Abstract
The brown planthopper, Nilaparvata lugens, is an economically important pest on rice in Asia. Chemical control is still the most efficient primary way for rice planthopper control. However, due to the intensive use of insecticides to control this pest over many years, resistance to most of the classes of chemical insecticides has been reported. In this article, we report on the status of eight insecticides resistance in Nilaparvata lugens (Stål) collected from China over the period 2012–2016. All of the field populations collected in 2016 had developed extremely high resistance to imidacloprid, thiamethoxam, and buprofezin. Synergism tests showed that piperonyl butoxide (PBO) produced a high synergism of imidacloprid, thiamethoxam, and buprofezin effects in the three field populations, YA2016, HX2016, and YC2016. Functional studies using both double-strand RNA (dsRNA)-mediated knockdown in the expression of CYP6ER1 and transgenic expression of CYP6ER1 in Drosophila melanogaster showed that CYP6ER1 confers imidacloprid, thiamethoxam and buprofezin resistance. These results will be beneficial for effective insecticide resistance management strategies to prevent or delay the development of insecticide resistance in brown planthopper populations.
Similar content being viewed by others
Introduction
The brown planthopper (BPH), Nilaparvata lugens (Stål) (Hemiptera: Delphacidae), is a serious pest on rice in Asia1. This monophagous pest causes severe damage to rice plants through direct sucking often causing “hopper burn”, ovipositing and virus disease transmission during its long-distance migration1,2. In recent years, N. lugens outbreaks have occurred more frequently in China and other Asian countries3. The damage to the rice crop can result in a significant loss of yield in susceptible rice varieties1. The losses to rice production caused by N. lugens in Asia have been estimated as more than $300 million annually4.
The control of BPH has for many years predominantly relied on the use of synthetic chemicals5. However, due to the large scale and intensive use of insecticides, BPH has evolved high levels of resistance to many of the major classes of insecticides including organophosphates, carbamates, pyrethroids, neonicotinoids, insect growth regulators, and phenylpyrazoles3,6. Since the early 1990s, the neonicotinoid insecticide imidacloprid was introduced into Asia for BPH control7. Resistance to this insecticide emerged in field populations across Asia over the period 2005–20127,8. Following suspension of imidacloprid in 2006, banning of fipronil and buprofezin in 2009 and 2013, respectively, for controlling N. lugens in China, pymetrozine, thiamethoxam, flufiprole, nitenpyram, dinotefuran, sulfoxaflor and chlorpyrifos have been commonly used for controlling this pest insect in China in recent years9. BPH has developed resistance to 29 compounds in the world10. Therefore, resistance monitoring is a key to understand the current status of susceptibility of the field population of N. lugens to various insecticides6. Early detection of changes in resistance/susceptibility can prompt adoption of alternative control measures, which are essential for the successful management of this pest3.
The mechanisms of insect resistance to insecticides involve the over-expression or mutations of detoxifying enzyme genes and amino acid mutations of targeted genes10,11. The molecular mechanism(s) underlying resistance to imidacloprid have been characterised in the BPH. Although target-site resistance to this compound was described in a laboratory-selected strain of BPH, this mutation site has never been detected in filed-collected population5,12,13,14. In contrast, it was generally accepted that, changes in detoxifying enzymes, especially enhanced cytochrome P450 monooxygenase (P450) activity contributes to the neonicotinoid resistance of field-collected populations of BPH5,15,16,17,18,19. This detoxification mechanism was initially implicated by use of the metabolic enzyme inhibitor piperonyl butoxide (PBO) and the model substrate 7-ethoxycoumarin13,14. Although many P450 genes were over-expressed in resistant strains or field populations of BPH, two candidate P450 gens, CYP6ER1 and CYP6AY1, has been linked with imidacloprid resistance5,16,18,20. It was reported that CYP6AY1 was identified as the highest level of overexpression in the resistant strain compared with the susceptible strain and functional expression of CYP6AY1 proved that CYP6AY1 has the capacity to metabolise imidacloprid15. While one study showed that CYP6AY1 was not over-expressed even down-regulated in most of imidacloprid resistance field populations5. Recent studies showed that CYP6AY1 metabolized imidacloprid more efficiently and CYP6ER1 gene could be up-regulated by imidacloprid at a higher level and metabolic imidacloprid resistance in BPH relies on multiple P450 enzymes16,21. It was reported that the cross-resistance between imidacloprid and thiamethoxam was presented in the imidacloprid-resistant strains of N. lugens22,23. However, little is known about the resistance mechanism of BPH to thiamethoxam and buprofezin.
The aim of this study was to provide the changing levels of resistance to eight insecticides in N. lugens field strains collected from eight provinces in China from 2012 to 2016, and to study the relative roles of the cytochrome P450 monooxygenase (P450), CYP6ER1 and CYP6AY1, in determining their resistance phenotype to imidacloprid, thiamethoxam and buprofezin.
Results and Discussion
Variations in resistance ratios to eight insecticides
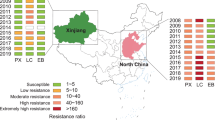
Resistance to eight insecticides were monitored in sixty-nine field populations of N. lugens collected from eight Chinese provinces in the period 2012–2016 (Supplementary Table S1 and Fig. 1). Pooled resistance ratio (RR) data from all populations of the same year indicated that substantially different resistance levels were developed in N. lugens to eight different insecticides (Fig. 2). The increase in RR over years was observed in all of insecticides. Our results indicated that N. lugens have developed high resistance levels to imidacloprid (mean RR value in 2016, 2104-fold), buprofezin (mean RR value in 2016, 1736-fold), thiamethoxam (mean RR value in 2016, 222-fold), pymetrozine (mean RR value in 2016, 125-fold) and flufiprole (mean RR value in 2016, 160-fold), a medium resistance level to chlorpyrifos (mean RR value in 2016, 30-fold) and sulfoxaflor (mean RR value in 2016, 12-fold) and a low resistance levels to nitenpyram (mean RR value in 2016, 5-fold). The results showed that resistance levels of six insecticides imidacloprid, thiamethoxam, nitenpyram, sulfoxaflor, buprofezin and flufiprole have a dramatic increase in 2015 and 2016 compared with former three years (Supplementary Table S4 and Fig. 2) and previously reported6,24.
Collection sites of Nilaparvata lugens populations during 2012–2016. The map in this figure were generated by software Adobe Photoshop CS5 version (San Jose, CA, http://www.adobe.com/products/photoshop.html) based on our own data.
Nicotinc acetylcholine receptor (nAChR) competitive modulators: imidacloprid, thiamethoxam, nitenpyram and sulfoxaflor
High to extremely high resistance levels to imidacloprid (RR: 132-8478-fold) and medium to high resistance levels to thiamethoxam (RR: 10-774-fold) were found in the field populations of N. lugens in 2012 to 2016 (Fig. 3A and B). The resistance ratio of N. lugens to imidacloprid and thiamethoxam showed an sharply increased in 2015. Surveys conducted in 1996–2007 showed that most populations of filed collected brown planthopper in China quickly developed medium to high level of resistance to imidacloprid in 2005–20067,19. Similar phenomenon was also observed by other studies5. Hence, suspension of imidacloprid for controlling N. lugens was carried out in China since 20067. However, the resistance of N. lugens to imidacloprid did not decrease in recent years. One reason is that imidacloprid is being used for controlling white-backed planthopper in China25. This will make BPH contact with imidacloprid. Another possible reason is, in other Southeast Asian countries, the intensive use of imidacloprid against N. lugens, which subsequently migrate to China22. Since N. lugens has evolved high level of resistance to imidacloprid, thiamethoxam has been used for BPH control instead of imidacloprid. Besides its extensive and intensive use, the main reason for the rapid resistance of N. lugens development to thiamethoxam might be the cross-resistance between imidacloprid and thiamethoxam in the imidacloprid-resistant strains of N. lugens22,23.
Most field populations developed a low to medium resistance level to nitenpyram (5–12.4-fold), except five populations, which were still susceptible or slightly resistant to the chemical in 2015 (Fig. 3C). However, one population (JS2016) have developed medium resistance levels (15.5-fold) to nitenpyram. One recent study also showed that nitenpyram resistance has appeared in some field populations of N. lugens in China and cytochrome P450 monooxygenase is likely a contributing factor to this insecticide resistance26. Sulfoxaflor, a new developed chemical by Dow AgroSciences, exhibits broad-spectrum control of many sap-feeding pests, including planthoppers, aphids, whiteflies and true bugs27,28. The susceptibility of N. lugens to sulfoxaflor was only investigated in three years. However, to our surprise, the results showed that the populations of N. lugens collected in 2015 and 2016 have developed low to medium resistance levels to sulfoxaflor (7.8–25.9-fold) (Fig. 3D). Liao et al. also showed that field populations of N. lugens collected in 2016 have developed a low level of resistance to sulfoxaflor29. These results imply that these two neonicotinoid insecticides, nitenpyram and sulfoxaflor, might have a potentially risk of resistance to N. lugens30.
Inhibitors of chitin biosynthesis: buprofezin
Of the 14 field populations of BPH collected in 2012 in China, 11 field populations were at medium resistance level to buprofezin (24–93-fold) except SH2012 (Shanghai) and YA2012 (Yongan) populations, which had reached to high resistance level (102 and 154-fold). And there was one population (GL2012) maintained susceptibility (RR < 5-fold) to the insecticide. However, in 2016, although the resistance ratios were substantially different among the populations, all populations had reached to high resistance level (839–3241.5-fold) to buprofezin (Fig. 4A). Buprofezin has been used for many years to control N. lugens in China. Most field populations of N. lugens were susceptible before 2004. However, resistance to buprofezin had been found in some field populations of N. lugens after 200419. And medium to high level of resistance was observed in 2010–2012 in China9,24. Our monitoring results suggested that the field populations of N. lugens have developed high levels of resistance to buprofezin. For this reason, it was suspended for the control of N. lugens in 2014 by the National Agro-tech Extension and Service Center, Ministry of Agriculture of China based on our results. However, the main reason why the resistance to buprofezin sharply increased in 2015 and 2016 need addressed in the future.
Acetylcholinesterase (AChE) Inhibitors: chlorpyrifos
The results showed that populations collected in 2012 kept susceptibility to chlorpyrifos except YA2012 population, which developed low resistance levels to chlorpyrifos. The surveys conducted in the next three years showed that the resistance ratio relatively increased every year. In 2015, all of the populations developed medium resistance levels to chlorpyrifos (RR 13.5–64.3-fold) except one population (HX2015), which developed high resistance level to chlorpyrifos (110-fold) (Fig. 4B). Because N. lugens had developed high level of resistance to insecticides such as imidacloprid, buprofezin, and thiamethoxam in the field in recent years, some farmers chose organophosphorus and carbamates insecticides for controlling brown planthopper6,31. Therefore, the resistance in this insect to chlorpyrifos increased every year in China6.
Chordotonal organ Transient Receptor Potential Vanilloid (TRPV) channel modulators: pymetrozine
The resistance ratios of N. lugens to pymetrozine in 2012 had a greater difference (4–1168-fold). In 2013, 68.8% of populations had a high resistance level (118–313-fold) and the others maintained a medium resistance level (67–98-fold), consistent with previously reported9,24. In 2014 and 2015, pymetrozine resistance of most populations decreased compared to the level in the previous two years. In 2015, there are two populations showed low resistance level (QS2015, 5-fold) or susceptible to pymetrozine (XG2015, 3.5-fold). Three populations were at high levels to pymetrozine (160, 223 and 340-fold) (Fig. 4C). However, in 2016, most of populations except YA2016, have developed high level of resistance to pymetrozine (Fig. 4C). Pymetrozine disrupt coordination and feeding of plant-sucking insects and are effective against insects that have developed to other insecticides32. Its molecular target have been identified as a transient receptor potential (TRP) ion channel complex33. The pymetrozine resistance was associated with increasing use of this insecticide against brown planthopper in China and Southeast Asian countries. As reported in the greenhouse whitefly, another possible reason was cross-resistance between pymetrozine and neonicotinoid insecticides34.
GABA-gated chloride channel blockers: flufiprole
Flufiprole is a novel kind of phenylpyrazole insecticide developed by Dalian Raiser Pesticide Co., Ltd., China35. Most populations of N. lugens collected in 2012 were susceptible to flufiprole except SH2015 (Shanghai) and TH2015 (Taihe) populations, which developed low level of resistance. However, there were clear increase of resistance in the next four years compared to results of 2012. In 2016, all of the populations have evolved medium to high level of resistance to flufiprole (88–281-fold) (Fig. 4D). Flufiprole have not been used extensively to control N. lugens in the rice-growing areas in China. In 2012–2013, field populations of N. lugens remain susceptible to flufiprole. However, it was recommended for N. lugens control instead of those insecticides to which the pest has developed high resistance levels, such as imidacloprid or buprofezin. Hence, in 2014–2015, N. lugens has developed low to moderate levels of resistance to flufiprole. The other possible reason was the cross-resistance between flufiprole and ethiprole or fipronil, which were used intensively and extensively against N. lugens in immigrant source areas such as Thailand5,6,36.
Synergists assessment
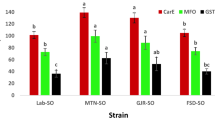
The synergistic effects of three representative synergists, piperonyl butoxide (PBO), triphenyl phosphate (TPP) and diethyl meteate (DEM), on imidacloprid, thiamethoxam, and buprofezin toxicities in three field populations were tested (Table 1). In the susceptible strain (SS), none of three synergists showed significant synergism on imidacloprid, thiamethoxam, and buprofezin. However, PBO significantly synergized imidacloprid, thiamethoxam, and buprofezin in field populations with the synergism ratio (SR) of 4.1-, 2.8-, and 3.5-fold, respectively, suggesting cytochrome P450s are involved in the resistance of BPH to imidacloprid, thiamethoxam, and buprofezin (Table 1). Cytochrome P450s have been reported that are the primary enzyme system involved in the resistance of brown planthopper to imidacloprid5,16,21,37. Previous studies have proved that insecticides cross-resistance was presented between imidacloprid, thiamethoxam, and buprofezin in field populations of BPH6,23, peach-potato aphid38 and Colorado potato beetle39. Hence, the results suggested that cytochrome P450 monooxygenases might play an important role in the resistance of N. lugens to imidacloprid, thiamethoxam, and buprofezin.
Association of overexpression of CYP6ER1 with resistance to imidacloprid, thiamethoxam and buprofezin
Two cytochrome P450s (CYP6ER1 and CYP6AY1) have previously been linked with imidacloprid resistance in a number of brown planthopper laboratory and field populations5,15,16,17. However, little is known about these two genes contributed to other insecticides resistance. In this present study, the mRNA levels of CYP6ER1 and CYP6AY1 were examined and compared in susceptible strain and five field populations collected from five areas in China that exhibited clear resistance to imidacloprid, thiamethoxam and buprofezin in discriminating dose bioassays and one buprofezin-resistance population (Bup-R, 533-fold). The results showed that CYP6ER1 in all field populations and Bup-R strain were significantly overexpressed, when compared with a lab susceptible strain, with fold changes ranging from seven- to 24-fold. However, CYP6AY1 was underexpressed in five of the populations compared with the same susceptible strain, and was only significantly overexpressed (13.5-fold) in a single population (SG2014) (Fig. 5).
To see whether selection of the field strains with thiamethoxam caused any increase in the expression levels of CYP6ER1 or CYP6AY1, three field strains (HX2015, DY2015 and XG2015) were selected with thiamethoxam up to final concentrations of 40, 80 and 80 mg L−1 thiamethoxam, respectively. When the expression levels of CYP6ER1 were compared between DY2015 (unselected) and DY2015-80 (selected), the expression level was found to have significantly increased two-fold after selection, rising from ∼11-to 23.2-fold. Similar effects were seen for HX2015 (unselected) versus HX2015-40 (selected) and XG2015 (unselected) versus XG2015-80 (selected). Besides this, the variation in the level of expression of CYP6ER1 among individual biological replicates decreased considerably after selection (as showed by significantly reduced 95% fiducial limits – see Fig. 5). This indicated that thiamethoxam selection has reduced genetic heterogeneity in these strains and that all replicates overexpress CYP6ER1 at a universally high level5. After selection, to our surprise, CYP6AY1 expression significantly decreased from 0.27 in HX2015 to 0.14 in HX2015-40, from 0.74 in DY2015 to 0.20 in DY2015-80, and from 0.60 in XG2015 to 0.20 in XG2015-80 (Fig. 5).
Overexpression of CYP6ER1 is associated with resistance to imidacloprid in the BPH5,18,21. Our results provide evidence that overexpression of CYP6ER1 also was associated with thiamethoxam resistance in BPH of China. CYP6AY1 was found to be the most highly expressed gene of imidacloprid resistance BPH strain and functional studies provided evidence that the over-expression of CYP6AY1 was contributing to imidacloprid resistance in the laboratory selected resistance strain15. However, using the same primers as previously reported, we found that CYP6AY1 was down-regulated in many strains especially in thiamethoxam selected populations5,15. Screening of five field populations from China showed that CYP6AY1 was only significantly overexpressed in one field population (SG2014, 13.5-fold, Fig. 5). Recent studies also showed that CYP6AY1 was underexpressed in most of field collected populations compared with a lab susceptible strain5. It is possible that CYP6AY1 is overexpressed in some N. lugens field populations in China and not the other places. Besides this, our study used a single reference lab susceptible strain to compared with obtain BPH field strains. Further experiments to investigate the relative roles of CYP6ER1 and CYP6AY1 in thiamethoxam and buprofezin resistance by comparing resistant strains with additional susceptible laboratory or field strains is required to confirm our findings.
Transgenic expression of the potential resistance genes CYP6AY1 and CYP6ER1 in Drosophila melanogaster
To identify whether the expression of CYP6AY1 and CYP6ER1 is sufficient to confer imidacloprid, thiamethoxam and buprofezin resistance, we used a transgenic approach utilizing the GAL4/UAS system of D. melanogaster. We confirmed the expression of the transgene in the Da > CYP6ER1 and Da > CYP6AY1 by RT-PCR (Fig. 6A and B). Bioassays showed that Da > CYP6ER1 line was resistant to imidacloprid, thiamethoxam and buprofezin at a treated dose of 1 mg/L imidacloprid and thiamethoxam or 400 mg/L buprofezin, with significant higher survival rate than the control lines (Fig. 6C,E and G). However, Da > CYP6AY1 line showed no significant resistance to imidacloprid, thiamethoxam and buprofezin (Fig. 6D,F and H). These data suggest that the expression of CYP6ER1 is sufficient for causing imidacloprid, thiamethoxam and buprofezin resistance. However, the expression of CYP6AY1 might play little roles in imidacloprid, thiamethoxam and buprofezin resistance.
Transgenic expression of CYP6ER1 (A,C,E and G) and CYP6AY1 (B,D,F and H) in D. melanogaster and their effects on imidacloprid, thiamethoxam and buprofezin resistance. (A and B) The expressions of CYP6ER1 and CYP6AY1 were confirmed by RT-PCR in two control lines and transgenic line. Three biological replicates of Da-GAL4, flies with genetic background correspond to Da-GAL4; Three biological replicates of UAS-CYP6ER1 (A) or UAS-CYPAY1 (B), flies not expressing the CYP6ER1 or CYP6AY1; Three biological replicates of Da > CYP6ER1 (A) or Da > CYP6AY1 (B), transgenic flies expressing the CYP6ER1 or CYP6AY1. (C,D,E,F,G and H). The comparison between survival rates of two control lines and transgenic line exposed to 1 mg/ml imidacloprid (C and D), thiamethoxam (E and F) and 400 mg/ml buprofezin (G and H). The data shown are the mean ± s.e.m. (n = 3). **P < 0.01, ***P < 0.001 (Chi-squared Test), ns (no significant).
RNA interference of CYP6ER1
Recent work have showed that when CYP6ER1 mRNA levels in imidacloprid-resistance strain was reduced by RNA interference (RNAi), imidacloprid susceptibility was recovered20. To further evaluate the contributions of CYP6ER1 in thiamethoxam and buprofezin resistance in vivo, we designed dsRNA to silence CYP6ER1 in 2nd nymph from the FQ2016 and Bup-R strain, respectively. Our RNAi experiment showed that the CYP6ER1 mRNA levels decreased by >80% or >60% in FQ2016 or buprofezin-resistance strains at 3 days after injection of CYP6ER1 dsRNA, respectively, indicating that this gene was successfully silenced by RNAi (Fig. 7A and C). The mortality of the dsCYP6ER1 sample was 60% and 20% higher than those of the dsGFP control sample after thiamethoxam and buprofezin treatment, respectively (Fig. 7B and D). These data suggests that the expression of CYP6ER1 is required for the high level of thiamethoxam and buprofezin resistance in BPH.
Knockdown in the expression of CYP6ER1 in N. lugens field-resistance strain reduced its resistance to thiamethoxam and buprofezin. (A and C) The mRNA levels of CYP6ER1 were quantified by qRT-PCR at three days after dsRNA injection in FQ2016 (A) or buprofezin-resistance (C) strains, respectively. The data shown are mean + SEM (n = 3). Statistical significance of the gene expression between two samples was calculated using Student’s t test. *P < 0.05, **P < 0.01 and ***P < 0.001. (B and D) Effects on the mortality rate after injection of dsGFP or dsCYP6ER1 and followed by application of thiamethoxam (5 mg/L) (B) or buprofezin (50 mg/L) (D).
Conclusions
In summary, this study characterized resistance monitoring of brown planthopper populations of different geographic areas to eight insecticides using rice stem dipping method. The results provide current resistance status of insecticides against this economically important insect in China. There is evidence that individual planthoppers may exhibit multiple resistance to the different insecticide modes of action. Our results reveal that metabolism mediated by cytochrome P450 monooxygenases and overexpression of the cytochrome P450 CYP6ER1 is associated with imidacloprid, thiamethoxam, and buprofezin resistance in N. lugens populations in China. Besides confirming this conclusion, we found that CYP6ER1 was also involved in thiamethoxam and buprofezin resistance. Finally, although our studies provide evidence of a role for CYP6ER1 in thiamethoxam and buprofezin resistance, functional characterisation of this P450 to confirm its ability to detoxify thiamethoxam and buprofezin is emergent required.
Methods
Insects
Seventy field populations of the brown planthopper were collected from seven provinces and the City of Shanghai in China from 2012 to 2016 (Fig. 1 and Supplementary Table S1). The collected insects were reared on rice seedlings under standard conditions of 27 ± 1 °C and 70–80% relative humidity with a 16-h light/8-h dark photoperiod. The field-collected brown planthoppers were mass mated, and the third-instar nymphs of the first (F1) or second (F2) generation were used for the susceptibility bioassay.
Insecticides and synergists
Technical grade insecticides except sulfoxaflor were used in this study. Chlorpyrifos (96.5%), imidacloprid (97%) and nitenpyram (95%) were both supplied by Nanjing Red Sun Co. Ltd. (Jiangsu, China). Buprofezin (97%), thiamethoxam (98%), flufiprole (90%), pymetrozine (97.4%) and sulfoxaflor (22% SC) were supplied by Changlong Chemical Industrial Group Co. Ltd. (Jiangsu, China), Syngenta Investment Co. Ltd. (Shanghai, China), Dalian Ruize Chemicals Co. Ltd. (Liaoning, China), Aijin Agrochemical Co., Ltd. (Nanjing, China), and Dow AgroSciences China Ltd. (Shanghai, China), respectively; diethylmaleate (DEM) and triphenyl phosphate (TPP) by Shanghai Chemical Factory (Shanghai, China); and piperonyl butoxide (PBO) by Koch-Light Laboratories Co., Ltd. (UK). The technical grade insecticides were dissolved in acetone (N,N-dimethylformamide for pymetrozine) as stock solution and then diluted in a series of 5–6 concentration gradients with water containing 0.1% of Triton X-100 as described previously3.
Bioassays
The dose-responses of BPH to different insecticides were measured using the rice-stem dipping method19. Rice plants at the tillering to early booting stage were pulled out from the soil, washed thoroughly, cut into an approximately 10-cm-long rice stem with roots and air-dried. Three rice stems were grouped together and dipped into appropriate insecticide solutions for 30 s and then air-dried at room temperature for at least 30 min. The rice stems with roots were wrapped with moistened cotton and put into 500 mL plastic cups. The third instar nymphs were collected with a homemade aspirating device, and twenty nymphs were transferred onto rice stems into a plastic cup for each replicate. There were three replicates for each concentration and 5–6 doses for each insecticide. Control rice stems were treated with 0.1% Triton X-100 water solution only. All treatments were maintained at a temperature of 27 ± 1 °C and 70–80% relative humidity with a 16-h light/8-h dark photoperiod. Mortality was recorded 72 h after treatment for chlorpyrifos, 96 h for imidacloprid, thiamethoxam, sulfoxaflor, flufiprole, nitenpyram, 120 h for buprofezin, and 168 h for pymetrozine according to the speed of kill of the insecticides. The nymphs were considered dead if they were unable to move after a gentle prodding with a fine brush.
For synergism bioassays of BPH to imidacloprid, thiamethoxam and buprofezin, third-instar nymphs were first exposed to rice stems treated with acetone solutions of the selected detoxification enzyme inhibitors at the highest possible concentrations (no adverse effect on the insects; PBO, P450-monoxygenase inhibitor at 20 mg/L; TPP, esterase inhibitor at 100 mg/liter; and DEM, glutathione S-transferase inhibitor at 50 mg/L determined by preliminary testing) for 2 h. The rest of the procedures were kept the same with above mentioned bioassay methods.
Three of the field strains, HX2015, DY2015 and XG2015, demonstrating relatively high levels of resistance to thiamethoxam, were placed directly onto rice plants treated with 40, 80 and 80 mg/L thiamethoxam and selected one generation in the laboratory.
Data analysis
The mortality rates at diagnostic concentrations were subjected to Abbott’s formula. Lethal concentration values (LC50) and their 95% fiducial limits (F.L.) were estimated using POLO-Plus program (Version 2.0) (LeOra Software 2008). The resistance ratio (RR) was calculated by dividing the LC50 value of a field population by the corresponding LC50 value of the susceptible baseline (Supplementary Table S2). Synergistic ratio (SR) was calculated by dividing the LC50 value without a synergist treatment by the LC50 with a synergist treatment. Moreover, the LC50 value of the susceptible baseline for sulfoxaflor to N. lugens was established in the present study using a susceptible strain of N. lugens, which had been provided from the Zhejiang Chemical Industrial Group Co. Ltd. (Hangzhou, Zhejiang, China) and reared on rice-seedlings in the laboratory without exposure to any insecticide for many years. Insecticide resistance of the field strains was classified as: RR < 5-fold as susceptible, RR = 5~10-fold as low resistance level, RR = 10~100-fold as medium resistance level and RR > 100-fold as high resistance level25.
Real-time quantitative RT PCR
Real-time quantitative RT PCR (qRT-PCR) was performed to determine the mRNA levels of P450s using the SsoFast Eva Green Supermix with Low Rox (Bio-Rad, Hercules, CA) and Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems by Life Technologies, Carlsbad, CA) following the manufacturer’s instructions. Each qRT-PCR experiment was performed at least four independent biological replicates and analyzed in three technical replications. Data were analysed according to the 2−ΔΔCT method40. The primers used in this study are designed based on previous report (Supplementary Table S3)15,18.
Construction of transgenic Drosophila and bioassays
The CYP6ER1 and CYP6AY1 CDS cloned were inserted into a pJFRC-MUH vector to prepare UAS-CYP6ER1 and UAS-CYP6AY1 constructs, respectively. Subsequently, the F1 of “BDSC#8622 (y1w67c23; P(CaryP) attP2) male crossed to PhiC31 source virgin” were used for injection using standard techniques in the UniHuaii Co., Ltd. (Zhuhai, Guangdong, P.R. China). The transformed lines were backcrossed with W1118 line with several generations and then were crossed with the Da-GAL4 (Daughterless-GAL4, which expressed in all of the cells) line for expression of the CYP6ER1 and CYP6AY1 gene, respectively. The genotype of the crosses were Da-GAL4 > CYP6ER1 and Da-GAL4 > CYP6AY1. For use as a control, the transformed line and the Da-GAL4 line were crossed with W1118 line, the progenies of which did not express CYP6ER1 and CYP6AY1 gene. RT-PCR was used to confirm the expression of the CYP6ER1 and CYP6AY1 gene in transgenic Drosophila using primers specific for the CYP6ER1 and CYP6AY1 gene and the reference housekeeping gene DmActin (Supplementary Table S3)20. Insecticide bioassays for imidacloprid and thiamethoxam were performed using previously described techniques41. In brief, 50 first instar larvae were placed in vials with 10 ml corn meal medium containing 1 mg/L imidacloprid and thiamethoxam or 400 mg/L buprofezin. Three to five replicates were performed for each assay. Adult emergence of 50 first instar larvae per vial was scored. The survival rates were calculated and analyzed using the Chi-squared test.
RNA interference and bioassays
The coding sequences of CYP6ER1 and green fluorescent protein (GFP) (negative control) were cloned into the pGEM®-T easy vector (Promega) vector. To minimize non-target silencing, a unique region of CYP6ER1 gene was chosen for the design of dsRNAs. PCR-generated DNA templates were then used to synthesize dsRNA, which contains T7 promoter sequences at each end (Supplementary Table S3). We used a MEGAscript T7 transcription kit (Ambion, Austin, TX, USA) to produce the specific dsRNA of each gene as the manufacturer’s instruction. The quality and size of the dsRNA products were verified by 1% agarose gel electrophoresis. The 2nd instar nymph were used for microinjection with 30 nymphs used for each gene treatment. Approximately 30 nl of purified dsRNA (5 ng/nl) was injected into the mesothorax of the nymph. A set of six to 10 insects at 3 days after injection for each treatment was selected to verify dsRNA knockdown efficiency by qRT-PCR. Thirty nymphs were placed in rice seedlings soaked in a solution of 5 mg/L thiamethoxam and 50 mg/L buprofezin for 30 seconds. The rest of the procedures were kept the same with above mentioned bioassay methods. Each experiment was repeated in triplicate.
References
Heong, K. L., Cheng, J. & Escalada, M. M. Rice Planthoppers: Ecology, Management, Socio Economics and Policy. Zhejiang University Press 15, 1130–1139 (2014).
Hibino, H. Biology and epidemiology of rice viruses. Annu. Rev. Phytopathol. 34, 249–274 (1996).
Wang, Y. et al. Susceptibility to neonicotinoids and risk of resistance development in the brown planthopper, Nilaparvata lugens (Stål) (Homoptera: Delphacidae). Pest Manag. Sci. 64, 1278–1284 (2008).
Min, S., Lee, S. W., Choi, B.-R., Lee, S. H. & Kwon, D. H. Insecticide resistance monitoring and correlation analysis to select appropriate insecticides against Nilaparvata lugens (Stål), a migratory pest in Korea. J. Asia Pac. Entomol. 17, 711–716 (2014).
Garrood, W. T. et al. Field-evolved resistance to imidacloprid and ethiprole in populations of brown planthopper Nilaparvata lugens collected from across South and East Asia. Pest Manag. Sci. 72, 140–149 (2016).
Zhang, X. et al. Insecticide resistance monitoring and correlation analysis of insecticides in field populations of the brown planthopper Nilaparvata lugens (stål) in China 2012–2014. Pestic. Biochem. Physiol. 132, 13–20 (2016).
Wang, Y. H. et al. Imidacloprid susceptibility survey and selection risk assessment in field populations of Nilaparvata lugens (Homoptera: Delphacidae). J. Econ. Entomol. 101, 515–522 (2008).
Matsumura, M. et al. Species-specific insecticide resistance to imidacloprid and fipronil in the rice planthoppers Nilaparvata lugens and Sogatella furcifera in East and South-east Asia. Pest Manag. Sci. 64, 1115–1121 (2008).
Zhang, X. et al. Field evolution of insecticide resistance in the brown planthopper (Nilaparvata lugens Stål) in China. Crop Prot. 58, 61–66 (2014).
Sparks, T. C. & Nauen, R. IRAC: Mode of action classification and insecticide resistance management. Pestic. Biochem. Physiol. 121, 122–128 (2015).
Heckel, D. G. Insecticide resistance after silent spring. Science 337, 1612–1614 (2012).
Liu, Z. et al. A nicotinic acetylcholine receptor mutation conferring target-site resistance to imidacloprid in Nilaparvata lugens (brown planthopper). Proc. Natl. Acad. Sci. USA 102, 8420–8425 (2005).
Wen, Y., Liu, Z., Bao, H. & Han, Z. Imidacloprid resistance and its mechanisms in field populations of brown planthopper, Nilaparvata lugens Stål in China. Pestic. Biochem. Physiol. 94, 36–42 (2009).
Puinean, A. M., Denholm, I., Millar, N. S., Nauen, R. & Williamson, M. S. Characterisation of imidacloprid resistance mechanisms in the brown planthopper, Nilaparvata lugens Stål (Hemiptera: Delphacidae). Pestic. Biochem. Physiol. 97, 129–132 (2010).
Ding, Z. et al. Biochemical mechanisms of imidacloprid resistance in Nilaparvata lugens: Over-expression of cytochrome P450 CYP6AY1. Insect Biochem. Mol. Biol. 43, 1021–1027 (2013).
Bao, H. et al. The roles of CYP6AY1 and CYP6ER1 in imidacloprid resistance in the brown planthopper: Expression levels and detoxification efficiency. Pestic. Biochem. Physiol. 129, 70–74 (2016).
Pang, R. et al. Identification of promoter polymorphisms in the cytochrome P450 CYP6AY1 linked with insecticide resistance in the brown planthopper. Nilaparvata lugens. Insect Mol. Biol. 23, 768–778 (2014).
Bass, C. et al. Overexpression of a cytochrome P450 monooxygenase, CYP6ER1, is associated with resistance to imidacloprid in the brown planthopper. Nilaparvata lugens. Insect Mol. Biol. 20, 763–773 (2011).
Wang, Y. et al. Buprofezin susceptibility survey, resistance selection and preliminary determination of the resistance mechanism in Nilaparvata lugens (Homoptera: Delphacidae). Pest Manag. Sci. 64, 1050–1056 (2008).
Pang, R. et al. Functional analysis of CYP6ER1, a P450 gene associated with imidacloprid resistance in Nilaparvata lugens. Scientific Reports 6 (2016).
Zhang, Y., Yang, Y., Sun, H. & Liu, Z. Metabolic imidacloprid resistance in the brown planthopper, Nilaparvata lugens, relies on multiple P450 enzymes. Insect Biochem. Mol. Biol. 79, 50–56 (2016).
Matsumura, M. et al. Insecticide susceptibilities in populations of two rice planthoppers, Nilaparvata lugens and Sogatella furcifera, immigrating into Japan in the period 2005–2012. Pest Manag. Sci. 70, 615–622 (2014).
Wang, Y. H. et al. Dynamics of imidacloprid resistance and cross-resistance in the brown planthopper. Nilaparvata lugens. Entomol. Exp. Appl. 131, 20–29 (2009).
Wang, P. et al. Resistance monitoring to conventional insecticides in brown planthopper, Nilaparvata lugens (hemiptera: delphacidae) in main rice growing regions in China. Chin. J. Rice Sci. 27, 191–197 (2013).
Zhang, K. et al. Susceptibility of Sogatella furcifera and Laodelphax striatellus (Hemiptera: Delphacidae) to six insecticides in China. J. Econ. Entomol. 107, 1916–1922 (2014).
Zhang, X. et al. The role of detoxifying enzymes in field-evolved resistance to nitenpyram in the brown planthopper Nilaparvata lugens in China. Crop Prot. 94, 106–114 (2017).
Babcock, J. M. et al. Biological characterization of sulfoxaflor, a novel insecticide. Pest Manag. Sci. 67, 328–334 (2011).
Nugent, B. M. et al. Expanding the structure–activity relationship of sulfoxaflor: the synthesis and biological activity of N-heterocyclic sulfoximines. Pest Manag. Sci. 71, 928–936 (2015).
Liao, X. et al. Temporal variability and resistance correlation of sulfoxaflor susceptibility among Chinese populations of the brown planthopper Nilaparvata lugens (Stål). Crop Prot. 102, 141–146 (2017).
Cutler, P. et al. Investigating the mode of action of sulfoxaflor: a fourth-generation neonicotinoid. Pest Manag. Sci. 69, 607–619 (2013).
Mu, X. C. et al. Resistance monitoring and cross-resistance patterns of three rice planthoppers, Nilaparvata lugens, Sogatella furcifera and Laodelphax striatellus to dinotefuran in China. Pestic. Biochem. Physiol. 134, 8–13 (2016).
Maienfisch, P. Selective feeding blockers: pymetrozine, flonicamid, and pyrifluquinanzon. W. Krämer, U. Schirmer, P. Jenschke, M. Witschel (Eds.), Modern Crop Protection Compounds, John Wiley and Sons, New York,1327–1346 (2012).
Nesterov, A. et al. TRP channels in insect stretch receptors as insecticide targets. Neuron 86, 665–671 (2015).
Karatolos, N., Denholm, I., Williamson, M., Nauen, R. & Gorman, K. Incidence and characterisation of resistance to neonicotinoid insecticides and pymetrozine in the greenhouse whitefly, Trialeurodes vaporariorum Westwood (Hemiptera: Aleyrodidae). Pest Manag. Sci. 66, 1304–1307 (2010).
Li, J. et al. Simultaneous enantioselective determination of phenylpyrazole insecticide flufiprole and its chiral metabolite in paddy field ecosystem by ultra-high performance liquid chromatography/tandem mass spectrometry. J. Pharm. Biomed. Anal. 121, 261–270 (2016).
Zhao, X. H. et al. Differential resistance and cross-resistance to three phenylpyrazole insecticides in the planthopper Nilaparvata lugens (Hemiptera: Delphacidae). J. Econ. Entomol. 104, 1364–1368 (2011).
Zhang, J. et al. Expression induction of P450 genes by imidacloprid in Nilaparvata lugens: A genome-scale analysis. Pestic. Biochem. Physiol. 132, 59–64 (2016).
Foster, S. P., Cox, D., Oliphant, L., Mitchinson, S. & Denholm, I. Correlated responses to neonicotinoid insecticides in clones of the peach-potato aphid, Myzus persicae (Hemiptera: Aphididae). Pest Manag. Sci. 64, 1111–1114 (2008).
Mota-Sanchez, D., Hollingworth, R. M., Grafius, E. J. & Moyer, D. D. Resistance and cross-resistance to neonicotinoid insecticides and spinosad in the Colorado potato beetle, Leptinotarsa decemlineata (Say) (Coleoptera: Chrysomelidae). Pest Manag. Sci. 62, 30–37 (2006).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Daborn, P. J. et al. Evaluating the insecticide resistance potential of eight Drosophila melanogaster cytochrome P450 genes by transgenic over-expression. Insect Biochem. Mol. Biol. 37, 512–519 (2007).
Acknowledgements
This work was financed by the National Natural Science Foundation of China (31471804 & 31672068) and The National Agro-Tech Extension and Service Center (NATESC) the research project of the Resistance Monitoring and Control on Rice Pests. We are grateful to Plant Protection Stations of Shanghai City, Anhui Province, Hubei Province, Jiangsu Province, Jiangxi Province, Guangxi Province, and Zhejiang Province for helping in collecting tested populations of N. lugens.
Author information
Authors and Affiliations
Contributions
S.W., S.Z., C.G. and J.S. designed the conception and reviewed the manuscript. S.W., B.Z., C.Z., X.M., Y.Z., and J.H. performed the research and wrote the paper. S.W., C.G., B.Z., and C.Z. analyzed the data. All of the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, SF., Zeng, B., Zheng, C. et al. The evolution of insecticide resistance in the brown planthopper (Nilaparvata lugens Stål) of China in the period 2012–2016. Sci Rep 8, 4586 (2018). https://doi.org/10.1038/s41598-018-22906-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-22906-5
This article is cited by
-
Baseline susceptibility of brown planthopper (Nilaparvata lugens) populations to mesoionic insecticide triflumezopyrim in Southern India
International Journal of Tropical Insect Science (2024)
-
Transgenerational hormesis effects of nitenpyram on fitness and insecticide tolerance/resistance of Nilaparvata lugens
Journal of Pest Science (2023)
-
A chitin synthase mutation confers widespread resistance to buprofezin, a chitin synthesis inhibitor, in the brown planthopper, Nilaparvata lugens
Journal of Pest Science (2023)
-
Dinotefuran resistance in Nilaparvata lugens: resistance monitoring, inheritance, resistance mechanism and fitness costs
Journal of Pest Science (2023)
-
Bio-efficacy of novel chemicals and tribal pesticide-based integrated modules against brown planthopper in rice
International Journal of Tropical Insect Science (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.