitatively (P = 0.001 and P = 0.001, respectively) (Fig. 2), while WD + G also differed significantly from LF at both eight and 20 weeks, but only qualitatively (P = 0.008 and P = 0.002) (Fig. 2). As revealed by the un-weighted UniFrac distance metrices analysis, there were significant differences among less abundant species, such as Parabacteroides and Eubacterium (Fig. 3).
Gut microbiota composition changes in response to Western (WD) or low fat (LF) diet and gluten (G) content. (A–D) PCoA plots generated based on unweighted (A) and weighted (B) distance metrics visualizing the level of qualitative and quantitative similarities between the gut microbial composition of young (8 weeks of age), and adult rats (20 weeks of age) - unweighted (C) and weighted (D). Tables below the plots present results of ANOSIM analysis performed for randomly selected unweighted (regular font) or weighted (bold font) rarefied uniFrac distance metrics.
Western Diet (WD) and/or gluten feeding of Apoe−/− rats induces changes in composition of low abundant microbiota taxa. Dietary gluten reduced the abundance of the genus Parabacteroides (50%, 0.2%, 9%, and 0.2%) in the Western Diet without gluten (WD), Western Diet with gluten (WD + G), low fat control without gluten (LF), and low fat control with gluten (LF + G) category, respectively), the relative abundance of which was increased in eight week old WD-fed compared to LF-fed rats (Bonferroni corrected P = 0.000). In the 20 weeks old animals, the genus Eubacterium (greengenes13.8 unofficial name representing the Erysipelotrichaceae family) was significantly increased in abundance (Bonferroni corrected P = 0.014) in the WD group (0.4%, 0.1%, 0.03%, and 0.02% in the WD, WD + G, LF, and LF + G category, respectively). No significant qualitative differences in genera distribution between tested categories were found with G-test. Worth noting is it that the number of estimated species in the LF + G group was significantly reduced at eight weeks of age compared to the WD and WD + G categories (P = 0.042 and P = 0.012 respectively; data not shown).
Western diet-fed Apoe −/− rats develop small early fatty streaks in the aortic sinus but not in the thoracic aorta
By the method described by Tangirala and reviewed by Meir1,31, we found evidence of very early fatty streak lesions in the aortic sinus. WD-fed rats exhibited small early fatty streak lesions, whereas no lesions were present in LF-fed rats (Fig. 4A,B). The oil red O-positive area out of total cross-sectional aortic sinus intimal area was significantly lower in LF-fed compared to WD-fed rats, but the effect of WD-feeding was very low compared to previous findings in the Apoe−/− mouse at a similar age (P = 0.000; 0.14 ± 0.09% in LF-fed rats, 0.18 ± 0.12% in LF + G-fed rats, 2.34 ± 0.40% in WD-fed rats, and 1.38 ± 0.27% in WD + G-fed rats) (Fig. 4C). Furthermore, the lesions were far less severe32. We could not show any accelerating effect of gluten on fatty streak size in WD-fed rats, and although it might seem as if there was an alleviating effect of gluten, power and study design does not allow such conclusions. In the thoracic aortas of 20 week-old rats, no lesions could be detected macroscopically in either WD-fed or LD-fed rats (data not shown), consistent with the previous study on Apoe−/− rats by Wei et al.30. Also, no evidence of increased presence of gene markers of macrophages and monocytes (Cd68, Adgre1), markers of lipid accumulation (Abca1, Abcg1), cytokines (Il1b, Il6, Tnfa), chemokines (Ccl2), or adhesion molecules (Icam1, Vcam1) were detected in aortas from WD-fed Apoe−/− rats, further supporting the conclusion that even early fatty streak lesions are absent in the thoracic part of aorta at the 20-week time-point (Fig. S1).
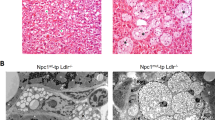
Western Diet (WD)-fed Apoe−/− rats exhibit small fatty streak lesions of atherosclerosis limited to the aortic sinus. (A) Histological slides of the aortic sinus in an Apoe−/− rat fed the LF diet without detectable plaque (40X, Stained with Oil Red O), and Apoe−/− rat fed the WD with an early fatty streak-type lesion (40X, Stained with Oil Red O). (B) Higher magnification of the fatty streak lesion in A. The lesion is indicated by an arrow. (C) Graphical representation of the lesion area measurements at 20 weeks of age showed significantly higher plaque burden in animals fed Western diets (groups 3 and 4) compared with animals fed the low fat control diet. Data are expressed as mean ± SEM. n = 14 in the low fat (LF) and low fat + gluten (LF + G) groups, n = 11 in the Western Diet (WD) group and n = 7 in the Western Diet + gluten (WD + G) group.
Western diet-fed Apoe −/− rats have a human-like lipoprotein profile with elevated levels of blood lipids and liver enzymes
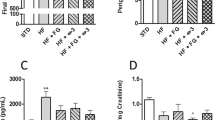
Total plasma cholesterol levels were significantly higher in the WD-fed rats at 8, 14 and 20 weeks of age (Fig. 5A) compared to LF-fed rats (P = 0.000). A plasma cholesterol level of 30–35 mmol/L after 20 weeks on WD is similar to levels in TALEN-generated Apoe−/− rats fed a high fat diet for 12 weeks30. In the present study, the Apoe−/− rats had human-like plasma profiles with LDL-fractions not dropping below 50% of the total plasma cholesterol at any time point regardless of diet (Fig. 5B–D and Table 1). Both total VLDL and the relative level of VLDL were significantly increased after WD feeding as compared to LF feeding (P = 0.000) (Fig. 5B–E). As expected, total LDL levels (Fig. 5F) were significantly increased with WD feeding (P = 0.000), but the relative levels of LDL were stable between experimental groups. Furthermore, even though total HDL levels were significantly increased in WD-fed rats (P = 0.000) (Fig. 5G), the relative contribution of HDL was reduced with WD feeding, especially later on during the study (Fig. 5B–D, Table 1). Consistent with elevated VLDL levels, WD-feeding also resulted in significantly elevated plasma triglyceride levels (P = 0.000) (Fig. 5H). The addition of gluten to the diets had no effect on lipid levels. WD-fed compared to LF-fed rats had significantly higher plasma levels of the liver enzymes ALT and AST (P = 0.000) (Fig. 5I,J).
Western diet (WD)-fed Apoe−/− rats exhibit elevated blood lipid levels and human-like lipoprotein profiles. (A) At 8, 14 and 20 weeks of age low fat diet (LF) fed groups had significantly lower total plasma cholesterol levels than Western Diet (WD) fed groups (n = 4/group). (B–D) Graphs showing the percentagewise distribution of the lipoprotein fractions HDL, LDL and VLDL (n = 4/group). See also Table 1. (E–G) Data points for VLDL, LDL and HDL cholesterol represent four samples, each consisting of pooled plasma from two rats. There was a significant impact of diet on VLDL and LDL at 8 weeks (P = 0.000 and P = 0.026, respectively), on VLDL also at 14 weeks (P = 0.000) and 20 weeks (P = 0.000) and on HDL at 14 weeks (P = 0.000) and 20 weeks (P = 0.000). (H) Triglyceride levels were measured at 20 weeks of age and showed a significant difference between the groups related to dietary fat content (n = 16 in the LF and LF + G groups, n = 12 in the WD group, n = 8 in the WD + G group). (I,J) The liver enzymes ALT and AST were both significantly elevated in the WD compared to LF groups (n = 16 in the LF group, n = 14 in the LF + G group, n = 13 in the WD group, n = 8 in the WD + G group). Gluten did not have any significant influence on blood lipids and liver enzymes. Data are expressed as mean ± SEM.
Western diet-fed Apoe −/− rats gain more weight than low fat-fed Apoe −/− rats and are glucose intolerant
WD-fed rats exhibited a modest, albeit significant, body weight gain, as compared to LF-fed rats (P = 0.000), while dietary gluten did not affect weight gain (Fig. 6A,B). Consistent with these observations, WD-fed rats showed an overall impaired glucose tolerance at 14 and 20 weeks of age (P = 0.000), as compared with the LF-fed rats (Fig. 6C–E), and at 8, 14 and 20 weeks of age, their rise in insulin levels during the first 30 minutes of OGTTs was higher (P = 0.000) (Fig. 6F–H). However, this did not result in elevated levels of glycated hemoglobin (HbA1c) (Fig. S2).
Western Diet (WD)-fed Apoe−/− rats gain more weight and are glucose intolerant. (A) Dietary fat content related weight differences (Area under curve (AUC)). Low fat (LF) (n = 16), low fat + gluten (LF + G) (n = 14), Western Diet (WD) (n = 13) and Western Diet + Gluten (WD + G) (n = 8). (B) Dietary fat content weight gain differences (AUC). (C) Oral glucose tolerance tests (OGTT) at 8 weeks did not differ significantly over all (P = 0.0651). (LF (n = 16), LF + G (n = 15), WD (n = 14), WD + G (n = 8)). (D,E) OGTTs at 14 and 20 weeks differed significantly related to dietary fat content (P = 0.000 and P = 0.000 respectively). Week 14: LF (n = 16), LF + G (n = 15), WD (n = 13), WD + G (n = 8); Week 20: LF (n = 16), LF + G (n = 14), WD (n = 13), WD + G (n = 8). Gluten did not influence OGTTs. (F–H) ΔInsulin (t = 30 minutes − t = 0 minutes) differed significantly between groups related to dietary fat content at all three time points (LF (n = 15), LF + G (n = 16), WD (n = 13), WD + G (n = 8)). Data are expressed as mean ± SEM.
Western diet-fed Apoe −/− rats exhibit elevated markers of oxidative stress
Oxidative stress might be elevated by WD feeding, so two markers of oxidative stress were measured. In the WD-fed rats, lower levels of reduced BH4 compared to its oxidation product BH2 may be expected as a result of increased oxidative stress. Supporting this hypothesis, we found that the LF-fed animals had significantly higher plasma BH4 and lower BH2 levels compared to the WD-fed animals (BH4: P = 0.014; BH2 P = 0.039; BH2/BH4 P = 0.000), and also that gluten increased the BH2 levels and thereby the BH2/BH4 ratio significantly (BH2 P = 0.025; BH2/BH4 P = 0.046) (Fig. 7A–C). Malondialdehyde (MDA) is an end-product of lipid oxidation and has been detected in oxidized LDL33. Plasma MDA levels were significantly higher in WD-fed compared to LF-fed rats, indicating elevated oxidative stress (P = 0.002) (Fig. 7D). Gut microbiota composition correlated with MDA levels in both young and adult rats in this study. Thus, in the eight weeks old rats, genera Roseburia, Coprococcus, and two unclassified genera belonging to the Lachnospiraceae family, correlated positively with the MDA level [P = 0.007 (r = 0.572), P = 0.009 (r = 0.562), P = 0.010 (r = 0.559) and P = 0.040 (r = 0.512), respectively, Bonferroni corrected]. In the 20 weeks old rats, genus Dorea, Clostridium and an unknown genus belonging to the Erysipelotrichaceae family also correlated positively with the MDA level [P = 0.002 (r = 0.638), P = 0.010 (r = 0.587) and P = 0.009 (r = 0.593) respectively, Bonferroni corrected]. The increased oxidative stress did not correlate with plasma levels of IL-1α, CCL2, IL-4, IFNγ, and TNF-α, but these cytokine levels were below detection in most animals (Fig. S3).
Biomarkers of oxidative stress are increased in Western (WD) compared to low fat (LF) diet-fed Apoe−/− rats. (A,B) At 20 weeks of age tetrahydrobiopterin (BH4) levels in plasma showed significant difference between the groups due to fat content in the diets, whereas dihydrobiopterin (BH2) showed no group differences (n = 16 in the LF group, n = 14 in the LF + G group, n = 13 in the WD group and n = 8 in the WD + G group). (C) The BH2/BH4 ratio also showed a significant difference between the groups due to fat content in the diets. (D) At all three time points a difference in plasma levels of malondialdehyde (MDA) between the groups due to fat content in the diets could be demonstrated (data from all time points from n = 16 in the LF group, n = 14 in the LF + G group, n = 13 in the WD group and n = 8 in the WD + G group).
Discussion
In the present study, we show that female Apoe−/− rats exhibit early lesions of atherosclerosis in the aortic sinus, when fed a Western high-fat diet, as compared with a low fat diet. In relation to dietary gluten, the study design only allowed us to conclude that there was a minor impact on the oxidative stress marker BH2, and an early life change in gut microbiota composition, which is in agreement with previous data in Apoe−/− mice23. The abundance of Parabacteroides was decreased by gluten feeding at 8 weeks. This species is one of the most abundant genera of bacteria in the human gastrointestinal tract34, it is increased by prenatal exposure to metformin35, and it has been shown to be reduced in abundance in patients with irritable bowel syndrome or ulcerative colitis36. However, although the reduced abundance of Parabacteroides as a response to gluten feeding fits the expectation that gluten feeding might exert pro-inflammatory effects, it is not possible from our data to conclude whether gluten might have had effects on early or more advanced atherosclerosis.
The abundance of Erysipelotrichaceae was increased after 20 weeks of Western diet feeding in our study, and this has also been observed as a response to high fat feeding in mice37, while prenatal exposure to metformin has been shown to reduce Erysipelotrichaceae35. Different microbial taxa correlated with MDA at different ages, which emphasizes findings in previous studies showing age differences in the gut microbiota composition and also the presence of a window of opportunity for altering the gut microbiota18,38,39.
The most promising characteristics of WD-fed Apoe−/− rats seem to be the more human-like LDL dominated lipid profile, the increase in liver enzymes indicating a potential for developing steatohepatitis as a late complication, and obesity, which is linked to an increased oxidative stress level. The presence of markedly elevated LDL, VLDL and elevated triglycerides in female Apoe−/− rats is consistent with an earlier study in male rats30.
On the other hand, the manifestations of obesity, type 2 diabetes and atherosclerosis are modest in this new rat model, although it is possible that atherosclerosis severity would have been different in male rats. Compared to results from Apoe−/− mice, a weight gain of less than 15% in WD-fed rats compared to LF-fed rats is at the low end40,41. HbA1c remains unaffected, and at any time point the increased glucose intolerance is counteracted by an increased insulin response, so after 20 weeks of WD-feeding, these rats still could not be characterized as type 2 diabetic, but rather represented a pre-diabetic state. Also, cardiovascular changes were observed only as fatty streak lesions in the aortic sinus with both lower area and less severity, i.e. the mean percentage of the lesions of WD fed rats was about 2.5% of the total cross-sectional aortic sinus intimal area after 20 weeks, while in previous observations in Apoe−/− mice it was about 8%23. This indicates that a longer study may be required to induce larger and more advanced lesions in Apoe−/− rats. Therefore, the present study gives an unfavorable impression of the Apoe−/− rat as a model to investigate atherosclerosis, as compared with Apoe−/− mice. The Apoe−/− rat is on the Sprague-Dawley (SD) background. Inter-colony genetic differences occur within the same stock of rodents42,43,44,45, and also gut microbiota differences, which may account for phenotypical differences38, exist between populations46. This makes it impossible to select a proper wild-type control, if the exact background stock, as in this case, is unavailable from the same vendor facility. Schemmel et al. investigated dietary obesity in seven different rat strains and found that high-fat-fed SD rats gained only approximately 25% more body weight than grain-fed controls of the same age and sex, but the increase in body weight could partially be ascribed to an increase in body fat47. Levin et al. found that only around 50% of SD rats respond to high fat diet by becoming obese whereas the remainders are resistant to diet-induced obesity48. The group also demonstrated the ability to overcome this issue by selectively breeding the highest and lowest responders leading to two phenotypically different groups of animals by the F3 generation with only 2% and 14% overlap in weight in females and males, respectively48. So, not only is it generally problematic to use an outbred background for a genetically modified model, but in this case, the specific choice of SD as the background may also be unfavourable.
In conclusion, the Apoe−/− rat responds to 20 weeks of WD feeding by developing dyslipidaemia characterized by high levels of LDL and VLDL cholesterol in plasma, glucose intolerance, and oxidative stress. Atherosclerotic lesions are small, i.e. early fatty streaks limited to the aortic sinus, and in general, all disease symptoms are rather modest and, as in Apoe−/− mice, they are not alleviated by a gluten-free feeding. The Apoe mutation should be backcrossed to one or several inbred strains to elucidate whether a more disease-prone phenotype could be achieved, and to allow comparison with a proper wild-type control.
Methods
Animals and study design
Experiments were in accordance with the EU directive 2010/63/EU and the Danish Animal Experimentation Act (LBK 1306 from 23/11/2007 with 2011 amendments) and approved by the Animal Experiments Inspectorate, Ministry of Environment and Food, Denmark. Sixty female SD-Apoetm1sage (Apoe−/−) rats from Sigma Advanced Genetic Engineering (SAGE) Labs (Boyertown, USA) were randomly divided into four groups housed as 3–5 individuals per cage in a 12-hour light interval at room temperature (Novo Nordisk A/S, Denmark) (Fig. 1A). To avoid gliadin transfer with maternal milk, the experimental diets (all Research Diets Inc., New Brunswick, NJ, USA) were fed ad libitum to the mothers from approximately 10 days pre partum, and subsequently to the offspring until 20 weeks of age. Group LF was fed a gluten-free low fat control diet (Cat. #98121701, 10% energy from fat and no cholesterol); group LF + G was fed similarly (Cat. #98121704) with 3.5% added gluten (Cat. #G5004, Sigma, St. Louis, USA), i.e the concentration in ordinary barley-based rodent chow49; group WD was fed a gluten-free Western Diet (Cat. #D12079B, 41% energy from fat and 0.21% cholesterol); and group WF + G was fed similarly with added gluten (Cat. #D11061501–2) (Fig. 1B). LF diet was iso-calorimetric with the LF + G diet and the WD diet was iso-calorimetric with the WD + G diet. Prior to euthanasia by cardiac perfusion with a cooled physiological saline solution, animals were anaesthetized with a Hypnorm/Dormicum mixture (1:1:2 water solution, Vetapharm Ltd, Sherburn in Elmet, Leeds, UK; Roche A/S, Hvidovre, Denmark) subcutaneously. Animals were subjected to daily visual controls, and a veterinarian was consulted if signs of illness or misthriving appeared, leading to the euthanasia of nine animals. Two rats were euthanized in the LF + G group due to an eye injury or an extensive wound, while three rats in the WD group and four rats in the WD + G group died due to acute respiratory distress, i.e. there was a higher incidence of fatal acute respiratory distress in the WD-fed groups (P = 0.003), while this was not significantly influenced by gluten. However, it is questionable whether the mortality was due to the WD feeding, as all of these rats died very early in the study (see legend of Fig. 1).
Study parameters
Animals were weighed weekly. Oral glucose tolerance tests (OGTT) were performed at 8, 14 and 20 weeks of age using a glucose solution (2 g/kg, Fresenius Kabi, Copenhagen, Denmark; concentration 500 g/l) following four hours of fasting (Fig. 1A). Blood glucose levels were measured at t = 0, 30, 60, 90, 120 and 180 minutes by transferring 10 µl whole-blood to 500 µl Glucose/Lactate System Solution (EKF Diagnostic GmbH, Barleben, Germany) followed by analysis using a Biosen glucose auto analyser (Eppendorf, Hamburg, Germany) as per the manufacturer’s instructions. Insulin levels were measured during the OGTT’s at t = 0 and t = 30 minutes on EDTA-stabilized plasma samples using the Ultra-sensitive rat insulin ELISA kit (Crystal Chem, Downer’s Grove, USA) with the modification that in-house rat insulin standards, prepared using heat-treated rat plasma, were used.
Total plasma cholesterol (TPC) and cholesterol in lipoprotein fractions were measured on plasma samples obtained in relation to fasting and OGTT (Department of Pathology/Lipid Sciences, Wake Forrest University School of Medicine (Winston-Salem, NC, USA)). TPC levels were measured using colorimetric enzymatic assays as previously described50,51,52. For lipoprotein measurements, an aliquot of plasma containing approximately 20 µg of TPC was diluted in phosphate buffered saline (PBS) into a final volume of 400 µl. After centrifugation to remove protein precipitates, samples were injected onto a Superose 6 HR 10/30 (Amersham Pharmacia, NJ, USA) chromatography column subsequently run at 0.4 ml/min. The signal was integrated using Chrom Perfect Spirit Software (Justice Laboratory Software, NJ, USA). VLDL-, LDL-, and HDL-cholesterol were determined by multiplying the TPC concentration by the cholesterol percentage within the elution region for each lipoprotein class. For TPC and lipoprotein measurements plasma samples were pooled; two animals per pool and four pools corresponding to a total of eight animals per group were measured. Triglyceride (TG) and the liver enzymes ALT and AST were measured at 20 weeks of age, whereas total plasma cholesterol (TPC) levels and lipoprotein fractions were measured at 8, 14 and 20 weeks of age. TG, ALT and AST were measured on a Hitachi 912 analyser (Roche A/S Diagnostics, Mannheim, Germany) according to the manufacturer’s instructions.
Malondialdehyde (MDA) was assessed in plasma as an index of lipid oxidation by high-performance liquid chromatography (HPLC) as previously described53. Furthermore, tetrahydrobiopterin (BH4) providing reducing equivalents for eNOS54 as well as its oxidation products dihydrobiopterin (BH2) and biopterin were measured in plasma as indicators for vascular oxidative stress using HPLC with fluorescence detection employing iodine oxidation as described by Fukushima and Nixon55. To prevent ex vivo oxidation when measuring biopterins, 1 ml of whole blood was drawn with a K3-EDTA flushed syringe and immediately added to a microcentrifuge tube containing 25 µl of 2.5% (w/v) dithioerythriol (DTE) in Milli-Q (18.2 MΩ) water56. The sample was then centrifuged (4 °C; 15000 × g; 1 minute) to obtain plasma, which was immediately stored at −80 °C.
After trans-cardiac perfusion with 5 × 10 ml 0.9% NaCl, the heart and 1–2 mm of the aortic sinus were fixed in 10% buffered formalin for 24 hours and transferred to 20% sucrose in phosphate buffered saline (PBS) for 24 hours. The hearts were trimmed by removing the lower part of the heart with the cutting axis perpendicular to the aortic root. The trimmed hearts were placed in a cryo mold with the cut surface at the bottom of the mold, cryo fixated in OCT compound (Tissue-Tek, Sakura Fineteck, Værløse, Denmark) and frozen on dry ice. Sections were cut and discharged until the appearance of the aortic valve. Hereafter, all sections were cut at 10 µm thickness and were collected and placed on numbered glass slides (SuperFrost Plus, Hounisen, Denmark), two on each glass slide with a distance of 150 µm between the sections. Sections were stained with oil red O (Sigma-Aldrich, MO, USA) and scanned on a digital slide scanner (Hamamatsu NanoZoomer 2.0 HT, Japan) at a magnification of 20X. The oil red O stained area was measured using an image analysis tool (Visio-morph) and expressed as the mean of the oil red O-positive area out of total cross-sectional aortic sinus intimal area.
Gut microbiota profiles were determined using tag-encoded 16S rRNA gene MiSeq (Illumina, CA, USA) sequencing. Cellular DNA was extracted using a QIAamp DNA Stool Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions, but with the addition of an initial bead-beating step (FastPrep, MP Biomedicals, CA, USA) to increase cell lysis. Extracted DNA was stored at −40° until analysis. Amplicons (~460 bp) including the V3 and V4 regions of the 16S rRNA gene were amplified using primers containing overhangs compatible with Nextera Index Kit (Illumina, CA, USA), tagged and sequenced (250 bp paired end) as previously described57.
Statistical analyses
Data were analysed by two-tailed, two-way ANOVA in either GraphPad Prism version 6 or 7 (GraphPad Software, CA, USA; OGTT (area under the curve (AUC)), weight curves (AUC), insulin levels, blood lipid levels, lipoprotein fractions, histological data, HbA1c, plasma cytokines and gene expression levels in the aortic arch) or Minitab 17 (Minitab Ltd, Coventry, UK; oxidative stress markers (MDA and biopterins). Mortality rates were compared by Fisher’s Exact Test in GraphPad Prism version 7. Data were considered significant when having a p-value of less than 0.05 and P = 0.000 was set as lowest reported p-value. Values below detectable limits (insulin and cytokine measurement kits) were given the value of half the lover limit of quantification (½LLOQ).
Gut microbiota data were analysed as previously described57. In brief, the raw dataset was trimmed using CLC Genomic Workbench (CLC bio, Aarhus, Denmark) with merging of overlapped reads and using the Quantitative Insight Into Microbial Ecology (QIIME) tool (version. 1.7.0; Open source software)58. USEARCH was used to purge the dataset from chimeric reads, while the usearch61 method was used for Operational Taxonomic Units (OTU) selection59 with the Greengenes (version 12.10) 16S rRNA gene database as reference60. Principal Coordinate Analysis (PCoA) plots were generated with the Jackknifed Beta Diversity workflow based on 10 distance metrics calculated using 10 subsampled Operational Taxonomic Units (OTU) tables taking 85% of the sequence number within the most indigent sample for each jackknifed subset. Analysis of similarities (ANOSIM) was used to evaluate group differences using weighted and unweighted uniFrac distance metrics generated based on rarefied (25,000 reads per sample) OTU tables. Alpha diversity measure expressed with an observed species (sequence similarity 97% OTUs) value was computed for rarefied OTU tables (25,000 reads per sample) using alpha rarefaction workflow. Testing on the differences in alpha diversity was conducted using t-test employing the non-parametric (Monte Carlo) method (999 permutations) implemented in the compare alpha diversity workflow. Determining differences in taxa composition between categories was performed using otu_category_significance.py script implemented in QIIME 1.7.0. The G test of independence (G-test) and ANOVA determining respectively: qualitative (presence/absence) and quantitative (relative abundance) association of OTUs in the given category were calculated based on 999 subsampled OTU-tables rarefied to an equal number of reads (25,000 per sample).
Correlations between the metabolic parameters, biomarkers of oxidative stress, atherosclerosis quantification measures, and taxa relative abundance summarized to the genus level in samples verified at 8 or 20 weeks of age were tested with the Pearson’s product-moment correlation coefficient implemented in the otu_category_significance.py script (QIIME 1.7.0). Testing was performed based on 999 rarefied OTU tables unified to an equal number of reads per sample (25,000).
Equipment and settings for figures
All statistical graphs (Figs 4–7) were computed by GraphPad Prism versions 6 and 7, while microbiota graphs (Figs 2 and 3) were computed by QUIIME. All Figures, except for Fig. 3, which was based upon a PDF file, were plotted into Power Point (Microsoft, WA, USA), fitted in size and appearance. Subsequently all figures were saved as TIFF images. The images in Fig. 4 have not been edited in other forms of software.
Data availability
Data and associated protocols are stored on the University of Copenhagen’s backup servers, and can be made available for other non-commercial research groups along with any remaining study materials, as far as still in stock, by communication with the corresponding author.
References
Meir, K. S. & Leitersdorf, E. Atherosclerosis in the apolipoprotein-E-deficient mouse: a decade of progress. Arterioscler Thromb Vasc Biol 24, 1006–1014, https://doi.org/10.1161/01.ATV.0000128849.12617.f4 (2004).
Zaragoza, C. et al. Animal models of cardiovascular diseases. J Biomed Biotechnol 2011, 497841, https://doi.org/10.1155/2011/497841 (2011).
Zechner, R., Moser, R., Newman, T. C., Fried, S. K. & Breslow, J. L. Apolipoprotein E gene expression in mouse 3T3-L1 adipocytes and human adipose tissue and its regulation by differentiation and lipid content. J Biol Chem 266, 10583–10588 (1991).
Heeren, J., Beisiegel, U. & Grewal, T. Apolipoprotein E recycling: implications for dyslipidemia and atherosclerosis. Arterioscler Thromb Vasc Biol 26, 442–448, https://doi.org/10.1161/01.ATV.0000201282.64751.47 (2006).
Dallinga-Thie, G. M. et al. The metabolism of triglyceride-rich lipoproteins revisited: new players, new insight. Atherosclerosis 211, 1–8, https://doi.org/10.1016/j.atherosclerosis.2009.12.027 (2010).
Hussain, M. M. et al. Clearance of chylomicron remnants by the low density lipoprotein receptor-related protein/alpha 2-macroglobulin receptor. J Biol Chem 266, 13936–13940 (1991).
Chappell, D. A. & Medh, J. D. Receptor-mediated mechanisms of lipoprotein remnant catabolism. Prog Lipid Res 37, 393–422 (1998).
Cooper, A. D. Hepatic uptake of chylomicron remnants. J Lipid Res 38, 2173–2192 (1997).
Ishibashi, S. et al. Hypercholesterolemia in low density lipoprotein receptor knockout mice and its reversal by adenovirus-mediated gene delivery. J Clin Invest 92, 883–893, https://doi.org/10.1172/JCI116663 (1993).
Zhang, S. H., Reddick, R. L., Burkey, B. & Maeda, N. Diet-induced atherosclerosis in mice heterozygous and homozygous for apolipoprotein E gene disruption. J Clin Invest 94, 937–945, https://doi.org/10.1172/JCI117460 (1994).
Vinje, S., Stroes, E., Nieuwdorp, M. & Hazen, S. L. The gut microbiome as novel cardio-metabolic target: the time has come! Eur Heart J 35, 883–887, https://doi.org/10.1093/eurheartj/eht467 (2014).
Wang, Z. et al. Non-lethal Inhibition of Gut Microbial Trimethylamine Production for the Treatment of Atherosclerosis. Cell 163, 1585–1595, https://doi.org/10.1016/j.cell.2015.11.055 (2015).
Luck, H. et al. Regulation of obesity-related insulin resistance with gut anti-inflammatory agents. Cell Metab 21, 527–542, https://doi.org/10.1016/j.cmet.2015.03.001 (2015).
Ley, R. E. et al. Obesity alters gut microbial ecology. Proc Natl Acad Sci USA 102, 11070–11075, https://doi.org/10.1073/pnas.0504978102 (2005).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031, https://doi.org/10.1038/nature05414 (2006).
Cani, P. D. et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 57, 1470–1481, https://doi.org/10.2337/db07-1403 (2008).
Arumugam, M. et al. Enterotypes of the human gut microbiome. Nature 473, 174–180, https://doi.org/10.1038/nature09944 (2011).
Rune, I. et al. Ampicillin-improved glucose tolerance in diet-induced obese C57BL/6NTac mice is age dependent. J Diabetes Res 2013, 319321, https://doi.org/10.1155/2013/319321 (2013).
Hansen, C. H. F. et al. A maternal gluten-free diet reduces inflammation and diabetes incidence in the offspring of NOD mice. Diabetes 63, 2821–2832 (2014).
Hansen, A. K. et al. Diabetes preventive gluten-free diet decreases the number of caecal bacteria in non-obese diabetic mice. Diabetes-Metabolism Research & Reviews 22, 220–225 (2006).
Zhang, L. et al. Effects of Gliadin consumption on the Intestinal Microbiota and Metabolic Homeostasis in Mice Fed a High-fat Diet. Scientific Reports 7, 44613, doi:https://doi.org/10.1038/srep44613, http://www.nature.com/articles/srep44613#supplementary-information (2017).
Haupt-Jorgensen, M., Buschard, K., Hansen, A. K., Josefsen, K. & Antvorskov, J. C. Gluten-free diet increases beta-cell volume and improves glucose tolerance in an animal model of type 2 diabetes. Diabetes-Metabolism Research and Reviews 32, 675–684, https://doi.org/10.1002/dmrr.2802 (2016).
Rune, I. et al. Modulating the Gut Microbiota Improves Glucose Tolerance, Lipoprotein Profile and Atherosclerotic Plaque Development in ApoE-Deficient Mice. PLoS One 11, e0146439, https://doi.org/10.1371/journal.pone.0146439 (2016).
Fernandez, M. L. & Volek, J. S. Guinea pigs: a suitable animal model to study lipoprotein metabolism, atherosclerosis and inflammation. Nutr Metab (Lond) 3, 17, https://doi.org/10.1186/1743-7075-3-17 (2006).
Kris-Etherton, P. M., Ho, C. Y. & Fosmire, M. A. The effect of dietary fat saturation on plasma and hepatic lipoproteins in the rat. J Nutr 114, 1675–1682 (1984).
Pharma, M. C. Sigma Life Science broadens gene knockout rat models, https://www.manufacturingchemist.com/news/article_page/Sigma_Life_Science_broadens_gene_knockout_rat_models/70939 (2011).
Sung, Y. H. et al. Knockout mice created by TALEN-mediated gene targeting. Nat Biotech 31, 23-24, http://www.nature.com/nbt/journal/v31/n1/abs/nbt.2477.html#supplementary-information (2013).
Wang, H. Y. et al. One-Step Generation of Mice Carrying Mutations in Multiple Genes by CRISPR/Cas-Mediated Genome Engineering. Cell 153, 910–918, https://doi.org/10.1016/j.cell.2013.04.025 (2013).
Ekuni, D. et al. Occlusal disharmony accelerates the initiation of atherosclerosis in apoE knockout rats. Lipids in Health and Disease 13, 144, https://doi.org/10.1186/1476-511x-13-144 (2014).
Wei, S. et al. Apolipoprotein E-deficient rats develop atherosclerotic plaques in partially ligated carotid arteries. Atherosclerosis 243, 589–592, https://doi.org/10.1016/j.atherosclerosis.2015.10.093 (2015).
Tangirala, R. K., Rubin, E. M. & Palinski, W. Quantitation of atherosclerosis in murine models: correlation between lesions in the aortic origin and in the entire aorta, and differences in the extent of lesions between sexes in LDL receptor-deficient and apolipoprotein E-deficient mice. J Lipid Res 36, 2320–2328 (1995).
Nakashima, Y., Plump, A. S., Raines, E. W., Breslow, J. L. & Ross, R. ApoE-deficient mice develop lesions of all phases of atherosclerosis throughout the arterial tree. Arterioscler Thromb 14, 133–140 (1994).
Palinski, W. et al. ApoE-deficient mice are a model of lipoprotein oxidation in atherogenesis. Demonstration of oxidation-specific epitopes in lesions and high titers of autoantibodies to malondialdehyde-lysine in serum. Arterioscler Thromb 14, 605–616 (1994).
Hakansson, A. & Molin, G. G. Microbiota and Inflammation. Nutrients 3, 637–682, https://doi.org/10.3390/nu3060637 (2011).
Salomaki-Myftari, H. et al. Neuropeptide Y Overexpressing Female and Male Mice Show Divergent Metabolic but Not Gut Microbial Responses to Prenatal Metformin Exposure. Plos One 11, 19, https://doi.org/10.1371/journal.pone.0163805 (2016).
Noor, S. O. et al. Ulcerative colitis and irritable bowel patients exhibit distinct abnormalities of the gut microbiota. BMC Gastroenterology 10, 134, https://doi.org/10.1186/1471-230x-10-134 (2010).
Fleissner, C. K. et al. Absence of intestinal microbiota does not protect mice from diet-induced obesity. British Journal of Nutrition 104, 919–929, https://doi.org/10.1017/s0007114510001303 (2010).
Ellekilde, M. et al. Characterization of the gut microbiota in leptin deficient obese mice - Correlation to inflammatory and diabetic parameters. Res Vet Sci 96, 241–250, https://doi.org/10.1016/j.rvsc.2014.01.007 (2014).
Hansen, C. H. et al. Early life treatment with vancomycin propagates Akkermansia muciniphila and reduces diabetes incidence in the NOD mouse. Diabetologia 55, 2285–2294, https://doi.org/10.1007/s00125-012-2564-7 (2012).
Guo, J., Jou, W., Gavrilova, O. & Hall, K. D. Persistent Diet-Induced Obesity in Male C57BL/6 Mice Resulting from Temporary Obesigenic Diets. Plos One 4 (2009).
Toye, A. A. et al. A genetic and physiological study of impaired glucose homeostasis control in C57BL/6J mice. Diabetologia 48, 675–686 (2005).
Aldinger, K. A., Sokoloff, G., Rosenberg, D. M., Palmer, A. A. & Millen, K. J. Genetic Variation and Population Substructure in Outbred CD-1 Mice: Implications for Genome-Wide Association Studies. Plos One 4, 10, https://doi.org/10.1371/journal.pone.0004729 (2009).
Inoue, T., Coitinho, H., Pochintesta, E., Baltar, J. & Goto, N. Genetic relationships between CD-1 stocks of mice in Uruguay. Jarq - Jpn. Agric. Res. Q. 34, 265–270 (2000).
Nohynek, G. J., Longeart, L., Geffray, B., Provost, J. P. & Lodola, A. Fat, Frail And Dying Young - Survival, Body-Weight And Pathology Of The Charles River Sprague-Dawley-Derived Rat Prior To And Since The Introduction Of The Vafr Variant In 1988. Hum. Exp. Toxicol. 12, 87–98 (1993).
Wells, T., Windle, R. J., Peysner, K. & Forsling, M. L. Intercolony Variation In Fluid Balance And Its Relationship To Vasopressin Secretion In Male Sprague-Dawley Rats. Laboratory Animals 27, 40–46, https://doi.org/10.1258/002367793781082395 (1993).
Hufeldt, M. R., Nielsen, D. S., Vogensen, F. K., Midtvedt, T. & Hansen, A. K. Variation in the Gut Microbiota of Laboratory Mice Is Related to Both Genetic and Environmental Factors. Comparative Medicine 60, 336–342 (2010).
Schemmel, R., Mickelsen, O. & Gill, J. L. Dietary obesity in rats: Body weight and body fat accretion in seven strains of rats. J Nutr 100, 1041–1048 (1970).
Levin, B. E., Dunn-Meynell, A. A., Balkan, B. & Keesey, R. E. Selective breeding for diet-induced obesity and resistance in Sprague-Dawley rats. Am J Physiol 273, R725–730 (1997).
Funda, D. P., Kaas, A., Tlaskalova-Hogenova, H. & Buschard, K. Gluten-free but also gluten-enriched (gluten+) diet prevent diabetes in NOD mice; the gluten enigma in type 1 diabetes. Diabetes Metab Res Rev 24, 59–63, https://doi.org/10.1002/dmrr.748 (2008).
Lee, R. G. et al. Plasma cholesteryl esters provided by lecithin:cholesterol acyltransferase and acyl-coenzyme a:cholesterol acyltransferase 2 have opposite atherosclerotic potential. Circ Res 95, 998–1004, https://doi.org/10.1161/01.RES.0000147558.15554.67 (2004).
Brown, J. M. et al. Targeted depletion of hepatic ACAT2-driven cholesterol esterification reveals a non-biliary route for fecal neutral sterol loss. J Biol Chem 283, 10522–10534, https://doi.org/10.1074/jbc.M707659200 (2008).
Temel, R. E. et al. Intestinal cholesterol absorption is substantially reduced in mice deficient in both ABCA1 and ACAT2. J Lipid Res 46, 2423–2431, https://doi.org/10.1194/jlr.M500232-JLR200 (2005).
Lykkesfeldt, J. Determination of malondialdehyde as dithiobarbituric acid adduct in biological samples by HPLC with fluorescence detection: comparison with ultraviolet-visible spectrophotometry. Clin Chem 47, 1725–1727 (2001).
Mortensen, A. & Lykkesfeldt, J. Does vitamin C enhance nitric oxide bioavailability in a tetrahydrobiopterin-dependent manner? In vitro, in vivo and clinical studies. Nitric Oxide 36, 51–57, https://doi.org/10.1016/j.niox.2013.12.001 (2014).
Fukushima, T. & Nixon, J. C. Analysis of reduced forms of biopterin in biological tissues and fluids. Anal Biochem 102, 176–188 (1980).
Mortensen, A. & Lykkesfeldt, J. Kinetics of acid-induced degradation of tetra- and dihydrobiopterin in relation to their relevance as biomarkers of endothelial function. Biomarkers 18, 55–62, https://doi.org/10.3109/1354750X.2012.730552 (2013).
Pyndt Jorgensen, B. et al. A possible link between food and mood: dietary impact on gut microbiota and behavior in BALB/c mice. PLoS One 9, e103398, https://doi.org/10.1371/journal.pone.0103398 (2014).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336, https://doi.org/10.1038/nmeth.f.303 (2010).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200, https://doi.org/10.1093/bioinformatics/btr381 (2011).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6, 610–618, https://doi.org/10.1038/ismej.2011.139 (2012).
Acknowledgements
Christian Larsen, Lene Winther Takla, Line Rasmussen, Nicolai Hansen, Jane Larsen, Bettina Brandrup, Trine Pagh Ludvigsen, Cecilie Löe Licht, Joan Frandsen, Belinda Bringtoft and Annie Bjergby Kristensen are kindly thanked for their technical assistance. This study was carried out as a part of the 3G research program (Guts, grains and greens, (http://www.3g-center.dk/). The 3G program is supported by the Danish Ministry of Science, Technology and Innovation. Ida Rune and Jens Lykkesfeldt are supported by The LIFEPHARM Centre for In Vivo Pharmacology (www.lifepharm.dk). Research reported in this publication was also supported in part by the National Heart, Lung, and Blood Institute and the National Institute of Diabetes and Digestive and Kidney Diseases of the National Institutes of Health under award number R01HL062887, P01HL092969, R01HL126028 and P30DK017047 to Karin E. Bornfeldt.
Author information
Authors and Affiliations
Contributions
I.R., A.K.H., B.R., J.L. and B.C. designed the study. I.R., J.K., Ł.K., P.K., K.J. and A.M. conducted analysis. J.L., K.B., D.S.N., J.J.F. and R.K.K. provided special knowledge, equipment for specific analysis and assistance related to its use. I.R., K.B., J.K. and A.K.H wrote the manuscript and I.R., L.K., A.K.H. and J.K. prepared the figures. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rune, I., Rolin, B., Lykkesfeldt, J. et al. Long-term Western diet fed apolipoprotein E-deficient rats exhibit only modest early atherosclerotic characteristics. Sci Rep 8, 5416 (2018). https://doi.org/10.1038/s41598-018-23835-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-23835-z
This article is cited by
-
Mild hypercholesterolemia impacts achilles sub-tendon mechanical properties in young rats
BMC Musculoskeletal Disorders (2023)
-
Monobutyrin and monovalerin improve gut–blood–brain biomarkers and alter gut microbiota composition in high-fat fed apolipoprotein-E-knockout rats
Scientific Reports (2022)
-
The microbiome and rodent models of immune mediated diseases
Mammalian Genome (2021)
-
Endothelial dysfunction in small arteries and early signs of atherosclerosis in ApoE knockout rats
Scientific Reports (2020)
-
Apolipoprotein E deficient rats generated via zinc-finger nucleases exhibit pronounced in-stent restenosis
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.