Abstract
Sample pooling enabled by dedicated indexes is a common strategy for cost-effective and robust high-throughput sequencing. Index misassignment leading to mutual contamination between pooled samples has however been described as a general problem of the latest Illumina sequencing instruments utilizing exclusion amplification. Using real-life data from multiple tumour sequencing projects, we demonstrate that index misassignment can induce artefactual variant calls closely resembling true, high-quality somatic variants. These artefactual calls potentially impact cancer applications utilizing low allelic frequencies, such as in clonal analysis of tumours. We discuss the available countermeasures with an emphasis on improved library indexing methods, and provide software that can assist in the identification of variants that may be consequences of index misassignment.
Similar content being viewed by others
Introduction
Identification of somatic variants by next-generation sequencing has become an important technique in cancer research by pinpointing the genomic causes of tumour phenotypes. An increasing number of examples have further shown that genomic aberrations have prognostic value and can inform rational clinical deployment of targeted cancer drugs1,2. As of now, next-generation sequencing technologies enable affordable assaying of variation in the entire tumour genome within days, but despite the availability of specialized software tools, somatic variant calling continues to pose challenges. Notably, the inherent complexity of tumours, as exemplified by aneuploidy, tumour heterogeneity, and sample impurity, often leads to important somatic variants only being detectable in low allelic fractions (AFs)3,4,5,6,7. AFs can be further affected due to technical reasons, such as the inability to accurately represent allelic ratios at genomic loci with low coverage8. As a consequence of the necessarily high required sensitivity, somatic variant calling is susceptible to random noise, systematic artefacts, and sample contamination9,10. When unnoticed, false positive variant calls can contribute to high costs incurred by follow-up analyses and experiments, and in the worst case support inadequate therapeutic strategies.
As previously described in the context of RNA-seq applications11, when material from multiple samples is being pooled together before sequencing, the sequencing technology itself can be a source of noticeable sample cross-contamination. Sample pooling (also called sample multiplexing) is a standard means of dividing the throughput of a sequencing instrument among multiple samples, relying on index sequences that uniquely barcode the material of each involved sample. Sample index misassignment, a phenomenon most evident on Illumina instruments utilizing exclusion amplification chemistry (ExAmp) and patterned flow cell technology (i.e., HiSeq 3000/4000/X Ten and NovaSeq), effectively leads to transfer of individual sequencing reads between samples included into a common pool11.
Our analysis of high-coverage tumour sequencing data shows that index misassignment is a source of false positive somatic variant calls in a form of true variation obtained from co-multiplexed samples.
Results
We investigated tumour-normal sample pairs of three different tumour types: diffuse large B-cell lymphoma, follicular lymphoma, and sarcoma. All samples were collected from cancer patients in Norway, and deep exome sequencing (median coverages: 315X for tumour samples, 146X for controls) was carried out on local Illumina instruments: three HiSeq 2000/2500 instruments utilizing bridge amplification (generating 42 diffuse large B-cell lymphoma samples and 31 sarcoma samples) and a HiSeq 4000 employing ExAmp (generating 81 follicular lymphoma samples and 77 sarcoma samples). All samples were subject to standardized library preparation, sequencing, and bioinformatics analysis, as adopted by the Norwegian Cancer Genomics Consortium (NCGC, http://cancergenomics.no) (Methods, Supplementary Information).
In our assessment of index misassignment, we have examined the consequences for calling of somatic single nucleotide variants (SSNVs). For increased reliability, all tests and analyses were limited to SSNVs agreed upon by two independent variant callers (MuTect and Strelka)3,12. Sample-wise contamination estimates, our primary measure of contamination, were generated by Conpair13, based on the allelic composition on several thousand genomic marker sites in a given matched sample pair. Our tests on simulated data suggest that Conpair provides consistent quantifications, despite apparent progressive underestimation dependent on the number of contributing contamination sources (Supplementary Tables 1 and 2, Supplementary Information). We attribute this underestimation to decreasing contaminant variant AFs in sample pools of increasing size (Supplementary Fig. 1) paired with the fact that Conpair’s contamination model is intended for two-sample mixtures only.
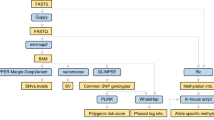
A schematic overview of all conducted experiments and their data dependencies is available on Fig. 1.
Contamination rates
During the analysis of hundreds of sequenced tumour-normal matched pairs, we noted that data generated on the ExAmp instrument showed significantly higher sample-wise contamination levels in comparison to data from bridge amplification instruments (median per-sample contamination estimates: 0.839% vs. 0.187%, p-value < 0.001, Fig. 2a, Supplementary Fig. 2). To specifically assess the role of multiplexing on the ExAmp instrument, we sequenced 16 selected sample libraries both in pools and individually (i.e., one sample library per flow cell lane), observing significantly increased contamination rates in sequencing output from the multiplexed libraries (median per-sample contamination estimates: 0.644% vs. 0.0465%, p-value < 0.001, Supplementary Table 3). Removal of free adapters/primers prior to sequencing was in Illumina’s report identified as a key measure for mitigating the cross-contamination rates (https://www.illumina.com/science/education/minimizing-index-hopping.html), but for the 16 libraries included in our testing, performing a gel-based library purification step in combination with bead purification did not provide improvements over bead purification alone (median per-sample contamination estimates: 0.671% vs. 0.6115%, p-value = 0.159). In accordance with previous reports by Illumina and Sinha et al., we concluded that ExAmp chemistry is the cause of sample contamination. The dependency on sample pooling indicated that co-multiplexed samples serve as the contaminants.
Amplification chemistry and its relationships to (a) sample contamination estimates and (b) variant counts and PC-AF values. All values are plotted separately for each combination of tumour type and amplification chemistry represented in the analysis. The colours distinguish between variants called in “high-contamination” samples (Conpair contamination estimate > = 0.5%) and variants coming from samples with “low contamination” (Conpair contamination estimate < 0.5%). BrAmp: bridge amplification; FL: follicular lymphoma; SARC: sarcoma; DLBCL: diffuse large B-cell lymphoma; SCV: suspected contaminant variant.
Artefactual variant calls
In order to assess the impact of ExAmp-associated index misassignment on the detection of SSNVs, we first set out to identify SSNVs that were likely to originate from contamination. For each somatic variant found in a given tumour sample, we quantified the variant support in the expected source of contamination - the corresponding “pool complement” consisting of reads from all co-multiplexed samples from other individuals. We thus calculated two allelic fractions for each somatic variant: the standard sample AF and an AF derived from the pool complement (PC-AF) (Supplementary Fig. 3, Supplementary Information). Two distinct classes of variants can be identified when variant AFs and PC-AFs are plotted against each other: (1) apparently true somatic variants, consisting of variants not present in the Norwegian population (i.e., absent from NCGC’s cohort of normal samples) and lacking support in their respective pool complements (PC-AF < 0.01); and (2) suspected contaminant variants, consisting of common Norwegian germline variants (> = 5% allele frequency in NCGC’s cohort of normal samples) with a considerable support in their respective pool complements (PC-AF > = 0.2) (Supplementary Fig. 4). We classify the remaining variants as ambiguous.
In comparison to bridge amplification, ExAmp leads to higher occurrence of SSNVs that coincide with germline variation common in the Norwegian population (Fig. 2b). At the same time, PC-AFs of these variants are significantly higher in the ExAmp datasets (median values: 0.508 vs. 0.125, p-value < 0.001), showing a much better correspondence to the suspected contamination source. Samples sequenced on the ExAmp instrument have significantly higher counts of suspected contaminant variants than samples sequenced on bridge amplification instruments (per-sample median counts: 4 vs. 0, p-value < 0.001) and show significant correspondence between the number of suspected contaminant variants and the estimated contamination (p-values < 0.001, Fig. 3).
Per-sample counts of apparently true somatic variants (a) and suspected contaminant variants (b) plotted against contamination estimates. All values, including Spearman’s rank correlation coefficients and their associated p-values, are plotted separately for each combination of tumour type and amplification chemistry represented in the analysis. The colours distinguish between “high-contamination” samples (Conpair contamination estimate > = 0.5%) and samples with “low contamination” (Conpair contamination estimate < 0.5%). BrAmp: bridge amplification; FL: follicular lymphoma; SARC: sarcoma; DLBCL: diffuse large B-cell lymphoma.
Discussion
We have focused on exploring the general link between tumour sample cross-contamination and suspected contaminant variants of germline origin, but the effects of index misassignment require further consideration (Table 1). Firstly, cross-sample contamination by recurrent somatic variants would represent a class of potential false positives that would be more difficult to recognize, and may have stronger clinical implications. Concern for such cases of contamination gains relevance when several samples containing identical somatic variants are being co-multiplexed and thereby jointly contribute to elevated pool complement support (e.g., pooling serial patient biopsies or screening multiple samples likely to harbour identical hotspot mutations, particularly in sensitive analyses of high-purity samples). Secondly, when co-multiplexing tumour and control samples, false negative variant rates might be increased due to somatic variation contaminating the controls. Lastly, contamination rates within a given sample can be influenced by a combination of independent factors, such as the fraction of the pool that is constituted by the pool complement14 or a sample’s copy number profile - in copy number loss regions of high-purity tumour samples, we expect both the average AF and the number of contaminant variants to be increased due to the local underrepresentation of non-contaminant reads.
Several countermeasures have been suggested for the prevention of index misassignment, with sample storage conditions being shown by Illumina to influence the problem’s severity. Sequencing of a single sample per lane proved to be an effective solution, which however might be too costly for many applications. Rigorous gel- or bead-based library purification has been strongly recommended as a means of index misassignment mitigation in case of sample multiplexing. However, in our experience, both purification methods appear to be insufficient even in combination. Dual indexing has been suggested for circumventing the problem by using sample-specific index pairs rather than individual indexes, enabling recognition of reads with unexpected index combinations caused by index misassignment14. Dual indexes are becoming a part of best practices for multiplexed libraries sequenced with ExAmp chemistry, but their current availability may vary depending on the applied library preparation protocol. The possible solutions therefore need to be assessed in the context of each particular project.
For data that have already been generated, suitable variant post-processing should be chosen based on the type of contamination artefacts relevant for given project (Table 1). Commonly used allelic fraction thresholds and germline variant database filters are likely to remove the majority of false positive somatic variant calls caused by index misassignment. However, a more discriminative filtering approach would be preferable in settings where sensitive detection is the priority, such as in clonal analysis. Low-AF variation can, in multiple cancer types, harbour markers of prognostic and therapeutic value15,16,17. When available, pool complement information can help identifying variants that are unlikely to be contamination artefacts, thereby reducing the number of potentially important true positive somatic variants discarded due to low AF or germline database presence. The code used for PC-AF calculation in our analyses is available for use in other projects (Methods). If control samples have been contaminated by true somatic variation, it might be necessary to adopt variant caller settings more permissive to somatic allele evidence in the matched normal material.
We believe our findings to be of relevance to other cancer sequencing projects that utilize the ExAmp chemistry, even though the impacts of index misassignment on somatic variant calling may vary depending on the combination of employed sequencing instruments, library preparation protocols, and bioinformatics analyses. In general, we expect the effects of index misassignment to dampen as the sequencing depth decreases and the allelic fraction threshold for accepted somatic variants increases. On the other hand, we note that besides single nucleotide variants, other types of somatic variation (e.g. insertions and deletions) are likely to be affected.
Methods
Methods used for sample collection, DNA extraction, library preparation and sequencing, as well as full details of all bioinformatics analyses, are available in the methods section of Supplementary Information.
Bioinformatics analyses
All sequenced samples were pre-processed with BWA MEM18, Picard (http://broadinstitute.github.io/picard/) and GATK19 tools, before performing somatic calling with MuTect and Strelka (only consensus calls were considered in subsequent processing and testing). All analyses were performed using human reference genome build b37 with an added decoy contig and corresponding variation databases.
The pool complement of each individual tumour sample was formed by reads of all co-multiplexed samples originating from other individuals. For each identified somatic variant in a given tumour sample, all overlapping reads from the corresponding pool complement were extracted, and the pool complement allelic fraction (PC-AF, the fraction of reads supporting the variant within the pool complement) was calculated. Tumour samples without a matched control and/or tumour samples failing our quality metrics thresholds (Supplementary Information) were excluded from the analyses of potential contamination targets, but did serve as potential contamination sources in our PC-AF calculations.
For evaluating the role of multiplexing in ExAmp-associated index misassignment, tumour-normal sample pairs from 8 selected individuals were sequenced (i) individually (one sample per dedicated flow cell lane), (ii) in a pool of all 8 tumour/normal samples and (iii) in a pool of all 8 tumour/normal samples after applied gel purification. For generating contamination estimates with Conpair, the separately sequenced normal sample of each individual was used as a control for the five other samples of given individual.
Accuracy of Conpair contamination estimates was assessed with the help of artificially created exome-wide sample admixtures. Reads from multiple libraries were mixed to simulate contamination by 1, 2, 4 or 7 samples from as many different individuals, with the total contamination amounting to either ~2% or ~8% in each case, depending on the experiment. Conpair’s contamination estimates were compared to true contamination content, which was validated by library-specific depth of coverage calculations.
NCGC’s cohort of normal samples consisted of blood samples of 789 different individuals living in Norway. The samples were processed according to “Best Practices for Germline SNP & Indel Discovery in Whole Genome and Exome Sequence” developed by the Broad Institute, utilizing allele-specific (rather than site-specific) variant calling.
Statistics
All statistical tests were performed in R (http:/www.r-project.org) as two-tailed non-parametric tests (either Wilcoxon rank sum test with continuity correction or Wilcoxon signed rank test) of the identity of two populations. All input test data are available as Supplementary Data. R session information, utilized R commands together with their outputs, as well as median and interquartile range values of all the tested distributions are available in the Supplementary Note. All distributions have been plotted as Supplementary Figures.
Ethics approval and consent
All patients provided informed consent. The study was approved by the Regional Committee for Medical and Health Research Ethics of South-East Norway (reference numbers 2014/127, S-06133 and 2010/1244). All experiments were performed in accordance with relevant guidelines and regulations.
Data availability
All data that support the findings of this study and that do not compromise research participant privacy are available as Supplementary Data. All data that may compromise research participant privacy (e.g., raw sequence data) are available for inspection upon request to the corresponding author [EH] on non-disclosure terms.
Code availability
Python code used for calculating PC-AF values is available at GitHub (https://github.com/danielvo/IBPC). R code for generating Figures 2 and 3 is included in the Supplementary Note.
References
Garraway, L. A. Genomics-Driven Oncology: Framework for an Emerging Paradigm. J. Clin. Oncol. 31, 1806–1814 (2013).
Griffith, M. et al. CIViC is a community knowledgebase for expert crowdsourcing the clinical interpretation of variants in cancer. Nat. Genet. 49, 170–174 (2017).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Ryu, D., Joung, J.-G., Kim, N. K. D., Kim, K.-T. & Park, W.-Y. Deciphering intratumor heterogeneity using cancer genome analysis. Hum. Genet. 135, 635–642 (2016).
Lee, J.-K., Choi, Y.-L., Kwon, M. & Park, P. J. Mechanisms and Consequences of Cancer Genome Instability: Lessons from Genome Sequencing Studies. Annu. Rev. Pathol. 11, 283–312 (2016).
Cheng, D. T. et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): A Hybridization Capture-Based Next-Generation Sequencing Clinical Assay for Solid Tumor Molecular Oncology. J. Mol. Diagn. 17, 251–264 (2015).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Alioto, T. S. et al. A comprehensive assessment of somatic mutation detection in cancer using whole-genome sequencing. Nat. Commun. 6, 10001 (2015).
Chen, L., Liu, P., Evans, T. C. Jr. & Ettwiller, L. M. DNA damage is a pervasive cause of sequencing errors, directly confounding variant identification. Science 355, 752–756 (2017).
Sinha, R. et al. Index Switching Causes ‘Spreading-Of-Signal’ Among Multiplexed Samples In Illumina HiSeq 4000 DNA Sequencing. bioRxiv 125724 (2017).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Bergmann, E. A., Chen, B.-J., Arora, K., Vacic, V. & Zody, M. C. Conpair: concordance and contamination estimator for matched tumor-normal pairs. Bioinformatics 32, 3196–3198 (2016).
Costello, M. et al. Characterization and remediation of sample index swaps by non-redundant dual indexing on massively parallel sequencing platforms. bioRxiv 200790 (2017).
Landau, D. A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Schmitt, M. W., Loeb, L. A. & Salk, J. J. The influence of subclonal resistance mutations on targeted cancer therapy. Nat. Rev. Clin. Oncol. 13, 335–347 (2016).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Acknowledgements
We are grateful for the sequencing services provided by the Genomics Core Facility and the Norwegian High-Throughput Sequencing Centre, South East Health Region/Oslo University Hospital, and for technical assistance provided by Jinchang Sun. We thank Illumina, Inc. for providing sequencing reagents and flow cells necessary to conduct the experiment for evaluating the role of multiplexing in ExAmp-associated index misassignment. This work has been supported by the Research Council of Norway (RCN grants 218241, 221580 and 230817/F20).
Author information
Authors and Affiliations
Contributions
D.V. and S.L. designed data analysis. D.V., S.N. and L.B.A. conducted data analysis. D.V., S.L., L.A.M.-Z. and E.H. designed the sequencing experiment. D.V. and S.N. drafted the manuscript. S.L., L.B.A., O.M., H.H., B.B., L.A.M.-Z. and E.H. revised the manuscript. O.M., H.H. and B.B. acquired sample data. E.H. and L.A.M.-Z. supervised the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The research has been in part funded by Illumina, Inc., which provided sequencing reagents and flow cells necessary to conduct the experiment for establishing the role of multiplexing in ExAmp-associated index misassignment.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vodák, D., Lorenz, S., Nakken, S. et al. Sample-Index Misassignment Impacts Tumour Exome Sequencing. Sci Rep 8, 5307 (2018). https://doi.org/10.1038/s41598-018-23563-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-23563-4
This article is cited by
-
A cost-effective sequencing method for genetic studies combining high-depth whole exome and low-depth whole genome
npj Genomic Medicine (2024)
-
Low-complexity and highly robust barcodes for error-rich single molecular sequencing
3 Biotech (2021)
-
Reliable multiplex sequencing with rare index mis-assignment on DNB-based NGS platform
BMC Genomics (2019)
-
A novel high-throughput molecular counting method with single base-pair resolution enables accurate single-gene NIPT
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.