Abstract
Adult Cebpb KO mice incisors present amelogenin-positive epithelium pearls, enamel and dentin allopathic hyperplasia, fewer Sox2-positive cells in labial cervical loop epitheliums, and reduced Sox2 expression in enamel epithelial stem cells. Thus, Cebpb acts upstream of Sox2 to regulate stemness. In this study, Cebpb KO mice demonstrated cementum-like hard tissue in dental pulp, loss of polarity by ameloblasts, enamel matrix in ameloblastic layer, and increased expression of epithelial-mesenchymal transition (EMT) markers in a Cebpb knockdown mouse enamel epithelial stem cell line. Runx2 knockdown in the cell line presented a similar expression pattern. Therefore, the EMT enabled disengaged odontogenic epithelial stem cells to develop supernumerary teeth. Cebpb and Runx2 knockdown in the cell line revealed higher Biglycan and Decorin expression, and Decorin-positive staining in the periapical region, indicating their involvement in supernumerary tooth formation. Cebpb and Runx2 acted synergistically and played an important role in the formation of supernumerary teeth in adult incisors.
Similar content being viewed by others
Introduction
Alterations in tooth development have enabled us to study a variety of dental anomalies, with supernumerary teeth (extra teeth) and tooth agenesis (missing teeth) being among the most common ones in humans. That said, numerous mouse mutant models have provided insights into the formation of supernumerary teeth1,2,3,4,5,6,7,8,9,10,11,12,13. We previously showed that inhibition of apoptosis can lead to the successive development of rudimentary maxillary incisors in uterine sensitization-associated gene-1 (Usag-1) null mice5,14,15. We also demonstrated that CCAAT/enhancer binding protein (C/EBP) beta (Cebpb) deficiency was related to the formation of supernumerary teeth. A total of 66.7% of Cebpb−/− 12-month-old animals presented supernumerary teeth and/or odontomas1. We found mineralized tissue formation typical of supernumerary teeth, around the labial cervical loop epithelium in 3-month-old adult mice, whereas no phenotypic changes were observed during embryonic tooth development. However, the mechanism of supernumerary tooth formation around the labial cervical loop epithelium in adulthood remains unclear.
We recently found interesting phenotypes in Usag-1/runt-related transcription factor 2 (Runx2) double knockout mice: the prevalence of supernumerary teeth was lower than that of Usag-1 null mice and the frequency of molar lingual buds were lower than that of Runx2 null mice. Hence, we suggested that Runx2 and Usag-1 act in an antagonistic manner in supernumerary tooth formation16, with Runx2 negatively regulating the process.
Endogenous Cebpb is expressed in ameloblasts and odontoblasts of 6-month-old wild-type mouse incisors17. Runx2 is expressed in the dental epithelium and/or mesenchyme of both the incisors and molars, and exhibits distinct temporospatial patterns18,19,20,21. Mutations in Runx2 are responsible for inherited cleidocranial dysplasia (CCD), an autosomal-dominant disorder characterized by elevated supernumerary tooth formation22,23. Interestingly, both heterozygous and null Runx2 mice possess a lingual epithelial bud, which represents a putative successional tooth associated with the upper molars and incisors; however, supernumerary tooth formation has never been observed in Runx2+/− mice24,25. The dental abnormalities in CCD suggest that Runx2 plays an important role during dental formation. Although Runx2 has been considered a determinant of CCD, some CCD patients do not present Runx2 mutations. We demonstrated previously in mice that prospective signs of CCD could be associated with Cebpb deficiency26. These could provide an additional etiological factor of CCD. Cebpb−/−Runx2+/− mice have shown more severe dwarfism than their Cebpb−/− littermates during the perinatal period and remained smaller even 12 weeks after birth. Thus, Cebpβ and Runx2 collaboratively control skeletal growth and osteoarthritis during development27, however their combined role in tooth morphogenesis remains unknown.
Given the above evidence, we hypothesized that Cebpb and Runx2 acted synergistically and played an important role in tooth morphogenesis and the formation of supernumerary teeth around the labial cervical loop epithelium in adult incisors. To test our hypothesis and study the in vivo relationship between Cebpb and Runx2 we established double knockout mice.
Results
Cebpb is associated with the maintenance of stemness in odontogenic epithelial stem cells and the labial cervical loop epithelium of adult maxillary incisors
To investigate the mechanisms underlying mineralized tissue formation around the labial cervical loop epithelium, such as those seen in supernumerary tooth structures, we first performed a detailed histological evaluation of adult Cebpb−/− mice (129 Sv background) (Fig. 1A–C). We observed numerous amelogenin-positive epithelium pearls near the periapical tissue (Fig. 1D), enamel and dentin ectopic hyperplasia in the periapical tissue (Fig. 1B–E,H), as well as miniaturization of the labial cervical loop epithelium that is specialized epithelial structure for stem cell niche (Fig. 1G,K–N). Based on these results, we concluded that odontogenic epithelial stem cells (OESCs) within the labial cervical loop epithelium lost their stemness in Cebpb−/− mice. Consequently, these OESCs differentiated into ameloblasts and secrete enamel matrix protein and enamel ectopically. Ectopic enamel formation was observed mainly in the maxilla, and was also observed in the mandible with low frequency (in a 129 Sv background) (Fig. 1H,I)1. We therefore investigated SRY (sex determining region Y)-box 2 (Sox2) expression in the labial cervical loop epithelium of maxillary incisors in Cebpb−/− mice. Sox2 localized to the inner and outer enamel epithelium, particularly the transit-amplifying zone, as well as the stellate reticulum (Fig. 1J), confirming a previous report28. There were significantly fewer Sox2-positive cells in the labial cervical loop epithelium of adult Cebpb−/− mouse incisors than in wild-type animals (Fig. 1J–N). These findings suggest that Cebpb maintains Sox2-positive OESCs in the labial cervical loop epithelium during postnatal life.
Phenotypes of incisors in Cebpb−/− mice maintained in a 129 Sv background. (A) Three-dimensional image of the skull. The left incisor is short. Scale bar: 10 mm. (B) Micro-computed tomography (CT) axial section image at the level of the maxillary incisor. Left incisors are deformed. (C) Sagittal hematoxylin-eosin (H&E)-stained section of the left upper incisor. Ectopic marked growth of dentin and enamel is observed in the apical region of the left incisor. Scale bar: 1 mm, 40×. (D) Amelogenin immunostaining (red) and Hoechst nuclear staining (blue). Amelogenin is observed in the hypocalcification site. Mature enamel was removed from this decalcified section leaving a clear enamel space (ES). Arrowheads indicate epithelial pearls. Scale bar: 100 μm, 100×. (E) Many dental tubules can be seen in dentin, which is also more abundant. Derangement of ameloblasts are observed near enamel matrix. Scale bar: 100 μm, 200×. (F) Magnified image of dental tubules in dentin. Scale bar:200 μm. 400× Inset: 1000×. (G) Nearly normal development of the right incisor. Ameloblasts, enamel-producing cells, are arranged in a single-cell row. Odontoblasts, dentin-producing cells, line the pulp cavity. Increased dentin is observed in both the labial and lingual side. Miniaturization of the labial cervical loop epithelium is seen. Scale bar: 1 mm, 40×. (H) Sagittal H&E-stained section of the left mandibular incisor of a 10-month-old mouse. Scale bar: 2 mm, 20×. (I) The boxed region in H. A mature tooth structure with pulp enclosed with dentin in a different direction is observed in a periapical region of a mandibular incisor. Scale bar: 500 μm, 100×. (J) SRY (sex determining region Y)-box 2 (Sox2)-Positive cells in a maxillary incisor labial cervical loop epithelium of 1-month-old wild-type (WT) mice. Scale bar: 100 μm, 400×. (K) Sox2-Positive cells in a maxillary incisor labial cervical loop epithelium of 19-month-old Cebpb−/− mice. Scale bar: 100 μm, 400×. (L) Comparison between WT and Cebpb−/− mice. Y-axis indicates the labial cervical loop epithelium area; X-axis indicates genotypes. (M) Comparison between WT and Cebpb−/− mice. Y-axis indicates number of Sox2-positive cells in the labial cervical loop epithelium; the X-axis indicates genotypes. (N) Comparison between WT and Cebpb−/− mice. Y-axis indicates Sox2-positive area in the labial cervical loop epithetium area; X-axis indicates genotypes.
Abrogation of Cebpb and Runx2 works synergistically in supernumerary tooth formation around the labial cervical loop epithelium in adult Cebpb −/− Runx2 +/− mice
We had previously observed mineralized tissue, akin to that of supernumerary tooth structures, around the labial cervical loop epithelium in adult Cebpb−/− mice1. Although Runx2−/− mice die shortly after birth due to the absence of bone formation22, some Runx2+/− mice, showed lingual bud formation in the embryonic stage. Therefore, to determine whether Cebpb and Runx2 acted synergistically and played an important role in supernumerary tooth formation, we performed a series of histological studies of maxillary incisor formation in Cebpb+/+Runx2+/+, Cebpb−/−Runx2+/+, Cebpb+/+Runx2+/−, and Cebpb−/−Runx2+/− mice (Fig. 2A–D). We first investigated the synergistic action of Cebpb and Runx2 during lingual bud formation. We previously reported that Usag-1 and Runx2 acted in an antagonistic manner in lingual bud formation at embryonic day 15 (E15)16. Here, the Cebpb+/+Runx2+/− (F2 background) maxillary incisor germ displayed lingual budding at E15 (Fig. 2C); however this genotype was smaller incidence rate of budding (Fig. 2E) and was significantly fewer cell number of budding than Cebpb−/−Runx2+/− (Fig. 2F). These results support the hypothesis that Cebpb and Runx2 act synergistically in OESCs during embryonic development. In contrast, the lingual bud was highly enriched with Sox2-positive OESCs. At present, we could not find out how OESCs within the lingual bud lost their stemness during the embryonic stage. Next, we analyzed whether Cebpb and Runx2 acted synergistically during supernumerary tooth formation around the labial cervical loop epithelium in adult mice (Fig. 3). Five out of eight (62.5%) adult Cebpb−/−Runx2+/+ (F2 background) mice presented mineralized tissue around the labial cervical loop epithelium, akin to that of supernumerary tooth structures, roughly reflecting the value reported in Cebpb−/− (129 Sv background) mice (66.7%)1 (Table 1). In severe cases, no labial cervical loop epithelium structures were seen in the aberrant incisors. These results indicate that there is no phenotypic difference between F2 and 129 Sv mouse background. Four (33%) out of twelve Cebpb−/−Runx2+/− 3-month-old (F2 background) mice presented aberrant maxillary incisors, with developing or mature ectopic supernumerary teeth in the periapical tissue and dental pulp (Table 1, Fig. 3A–D,H–K). The incidence of short incisors was higher in Cebpb−/− mice (129 Sv or F2 background) than in Cebpb−/−Runx2+/− mice (F2 background), whereas the incidence of supernumerary teeth showed the opposite trend (Table 1). These data confirm the synergistic action of Cebpb and Runx2 in supernumerary tooth formation around the labial cervical loop epithelium during postnatal life. Moreover, this is the first report on supernumerary tooth formation in Cebpb−/−Runx2+/− mice.
Phenotype of maxillary incisor germ in the F2 generation at embryonic day 15. (A,B) Incisor germs of Cebpb+/+Runx2+/+ and Cebpb−/−Runx2+/+ mice. Scale bar: 100 μm, 400×. Left indicates the labial side. (C,D) Immunostaining for SRY (sex determining region Y)-box 2 (Sox2) in incisor germs of Cebpb+/+Runx2+/− and Cebpb−/−Runx2+/− mice. Scale bar: 100 μm, 400×. Left indicates labial side. Runx2+/− tooth germ displays lingual budding(arrowhead). (E) Penetrance of budding according to genotype. (F) Comparison between Cebpb+/+Runx2+/− and Cebpb−/−Runx2+/− mice lingual budding. The Y-axis on the left indicates the cell number of buddings in the maximal area of all sections; the X-axis on the bottom indicates genotypes.
Phenotypes of maxillary incisors in the F2 generation of 3-month-old Cebpb−/−Runx2+/− mice. All sections were decalcified. (A–G) Correspond to the same mouse, H corresponds to another animal, and (I–K) correspond to the other animal. (A) Three-dimensional image of the skull. Scale bar: 10 mm. (B) Micro-computed tomography (CT) axial image at the level of the maxillary incisor. (C) Sagittal hematoxylin-eosin (H&E)-stained section of the upper incisor. A box E indicates epithelial pearls. An arrowhead indicates a developing tooth. Developing teeth are characterized by odontoblasts, dentin, ameloblasts derangement, and surface roughness of the enamel matrix (hypocalcification site). Scale bar: 100 μm, 100×. (D) Amelogenin immunostaining (red) and Hoechst nuclear staining (blue). An arrowhead indicates epithelial pearls. Scale bar: 100 μm, 100×. (E) High-magnification image of epithelial pearls. Scale bar: 100 μm, 400×. (F) High-magnification image of epithelial pearls. Scale bar: 50 μm, 1000×. (G) Ameloblasts derangement and surface roughness of the enamel matrix are visible in an opposite incisor. Scale bar: 100 μm, 400×. (H) Two mature teeth and a developing tooth are visible. Scale bar: 1 mm, 40×. Inset: Higher magnification image of a developing tooth showing odontoblasts, ameloblasts, and dentin 400×. (I) Developing and mature teeth are visible. Scale bar: 1 mm, 40×. (J) A multipartite mature tooth (arrowhead) is observed in the dental pulp. Scale bar: 100 μm, 100×. (K) Many developing teeth (arrowheads) are visible near an extra mature tooth. Scale bar: 100 μm, 200×.
Reduction of Cebpb and Runx2 expression contributes to epithelial-mesenchymal transition (EMT) in odontogenic epithelial cells of the maxillary incisor
With an aim to examine the effect of Runx2+/− on postnatal supernumerary tooth formation, we performed a detailed histological evaluation around the labial cervical loop epithelium. Compared to wild-type mice (Fig. 4A), differentiated ameloblasts in the maxillary incisor were deranged and lost their cell polarity in adult Cebpb+/+Runx2+/− animals (F2 background) (Fig. 4B). Furthermore, loss of cell polarity and surface roughness of the enamel matrix were visible also in Cebpb−/−Runx2+/+ and Cebpb−/−Runx2+/− mice (Fig. 4C–E). Concomitant to the EMT process is the disappearance of the apical-basal polarity of epithelial cells29. Therefore, we performed immunostaining for the EMT marker-N cadherin-in the maxillary incisors of 3-month-old Cebpb+/+Runx2+/− and Cebpb−/−Runx2+/− mice. Non-polarized differentiated ameloblasts and the cell component surrounding ectopic enamel formation and cementum-like hard tissue within the dental pulp in the periapical region were positive for presence of N-cadherin (Fig. 4F–H). Thus, based on these morphological changes, we suggest that Cebpb and Runx2 knockdown contributed to EMT in odontogenic epithelial cells of the maxillary incisor. Taken together, supernumerary tooth formation around the labial cervical loop epithelium of adult maxillary incisors is dependent on both Cebpb knockdown-induced loss of stemness in OESCs and EMT of odontogenic epithelial cells in Runx2+/− and/or Cebpb−/− mice.
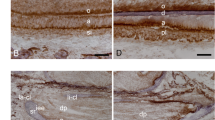
Analysis of differentiated ameloblasts in maxillary incisors of 3-month-old mice (F2 generation). All sections were decalcified. These insets in (A–C,E) are 1000×. (F–H) N-cadherin immunostaining. All scale bars: 200 μm, 400×. (A) Highly polarized secretory ameloblasts are visible in wild-type mice. (B) Differentiated ameloblasts are seen to have lost their cell polarity in Cebpb+/+Runx2+/− mice. (C) Non-polarized differentiated ameloblasts and surface roughness of the enamel matrix are visible in Cebpb−/−Runx2+/+ mice. (D) Amelogenin immunostaining (red) and Hoechst nuclear staining (blue). Free blobs containing amelogenin are observed in Cebpb−/−Runx2+/+ mice. (E) Derangement of ameloblasts and surface roughness of the enamel matrix. in Cebpb−/− Runx2+/− mice. (F) Positive staining in ameloblasts containing non-polarized differentiated ameloblasts and labial cervical loop epithelium in Cebpb+/+Runx2+/− mice incisor. Scale bar: 500μm, 100×. The inset: High-magnification image of non-polarized differentiated ameloblasts. 1000×. (G) Ectopic enamel formation and cementum-like hard tissue within the dental pulp in the periapical region of Cebpb−/− Runx2+/− mice incisor. Scale bar: 2 mm, 40×. (H) Positive staining of the cell component surrounding ectopic enamel formation and cementum-like hard tissue are visible. Scale bar: 200 μm, 400×. The inset: High-magnification image of the cementum-like hard tissue 1000×.
Reduced Cebpb mRNA expression leads to a decrease of Sox2, Runx2 knockdown inhibits adhesion molecules and EMT in OESCs
To confirm the above-mentioned function of Cebpb and Runx2 in odontogenic epithelial cells of the adult labial cervical loop epithelium, in vitro knockdown experiments were performed using mHAT9d cells. These cells are derived from the labial cervical loop epithelium of a mouse incisor and could undergo EMT30. Transfection efficiency was checked using a fluorescence-activated cell sorter (FACS) and was determined to be approximately 60% 24 h after addition of 1.5 µL Lipofectamine RNAiMAX and control stealth siRNA (Supplement Fig. 1). We used a semi-quantitative reverse transcription polymerase chain reaction (sqRT-PCR) to evaluate Cebpb and Runx2 type1 knockdown by stealth siRNA in mHAT9d cells 48 h after transfection (Fig. 5). The specific stealth siRNA effectively abolished Cebpb and Runx2 mRNA and protein levels (Fig. 5A,B). Furthermore, to confirm the specificity for the Runx2 type1 knockdown, we also investigated the expression of Runx1, other Runx2 isoforms, and Runx3 mRNA. We found that expression of other Runx2 isoforms and Runx3 declined, whereas that of Runx1 increased. Additionally, we showed that Cebpb acted upstream of Sox2 to maintain stemness of mHAT9d cells, as indicated by reduced Sox2 mRNA expression following Cebpb knockdown (Fig. 5A). However, expression of two ameloblast differentiation markers, amelogenin (Amelx) and ameloblastin (Ambn), could not be detected (Supplement Fig. 2). These results indicate that Cebpb alone is not sufficient for maintaining stemness and differential inhibition in mHAT9d cells in vitro. Next, we analyzed whether Runx2 type1 knockdown showed EMT in mHAT9d cells. SqRT-PCR analysis of Runx2 type1 knockdown by stealth siRNA demonstrated decreased mRNA expression of adhesion molecules (Fig. 5C). Expression of EMT markers snail family zinc finger 2 (Snai2) increased, as did that of the mesenchymal marker N-cadherin (Cdh2), whereas another mesenchymal marker, vimentin (Vim), and the epithelial marker E-cadherin declined. Similarly, Cebpb knockdown showed increased expression of Snai2, but a reduction of Vim and E-cadherin (Fig. 5C). Biglycan (Bgn), decorin (Dcn), and bone morphogenetic protein 4/6/7 (Bmp4/6/7) mRNA expression was seen to increase in Runx2 type1 knockdown, and even more in Cebpb and Runx2 type1 knockdown (Fig. 5D). Additionally, bone morphogenetic protein (Bmp2/11) mRNA expression decreased in Runx2 type 1 knockdown and Cebpb and Runx2 type 1 knockdown (Fig. 5D). Because Dcn, encoding a small leucine-rich proteoglycan was particularly up-regulated in double knockdown, we immunostained for decorin in the incisors of 3-month-old F2-generation Cebpb−/−Runx2+/− mice (Fig. 6). Positive staining was observed in the enamel epithelial pearl and extracellular matrix near the supernumerary tooth in the periapical region, as well as the extracellular matrix of the cell component surrounding cementum-like hard tissue in the dental pulp. These in vitro experiments indicated that knockdown of both Cebpb and Runx2 contributed to EMT in a synergistic manner, whereas Cebpb knockdown alone caused down-regulation of Sox2.
Semi-quantitative reverse transcription polymerase chain reaction (RT-PCR) and Western blotting results in mHAT9d cells 48 h after transfection with Cebpb and Runx2 type1 stealth siRNA. (A) Changes in the expression of Cebpb, members of the Runx family, and SRY (sex determining region Y)-box 2 (Sox2) in mHAT9d cells 48 h after transfection with Cebpb and Runx2 type1 stealth siRNA. Cebpb knockdown (N(negative control) + 925) shows slightly reduced expression of Sox2. Runx2 knockdown (N + 1623) shows increased expression of Sox2 and Runx1, but decreased expression of other Runx2 isoforms and Runx3. Concomitant Cebpb and Runx2 type1 siRNA knockdown (925 + 1623) was also performed. (B) Western blotting results. N indicates negative control stealth siRNA, 925 and 1623 indicate Cebpb and Runx2 type1 siRNA, respectively (final concentration 10 nM). (C) Changes in the expression of adhesion molecules, epithelial-mesenchymal transition (EMT) markers, and mesenchymal markers in mHAT9d cells 48 h after transfection with Cebpb and Runx2 type1 stealth siRNA. Cebpb knockdown (N(Negative control) + 925) caused increased expression of EMT markers snail family zinc finger 2 (Snai2) and decreased expression of the mesenchymal marker vimentin (Vim) and the epithelial marker E-cadherin (Cdh1). Runx2 type1 knockdown (N + 1623) caused decreased expression of adhesion molecules Cdh1 and integrin alpha 6 (Itga6), decreased expression of Vim, and increased expression of Snai2. Concomitant Cebpb and Runx2 type1 knockdown (925 + 1623) caused decreased expression of Cdh1, Itga6 and VIm, and increased expression of mesenchymal marker N-cadherin (Cdh2) and Snai2. (D) Changes in biglycan (Bgn), decorin (Dcn), bone morphogenic proteins (Bmps) in mHAT9d cells 48 h after transfection with Cebpb and Runx2 type1 stealth siRNA. Concomitant Cebpb and Runx2 type1 siRNA knockdown (925 + 1623) shows a markedly increased expression of Bgn and Dcn (encoding two small leucine-rich peptidogycans), and increased expression of Bmp4/6.
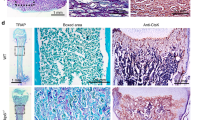
Decorin immunostaining in the incisors of 3-month-old Cebpb−/−Runx2+/− mice (F2 generation). (A,C) Wild-type (WT)/WT mice Scale bar: 1 mm, 40×. (B,D,E) Cebpb−/−Runx2+/− mice. (B) Positive staining of cells in the dental pulp, except for cementum-like hard tissue, and the extracellular matrix. Scale bar: 1 mm, 40×. (D) Positive staining in the enamel epithelial pearl and extracellular matrix near the supernumerary tooth in the periapical region. Scale bar: 100 μm, 400×. (E) Positive staining in the extracellular matrix surrounding the cementum-like hard tissue in the dental pulp. Scale bar: 100 μm, 400×.
Discussion
The Bmp signaling network may provide a niche supporting transient Sox2-positive dental epithelial stem cells in murine incisors31. In undifferentiated mesenchymal stem cells (MSCs), CEBPB acts as a repressor, helping maintain stemness. Upon adipogenic, chondrogenic, or osteogenic stimulation, different signaling pathways activate Smad proteins, these bind to CEBPB and remove it from its target regulatory elements, thus converting the repressor into an activator32,33. Next, CEBPB binds to and activates its differentiation target genes and, together with other lineage-specific transcription factors, promotes mesenchymal differentiation into adipocytes, osteoblasts or chondrocytes. Here, part of the dental pulp caused ectopic cementogenesis in the maxillary incisor of Cebpb−/− mice (F2 background) (Fig. 3C,H,J and Table 1, Figure Supplement 3A,B). The phenotype was thought to arise from transdifferentiation of OESCs, as these cells invaded the incisor dental pulp rather than its root analogue. Some cementum-producing cells (cementoblasts) may originate from dental epithelial cells34. Cebpb knockout promotes the differentiation of crown epithelia into the root lineage or even cementoblast-like cells rather than ameloblasts35 (Figure Supplement 3C–J). Taken together, these findings suggest that Cebpb maintains OESCs in the labial cervical loop epithelium and dental pulp stem cells (DPSCs) in murine incisors (Fig. 7).
Schematic representation of the role played by Cebpb and Runx2 in odontogenic epithelial stem cells (OESCs). Cebpb acts upstream of SRY (sex determining region Y)-box 2 (Sox2). In cooperation with Oct 3/4 protein (Pou5f1), Sox2 inhibits genes related to differentiation and maintains stemness in OESCs. Cebpb and Runx2 prevent epithelial-mesenchymal transition (EMT) of OESCs via snail family zinc finger 2 (Snai2). Both also inhibit biglycan (Bgn), decorin (Dcn) and Bmp4. EMT promotes expression of Sox2 and thus stemness of OESCs. EMT enables disengaged OESCs to develop supernumerary teeth by interaction with various mesenchymal stem cells such as dental MSCs.
In Runx2 heterozygous and null mice, budding was observed in maxillary incisors at E15 (Fig. 2C,D). Both Runx2 mutants displayed lingual buds in front of the maxillary molars, which are in line with Runx2 preventing the formation of buds for successional teeth24,25. There is a difference between the phenotypes of OESCs in Cebpb−/−Runx2+/+ and Cebpb−/−Runx2+/− incisors (F2 background) between E15 and adult animals. No buddings were observed at E15 in Cebpb−/− mice (F2 background) (Fig. 2B), however they were seen at E15 in Cebpb+/+Runx2+/− and Cebpb−/−Runx2+/− mice (Fig. 2C,D) and disappeared on postnatal day 7 (Figure supplement 4). Meanwhile, some adult Cebpb−/− mice (129 Sv background) possessed an unusual incisor that presented ectopic hyperplasia of enamel and dentin in the periapical tissue. Moreover, 33% of 3-month-old Cebpb−/−Runx2+/− mice (F2 background) had aberrant incisors, characterized by developing or mature ectopic supernumerary teeth in the periapical tissue and dental pulp (Table 1, Figs 1B–D and 3A–D,H–K). Indeed, in humans, supernumerary teeth are less common in deciduous dentition (first generation of teeth) than in permanent dentition (second generation of teeth)36. In mice, the difference may be linked to stem cell aging in the incisor. Common contributing factors of aging in different organisms, but particularly in mammals, are genomic instability, telomere attrition, epigenetic alterations, loss of proteostasis, deregulated nutrient sensing, mitochondrial dysfunction, cellular senescence, stem cell exhaustion, and altered intercellular communication37. As another example of epithelial-mesenchymal interactions, hair graying is the most obvious sign of aging in mammals. Irreparable DNA damage, as that caused by ionizing radiation, abolishes renewal of MSCs in mice and results in hair graying inasmuch as it also triggers MSC differentiation into mature melanocytes in the niche38. The hallmarks of OESCs can change according to aging.
Differentiated ameloblasts lost their cellular polarity and derangement of ameloblasts was observed in Cebpb+/+Runx2+/−, Cebpb−/−Runx2+/+, and Cebpb−/−Runx2+/− mice (F2 background) (Fig. 4B–E). Furthermore, we observed the positive staining of N-cadherin during EMT within the dental epithelium in the periapical region (Fig. 4F,G). SqRT-PCR analysis of Runx2 knockdown by siRNA in mHAT9d cells resulted in decreased expression of the epithelial marker E-cadherin (Fig. 5C). The same cells exhibited also increased expression of EMT markers Cdh2 (N-cadherin) and Snai2, but not the other mesenchymal marker Vim. It should be noted that Cebpb knockdown confirmed increased expression of Snai2, as well as a decline in Vim (Fig. 5C). This in vitro knockdown experiment may be unable to reproduce in vivo conditions sufficiently and may only reflect the initial state of EMT, carried out by epithelial cell rests of Malassez (ERM). Nevertheless, these results indicate the additive effect of Cebpb and Runx2 type1 knockdown on EMT. Rodent incisors are a continuation of Hertwig’s epithelial root sheath (HERS), which consists of two epithelial layers and plays a role in inducing odontogenesis during root development, becoming fragmented after that. The fragmentation of HERS cells is associated with EMT39,40,41. Growth inhibition of the inner enamel epithelium and the active proliferation of the outer enamel epithelium and/or stellate reticulum result in HERS formation42. ERM are capable of undergoing EMT under osteogenic conditions, when Vim is down-regulated43, as reported also in this study. Strong Bmp signaling in crown/labial dental epithelia acts in synchrony with weak Wnt signaling to trigger amelogenesis. The balance between strong Wnt signaling and weak Bmp signaling in apical/lingual dental epithelia promotes changes in adhesion properties and thus the formation of HERS/ERM. Cessation of Bmp signaling and up-regulation of Wnt signaling in dental epithelia stimulates the precocious formation of HERS/ERM and epithelium-originated cementogenesis via EMT34. Runx2 is the direct downstream target of Bmp signaling44. Down-regulation of Runx2 promotes a similar phenotypic change, as indicated by the onset of EMT in BMPR-1A-depleted tissues during the differentiation stage35. Cebpb might also be regulated by Bmp signaling45. Knockout of Cebpb mimics a situation characterized by strong Wnt signaling and no Bmp signaling, thereby leading to EMT and the cementum-like structure of ameloblasts. Therefore, we believe that EMT enabled disengaged OESCs bud and develop supernumerary teeth following induction of DPSCs, MSCs originating from OESCs, or those present in connective tissue near the periapical region. These MSCs may have interacted with free ameloblasts, stimulating their differentiation and the formation of amelogenin-positive enamel epithelial pearls and free blobs. SqRT-PCR analysis of Runx2 knockdown by siRNA in mHAT9d cells showed that Sox2 increased during EMT, allowing cells to gain stemness (Fig. 5A). Cebpb knockdown and Runx2 type1 knockdown act on stemness in an opposing manner (Fig. 5A). This behavior has been reported to indicate a direct link between EMT and acquisition of epithelial stem cell properties including self proliferation46,47. SqRT-PCR analysis demonstrated increased expression of Bmp4/6 in Runx2 type1 knockdown cells and even more so in Cebpb and Runx2 type1 knockdown cells (Fig. 5D). Bmp signaling plays a role in tumor generation and progression by affecting tumor EMT and stemness, thus increased Bmp4/6 expression could induce EMT and increased Sox2 expression in vitro48. Taken together, Runx2 inhibits EMT, differentiation, and branching of undifferentiated enamel epithelial cells, whereas Cebpb and Runx2 knockdown contribute to EMT additively, with EMT promoting Sox2 mRNA expression and acquisition of stemness (Fig. 7).
Decorin and biglycan are small leucine-rich proteoglycans distributed intracellularly, on cell surfaces, and within extracellular matrices. Biglycan and decorin regulate early stages of tooth formation and mediate mineralization of dental hard tissues. Both Bgn and Dcn null mice exhibited hypomineralized dentin, with the effect being more conspicuous in the latter. Enamel formation increased strikingly in newborn Bgn null mice and was retarded in the case of Dcn deletion49. The expression of Bgn in odontoblasts is highly differentiation stage-dependent. Dcn mRNA is expressed by odontoblasts abutting the premineralized matrix and predentin, and is synchronously related to expression of type I collagen. Bgn mRNA is expressed by pre-secretory and secretory ameloblasts50. Bgn may be associated with odontoblast and ameloblast differentiation, enamel formation, and restraining of collagen fibrillogenesis in predentin. Meanwhile, Dcn may be committed to dentin formation as a regulator of collagen mineralization, possibly through its specific association with type I collagen fibrils49,50,51. SqRT-PCR analysis demonstrated increased expression of Bgn, Dcn, and Bmp4/6 in Runx2 type1 knockdown cells and even more so in Cebpb and Runx2 type1 knockdown cells (Fig. 5D). From the above results, it appears that increased Bgn and Dcn expression in OESCs is likely involved in promoting the development of supernumerary teeth in Runx2+/−Cebpb−/− mice. BGN is essential for Bmp4-induced osteoblast differentiation in neonatal murine calvarial cells52. BGN may drive tooth formation by cooperating with Bmp4. DCN inhibits multiple growth factors, such as Tgfb1, and directly antagonizes several members of the receptor tyrosine kinase (RTK) family, including the epidermal growth factor receptor, the hepatocyte growth factor receptor, and the insulin-like growth factor receptor I53. Soluble decorin has been characterized as a pan-RTK inhibitor, with a simultaneous powerful downstream signaling inhibitory effect54. Higher Dcn expression enabled the collagen fibers to be markedly thinner and caused no mineralized matrices in vitro55. These functions of decorin may prevent proliferation of dental epithelial cells and mineralization of mesenchymal cells in the dental pulp, and help the development of a nearly normal supernumerary tooth conformation in the periapical tissue of Cebpb−/−Runx2+/− mice (F2 background) (Figs 3C,D and 6, and Table 1).
Molecular targeted therapy uses small molecules, monoclonal antibodies, or other substances to identify and attack specific types of cells by interfering with cellular targets. It could be used for de novo whole tooth regeneration by stimulating arrested tooth germs in animal models with congenital tooth agenesis16. Tooth development is under genetic control and is the result of reciprocal and reiterative signaling between oral ectoderm-derived dental epithelium and cranial neural crest cell-derived dental mesenchyme56. Two different types of cell components (epithelial and mesenchymal cells) are necessary and essential for whole tooth regeneration. Our results demonstrate that OESCs had the potential to differentiate odontoblasts by EMT and regenerate de novo whole teeth (supernumerary teeth) by synergistic abrogation of Cebpb and repression of Runx2. In recent years, there have been many studies on stem cells in relation to the development of regenerative therapies for various organs. This has led to the discovery that adult zebrafish regenerate nephrons (the functional units of the kidney) throughout their lives. The zebrafish kidney contains renal stem cells, which differentiate into epithelial or mesenchymal cells of the nephrons57. The EMT in Sox2-expressing stem/progenitor cells contributes to the regeneration of the pituitary gland58. It has been confirmed that Sox2, which is one of the molecular marker of OESCs, localizes to the dental lamina of developing human primary molars59. OESCs are suitable for molecular targeted therapy using Cebpb and/or Runx2 (Fig. 7). However, from our analysis, it remains unclear whether OESCs differentiate into odontogenic epithelia and mesenchyme directly or through other cells. Future studies tracking individual cells are needed to demonstrate cell dynamics during de novo whole tooth regeneration.
Materials and Methods
Production and analysis of Cebpb and Runx2 mutant mice
This study received appropriate approval from the Animal Research Committee of Kyoto University (Reference Number: Med Kyo 11518), as well as from the Recombinant DNA Experiment Safety Committee of Kyoto University to perform recombinant DNA experiments. Experiments were carried out in accordance with approved guidelines.
Cebpb+/− and Runx2+/− mice were gifts from Shizuo Akira (Osaka University) and Toshihisa Komori (Nagasaki University), respectively22,60. Cebpb+/− mice were maintained in a 129 Sv background. Runx2+/− mice were maintained in a C57BL/6 background. We interbred heterozygous Cebpb and Runx2 mice and analyzed the F2 generation. Runx2 null mice were embryonic lethal. Day E0 was taken to be midnight prior to finding a vaginal plug. Wild-type mice were used as control.
PCR amplification was performed with KOD FX NEO polymerase (KFX-201, TOYOBO, Osaka, Japan) and specific primers.
Three-dimensional (3D) computed tomography (CT) scans (SMX-100XT-SV3; Shimadzu, Kyoto, Japan) were performed to the heads of adult mice. We converted CB to TIFF files. CB files have 512 × 512 pixels, 8 bits, and voxel size x:y:z = 1:1:1 (approx. 0.06 mm per side). 3D images were reconstructed and analyzed using computer imaging software (INTAGE Realia and Volume Player; KGT Inc., Tokyo, Japan)61.
Immunohistochemistry
Heads of adult mice were decalcified with Kalkitox (Wako, Osaka, Japan) overnight at 4 °C and neutralized with 5% sodium sulfate solution (Wako) for more than 2 h. Paraffin-embedded sections were subjected to immunostaining with primary anti-Sox2 rabbit monoclonal antibody (1:100) (ab92494; Abcam, Cambridge, UK), and secondary anti-rabbit antibody (414341; Nichirei Bioscience, Tokyo, Japan), and were counterstained with hematoxylin (Wako). Some sections were immunostained with a primary anti-amelogenin antibody (1:500) (Hokudo, Sapporo, Japan) and a secondary Alexa Fluor 568 goat anti-rabbit antibody (A11036; Thermo Fisher Scientific, Waltham, MA, USA). Subsequently, sections were counterstained with Hoechst 33258 (H3569; Thermo Fisher Scientific). Other sections were immunostained with a primary anti-decorin goat polyclonal antibody (N-15, sc-22613; Santa Cruz Biotechnology, Dallas, TX, USA), and a secondary anti-goat antibody (414351; Nichirei Bioscience); and diaminobenzidine (415171; Nichirei Bioscience). The other sections were immunostained with a primary anti-N-cadherin Rabbit polycloneal antibody (1:100) (ab18203; Abcam) and a EnVision System HRP-labeled polymer (rabbit, K4003; Dako) with diaminobenzidine (K3468; Dako). They were also counterstained with hematoxylin.
Cebpb and Runx2 siRNA in a mouse enamel epithelial stem cell line
Cell culture
mHAT9d is a dental epithelial stem cell line derived from the labial cervical loop epithelium of a mouse incisor. Cells were cultured in Dulbecco’s modified Eagle’s medium/F12 (Thermo Fisher Scientific) containing B-27 supplement (Thermo Fisher Scientific), basic fibroblast growth factor (25 ng/mL; R&D Systems, Minneapolis, MN, USA), epidermal growth factor (100 g/mL; R&D Systems), and penicillin-streptomycin (1%). Cells were cultured on Primaria tissue culture plates (BD Biosciences, Franklin Lakes, NJ, USA). Mycoplasma detection was performed using the PCR Mycoplasma Detection Set (TaKaRa Bio, Shiga, Japan).
Examination of transfection efficiency using fluorescent oligos
For each transfection reaction on a 24-well plate, fluorescent oligo-Lipofectamine® RNAiMAX (Thermo Fisher Scientific) complexes were prepared as follows. BLOCK-iTTM Alexa Fluor Red Fluorescent Control (Thermo Fisher Scientific) (6 pmol) was diluted in 100 μL of Opti-MEM® I Reduced Serum Medium (Thermo Fisher Scientific). Lipofectamine® RNAiMAX (0.5, 1.0, and 1.5 μL) was added to the mixture and incubated at room temperature for 15 min. The cell suspension (500 μL) was diluted with the above-mentioned medium and was added to a well containing the fluorescent oligo and Lipofectamine® RNAiMAX mixture. This gave a final fluorescent oligo concentration of 10 nM, a final volume of 600 μL, and 2 × 105 cells. The cells were incubated at 37 °C in a CO2 incubator until they were ready for harvesting and assayed for the gene of interest. Transfection efficiency and thus target gene knockdown was measured using a FACS calibur (BD Biosciences) 24, 48, and 72 h after transfection.
Gene knockdown using stealth RNAi siRNA in mHAT9d cells
Stealth RNAi siRNA (120 pmol) was diluted in 100 μL of Opti-MEM® I Reduced Serum Medium in a 10-cm dish. Stealth siRNA was as follows: Stealth RNAiTM siRNA Negative Control Med GC Dup, Cebpb (NM_001287739.1_stealth_925, Sense: 5′-GAGAGCUCAGCACCCUGCGGAACUU-3′, Anti-Sense: 5′-AAGUUCCGCAGGGUGCUGAGCUCUC-3′), Runx2 type1 (NM_001145920.2_stealth_1623, Sense: 5′-GGCCACUUACCACAGAGCUAUUAAA-3′, Anti-Sense: 5′-UUUAAUAGCUCUGUGGUAAGUGGCC-3′). Stealth siRNA was mixed with 30 µL Lipofectamine® RNAiMAX and 10 mL of cell suspension diluted with culture medium. This gave a final fluorescent oligo concentration of 10 nM, a final volume of 12 mL, and 4 × 106 cells. mRNA and protein expression at 48 h post transfection were measured using semi-quantitative RT-PCR and western blotting, respectively.
SqRT-PCR
RT-PCR was performed as follows. Total RNA from 70% confluent mHAT9d cells was extracted using TRIzol reagent (Thermo Fisher Scientific), 0.2 mL chloroform, and 0.2 mL phenol chloroform per 1 mL of TRIZOL. The RNA was precipitated from the aqueous phase by mixing with isopropyl alcohol and 16.7 mg/mL glycogen or with NucleoSpin® RNA Clean-up XS columns (U0903, TaKaRa Bio). Total RNA (3 μg) was reverse transcribed using the SuperScript ®IV First-Strand Synthesis System (Thermo Fisher Scientific).
cDNA was serially diluted and PCR amplification was performed using Ex Taq (RR001; TaKaRa Bio), KOD FX (KFX-101; TOYOBO), and specific oligonucleotide primers (Supplementary Table 1, 2). All PCR products were examined with serial dilutions of the cDNA by electrophoresis on 2% agarose gels with ethidium bromide staining. The resulting bands were quantified with a Bio-image analyzer (FAS-IV; NIPPON Genetics, Tokyo, Japan). Experiments were carried out in triplicate. To characterize the mHAT9d cell line, we used RT-PCR analysis to confirm the expression of dental epithelial cell markers: cytokeratin 14 (Krt14), notch 1 (Notch1), jagged 1 (Jag1), and p63 (Tprg); core binding factor beta 1 (Cbfb1),Polycomb complex protein BMI-1 (Bmi1), ras homolog family member A (Rhoa), and fibronectin (Fn1); small integrin-binding ligands of the N-linked glycoprotein family members dentin matrix protein1 (Dmp1) and osteopontin (Spp1); and the neural crest marker proteolipid protein 1 (Plp1)62; P-cadherin (Cdh3); transforming growth factor beta (Tgfb)1/2/3; the Bmp antagonist-sclerostin domain containing 1 (Sostdc1) (Supplemental Fig. 5, Supplementary Table 2).
Each siRNA was the same in quantity (10 nM each) when mixed. 1/10, 1/30, and 1/90 indicates dilution ratio. 925 and 1623 indicate Cebpb and Runx2 type1 siRNA, respectively. RT-PCR analysis of Cebpb and Runx2 type1 expression after knockdown with negative control stealth siRNA or the corresponding siRNA (final concentration 20 nM). All experiments were performed at least thrice, and representive data are shown. GAPDH was used as an internal control for RT-PCR. The bar graphs indicate relative mRNA level of genes, which were calculated by image j software. Values of them were normalized to GAPDH and mRNA expression data on negative control stealth siRNA were used as denominator of ratios.
Western blotting
Western blotting of mHAT9d cells transfected with Cebpb and Runx2 type1 stealth siRNA was performed using the following antibodies: anti-Gapdh primary rabbit polyclonal (1:1000, sc-150; Santa Cruz Biotechnology), secondary anti-rabbit IgG (1:5000, #7074; Cell Signaling Technology), anti-C/EBP beta primary rabbit polyclonal (1:200, #3087; Cell Signaling Technology), anti-Runx2 primary rabbit polyclonal (1:200, M-70 and sc-10758; Santa Cruz Biotechnology), and secondary anti-rabbit IgG (1:1000, #7074; Cell Signaling Technology). Ez West Blue (AE-1490; ATTO, Tokyo, Japan) was used to stain the membranes and detect the proteins of interest.
Statistical analysis
Data are presented as mean ± standard deviation. Statistical significance was assessed by analysis of variance in R. Gaussian distribution was determined with an Anderson-Darling normality test.
To analyze the labial cervical loop epithelium area, the number of Sox2-positive cells and Sox2-positive area/Labial cervical loop epithelium area, which was measured using the imageJ software, in the maximal area of all sections in Cebpb genetically modified mice (129 Sv), statistical significance was determined using a Mann-Whitney U test. Sox2-positive area (diaminobenzidine coloring) was extracted with Photoshop elements 12 (Adobe Systems, San Jose, CA, USA). To analyze the area and the cell number of lingual buds in the maximal area of all sections in Cebpb+/+Runx2+/− and Cebpb−/−Runx2+/− F2 (129 Sv/C57BL/6) mice, statistical significance was determined using a Mann-Whitney U test.
References
Huang, B. et al. Phenotypes of CCAAT/enhancer-binding protein beta deficiency: hyperdontia and elongated coronoid process. Oral diseases 19, 144–150 (2013).
Järvinen, E. et al. Continuous tooth generation in mouse is induced by activated epithelial Wnt/beta-catenin signaling. Proceedings of the National Academy of Sciences of the United States of America 103, 18627–18632 (2006).
Kaufman, M. H., Chang, H. H. & Shaw, J. P. Craniofacial abnormalities in homozygous Small eye (Sey/Sey) embryos and newborn mice. Journal of anatomy 186(Pt 3), 607–617 (1995).
Klein, O. D. et al. Sprouty genes control diastema tooth development via bidirectional antagonism of epithelial-mesenchymal FGF signaling. Developmental cell 11, 181–190 (2006).
Murashima-Suginami, A. et al. Rudiment incisors survive and erupt as supernumerary teeth as a result of USAG-1 abrogation. Biochemical and biophysical research communications 359, 549–555 (2007).
Mustonen, T. et al. Stimulation of ectodermal organ development by Ectodysplasin-A1. Developmental biology 259, 123–136 (2003).
Nakamura, T. et al. Transcription factor epiprofin is essential for tooth morphogenesis by regulating epithelial cell fate and tooth number. The Journal of biological chemistry 283, 4825–4833 (2008).
Ohazama, A. et al. Lrp4 modulates extracellular integration of cell signaling pathways in development. PloS one 3, e4092 (2008).
Pispa, J. et al. Tooth patterning and enamel formation can be manipulated by misexpression of TNF receptor Edar. Developmental dynamics 231, 432–440 (2004).
Tucker, A. S., Headon, D. J., Courtney, J. M., Overbeek, P. & Sharpe, P. T. The activation level of the TNF family receptor, Edar, determines cusp number and tooth number during tooth development. Developmental biology 268, 185–194 (2004).
Zhang, Q. et al. Loss of the Tg737 protein results in skeletal patterning defects. Developmental dynamics 227, 78–90 (2003).
Zhang, Z., Lan, Y., Chai, Y. & Jiang, R. Antagonistic actions of Msx1 and Osr2 pattern mammalian teeth into a single row. Science 323, 1232–1234 (2009).
Zhou, P., Byrne, C., Jacobs, J. & Fuchs, E. Lymphoid enhancer factor 1 directs hair follicle patterning and epithelial cell fate. Genes & development 9, 700–713 (1995).
Kiso, H. et al. Interactions between BMP-7 and USAG-1 (uterine sensitization-associated gene-1) regulate supernumerary organ formations. PloS one 9, e96938 (2014).
Murashima-Suginami, A. et al. Enhanced BMP signaling results in supernumerary tooth formation in USAG-1 deficient mouse. Biochemical and biophysical research communications 369, 1012–1016 (2008).
Togo, Y. et al. Antagonistic Functions of USAG-1 and RUNX2 during Tooth Development. PloS one 11, e0161067 (2016).
Savage, T. et al. Mandibular phenotype of p20C/EBPbeta transgenic mice: Reduced alveolar bone mass and site-specific dentin dysplasia. Bone 39, 552–564 (2006).
Chen, S. et al. Spatial expression of Cbfa1/Runx2 isoforms in teeth and characterization of binding sites in the DSPP gene. Connective tissue research 43, 338–344 (2002).
D’Souza, R. N. et al. Cbfa1 is required for epithelial-mesenchymal interactions regulating tooth development in mice. Development 126, 2911–2920 (1999).
Jiang, H. et al. Expression of core binding factor Osf2/Cbfa-1 and bone sialoprotein in tooth development. Mechanisms of development 81, 169–173 (1999).
Kobayashi, I. et al. Type II/III Runx2/Cbfa1 is required for tooth germ development. Bone 38, 836–844 (2006).
Komori, T. et al. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell 89, 755–764 (1997).
Mundlos, S. et al. Mutations involving the transcription factor CBFA1 cause cleidocranial dysplasia. Cell 89, 773–779 (1997).
Aberg, T. et al. Phenotypic changes in dentition of Runx2 homozygote-null mutant mice. The journal of histochemistry and cytochemistry 52, 131–139 (2004).
Wang, X. P. et al. Runx2 (Cbfa1) inhibits Shh signaling in the lower but not upper molars of mouse embryos and prevents the budding of putative successional teeth. Journal of dental research 84, 138–143 (2005).
Huang, B. et al. Prospective signs of cleidocranial dysplasia in Cebpb deficiency. Journal of biomedical science 21, 44 (2014).
Hirata, M. et al. C/EBPbeta and RUNX2 cooperate to degrade cartilage with MMP-13 as the target and HIF-2α as the inducer in chondrocytes. Human molecular genetics 21, 1111–1123 (2012).
Juuri, E. et al. Sox2+ stem cells contribute to all epithelial lineages of the tooth via Sfrp5+ progenitors. Developmental cell 23, 317–328 (2012).
Moreno-Bueno, G., Portillo, F. & Cano, A. Transcriptional regulation of cell polarity in EMT and cancer. Oncogene 27, 6958–6969 (2008).
Otsu, K., Kishigami, R., Fujiwara, N., Ishizeki, K. & Harada, H. Functional role of Rho-kinase in ameloblast differentiation. Journal of cellular physiology 226, 2527–2534 (2011).
Li, J. et al. BMP-SHH signaling network controls epithelial stem cell fate via regulation of its niche in the developing tooth. Developmental cell 33, 125–135 (2015).
Dingwall, M., Marchildon, F., Gunanayagam, A., Louis, C. S. & Wiper-Bergeron, N. Retinoic acid-induced Smad3 expression is required for the induction of osteoblastogenesis of mesenchymal stem cells. Differentiation 82, 57–65 (2011).
Nerlov, C. C/EBPs: recipients of extracellular signals through proteome modulation. Current opinion in cell biology 20, 180–185 (2008).
Huang, X., Bringas, P. Jr., Slavkin, H. C. & Chai, Y. Fate of HERS during tooth root development. Developmental biology 334, 22–30 (2009).
Yang, Z. et al. Cessation of epithelial Bmp signaling switches the differentiation of crown epithelia to the root lineage in a beta-catenin-dependent manner. Molecular and cellular biology 33, 4732–4744 (2013).
Ravn, J. J. Aplasia, supernumerary teeth and fused teeth in the primary dentition. An epidemiologic study. Scandinavian journal of dental research 79, 1–6 (1971).
López-Otín, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217 (2013).
Inomata, K. et al. Genotoxic stress abrogates renewal of melanocyte stem cells by triggering their differentiation. Cell 137, 1088–1099 (2009).
Akimoto, T. et al. Establishment of Hertwig’s epithelial root sheath cell line from cells involved in epithelial-mesenchymal transition. Biochemical and biophysical research communications 404, 308–312 (2011).
Chen, J. et al. TGF-β1 and FGF2 stimulate the epithelial-mesenchymal transition of HERS cells through a MEK-dependent mechanism. Journal of cellular physiology 229, 1647–1659 (2014).
Lee, J. H. et al. Upregulation of GM-CSF by TGF-beta1 in epithelial mesenchymal transition of human HERS/ERM cells. In vitro cellular & developmental biology animal 50, 399–405 (2014).
Yokohama-Tamaki, T. et al. Cessation of Fgf10 signaling, resulting in a defective dental epithelial stem cell compartment, leads to the transition from crown to root formation. Development 133, 1359–1366 (2006).
Xiong, J., Mrozik, K., Gronthos, S. & Bartold, P. M. Epithelial cell rests of Malassez contain unique stem cell populations capable of undergoing epithelial-mesenchymal transition. Stem cells and development 21, 2012–2025 (2012).
Lee, K. S. et al. Runx2 is a common target of transforming growth factor beta1 and bone morphogenetic protein 2, and cooperation between Runx2 and Smad5 induces osteoblast-specific gene expression in the pluripotent mesenchymal precursor cell line C2C12. Molecular and cellular biology 20, 8783–8792 (2000).
Nerlov, C. The C/EBP family of transcription factors: a paradigm for interaction between gene expression and proliferation control. Trends in cell biology 17, 318–324 (2007).
Kong, D., Li, Y., Wang, Z. & Sarkar, F. H. Cancer Stem Cells and Epithelial-to-Mesenchymal Transition (EMT)-Phenotypic Cells: Are They Cousins or Twins? Cancers 3, 716–729 (2011).
Mani, S. A. et al. The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133, 704–715 (2008).
Zhang, L. et al. BMP signaling and its paradoxical effects in tumorigenesis and dissemination. Oncotarget 7, 78206–78218 (2016).
Goldberg, M. et al. Targeted disruption of two small leucine-rich proteoglycans, biglycan and decorin, excerpts divergent effects on enamel and dentin formation. Calcified tissue international 77, 297–310 (2005).
Matsuura, T., Duarte, W. R., Cheng, H., Uzawa, K. & Yamauchi, M. Differential expression of decorin and biglycan genes during mouse tooth development. Matrix biology 20, 367–373 (2001).
Haruyama, N. et al. Genetic evidence for key roles of decorin and biglycan in dentin mineralization. Matrix biology 28, 129–136 (2009).
Chen, X. D., Fisher, L. W., Robey, P. G. & Young, M. F. The small leucine-rich proteoglycan biglycan modulates BMP4-induced osteoblast differentiation. FASEB J 18, 948–958 (2004).
Iozzo, R. V. & Sanderson, R. D. Proteoglycans in cancer biology, tumour microenvironment and angiogenesis. Journal of cellular and molecular medicine 15, 1013–1031 (2011).
Neill, T., Schaefer, L. & Iozzo, R. V. Decorin: a guardian from the matrix. The American journal of pathology 181, 380–387 (2012).
Mochida, Y. et al. Decorin modulates collagen matrix assembly and mineralization. Matrix biology 28, 44–52 (2009).
Thesleff, I. The genetic basis of tooth development and dental defects. American journal of medical genetics. Part A. 140, 2530–2535 (2006).
Diep, C. Q. et al. Identification of adult nephron progenitors capable of kidney regeneration in zebrafish. Nature 470, 95–100 (2011).
Yoshida, S., Kato, T. & Kato, Y. EMT Involved in Migration of Stem/Progenitor Cells for Pituitary Development and Regeneration. Journal of clinical medicine 5, 43 (2016).
Juuri, E., Isaksson, S., Jussila, M., Heikinheimo, K. & Thesleff, I. Expression of the stem cell marker, SOX2, in ameloblastoma and dental epithelium. European journal of oral sciences 121, 509–516 (2013).
Tanaka, T. et al. Targeted disruption of the NF-IL6 gene discloses its essential role in bacteria killing and tumor cytotoxicity by macrophages. Cell 80, 353–361 (1995).
Ito, H. et al. Three-dimensional computed tomography analysis of non-osteoarthritic adult acetabular dysplasia. Skeletal radiology 38, 131–139 (2009).
Kaukua, N. et al. Glial origin of mesenchymal stem cells in a tooth model system. Nature 513, 551–554 (2014).
Acknowledgements
This study was supported by Grants-in-Aid for Scientific Research(C) Nos 22592213 and 25463081. Cebpb+/− mice were a kind gift from Dr. Shizuo Akira of Osaka University.
Author information
Authors and Affiliations
Contributions
K. Takahashi and M. Sugai designed the research plan. K. Saito preformed all the experiments. Analysis and interpretation of data were performed by Boyen Huang, Masakazu Asahara, Honoka Kiso, Yumiko Togo, Hiroko Tsukamoto, Sayaka Mishima, Masaki Nagata, Machiko Iida, Yoshihito Tokita, Masato Asai, Akira Shimizu, Toshihisa Komori, Hidemitsu Harada, Mary MacDougall and Kazuhisa Bessho. K. Saito and K. Takahashi wrote the main manuscript text. K. Saito and Masato Asai prepared all figures. All authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saito, K., Takahashi, K., Huang, B. et al. Loss of Stemness, EMT, and Supernumerary Tooth Formation in Cebpb−/−Runx2+/− Murine Incisors. Sci Rep 8, 5169 (2018). https://doi.org/10.1038/s41598-018-23515-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-23515-y
This article is cited by
-
Decoding cellular plasticity and niche regulation of limbal stem cells during corneal wound healing
Stem Cell Research & Therapy (2024)
-
Tooth number abnormality: from bench to bedside
International Journal of Oral Science (2023)
-
Local application of Usag-1 siRNA can promote tooth regeneration in Runx2-deficient mice
Scientific Reports (2021)
-
Odontogenesis-related candidate genes involved in variations of permanent teeth size
Clinical Oral Investigations (2021)
-
Expression dynamics of integrin α2, α3, and αV upon osteogenic differentiation of human mesenchymal stem cells
Stem Cell Research & Therapy (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.