Abstract
We propose a simple 3-parameter model that provides very good fits for incidence curves of 18 common solid cancers even when variations due to different locations, races, or periods are taken into account. From a data perspective, we use model selection (Akaike information criterion) to show that this model, which is based on the Weibull distribution, outperforms other simple models like the Gamma distribution. From a modeling perspective, the Weibull distribution can be justified as modeling the accumulation of driver events, which establishes a link to stem cell division based cancer development models and a connection to a recursion formula for intrinsic cancer risk published by Wu et al. For the recursion formula a closed form solution is given, which will help to simplify future analyses. Additionally, we perform a sensitivity analysis for the parameters, showing that two of the three parameters can vary over several orders of magnitude. However, the shape parameter of the Weibull distribution, which corresponds to the number of driver mutations required for cancer onset, can be robustly estimated from epidemiological data.
Similar content being viewed by others
Introduction
Cancers arise after accumulating epigenetic and genetic aberrations1,2,3. Earlier studies established a power law model on the basis of multi-stage somatic mutation theory to explain age-dependent incidences4,5,6 for several cancer types. As noted by Hornsby et al.7 in the context of classical epidemiological studies most cancers occur with the same characteristic pattern of incidence, and the simplicity of this pattern is in contrast to the perceived complexity of carcinogenesis. Orthogonal to these age stratification of different cancer types, Tomasetti and Vogelstein8 (with follow-ups9,10) reported a significant association between life time caner risk and stem cell divisions and concluded the latter substantially contributes to the former. Challenging the conclusion of Tomasetti and Vogelstein8 of a high-intrinsic cancer risk Wu et al.11 subdivided cancer risk into extrinsic and intrinsic risk, arguing extrinsic factors contribute more to cancer risks than intrinsic factors do. Based on a mechanistic model of accumulated mutations, these authors provided a recursion formula for theoretical life time intrinsic risk (tLIR) parameterized by age a. This recursion formula has the closed form solution \({\rm{tLIR}}(a)=1-{\mathrm{(1}-{\mathrm{(1}-{\mathrm{(1}-r)}^{{\mathrm{log}}_{2}S+d\cdot a})}^{k})}^{S}\), where S can be interpreted as the numbers of stem cell, d as the stem cell division rate, k as number of driver events required for cancer onset and r as the mutation rate per division. They reported that tLIR goes outside of the plausible range of empirical cancer risks by studying several pairs of values for two parameters (mutation rate and driver gene mutations) concluding that there is a substantial contribution of extrinsic risk factors to cancer development. However, this conclusion only holds in the studied parameter space and when parameters for all cancer types are treated uniformly. By performing a systematic grid search in the space of biologically plausible parameter values we showed that tLIR can be close to empirical risk for different cancer types (R2 > 0.85). If the extrinsic risk factor is computed by simply setting it to a complement of 1 for the intrinsic risk factor as performed by Wu et al. it will be concluded that there is a possibility of high intrinsic risk, so that one of the presented arguments by Wu et al.11 is fallacious.
On a pure mathematical side, we show that a scaled Weibull function with 3 parameters approximates the 4-parameter mechanistic tLIR model. On an epidemiological data analytical side, this simple 3-parameter model excellently agrees with age-dependent cancer incidence curves among 18 common solid cancers even when variations due to different locations, races, or periods are taken into account. With this model, we study the relationship between cancer risk and stem cell divisions, the high correlation between the two entities reported by previous studies8,10 breaks down when considering age stratified data.
Results
Approximation of tLIR model by a scaled Weibull function
As is derived in the Materials and Methods the 4-parameter mechanistic tLIR model can be approximated by a scaled Weibull function with 3 parameters:
assuming that λ is defined by
Here \({\rm{Weibull}}(\lambda ,\,k)(a)=1-{e}^{-{(a/\lambda )}^{k}}\) is the cumulative distribution function of the Weibull distribution, and P is the number of independent parallel processes, which e.g. can be interpreted as cell population at risk12. Whether the total tissue cells or only a fraction of stem cells are susceptible for cancer risk is unclear13,14. If one sets P = S then rd = λ−1. However, other possible choices for P allow to account for other factors such as the selection of mutations15,16, the stem cell microenvironment17,18, and tissue architecture19,20,21,22, or effects of clonal expansion23,24,25. Models incorporating clonal expansion have additional parameters such as the number of clonal copies. Reducing the dimensions of such complexed models results in tLIR, in which S is interpreted as number of independent clusters after clonal expansion rather than the number of stem cells, r and d denote “net” mutation and division rate of independent clusters at average level rather than those of single cells. Whereas a precise analysis of models for clonal expansion will be the topic of future work, these considerations show that when using the scaled Weibull distribution, prior knowledge on the parameter ranges is not necessary. This is indeed one benefit of scaled Weibull function comparing to tLIR model which requires a biologically reasonable guessing on stem cell numbers, mutation rate, cell division rate and number of driver mutations. The Weibull distribution is a special case of the generalized extreme value distribution (GEV)26. The GEV distribution plays the same role within extreme value statistics as the normal distribution does in average value statistics. It results in the limit distribution being maximized over many independent and identically distributed random variables, thus becoming the default model for the accumulation of micro events which finally leads to a macro event. The GEV is the limit distribution when one takes the maximum (and not the sum) of many independent and identically distributed random variables, thus being the default model for the accumulation of micro events which finally lead to a macro event. Accordingly, the Weibull distribution is not just a distribution providing a good empirical fit, but can be seen as justifiable for use in a plausible causative model of cancer genesis.
Fitting empirical incidence rates with scaled Weibull function
We performed extensive simulations and parameter fittings for the empirical incidence cuminc c (a) of cancer type c at age a using the scaled Weibull function: cuminc c (a) ≈ P c ⋅ Weibull(λ c , k c )(a). The model agrees excellently with age-dependent age incidences of 18 common solid cancers (R2 > 0.99, Fig. 1).
Empirical cumulative cancer incidence data are consistent with the Weibull cumulative probability function in 18 cancers (data for ages up to 85 years old). Empirical (blue line) and Weibull function-fitted (red line) cancer cumulative incidence curves for 18 tissues, goodness of fit is reported in each subplot. The 18 cancers exhibit a good goodness of fit when using R2 between model-reported age incidence and the empirical cumulative cancer incidence are used as metrics.
Goodness of fit maintains when parameters P c and λ c , varying roughly two orders of magnitudes (Fig. 2). This finding suggests that many parameter combinations provide similar dynamics that are consistent with empirical data. So any interpretations of P c and λ c have to take into account this considerable uncertainty. Nevertheless, the estimates for P c are several orders of magnitude smaller than the realistic number of stem cells provided by Tomasetti and Vogelstein8, yielding evidence supporting the above statement that the number of independent local processes is not equal to the number of stem cells.
Sensitivity analysis of parameter estimates using the scaled Weibull function for exemplary 14 cancer types. Whereas the estimates P c for the cell population at risk and the scale parameter λ c can vary over two order of magnitude, the estimates of the shape parameter k c are within about ±1. Notice that the shape parameter allows interpretation as the number of limiting events.
The estimates of parameter k c , which corresponds to the number of driver events in the mechanistic model, are robust against variations of parameters P c and λ c (Fig. 2). Moreover, the estimates of k c are robust against race, sex, period and location (Fig. 3). In Supplemental Fig. 2 the best fits of shape parameters are plotted against the best fits of scale parameters for 694 time series.
Shape parameters estimated by fitting empirical cancer incidence data using the Weibull function (data with ages for up to 85 years). Cancer patients are grouped by year of diagnosis, race and registry. Cancers are ordered by median values of shape. Shapes are uniform regardless of risk factors, which is consistent with intuitive expectations: race and environmental changes are less likely to alter the number of driver events for cancer onsets.
Relationship between cancer incidence and stem cell divisions
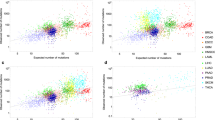
Tomasetti and Vogelstein8 suggested that the variation in cancer risk among tissues can be explained by the number of stem cell divisions. They reported that the tissue-specific cancer risk is strongly correlated (0.81) with life-time stem cell divisions (LSCD). These authors stated that the total number of stem cell divisions is a causative factor of cancer risk. This assumption yields a prediction on age structured data: for tissue type c the number of stem cell divisions up to age a, which we will denote by LCSD c (a), should then be strongly correlated with cuminc c (a). However, using age incidence data obtained from the SEER-database27 we found that the regression lines for most tissue types c for age data of 40, 50, 60, 70, and 80 years of cuminc c (a) plotted against LCSD c (a) in a log-log-scale are much steeper than the ones of the regression lines for different c and cuminc c (80)—using 80 as average life span as was done by Tomasetti and Vogelstein8 (see Fig. 4).
Relationship between cancer incidence and stem cell divisions among 30 cancer types. The lifetime cancer risk regression line is conceptually the same as that used by Wu et al.11.
This “life time cancer risk” moderately associates with age-dependent stem cell divisions, if one takes a life-time a that is less than 70 years (Fig. 5).
Overall, age-dependent stem cell divisions (using ages 40–80 years) is modestly correlated to age-dependent cancer risk for the 31 cancer types considered by Tomasetti and Vogelstein8 using the SEER database and the estimates of stem cell divisions given therein (Pearson correlation coefficient ρ = 0.51).
Hence, the strong correlation for (life-time) tissue-specific cancer risk with life-time stem cell divisions (LSCD) cannot be explained by the simple causative factor (involving the product of the number of stem cells and the number of divisions of each stem cell) suggested by Tomasetti and Vogelstein8. A causal explanation on cancer risk should at least shows that the association between cancer risk and risk factor observed at overall level is reproducible on age stratified data. However, one caveat to such explanation, co-factors of risk factors might not be appreciated.
In our 3-parameter model, which gives good fits for age dependent cancer risks, several relations between the model parameters and cancer risks at a certain age can be observed. For instance in our parameter estimates good fits are possible when taking the inverse of the lifetime cancer risk P c ≈ 1/cuminc c (85). However, we will not suggest that the number P c of cell population at risk is an explanation for the variation of cancer risks among tissues: as the range of P c yielding good fits varies by two orders of magnitude and independently determining this number is difficult to achieve, a corresponding hypothesis is difficult to verify or to falsify.
As the sensitivity analysis for the scale parameter λ c (Fig. 2) shows that this parameter varies over several order of magnitudes, still yielding very good fits (R2 > 0.99), the corresponding estimates for the mutation rate r in the tLIR model using the approximation (1) and relation (2) are also very uncertain, even when fixing values of S and d and leaving out the considerable uncertainty of these. Nevertheless, when using estimates of S and d taken from the literature8 the obtained ranges of values of r using relation (2) for several cancer types do not intersect the range [10−10, 10−6] of “plausible values” of r suggested by Wu et al.11. If we extend the analysis to allow “good fits” by setting a threshold R2 > 0.85, then good fit of the tLIR model with r ∈ [10−10, 10−6] are possible to achieve (Table 1).
Testing performance of our 3-parameter against other simpler model
For testing the performance of our 3-parameter model against other simpler models, we compared the fitting performance of the scaled Weibull function to that of 2-parameter power law model arising as the simplest instance from multistage theory5,7,12,13. The empirical time series for different locations, periods and races were fitted (694 time series all together) using both models, the power law model had a goodness of fit of R2 < 0.90 for 90 time series (13.0%), R2 < 0.95 for 257 time series (37.0%), and R2 < 0.98 for 366 time series (52.7%). In contrast, our 3-parameter scaled Weibull model resulted in R2 > 0.9 for all time series, R2 > 0.98 for 686 (=98.8%) of the time series, and R2 > 0.99 for 679 (=97.8%) of them (Fig. 6(a)).
We compare the fitting performance of the scaled Weibull function against that of the scaled Gamma function. Although both functions fit data equivalently well in most cases, the scaled Weibull function outperforms the scaled Gamma function in several time series. (Fig. 6(b)) displays R2 reporting goodness of fit for the two functions. We also calculate the Akaike information ciriterion(AIC)28, a likelihood based measurement. A lower AIC value indicates a better fit. Table 2 reports the AIC for 18 cancer types, the AIC values for Weibull function are lower than those of Gamma function in 15 cancer types.
Estimating the Number of Driver Mutations for Cancer Onset
In our model the shape parameter k c reflects the number of mutations required for cancer onset. The values of this parameter are, however, higher than the number of mutations estimated from sequencing data by Vogelstein et al.29. Vogelstein et al. suggested technical issues as an explanation for the inconsistency between estimates from epidemiological data and sequencing data. Notably, our k c estimates and the number of driver mutations estimated from a classical power law model are roughly in the same numerical range (Fig. 7). Since we obtain better and more robust fits than the power law model, we believe that our estimated driver mutation numbers are more trustworthy.
Discussion
In this study we connected the mechanism-based cancer development tLIR model to the Weibull distribution function. We tested its validity by fitting a 3-parameter Weibull function to data from 18 common solid cancer types, consisting of more than 600 time series. The scaled Weibull function fits well with age dependent incidence curves of all studied cancers and outperforms other models, such as the commonly used 2-parameter power law model and a 3-parameter scaled Gamma function model. With the scaled Weibull function, we can estimate the number of driver mutations required for cancer onset in individual cancer types. To our knowledge, this is the first work matching pan-cancer incidence curves with a statistical distribution function that is partially biologically informative.
Compared to the tLIR model developed by Wu et al.11 we see two technical benefits of our suggested approach: First, the scaled Weibull function involves less parameters than tLIR, but it remains to be biologically interpretable. The tLIR model includes several details of the multi-staged process of cancer development, e.g. the number of steps required for transforming a normal cell to malignancy, the number of stem cells in a tissue and division rate of stem cells. Although the tLIR model indeed provides useful insights into linking age-dependent somatic mutations to cancer risk, it has also limitations. For example, it ignores the effects of clonal expansion25. Another issue is that most parameters in the tLIR model are difficult to measure accurately in practice. Following Tomasetti and Vogelstein8 the number of stem cell divisions can be estimated, but the accuracy has been criticized30. In contrast, our suggested model requires less specific assumptions about the parameters to be measured in practice. Moreover, the Weibull distribution is a special case of the generalized extreme value distribution (GEV) which is well connected to classical statistical approaches to describe rare events26.
Our analysis results mostly agree with those provided by Wu et al.11. They defined intrinsic cancer risk as the probability that one tissue transforms from normal to tumor because of accumulated mutations, and extrinsic cancer risk as 1–intrinsic cancer risk. They quantified upper bounds to intrinsic cancer risk by tLIR but did not properly fit tLIR to epidemiological data, concluding intrinsic factor insignificantly contributes to cancer. According to our understanding their argument mainly results from insufficient exploration of parameter space and implicitly assumes that all tissues require the same number of driver mutation to initiate cancer. Our results suggest that the contribution of extrinsic factors to cancer is overestimated by Wu et al.11. However, one should note that the excellent agreement between the scaled Weibull distribution function and empirical data does not necessarily exclude that in addition to intrinsic there are further extrinsic and unknown risk factors. In that context it is worthwhile to mention that our estimated number of driver mutations required for cancer onset differs from tissue to tissue. Although the exact number is not validated by biological experiment, this observation is consistent with findings in genetic studies29.
One interesting observation is that all non-reproductive tissues have a similar cancer risk accumulation pattern. Cancer incidence rates increase dramatically at about 40–50 years, peaking at about 55–70 years and then decrease. This pattern matches findings reported by Podolskiy et al.31. A question for future work is whether mutation load agrees with the scaled Weibull function or age-specific mutational signatures32,33,34. Another interesting observation is that testicular germ cell cancer incidence peaks at younger age compared to other cancer types, which might be explained by accelerated aging of testis31. Altogether we believe that our suggested approach provides insights into cancer development by providing a link between empirical data and a mechanism-based model.
Methods
Fitting cumulative cancer incidence with a model for theoretical intrinsic cancer risk (tLIR)
Wu et al.11 provided the following recursion formula to compute the chance that a single stem cell acquires k mutation hits after g divisions given a mutation rate r.
given the initial cell state at generation 0:
Here X g is accumulated driver mutations at generation g, i and j represents accumulated driver mutations at generation g and g + 1, respectively. A fully developed tissue with S stem cells must go through \(n={\mathrm{log}}_{2}S+d\cdot a\) rounds of divisions, assuming division rate is d and age a. With this transition probability (3), the theoretical lifetime intrinsic cancer risk (tLIR) is formulated as
Although the recursion formula being dependent on more than one parameter cannot directly be solved in closed form by standard algorithmic techniques, it has nevertheless a simple closed form solution, which was derived by hand computations and verified by standard symbolic computations (using the computer algebra system Maple 2015.2):
The formula for the age-parameterized theoretical lifetime intrinsic cancer risk (tLIR) hence has the following simple closed form solution, which allows much faster and hence more extensive computations and extends the range of admissible values of k from the positive integers to the positive real numbers:
Notice that our result basically coincides with the one obtained by Calabrese and Shibata35 that was obtained by a direct probabilistic reasoning.
Relating the tLIR model to a scaled Weibull function
We found a connection between
and the scaled Weibull function
where P is the cell population at risk.
To see this connection, we assume that \(r\ll 1\). Then
and using the Taylor series for log and exp, we obtain
We have
Comparing (7) and (9) we observe that these relations coincide if
Since for small f0 > 0 we have 1 − f0 = exp(−f0), the last equation can rewritten as
Using that relation and (8) one finds
So we have obtained a shifted Weibull distribution. However, if we remove \({d}^{-1}{\mathrm{log}}_{2}S\) from the left hand side of the last equality assuming that
we obtain an unshifted one. This condition admits a transparent interpretation, namely, the number of stem cell divisions (for a fixed cell) should be more than the logarithm of stem cell number. Then we have that the tLIR incidence approximately equals to the scaled Weibull incidence if the parameters satisfy
Notice that using a Poisson approximation [12, p. 104] we finally obtain
Stem cell data
Tomasetti and Vogelstein8 collected stem cell information for 31 cancer types, including stem cell division rate, stem cell number, tissue total cell number. We excluded 6 from 31 cancer types due to lack of age incidence data: colorectal adenocarcinoma in familial adenomatous polyposis (FAP) patients, colorectal adenocarcinoma in patients with hereditary non-polyposis colorectal cancer (HNPCC, also called lynch syndrome), duodenal adenocarcinoma in FAP patients, head and neck squamous cell carcinoma with human papillomavirus (HPV), hepatocellular carcinoma with hepatitis C virus infection (HCV), lung adenocarcinoma in smokers. Among the 25 remaining cancer types, stem cell information were obtained from supplementary materials of Tomasetti and Vogelstein8. We discuss life time stem cell division (LSCD) hypothesis and extrinsic risk factor hypothesis for 25 remained cancers: AML, acute myeloid leukemia; BCC, basal cell carcinoma; CLL, chronic lymphocytic leukemia; COAD, colorectal adenocarcinoma; DUAD, duodenum adenocarcinoma; ESCA, esophageal squamous cell carcinoma; GBNPAD, gallbladder non papillary adenocarcinoma; GBM, glioblastoma; HNSC, head and neck squamous cell carcinoma; LHCA, hepatocellular carcinoma; LUAD, lung adenocarcinoma; MBM, medulloblastoma; SKCM, melanoma; OSARC, osteosarcoma; OSARCA, osteosarcoma of the arms; OSARCH, osteosarcoma of the head; OSARCL, osteosarcoma of the legs; OSARCP, osteosarcoma of the pelvis; OVGC, ovarian germ cell; PDAD, pancreatic ductal adenocarcinoma; PECA, pancreatic endocrine (islet cell) carcinoma; SIAD, small intestine adenocarcinoma; TGCC, testicular germ cell cancer; TPFC, thyroid papillary or follicular carcinoma; TMCA, thyroid medullary carcinoma.
Cancer incidence data
SEER-9 registries (1973–2013), SEER-4 registries (1992–2013), SEER-5 registries (2000–2013) data were downloaded from Surveillance, Epidemiology, and End Results Program (SEER) database27. SEER database covers about 28% USA population, involving more than 100 features such as race, sex, period, location, histology and ICD (international classification of disease) code. These data were stored in ASCII file, we used the SEERaBomb R package to parse them into sqlite file facilitating data manipulation.
Cancer names provided by Tomasetti and Vogelstein8 can not be directly mapped into those in SEER database. We addressed this difficulty by two steps: first, annotate tumor primary site to (international classification of disease-oncology 3) ICD-O-3 code based on the literal sense of site in Tomasetti and Vogelstein8; second, annotate histology to ICD-O-3 code based on the literal sense of cancer histology by Tomasetti and Vogelstein8. For instance, primary site of lung adenocarcinoma is lung, corresponding to ICD-O-3 site code: C340, C341, C342, C343, C348, C349; adenocarcinoma of lung cancer corresponds to ICD-O-3 histology code 8140, 8141, 8143, 8147, 8570, 8571, 8572, 8573, 8574, 8575, 8576. The dictionary needed for mapping step (we call it ICD dictionary) can be found in http://seer.cancer.gov/icd-o-3/. Osteosarcoma definition can be found in ICD dictionary, it is a subtype of malignant bone neoplasm, corresponding ICD-O-3 histology code: 9180–9189. However, the ICD dictionary does not differentiate between osteosarcoma detected in the head, leg, or arm. The ICD9Data database (http://www.icd9data.com/) defines bone cancer using ICD9 code 1700–1709, bone cancer in head, arms, legs, pelvis using ICD9 code 1700, 1704–1705, 1707–1708, 1706 respectively. Head and neck squamous cell carcinoma involves tumors located in many sites, ICD dictionary fails to provide its definition. Liao et al.36 provided ICD9 site code: 1400–1419, 1430–1499, 1600–1619, we then used ICD-O-3 histology code: 8070–8076, 8078 to select squamous cell carcinoma. More detailed cancer definitions using ICD code can be found in Table 3. Two hematopoietic cancers: acute myeloid leukemia and chronic lymphocytic leukemia, are defined using site recode ICD-O-3/WHO 2008 definition (http://seer.cancer.gov/siterecode/icdo3_dwhoheme/index.html).
Although we carefully annotated 25 cancer definitions using ICD code, we can not avoid misclassifications. because annotation needs several data sources of which information confidential levels differ from each other. The Cancer Genome Atlas (TCGA) program37 is a flag project of cancer research hosted by National Institutes of Health, it provides comprehensive, high-quality molecular and clinical data. Cancer definitions are well annotated using ICD code in TCGA clinic documents. We therefore assume TCGA cancer definitions are precise and extracted definitions of 18 solid tumors (Table 4). With 18 cancer definitions, we selected patients who were diagnosed with cancer after 2000 from SEER-9 registries, SEER-4 registries, SEER-5 registries data to form SEER-18 registries data. As the highest time resolution of SEER data is 1 year, for each year, we took middle age for fitting models, for example, 0 year-old is modified as 0.5 years-old.
For robustness analysis of parameter estimates we classified each cancer into subgroups based on location, period and race, data of subgroups were separately fitted to the mathematical models.
Fitting the models to empirical cancer incidence data
As was done in previous work38, empirical cancer incidence I(a) was calculated by
where p i is frequency of people diagnosed with caner at age i.
We performed grid search on an extensive parameter space to fit the tLIR model using \({R}^{2}={(\frac{\sum ({x}_{i}-\bar{x})({y}_{i}-\bar{y})}{\sqrt{\sum {({x}_{i}-\bar{x})}^{2}\sum {({y}_{i}-\bar{y})}^{2}}})}^{2}\) as the metrics for goodness of fit, where x i and y i is empirical and model-derived cancer incidence respectively, \(\bar{x}\) and \(\bar{y}\) respectively denotes mean value of x and y. Results of fits are given in Table 1 showing that there are biologically reasonable parameter combinations that can yield good fits of the tLIR model for most cancer types.
Data availability
All data used in this study are publicly available. The sources are detailed in the section on methods.
References
Feinberg, A. P., Ohlsson, R. & Henikoff, S. The epigenetic progenitor origin of human cancer. Nat Rev Genet 7, 21–33 (2006).
Feinberg, A. P., Koldobskiy, M. A. & Gondor, A. Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat Rev Genet 17, 284–299 (2016).
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719–724 (2009).
Nordling, C. O. A new theory on the cancer-inducing mechanism. British Journal of Cancer 7, 68–72 (1953).
Armitage, P. & Doll, R. The age distribution of cancer and a multi-stage theory of carcinogenesis. British Journal of Cancer 8, 1–12 (1954).
Fisher, J. Multiple-mutation theory of carcinogenesis. Nature 181, 651–652 (1958).
Hornsby, C., Page, K. M. & Tomlinson, I. P. What can we learn from the population incidence of cancer? Armitage and Doll revisited. Lancet Oncol. 8, 1030–1038, https://doi.org/10.1016/S1470-2045(07)70343-1 (2007).
Tomasetti, C. & Vogelstein, B. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science 347, 78–81, https://doi.org/10.1126/science.1260825 (2015).
Tomasetti, C., Li, L. & Vogelstein, B. Stem cell divisions, somatic mutations, cancer etiology, and cancer prevention. Science 355, 1330–1334, https://doi.org/10.1126/science.aaf9011 (2017).
Tomasetti, C. & Vogelstein, B. On the slope of the regression between stem cell divisions and cancer risk, and the lack of correlation between stem cell divisions and environmental factors-associated cancer risk. PLoS One 12, 2–5, https://doi.org/10.1371/journal.pone.0175535 (2017).
Wu, S., Powers, S., Zhu, W. & Hannun, Y. A. Substantial contribution of extrinsic risk factors to cancer development. Nature 529, 43–47, https://doi.org/10.1038/nature16166 (2016).
Frank, S. A. Dynamics of Cancer–Incidence, Inheritance, and Evolution http://www.ncbi.nlm.nih.gov/books/NBK1568/pdf/Bookshelf_NBK1568.pdf (2007).
Beerenwinkel, N., Schwarz, R. F., Gerstung, M. & Markowetz, F. Cancer evolution: Mathematical models and computational inference. Syst. Biol. 64, e1–e25, https://doi.org/10.1093/sysbio/syu081 (2014).
Beerenwinkel, N. et al. Genetic progression and the waiting time to cancer. PLoS Comput Biol 3, e225, https://doi.org/10.1371/journal.pcbi.0030225 (2007).
Loeb, K. R. & Loeb, L. A. Significance of multiple mutations in cancer. Carcinogenesis 21, 379–385, https://doi.org/10.1093/carcin/21.3.379 (2000).
Jackson, A. L. & Loeb, L. A. The mutation rate and cancer. Genetics 148, 1483–1490 (1998).
Albini, A. & Sporn, M. B. The tumour microenvironment as a target for chemoprevention. Nat Rev Cancer 7, 139–147, https://doi.org/10.1038/nrc2067 (2007).
Borovski, T., De Sousa E Melo, F., Vermeulen, L. & Medema, J. P. Cancer stem cell niche: The place to be. Cancer Research 71, 634–639, https://doi.org/10.1158/0008-5472.CAN-10-3220 (2011).
Vermeulen, L. et al. Defining stem cell dynamics in models of intestinal tumor initiation. Science 342, 995–998, https://doi.org/10.1126/science.1243148 (2013).
Frank, S. A. Somatic mutation: Early cancer steps depend on tissue architecture. Current Biology 13, R261–R263, https://doi.org/10.1016/S0960-9822(03)00195-7 (2003).
Nowak, M. A., Michor, F. & Iwasa, Y. The linear process of somatic evolution. Proceedings of the National Academy of Sciences of the United States of America 100, 14966–14969, https://doi.org/10.1073/pnas.2535419100 (2003).
Waclaw, B. et al. A spatial model predicts that dispersal and cell turnover limit intratumour heterogeneity. Nature 525, 261–264, https://doi.org/10.1038/nature14971 (2015).
Durrett, R. & Moseley, S. Evolution of resistance and progression to disease during clonal expansion of cancer. Theor. Popul. Biol. 77, 42–48, https://doi.org/10.1016/j.tpb.2009.10.008 (2010).
Yamamoto, K. N., Nakamura, A. & Haeno, H. The evolution of tumor metastasis during clonal expansion with alterations in metastasis driver genes. Sci. Rep. 5, 15886, https://doi.org/10.1038/srep15886 (2015).
Nowak, M. A. & Waclaw, B. Genes, environment, and “bad luck”. Science 355, 1266–1267, https://doi.org/10.1126/science.aam9746 (2017).
Coles, S. An Introduction to Statistical Modeling of Extreme Values (Springer, 2001).
National Cancer Institute, DCCPS, Surveillance Research Program, Surveillance Systems Branch. Surveillance, Epidemiology, and End Results (SEER) Program (www.seer.cancer.gov) Research Data (1973–2012) (2015).
Anderson, D. R. Model Based Inference in the Life Sciences (Springer, 2008).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558, https://doi.org/10.1126/science.1235122 (2013).
O’Callaghan, M. Cancer risk: Accuracy of literature. Science 347, 729–729, https://doi.org/10.1126/science.aaa6212 (2015).
Podolskiy, D. I., Lobanov, A. V., Kryukov, G. V. & Gladyshev, V. N. Analysis of cancer genomes reveals basic features of human aging and its role in cancer development. Nature Communications 7, 1–12, https://doi.org/10.1038/ncomms12157 (2016).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421, https://doi.org/10.1038/nature12477, 23945592[pmid] (2013).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat Genet 47, 1402–1407, https://doi.org/10.1038/ng.3441, 26551669[pmid] (2015).
Podolskiy, D. I. & Gladyshev, V. N. Intrinsic versus extrinsic cancer risk factors and aging. Trends in Molecular Medicine 22, 833–834, https://doi.org/10.1016/j.molmed.2016.08.001 (2016).
Calabrese, P. & Shibata, D. A simple algebraic cancer equation: calculating how cancers may arise with normal mutation rates. BMC Cancer 10, 3, https://doi.org/10.1186/1471-2407-10-3 (2010).
Liao, L.-J., Chou, H.-W., Wang, C.-T., Chung, C.-S. & Lai, M.-S. The impact of second primary malignancies on head and neck cancer survivors: A nationwide cohort study. PLoS ONE 8, e62116, https://doi.org/10.1371/journal.pone.0062116 (2013).
TCGA working group. TCGA data portal https://tcga-data.nci.nih.gov/tcga/dataAccessMatrix.htm (2016).
Michor, F., Iwasa, Y. & Nowak, M. A. The age incidence of chronic myeloid leukemia can be explained by a one-mutation model. Proceedings of the National Academy of Sciences 103, 14931–14934 (2006).
Author information
Authors and Affiliations
Contributions
X.Z., H.F. and A.W. conceived the experiments. All authors developed the models. X.Z. programmed the analysis software and performed the experiments. D.G., S.V. and J.Z. performed the mathematical analysis of the models. All authors analysed the results and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, X., Fröhlich, H., Grigoriev, D. et al. A Simple 3-Parameter Model for Cancer Incidences. Sci Rep 8, 3388 (2018). https://doi.org/10.1038/s41598-018-21734-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21734-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.