Abstract
Phytophthora capsici is a devastating oomycete that affects solanaceous, cucurbitaceous, fabaceous, and other crops in the United States (US) and worldwide. The release of the P. capsici genome allows for design of robust markers for genetic studies. We identified and characterized microsatellites in the P. capsici transcriptome. A subset of 50 microsatellites were assayed in a diverse set of P. capsici isolates and evaluated for polymorphism. Polymorphic microsatellites were confirmed by fragment analysis, and 12 were used for population characterization of 50 P. capsici isolates from different states, hosts, and mating types. Analysis of genetic relationship among isolates revealed significant geographic structure by state. Our findings highlight the usefulness of these 12 microsatellites to characterize the population structure of P. capsici and potential transferability to closely-related Phytophthora spp. since markers are located in coding regions. Our markers will facilitate genetic characterization and complement phenotypic studies of P. capsici populations, which may assist in deployment of disease management strategies.
Similar content being viewed by others
Introduction
The oomycete Phytophthora capsici Leonian is a devastating hemibiotrophic pathogen that causes severe epidemics over a broad host range of crops, including peppers, cucurbits, tomatoes, snap beans, eggplants, and many other plant species in the United States (US) and worldwide1,2,3,4,5,6. P. capsici can produce numerous symptoms depending on the host, including foliar blighting, damping-off, wilting, and root, stem, and fruit rot3,4. In the field, disease management relies primarily on the application of fungicides, in combination with cultural practices that are unfavorable for disease development such as raised beds, plastic mulch, and drip irrigation3. However, the broad host range of P. capsici and its capability to undergo sexual recombination and produce thick-walled oospores as persistent survival structures7 has limited the efficacy of control strategies such as crop rotation8,9. Sexual recombination between the A1 and A2 mating types of this heterothallic oomycete can result in novel genotypes with increased virulence, pathogenicity, and the ability to overcome host resistance and fungicides3. P. capsici sporangia are infrequently dispersed among fields by wind10. Instead, surface water sources for irrigation and movement of infected plant material or infested soil are known to be key factors in local spread of P. capsici and in the development of epidemics11. Because of limited long-distance dispersal of P. capsici and its ability to quickly overcome resistant varieties and fungicides due to frequent sexual reproduction, a better understanding of regional pathogen population structure would allow for more effective deployment of resistant varieties and breeding for durable host resistance12.
The amount and distribution of genetic diversity in P. capsici populations have been studied at a global scale within countries8,12,13,14,15 and at a local scale within states in the US10,16,17,18,19. Studies employed a wide range of molecular markers such as random amplified polymorphic DNA (RAPD)15,19,20, amplified fragment length polymorphism (AFLP)10,21,22, and single nucleotide polymorphism (SNP)8,13. Nonetheless, despite the benefits offered by microsatellites for population analyses23, only a few useful microsatellite loci have been developed to assess genetic diversity on P. capsici population studies14,18. Microsatellites, also known as simple sequence repeats (SSRs), are short DNA sequences consisting of tandemly repeated units, generally 1–6 base pairs in length24,25. Microsatellites can be found in either non-coding or coding regions and are highly polymorphic, making them desirable for population genetic analyses26,27. Microsatellites have been widely used in recent studies as a marker for diagnostics, population structure assessments, and epidemiological studies in oomycetes28,29,30,31,32,33. Genetic diversity in P. capsici has also been determined by microsatellites14,16,18,34,35, however, most of these studies relied on a set of microsatellites designed from a database of expressed sequence tags (ESTs)14,18. To date no microsatellite markers have been developed from the current P. capsici genome assembly, which could offer a set of standardized markers to be used by the scientific community35.
Traditionally, the methodology for microsatellite mining involved enrichment of genomic DNA libraries for a few targeted microsatellite motifs, followed by screening and sequencing of clones through Sanger sequencing. The entire process is labor intensive, resource consuming, and usually yields a small number of polymorphic markers36,37. The publication of the P. capsici genome, predicted transcriptome38, and the availability of microsatellite search tools, such as MISA (MIcroSAtellite identification tool)39, allow mining for microsatellites from whole genomes or transcriptomes. Identifying microsatellites in transcript sequences implies the identification of polymorphisms within coding regions, which represent annotated markers located in genes that may play important roles in pathogen virulence and survival40. This approach has been successfully applied to identify microsatellites in an increasing number of species, including Phytophthora species such as P. nicotianae28, P. ramorum41,42, P. sojae41,42, P. infestans42, P. plurivora32, P. multivora32, P. pini32, and plant pathogens such as Alternaria brassicicola43, Gaeumannomyces graminis44, Erysiphe necator45, Fusarium verticillioides46, F. oxysporum47, Anisogramma anomala48, and more recently Synchytrium endobioticum49. To date, development of microsatellite markers from transcript sequences in P. capsici has been unreported, despite the advantage of such studies and the economic importance of the pathogen. We aimed to: (i) identify and characterize microsatellites from the publicly available transcriptome of P. capsici using an in silico approach; (ii) determine microsatellite polymorphism in a diverse selection of P. capsici isolates and evaluate them in a population study of 50 isolates; and (iii) compare the distribution of microsatellites in the P. capsici transcriptome with the transcriptomes of two other plant pathogenic oomycetes (P. sojae and P. ramorum). The characterized polymorphic microsatellites in this study will help in understanding regional pathogen population structure, which will result in improved disease management strategies.
Results
Microsatellite identification and analysis
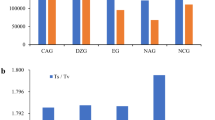
The in silico search for microsatellites using MISA (MIcroSAtellite identification tool) examined 20.36 Mb of the transcriptome of P. capsici (v. 1.1). 8.17% of the sequences contained microsatellites (Supplementary Table S3). A total of 1,855 microsatellites were identified and among those 75 were found more than once in a single gene. Trinucleotide repeats represented the most prevalent motif length in transcript sequences across the species (Table 2). Among the different types of repeats in P. capsici, trinucleotide repeats were the most common, accounting for 71.00% of all repeats, followed by tetranucleotides (13.98%), dinucleotides (9.33%), hexanucleotides (4.53%), and pentanucleiotides (1.62%). The P. capsici transcriptome is smaller than P. ramorum, and P. sojae, and it contains less microsatellites. The amount of microsatellites among these Phytophthora species seems to be highly correlated with their genome size (Pearson, r = 0.989, P > 0.05 true correlation is not equal to 0). When accounting for the differences in length of examined sequences between species, the P. capsici transcriptome exhibited the lowest relative abundance (microsatellite/Mb) and relative density (bp/Mb) (91.09 microsatellite/Mb and 1280.05 bp/Mb respectively), compared to P. ramorum (145.75 microsatellite/Mb, 2181.37 bp/Mb, P < 0.05) and P. sojae (196.04 microsatellite/Mb, 3190.98 bp/Mb, P < 0.05) (Supplementary Table S3). Overall, the P. capsici transcriptome consists of significantly fewer microsatellites compared to the other Phytophthora species.
The frequency, relative abundance, and relative density of each repeat unit in the P. capsici, P. ramorum, and P. sojae transcriptomes is presented in Table 2. Results indicated high similarity among the frequencies, relative abundances, and densities of the same repeat unit across the three species. For all three transcriptomes, the trinucleotide microsatellites were the most common motif length corresponding to around 70.00% of the microsatellites. P. capsici has higher frequency, relative abundance and density of tetranucleotide microsatellites when compared to P. ramorum. P. sojae has the highest relative abundance and density of all microsatellite motifs when compared to P. capsici and P. ramorum.
The frequency of the most abundant microsatellite motifs in the three Phytophthora species is shown in supplementary figure S1, each species harbored a predominant set of microsatellite motifs. The primary dinucleotide motif in P. capsici was AC/GT whereas CG/CG was more common for P. ramorum and P. sojae. The main trinucleotide motif included AAG/CTT for P. capsici and AGC/CTG for P. ramorum and P. sojae. In concordance with dinucleotide and tetranucleotide motifs frequencies, P. capsici presents a different primary tetranucleotide motif (ACGG/CCGT) when compared with P. ramorum and P. sojae (AGCG/CGCT). Pentanucleotide motifs and hexanucleotides motifs showed the lowest frequencies (<1%) across the three species. In all species, regardless the motif, the microsatellites tended to repeat 3, 4, and 5 times more frequently, and very few microsatellites repeat >10 times (Supplementary Fig. 2).
Primer3 successfully designed 1,491 primers pairs (80.37%) to flank microsatellites in the P. capsici transcriptome (Supplementary Table S4). Of the primers designed, 147, 1,015, 201, 21, and 59 belong to each of the motif lengths di-, tri-, tetra-, penta-, and hexa-nucleotide respectively. We randomly selected and synthesized 50 microsatellite primer pairs from 5 different motif lengths for agarose gel evaluation (Supplementary Table S2). Forty eight primers pairs (96.00%) successfully amplified and produced clear bands on 4% agarose gels. Thirty four (75.00%) out of 48 primer pairs yielded amplicons at the expected size. In total, 17 microsatellite primers exhibited polymorphism among the seven P. capsici isolates tested. Trinucleotide repeats yielded the highest number of polymorphic amplicons with nine primers exhibiting consistent polymorphic bands. Di-, Tetra-, Penta-, and Hexanucleotide primer pairs exhibited the lowest number of polymorphic bands. Figure 1 exemplifies allelic variation for Phyca_SSR14, Phyca_SSR20, and Phyca_SSR40 across 7 P. capsici isolates in 4% agarose gels. The fragment analysis profile for Phyca_SSR14 is presented for all tested isolates.
Amplification profile from 7 P. capsici isolates using the Phyca_SSR40, Phyca_SSR14, and Phyca_SSR20 primers. M: 100 bp marker, 1: NC19385, 2: R328, 3: RCZ-11, 4: WLB-8, 5: LT1534, 6: 12889, 7: SP98, and W: water. PCR products were resolved in 4% agarose gel. Fragment analysis profile for Phyca_SSR14 across the 7 P. capsici isolates, ordered as in the gel. The gels are cropped from different parts of different gels. Full-length gels are included in supplementary Figs S6, S7, and S8.
Population analysis with microsatellites
Among the 17 microsatellites considered polymorphic in the agarose gels, we selected the 12 top performing based on consistent amplification and clear polymorphism in their agarose profile for further examination via fragment analysis in a broader panel of isolates. The microsatellite primers selected were used to assess the genetic relationship of 50 P. capsici isolates from different states in the US, exhibiting both mating types, and collected from different hosts (Table 1).
The number of alleles per locus in the screened panel of isolates were, on average, 3.42 and ranged from 2 to 6 alleles. Heterozygosity varied from 0.38 to 0.79, with an average of 0.55. The evenness of alleles at each locus ranged from 0.63 to 1.0, with the mean evenness of 0.83 (Table 3). The panel of 50 P. capsici isolates were distributed across 50 multilocus genotypes. The genotype accumulation curve established 11 microsatellite markers as the minimum number of loci necessary to discriminate between the 50 P. capsici isolates (Supplementary Fig. S3). After clone correction, the standardized index of association (\({\bar{r}}_{d}\)) for all isolates (N = 50) was 0.019, which falls outside of the distribution expected under no linkage. The p value of 0.002 indicates significant support for rejecting the hypothesis that alleles are unlinked across loci, suggesting that the isolates used in this study are not recombining (Supplementary Fig. S4). Shannon-Wiener’s index was 3.912 indicating high genetic diversity in the panel of P. capsici isolates (N = 50). NC isolates showed the highest genetic diversity with Shannon-Weiner’s index of 3.091.
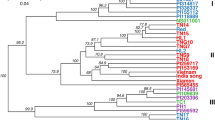
Analysis of genetic relationship among isolates using Provesti’s distance exhibited significant bootstrap support to separate the main group of isolates from North Carolina, which were collected from different fields across the state (Fig. 2). The other group comprised isolates from SC, TN, NY, GA, and MI. The genetic distance tree grouped some SC (WLB-232 and WLB-150) and NY (6180 and 0664-1) isolates closely with significant bootstrap support (>70%). Overall, the genetic distance tree separated isolates into two clusters, one with mainly NC isolates and the other with isolates from other states (Fig. 2). Population differentiation was revealed using pre-defined populations by states in the Discriminant Analysis of Principal Components (DAPC) (Fig. 3). DAPC is a multivariate statistical approach that partitions variance in the sample into a between-group and within-group component, in an effort to maximize discrimination between groups. P. capsici isolates from TN, NY, GA, SC, and MI overlapped in the first principal component, while NC isolates clustered separately from the other isolates. TN, NY, GA, and MI separated from SC in the second principal component (Fig. 3). The separation of NC isolates in the DAPC analysis supports the grouping observed in the Provesti’s genetic distance tree. Results from Analysis of Molecular Variance (AMOVA) revealed significant levels of differentiation among states and host (P < 0.005). Across states, nearly 13% of variation was attributed to differences among states, while more than 16% was attributed to variation between isolates within states, and 69% was attributed to variation within isolates. Only 6.38% of variation between isolates was attributed to differences due to host. Pairwise comparison (FST) between states also showed significant differentiation between P. capsici isolates from NC and all other states included in this study (Table 4). P. capsici isolates from TN, NY, and GA exhibited low levels of genetic differentiation between themselves, which confirms overlap observed in the DAPC analysis. MI isolates showed moderate and high levels of differentiation when compared with NC, SC, and NY (P < 0.005) (Table 4). The STRUCTURE analysis based on the Bayesian clustering method established the presence of two population clusters, Group 1 and Group 2 (Fig. 4). The delta K plot indicated a clear peak at K = 2 (Supplementary Fig. S5). Twenty one P. capsici isolates included here from NC fields were assigned to Group 1 with membership probabilities higher than 0.82 (Fig. 4). P. capsici isolates from TN, NY, GA, SC, and MI were allocated in Group 2 and unable to resolve by geography. The separation of NC isolates from TN, NY, GA, SC, and MI in the STRUCTURE analysis supports the clade identified in the genetic distance tree and the clustering observed in the DAPC analysis (Figs 2 and 3). None of the population structure analyses (Provesti’s distance tree, DAPC, or Structure) clustered the 50 P. capsici isolates by host or mating type.
UPGMA tree with 1,000 bootstrap replicates based on Provesti’s distance for 50 P. capsici isolates. Tip labels define the isolate information by isolate name, state (NC: North Carolina, SC: South Carolina, NY: New York, GA: Georgia, TN: Tennessee, and MI: Michigan), and host. Branches with bootstrap values greater than 50% are displayed.
Discussion
We exploited the P. capsici transcriptome to successfully identify 1,855 microsatellite loci using an in silico approach. The number of microsatellites detected in P. capsici is larger than those discovered in P. plurivora, P. multivora, P. pini, P. citricola, or P. alni using the 454-pyrosequencing-based method32,50,51. Our P. ramorum and P. sojae transcriptome survey and a P. nicotiana whole genome survey yielded higher numbers of microsatellites compared to what the P. capsici transcriptome yielded28. Relative abundance and relative density of microsatellites in P. capsici differs greatly to the observed values for P. ramorum and P. sojae; we hypothesize that such differences may be explained by the differences among genome sequencing approaches taken for these three species38,52. In fact, our results suggest high similarity among the frequencies, relative abundances, and densities of the same repeat unit across the three species. In the different transcriptome sequences analyzed here, trinucleotide microsatellites were the most common motif length, consistent with studies in other Phytophthora spp.28,41, fungal plant pathogens43,47,53, and plants54. The same studies also report lower abundance of di-, tetra-, and pentanucleiotides. Purifying selection for motifs that preserve reading frames explains the predominance of trinucleotide microsatellites in coding regions55.
We designed primer pairs for more than 80% of the identified microsatellite loci. Our results reveal information about the reliability and quality of the genome assembly. Garnica, et al.41 used transcriptome sequences of P. ramorum and P. sojae and had similar primer design success rate, greater than 80%, which is comparable with our findings. Insufficient flanking sequence adjacent to the microsatellite, such as a microsatellite close to the end of a contig, may explain the 20% primer design failure48. Gel electrophoresis screening of a set of 50 microsatellite primers in seven P. capsici isolates yielded 17 highly polymorphic microsatellite markers. We observed higher amplification efficiency (96.00%) than reported by Wang and Chilvers53 (85.00%) and Gagnon, et al.49 (79.03%) who examined whole genome sequences of F. virguliforme and S. endobioticum respectively. Amplification efficiency in our microsatellite primer screening might be attributed to the stability of coding regions with low selection pressure56. We identified a high number of polymorphic bands from trinucleotide microsatellites. Di-, tetra-, penta- and hexa-nucleotide microsatellites yielded the lowest number of polymorphic bands. The relatively low polymorphism implies that these repeat types are highly conserved across isolates used in this study in contrast with trinucleotides repeats, which may experience mutation and selection pressure for specific amino acids43. We further narrowed the list of microsatellites from 17 to 12 based on consistent amplification and polymorphism observed via gel electrophoresis, and confirmed the polymorphism via fragment length analysis.
Polymorphism indexes such as Nei’s expected heterozygosity (0.55) and evenness (0.83) indicated high potential for these markers to describe diversity in the panel of 50 P. capsici isolates. Our Nei’s expected heterozygosity values are similar to the ones reported by Gagnon, et al.49 and Wallace and Quesada-Ocampo57 when they characterized microsatellites for S. endobioticum and Pseudoperonospora cubensis respectively. In a recent study, Gagnon, et al.58 used microsatellites mined from whole genome sequence data of P. ramorum isolates to assess genetic diversity. The markers developed in their study had an average evenness of 0.943 and were able to detect temporal and spatial distribution of P. ramorum in Canadian nurseries. In our study, the MLG curve reached a plateau with 11 microsatellite markers out of 12, implying that these markers are sufficient to depict all 50 multilocus genotypes. The \({\bar{r}}_{d}\) calculated for all isolates after clone correction suggested non-recombination, which is probably due to the fact the our sampling contained isolates from different states and fields in the US. P. capsici populations are known to stratify by geography since long distance movement is unlikely12. The Shannon-Wiener’s index confirmed the presence of high genetic diversity in our sample of 50 P. capsici isolates. NC isolates displayed more genetic diversity than P. capsici populations reported from 12 different sites in four regions of New York State16. Sampling for NC isolates encompassed 7 different sites distributed in three regions of North Carolina. Our results for diversity measurements suggest that the 12 polymorphic microsatellites produced enough resolution for genotyping genetically distinct individuals.
As a first measure to detect population structure, we created a distance-based tree using Provesti’s distance59. The genetic distance tree detected some population structure based on geographic regions enabling us to differentiate isolates into two major clusters with 100% bootstrap support. One of the major clusters consisted of isolates mainly from NC while the other included isolates from all other states evaluated here. Such genetic differentiation can be explained by the fact that P. capsici exists in geographically isolated subpopulations in the US12,16. Our genetic distance tree was unable to separate isolates from SC, NY, TN, GA, and MI, therefore we performed a discriminatory analysis of principal components in order to infer the number of genetic clusters. Interestingly, the clustering of isolates based on the DAPC supported the hypothesis of the genetic distance tree. NC isolates collected across the state consistently formed a separate group relative to the other isolates in the DAPC. Unlike the genetic distance tree the relative position of SC isolates in the DAPC scatter plot suggests underlying population structure between SC and TN, NY, GA, and MI isolates. The clustering observed in the DAPC for NC and SC isolates showed significant genetic differentiation with FST values larger than 0.05 and in some cases larger than 0.25. Population differentiation statistics and the AMOVA analysis supported our hypothesis that the 12 microsatellite markers developed here are able to separate P. capsici isolates by geographic origin. Even though the AMOVA by host is significant, it only represents 6.38% of the variation between isolates. Two major clusters were detected in the STRUCTURE analysis using the admixture model. We observed some resemblance between the two major groups in the genetic distance tree and the two clusters revealed by STRUCTURE. All NC isolates clustered together in group 1 within the STRUCTURE analysis, supporting our hypothesis that the 12 microsatellite markers are robust and can differentiate isolates by state. We attribute the unresolved group 2 in the STRUCTURE analysis to the low number of isolates per field from SC, NY, TN, GA, and MI. Successful management of P. capsici requires a detailed knowledge of regional pathogen population structure and distribution3. Conducting surveys of local P. capsici populations and examining how they stratify can direct the deployment of management strategies in a more efficient way. The 12 microsatellite markers discovered here represent powerful and inexpensive tools for describing the population structure of P. capsici at a regional scale.
P. capsici is considered a pathogen with high evolutionary potential, because sexual reproduction increases the probability of developing genotypes that can overcome host resistance or are resistant to fungicides9. These novel genotypes could be leading to new epidemics that become isolated due to the soilborne dispersal mechanism of the pathogen3, and therefore understanding the phenotypic variation and genotypic structure in a particular region becomes highly desirable. Monitoring the genetic structure of the pathogen along with phenotypic monitoring of fungicide resistance, mating type, and virulence, allow for strategizing management. For example, if a genetic cluster was found to be more virulent to a particular host, it would be valuable to know the geographic location of isolates belonging to that genetic cluster so that one avoids planting a susceptible crop in that area60. Microsatellites as described by Cooke and Lees23 represent a powerful molecular marker for genetic analysis; they are affordable and reproducible among research groups. Even though genomic approaches are available now such as genotyping by sequencing, the P. infestans community still relies on microsatellite markers to understand the pathogen biology and its population dynamics61,62,63. In fact, Dunn, et al.16 emphasized the need for additional polymorphic markers to increase resolution and detect greater diversity in populations of P. capsici. Our study delivers a set of highly polymorphic microsatellite markers to the P. capsici community: these 12 microsatellites represent valuable tools to understand the structure and diversity of P. capsici populations, allowing for comparisons of datasets generated by different research groups. We foresee an additional application for our microsatellites as transferable markers across species complexes. Quesada-Ocampo, et al.12 used SNPs from several genomic regions to separate P. capsici and P. tropicalis in a STRUCTURE analysis, however, their SNP markers were unable to resolve an intermediate cluster containing both species. Microsatellites developed from coding regions are highly transferable to closely related species43,64,65. Since P. capsici is part of a species complex, we hypothesize that our microsatellites can distinguish among closely related species such as P. tropicalis and P. glovera. Further experiments should be completed to validate this hypothesis.
Taken together, our findings demonstrate that 12 microsatellites developed from the transcriptome sequences of P. capsici are highly polymorphic in a diverse panel of isolates, describe diversity of local populations, and may be transferable to species closely related to P. capsici. These microsatellites can be used to analyze the genetic structure of particular geographic regions and determine genotypes present in fields. The information generated by population studies using these markers may allow for better deployment of resistant cultivars if a correspondence between genetic cluster and virulence is found for a particular host. We are interested in using these markers to describe the until now unknown population structure of P. capsici in North Carolina and combining that knowledge with phenotypic information regarding fungicide resistance, mating type, and virulence.
Materials and Methods
Identification and analysis of microsatellites in the predicted transcriptomes of P. capsici, P. sojae, and P. ramorum
Predicted transcriptomes from genome sequences of isolates LT1534 of P. capsici38, Pr-102 of P. ramorum52, and P6497 of P. sojae52 were downloaded as FASTA files from publically available databases at http://genome.jgi.doe.gov/Phyca11/Phyca11.home.html, http://genome.jgi.doe.gov/ramorum1/ramorum1.home.html, and http://genome.jgi.doe.gov/Physo3/Physo3.home.html. The transcriptome data was searched for microsatellite motifs of one to six nucleotides in length using the program MISA (MIcroSAtellite identification tool)39. Search criteria were set for identification of at least 10 repeat units for mono-nucleotides, six for di-nucleotides and five for tri-, tetra-, penta-, or hexa-nucleotides. Results were compared among species for abundance, frequency, distribution, and relative density of each repeat unit in the transcriptomes as described previously46.
Microsatellite marker development
The MISA output files and Perl programming were used to extract the coordinates of identified microsatellites in the P. capsici transcriptome and generate input files for designing primers with the Primer3 program66. Primer3 identified primer pairs flanking each microsatellite loci with a melting temperature between 57 °C and 63 °C with an optimum at 60 °C, GC content between 20 and 80% with an optimum of 50%, and PCR products with expected length between 100 and 300 bp. The M13 sequence (5′-GACGGCCAGT-3′) was added at the 5′ end of each forward primer in order to allow for later fluorescent product labeling and fragment analysis as previously described67.
Isolates, culture conditions, and DNA extraction
A panel of seven single-spore P. capsici isolates were selected for primer validation via gel electrophoresis (Supplementary Table S1). This panel included one isolate from North Carolina (NC19385); one from New Jersey (R328); two from South Carolina (RCZ-11, WLB-8); two from Michigan (12889, SP98); and the isolate used in the P. capsici genome sequencing project (LT1534). All isolates were transferred from long term storage cultures to V8 juice broth as described previously12,68. Mycelia were harvested through filtration, immediately lyophilized, and stored at −80 °C.
Genomic DNA was isolated using the following phenol-chloroform protocol: for each isolate, 200 mg of lyophilized mycelium were ground on an OMNI Bead Ruptor 24 (Omni International, Kennesaw, GA) at a speed of 5 m/s for 60 s, adding 2 mm glass beads. Subsequently, 600 µl of extraction buffer (0.5 M Tris pH 8.0, 5 M NaCl, 0.5 M EDTA, 10% SDS) was added to the powdered tissue and homogenized by vortexing. Samples were treated with 4 µl of RNAase A (Invitrogen Life Technologies, Grand Island, NY) and incubated for 10 min at room temperature with occasional gentle mixing. 500 µl of phenol was added, and the mixture was centrifuged for 10 min at 21,130 g. The supernatant was transferred to a fresh tube and an equal volume of Phenol:Chloroform was added and mixed by gentle inversion; the mixture was centrifuged for 10 min at 21,130 g. The aqueous supernatant was transferred to a new tube and 0.8 volume of Isopropanol was added. Tubes were incubated at −20 °C overnight. After incubation, all samples were centrifuged for 15 min at 21,130 g, the supernatant was discarded and the pellet was precipitated with 80% Ethanol. The pellet was dissolved in 200 µl of sterile water. Quality and integrity of genomic DNA were estimated by measuring the 260/280 nm ratio in a NanoDrop ND-1000 spectrophotometer (Thermo Scientific, Wilmington, DE) and by electrophoresis in a 1% agarose gel in 0.5 × Tris-borate-EDTA buffer stained with ethidium bromide (5 µg/ml) for visualization.
Microsatellite evaluation
Ten microsatellite primer pairs each from di-, tri-, tetra-, penta-, and hexa-nucleotide motif length were randomly selected from the Primer3 output to detect polymorphism among 7 P. capsici isolates (Supplementary Table S2). Microsatellite primers selected were synthesized by Integrated DNA Technologies (Coralville, IA). Polymerase chain reactions (PCR) were executed in a T100 thermocycler (Bio-Rad, Hercules, CA). 20 µl of reaction volume contained 1 µl of genomic DNA at 50 ng/µl, 10 µl of GoTaq® Green Master Mix (2X GoTaq Green Master consisting of GoTaq Green Reaction Buffer, 400 µM of each dNTP, and 3 mM MgCl2; Promega, Madison, WI), 1 µl of each 10 µM primer, and 7 µl of sterile water. PCR cycling conditions included an initial denaturation at 94 °C for 3 min; followed by 35 cycles of denaturation at 94 °C for 30 s, annealing at 53 °C for 30 s and extension at 72 °C for 30 s; with a final extension at 72 °C for 5 min. PCR products were analyzed for polymorphism by electrophoresis in 4% agarose gels and visualized by ethidium bromide staining. A 100 bp DNA ladder (Invitrogen Life Technologies, Grand Island, NY) was used to estimate allele size.
Population analyses using the microsatellite markers
To determine the effectiveness of the microsatellite markers developed in this study for understanding population genetic structure and diversity, a subset of 12 microsatellite primers were selected on the basis of polymorphisms in agarose gels. These microsatellite markers were further analyzed across 50 P. capsici isolates from different states, hosts, and mating types via fragment analysis (Table 1). PCR products of the polymorphic microsatellite primers were subjected to a second round of PCR. In these PCRs the forward primer of each microsatellite was 5′ labelled with four fluorescent dyes (6-FAM, VIC, NED, and PET from Applied Biosystems, Foster City, CA). Prior to fragment analysis, 1 µl of PCR product from 4 microsatellite loci labeled with different fluorescent dyes were pooled together and diluted in ratio 1:100 with sterile dH2O. In order to accurately detect variability in microsatellite length among isolates, a genotyping reaction was performed suspending 1 µl of diluted PCR product in 8.5 µl HiDi Formamide and 0.5 µl of Gene Scan 600LIZ dye Size Standard (Applied Biosystems, Foster City, CA). Genotyping reactions were subsequently performed by the NCSU Genomic Science Laboratory (GSL, Raleigh, NC) on an ABI 3730xl DNA Analyzer (Applied Biosystems, Foster City, CA).
Allele sizes were called using the Microsatellite Plugin in Geneious version R9.0.5 (Biomatters, New Zealand). In order to reduce the risk of genotyping stutter peaks, we manually removed from the analysis peaks lower than 5% of the signal of the tallest peak at any given locus. We assumed two alleles to be present at each loci because P. capsici is a diploid organism belonging to the class Oomycota. Descriptive population statistics such as the number of alleles per locus (Na), heterozygosity (He), evenness, and the genotype accumulation curve were estimated using the functions “gac” and “locus_table” from the package poppr in RStudio (version 0.99.903 and R version 3.2.4; R Core Team, Austria)69,70,71.
We employed the function “poppr” using the stratified clonecorrected data (by states, host, and mating type) to calculate the number of multilocus genotypes, the Shannon-Wiener index of genetic diversity and other summary statistics. The standardized index of association (\({\bar{r}}_{d}\)) was calculated with the “ia” function72. We built a genetic distance tree to assess genetic relatedness between isolates using Provesti’s distance with 1,000 bootstrap replicates59. Because the set of microsatellite markers discovered in this study are intended to elucidate population structure of P. capsici, we assessed whether the 50 P. capsici isolates significantly structure by state, host, or mating type by implementing the Analysis of Molecular Variance (AMOVA) included in the poppr package. To test for differentiation between P. capsici isolates from different states, pairwise FST values were estimated, and significance levels were tested with 999 permutations using GenAlEx 6.573. The presence of an underlining structure was examined using the Discriminant Analysis of Principal Components (DAPC) with the R package adegenet74, and the Bayesian Markov Chain Monte Carlo (MCMC) clustering model using the software STRUCTURE v2.3.475. The values for length of burnin, chain replication, and lambda were set at 50,000, 250,000, and 1 respectively. Isolates membership was inferred for one to 10 clusters and the optimal K was chosen by computing ΔK using STRUCTURE HARVESTER v.0.6.9476. Population structure figure defining the state of origin was obtained using Microsoft Excel (version 15.20).
References
Foster, J. M. & Hausbeck, M. K. Resistance of pepper to Phytophthora crown, root, and fruit rot is affected by isolate virulence. Plant Disease 94, 24–30 (2010).
Foster, J. M., Naegele, R. P. & Hausbeck, M. K. Evaluation of eggplant rootstocks and pepper varieties for potential resistance to isolates of Phytophthora capsici from Michigan and New York. Plant Disease 97, 1037–1041 (2013).
Granke, L. L., Quesada-Ocampo, L. & Lamour, K. Advances in research on Phytophthora capsici on vegetable crops in the United States. Plant Disease 95, 1588–1600 (2012).
Lamour, K. H., Stam, R., Jupe, J. & Huitema, E. The oomycete broad-host-range pathogen Phytophthora capsici. Molecular plant pathology 13, 329–337, https://doi.org/10.1111/j.1364-3703.2011.00754.x (2012).
Naegele, R. P., Boyle, S., Quesada-Ocampo, L. M. & Hausbeck, M. K. Genetic diversity, population structure, and resistance to Phytophthora capsici of a worldwide collection of eggplant germplasm. PloS one 9, e95930, https://doi.org/10.1371/journal.pone.0095930 (2013).
Quesada-Ocampo, L. M. & Hausbeck, M. K. Resistance in tomato and wild relatives to crown and root rot caused by Phytophthora capsici. Phytopathology 100, 619–627 (2010).
Erwin, D. C. & Ribeiro, O. K. Phytophthora diseases worldwide. (American Phytopathological Society Press, 1996).
Gobena, D., Roig, J., Galmarini, C., Hulvey, J. & Lamour, K. Genetic diversity of Phytophthora capsici isolates from pepper and pumpkin in Argentina. Mycologia 104, 102–107, https://doi.org/10.3852/11-147 (2012).
McDonald, B. A. & Linde, C. Pathogen population genetics, evolutionary potential, and durable resistance. Annual review of phytopathology 40, 349–379, https://doi.org/10.1146/annurev.phyto.40.120501.101443 (2002).
Lamour, K. H. & Hausbeck, M. K. Investigating the spatiotemporal genetic gtructure of Phytophthora capsici in Michigan. Phytopathology 91, 973–980, https://doi.org/10.1094/PHYTO.2001.91.10.973 (2001).
Granke, L. L., Windstam, S. T., Hoch, H. C., Smart, C. D. & Hausbeck, M. K. Dispersal and movement mechanisms of Phytophthora capsici sporangia. Phytopathology 99, 1258–1264, https://doi.org/10.1094/PHYTO-99-11-1258 (2009).
Quesada-Ocampo, L. M., Granke, L. L., Mercier, M. R., Olsen, J. & Hausbeck, M. K. Investigating the genetic structure of Phytophthora capsici populations. Phytopathology 101, 1061–1073, https://doi.org/10.1094/PHYTO-11-10-0325 (2011).
Hulvey, J. et al. Genetic Diversity of the Pepper Pathogen Phytophthora capsici on Farms in the Amazonian High Jungle of Peru. American Journal of Plant Sciences 2, 461–466, https://doi.org/10.4236/ajps.2011.23054 (2011).
Meitz, J. C., Linde, C. C., Thompson, A., Langenhoven, S. & McLeod, A. Phytophthora capsici on vegetable hosts in South Africa: distribution, host range and genetic diversity. Australasian Plant Pathology 39, 431–439, https://doi.org/10.1071/AP09075 (2010).
Silvar, C., Merino, F. & Díaz, J. Diversity of Phytophthora capsici in Northwest Spain: analysis of virulence, metalaxyl response, and molecular characterization. Plant Disease 90, 1135–1142 (2006).
Dunn, A. et al. Population structure and resistance to mefenoxam of Phytophthora capsici in New York State. Plant Disease 94, 1461–1468, https://doi.org/10.1094/PDIS-03-10-0221 (2010).
French-Monar, R., Jones, J. & Roberts, P. Characterization of Phytophthora capsici associated with roots of weeds on Florida vegetable farms. Plant Disease 90, 345–350, https://doi.org/10.1094/PD-90-0345 (2006).
Wang, Z. et al. Development of an improved isolation approach and simple sequence repeat markers to characterize Phytophthora capsici populations in irrigation ponds in southern Georgia. Applied and environmental microbiology 75, 5467–5473, https://doi.org/10.1128/aem.00620-09 (2009).
Islam, S. Z., Babadoost, M., Lambert, K. N., Ndeme, A. & Fouly, H. M. Characterization of Phytophthora capsici Isolates from Processing Pumpkin in Illinois. Plant Disease 89, 191–197, https://doi.org/10.1094/PD-89-0191 (2005).
Sun, W. X., Jia, Y. J., O’Neill, N. R. & Feng, B. Z. H. Genetic diversity in Phytophthora capsici from eastern China. Canadian. Journal of Plant Pathology 30, 414–424, https://doi.org/10.1080/07060660809507539 (2008).
Bowers, J. H., Martin, F. N., Tooley, P. W. & Luz, E. Genetic and morphological diversity of temperate and tropical isolates of Phytophthora capsici. Phytopathology 97, 492–503 (2007).
Gevens, A. J., Donahoo, R. S. & Lamour, K. H. Characterization of Phytophthora capsici from Michigan surface irrigation water. Phytopathology 97, 421–428 (2007).
Cooke, D. E. L. & Lees, A. K. Markers, old and new, for examining Phytophthora infestans diversity. Plant Pathology 53, 692–704, https://doi.org/10.1111/j.1365-3059.2004.01104.x (2004).
Selkoe, K. A. & Toonen, R. J. Microsatellites for ecologists: a practical guide to using and evaluating microsatellite markers. Ecology Letters 9, 615–629, https://doi.org/10.1111/j.1461-0248.2006.00889.x (2006).
Ellegren, H. Microsatellites: simple sequences with complex evolution. Nature Reviews Genetics 5, 435–445, https://doi.org/10.1038/nrg1348 (2004).
Vukosavljev, M. et al. Efficient development of highly polymorphic microsatellite markers based on polymorphic repeats in transcriptome sequences of multiple individuals. Molecular Ecology Resources 15, 17–27, https://doi.org/10.1111/1755-0998.12289 (2014).
Zhou, Z. et al. Transcriptome sequencing of sea cucumber (Apostichopus japonicus) and the identification of gene‐associated markers. Molecular Ecology Resources 14, 127–138, https://doi.org/10.1111/1755-0998.12147 (2014).
Biasi, A., Martin, F. & Schena, L. Identification and validation of polymorphic microsatellite loci for the analysis of Phytophthora nicotianae populations. Journal of microbiological methods 110, 61–67, https://doi.org/10.1016/j.mimet.2015.01.012 (2015).
Li, Y., Cooke, D. E., Jacobsen, E. & van der Lee, T. Efficient multiplex simple sequence repeat genotyping of the oomycete plant pathogen Phytophthora infestans. Journal of microbiological methods 92, 316–322, https://doi.org/10.1016/j.mimet.2012.11.021 (2013).
Matasci, C. L., Jermini, M., Gobbin, D. & Gessler, C. Microsatellite based population structure of Plasmopara viticola at single vine scale. European Journal of Plant Pathology 127, 501–508, https://doi.org/10.1007/s10658-010-9615-z (2010).
Perumal, R. et al. Simple sequence repeat markers useful for sorghum downy mildew (Peronosclerospora sorghi) and related species. BMC genetics 9, 1–14, https://doi.org/10.1186/1471-2156-9-77 (2007).
Schoebel, C. N., Jung, E. & Prospero, S. Development of new polymorphic microsatellite markers for three closely related plant-pathogenic Phytophthora species using 454-pyrosequencing and their potential applications. Phytopathology 103, 1020–1027 (2013).
Yin, L., Zhang, Y., Hao, Y. & Lu, J. Genetic diversity and population structure of Plasmopara viticola in China. European Journal of Plant Pathology 140, 365–376, https://doi.org/10.1007/s10658-014-0470-1 (2014).
Hu, J. et al. Genetically diverse long-lived clonal lineages of Phytophthora capsici from pepper in Gansu, China. Phytopathology 103, 920–926 (2013).
Dunn, A. R., Bruening, S. R., Grünwald, N. J. & Smart, C. D. Evolution of an Experimental Population of Phytophthora capsici in the Field. Phytopathology 104, 1107–1117, https://doi.org/10.1094/PHYTO-12-13-0346-R (2014).
Dobrowolski, M. P., Tommerup, I. C., Blakeman, H. D. & O’Brien, P. A. Non-Mendelian inheritance revealed in a genetic analysis of sexual progeny of Phytophthora cinnamomi with microsatellite markers. Fungal genetics and biology: FG & B 35, 197–212, https://doi.org/10.1006/fgbi.2001.1319 (2002).
Prospero, S., Black, J. A. & Winton, L. M. Isolation and characterization of microsatellite markers in Phytophthora ramorum, the causal agent of sudden oak death. Molecular Ecology Notes 4, 672–674, https://doi.org/10.1111/j.1471-8286.2004.00778.x (2004).
Lamour, K. H. et al. Genome sequencing and mapping reveal loss of heterozygosity as a mechanism for rapid adaptation in the vegetable pathogen Phytophthora capsici. Molecular plant-microbe interactions: MPMI 25, 1350–1360, https://doi.org/10.1094/MPMI-02-12-0028-R (2012).
Thiel, T., Michalek, W., Varshney, R. & Graner, A. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theoretical and Applied Genetics 106, 411–422, https://doi.org/10.1007/s00122-002-1031-0 (2003).
Li, Y.-C. C., Korol, A. B., Fahima, T., Beiles, A. & Nevo, E. Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review. Molecular ecology 11, 2453–2465 (2002).
Garnica, D. P. et al. Survey and analysis of microsatellites from transcript sequences in Phytophthora species: frequency, distribution, and potential as markers for the genus. BMC genomics 7, 1–11, https://doi.org/10.1186/1471-2164-7-245 (2006).
Schena, L., Cardle, L. & Cooke, D. E. Use of genome sequence data in the design and testing of SSR markers for Phytophthora species. BMC genomics 9, 1–23, https://doi.org/10.1186/1471-2164-9-620 (2007).
Ruchi, S. et al. Identification and Characterization of Microsatellite from Alternaria brassicicola to Assess Cross-Species Transferability and Utility as a Diagnostic Marker. Molecular Biotechnology 56, 1049–1059, https://doi.org/10.1007/s12033-014-9784-7 (2014).
Li, W. et al. Analysis of simple sequence repeats in the Gaeumannomyces graminis var. tritici genome and the development of microsatellite markers. Current genetics 60, 237–245, https://doi.org/10.1007/s00294-014-0428-z (2014).
Frenkel, O. et al. Development of microsatellite markers from the transcriptome of Erysiphe necator for analysing population structure in North America and Europe. Plant Pathology 61, 106–119, https://doi.org/10.1111/j.1365-3059.2011.02502.x (2012).
Leyva-Madrigal, K. Y., Larralde-Corona, C. P. & Maldonado-Mendoza, C. L. C.-V. I. E. Genome distribution and validation of novel microsatellite markers of Fusarium verticillioides and their transferability to other Fusarium species. Journal of microbiological methods 101, 18–23, https://doi.org/10.1016/j.mimet.2014.03.011 (2014).
Mahfooz, S., Maurya, D. K., Srivastava, A. K., Kumar, S. & Arora, D. K. A comparative in silico analysis on frequency and distribution of microsatellites in coding regions of three formae speciales of Fusarium oxysporum and development of EST–SSR markers for polymorphism studies. FEMS Microbiology Letters 328, 54–60, https://doi.org/10.1111/j.1574-6968.2011.02483.x (2012).
Cai, G., Leadbetter, C. W., Muehlbauer, M. F., Molnar, T. J. & Hillman, B. I. Genome-Wide Microsatellite Identification in the Fungus Anisogramma anomala Using Illumina Sequencing and Genome Assembly. PLoS ONE 8, e82408, https://doi.org/10.1371/journal.pone.0082408 (2013).
Gagnon, M. C., van der Lee, T. A. J. & Bonants, P. J. M. Development of Polymorphic Microsatellite Loci for Potato Wart from Next-Generation Sequence Data. Phytopathology 106, 636–644 (2016).
Aguayo, J. et al. Strong genetic differentiation between North American and European populations of Phytophthora alni subsp. uniformis. Phytopathology 103, 190–199, https://doi.org/10.1094/PHYTO-05-12-0116-R (2013).
Abdelkrim, J., Robertson, B., Stanton, J. A. & Gemmell, N. Fast, cost-effective development of species-specific microsatellite markers by genomic sequencing. BioTechniques 46, 185–192, https://doi.org/10.2144/000113084 (2009).
Tyler, B. M. et al. Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science (New York, N.Y.) 313, 1261–1266, https://doi.org/10.1126/science.1128796 (2006).
Wang, J. & Chilvers, M. I. Development and characterization of microsatellite markers for Fusarium virguliforme and their utility within clade 2 of the Fusarium solani species complex. Fungal Ecology 20, 7–14 (2016).
Lawson, M. J. & Zhang, L. Distinct patterns of SSR distribution in the Arabidopsis thaliana and rice genomes. Genome biology 7, 1–11, https://doi.org/10.1186/gb-2006-7-2-r14 (2006).
Metzgar, D., Bytof, J. & Wills, C. Selection against frameshift mutations limits microsatellite expansion in coding DNA. Genome research 10, 72–80, https://doi.org/10.1101/gr.10.1.72 (2000).
Parida, S. K. et al. Informative genomic microsatellite markers for efficient genotyping applications in sugarcane. Theoretical and Applied Genetics 118, 327–338, https://doi.org/10.1007/s00122-008-0902-4 (2008).
Wallace, E. C. & Quesada-Ocampo, L. M. Analysis of microsatellites from the transcriptome of downy mildew pathogens and their application for characterization of Pseudoperonospora populations. PeerJ 5, e3266 (2017).
Gagnon, M. C. et al. Development and Validation of Polymorphic Microsatellite Loci for the NA2 Lineage of Phytophthora ramorum from Whole Genome Sequence Data. Plant Disease 101, 666–673 (2017).
Prevosti, A., Oca a, J. & Alonso, G. Distances between populations of Drosophila subobscura, based on chromosome arrangement frequencies. Theor Appl Genet 45, 231–241 (1975).
Granke, L. L., Quesada-Ocampo, L. M. & Hausbeck, M. K. Differences in virulence of Phytophthora capsici isolates from a worldwide collection on host fruits. European Journal of Plant Pathology 132, 281–296, https://doi.org/10.1007/s10658-011-9873-4 (2012).
Stroud, J. A., Shaw, D. S., Hale, M. D. & Steele, K. A. SSR assessment of Phytophthora infestans populations on tomato and potato in British gardens demonstrates high diversity but no evidence for host specialization. Plant Pathology 65, 334–341, https://doi.org/10.1111/ppa.12407 (2016).
Cooke, D. E. et al. Genome analyses of an aggressive and invasive lineage of the Irish potato famine pathogen. PLoS pathogens 8, e1002940, https://doi.org/10.1371/journal.ppat.1002940 (2012).
Tian, Y. et al. Population Structure of the Late Blight Pathogen Phytophthora infestans in a Potato Germplasm Nursery in Two Consecutive Years. Phytopathology 105, 771–777, https://doi.org/10.1094/PHYTO-03-14-0073-R (2015).
Liu, F. et al. Distribution, function and evolution characterization of microsatellite in Sargassum thunbergii (Fucales, Phaeophyta) transcriptome and their application in marker development. Scientific reports 6, 1–9, https://doi.org/10.1038/srep18947 (2016).
Wallace, E. C. Population Structure of Pseudoperonospora cubensis on Commercial and Non-commercial Cucurbits in North Carolina Determined by Simple Sequence … MS. in Plant Pathology thesis, North Carolina State University, (2016).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods in molecular biology 132, 365–386 (1999).
Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nature biotechnology 18, 233–234, https://doi.org/10.1038/72708 (2000).
Quesada-Ocampo, L. M., Fulbright, D. W. & Hausbeck, M. K. Susceptibility of Fraser Fir to Phytophthora capsici. Plant Disease 93, 135–141, https://doi.org/10.1094/PDIS-93-2-0135 (2009).
Kambar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 281, 1–14, https://doi.org/10.7717/peerj.281 (2014).
Nei, M. Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89, 583–590 (1978).
Grünwald, N. J., Goodwin, S. B., Milgroom, M. G. & Fry, W. E. Analysis of genotypic diversity data for populations of microorganisms. Phytopathology 93, 738–746 (2003).
Agapow, P. M. & Burt, A. Indices of multilocus linkage disequilibrium. Mol Ecol Resour 1, 101–102 (2001).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research–an update. Bioinformatics 28, 2537–2539 (2012).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC genetics 11, 1–15, https://doi.org/10.1186/1471-2156-11-94 (2010).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Earl, D. A. & vonHoldt, B. M. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genet Resour 4, 359–361 (2012).
Acknowledgements
We thank all the members of the Quesada lab for their valuable help; we specially thank colleagues listed in Table 1 for providing some of the isolates used in this study. This work was supported by NIFA/USDA under project number NC02418.
Author information
Authors and Affiliations
Contributions
C.H.P.-R. performed the whole experiment; C.H.P.-R. and L.M.Q.-O. conceived the experiment; C.H.P.-R. and L.M.Q.-O. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Parada-Rojas, C.H., Quesada-Ocampo, L.M. Analysis of microsatellites from transcriptome sequences of Phytophthora capsici and applications for population studies. Sci Rep 8, 5194 (2018). https://doi.org/10.1038/s41598-018-23438-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-23438-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.