Abstract
Renal ischemia-reperfusion (I/R) injury is a major cause of acute kidney injury (AKI). Non-coding RNAs are crucially involved in its pathophysiology. We identified hypoxia-induced long non-coding RNA Malat1 (Metastasis Associated Lung Adenocarcinoma Transcript 1) to be upregulated in renal I/R injury. We here elucidated the functional role of Malat1 in vitro and its potential contribution to kidney injury in vivo. Malat1 was upregulated in kidney biopsies and plasma of patients with AKI, in murine hypoxic kidney tissue as well as in cultured and ex vivo sorted hypoxic endothelial cells and tubular epithelial cells. Malat1 was transcriptionally activated by hypoxia-inducible factor 1-α. In vitro, Malat1 inhibition reduced proliferation and the number of endothelial cells in the S-phase of the cell cycle. In vivo, Malat1 knockout and wildtype mice showed similar degrees of outer medullary tubular epithelial injury, proliferation, capillary rarefaction, inflammation and fibrosis, survival and kidney function. Small-RNA sequencing and whole genome expression analysis revealed only minor changes between ischemic Malat1 knockout and wildtype mice. Contrary to previous studies, which suggested a prominent role of Malat1 in the induction of disease, we did not confirm an in vivo role of Malat1 concerning renal I/R-injury.
Similar content being viewed by others
Introduction
Ischemia-reperfusion (I/R) injury of the kidney is a major cause of acute kidney injury. Due to its high morbidity and mortality it represents a major socioeconomic health problem1. A variety of injurious insults in native kidneys may promote its development (e.g. during cardiac surgery). In addition, it is an unavoidable phenomenon during the kidney transplantation procedure2. We and others have previously shown that non-coding RNAs may contribute to the induction or resolution of this process2.
The number of RNA transcripts without protein-coding potential exceeds 98% of the human genome3. By arbitration these non-coding RNAs (ncRNAs) are divided into long ncRNAs (lncRNAs, ≥200 nucleotides) and small ncRNAs (≤200 nucleotides). Small RNAs such as microRNAs have previously been well characterized. Similar to microRNAs, lncRNAs have recently been shown to regulate gene expression4. The study of the function of lncRNAs is still in its infancy and currently a matter of intense research initiatives. LncRNAs may interact with all components of the cellular machinery, including protein, DNA and RNA4. The lncRNA metastasis-associated lung adenocarcinoma transcript 1 (Malat1) (alternative nomenclature: Nuclear enriched abundant transcript 2 (Neat2)) is nuclear enriched and controls alternative splicing by interacting with serine/arginine splicing factors in nuclear speckle domains5. It has been shown to regulate hyperglycemia induced inflammatory processes in endothelial cells6. It also regulates cell motility via the transcriptional and/or post-transcriptional regulation of motility-related genes7. It has been shown to regulate endothelial cell function and vessel growth8. In addition, it was recently demonstrated to be induced in kidneys of hypoxic mice, where it was mainly enriched in hypoxic proximal tubular epithelial cells9. Moreover, a major initial study found Malat1 to be one of the most highly regulated non-coding transcripts by hypoxia in a breast cancer cell line10. To our knowledge, the in vivo role and downstream mechanism of Malat1 in renal I/R-injury has never been investigated. In the present study Malat1 was found to be highly induced in ischemic kidneys of mice and humans as well as plasma samples of patients with AKI. Malat1 was demonstrated to be enriched in sorted proximal tubular epithelial and endothelial cells ex vivo following murine renal I/R-injury. We investigated the effects of Malat1 modulation on dysregulation of signalling pathways as well as kidney function and survival by subjecting Malat1 knockout mice to renal I/R-injury. Moreover, the downstream mechanisms were analysed in proximal tubular epithelial and endothelial cells. Even though previous studies have suggested a prominent role of Malat1 in the induction of disease, we did not confirm an effect of Malat1 loss on the progression of renal I/R-injury.
Results
LncRNA Malat1 is increased in hypoxic/ischemic kidney injury in humans, mice and cells
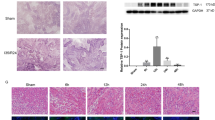
We first assessed the concentration of Malat1 in plasma samples of patients with acute kidney injury (AKI) and found it to be highly increased compared to healthy controls (Fig. 1A). In addition, the expression of Malat1 in kidney biopsies of transplant patients with long compared to short cold ischemia time was strongly increased (Fig. 1B). Patient characteristics are shown in Table 1 and Table 2. Moreover, Malat1 was highly enriched in hypoxic kidneys of mice following induction of I/R-injury (30 minutes of ischemia, 24 h of reperfusion; Fig. 1C). In order to identify the cellular origin of Malat1 we isolated LTA+/KIM1− proximal tubular epithelial cells and Cd31+ endothelial cells by fluorescence-associated cell sorting using specific antibodies and lectins from mouse kidneys following I/R-injury. Intriguingly, Malat1 was found to be enriched in both cell types (Fig. 1D,E). In line with this finding Malat1 was found to be induced in cultured HUVECs and HK-2 cells by hypoxia/reoxygenation (Fig. 1F,G). We performed a subcellular fractionation in HUVECs and HK-2 cells and found that Malat1 is mostly nuclear chromatin-associated in both cell types. However, in HK-2 cells Malat1 was also partly detected in the cytosolic fraction (Fig. 1H,I).
LncRNA Malat1 is increased in hypoxic/ischemic kidney injury in humans, mice and cells. Malat1 expression in plasma samples of patients with acute kidney injury (A). Malat1 expression in human kidney biopsies with short- versus long cold ischemia time (CIT) (n = 5) (B). Malat1 expression in mouse kidney following I/R-injury (n = 6) (C). Malat1 expression in sorted LTA+ and KIM1− cells of murine kidney after I/R injury (n = 5) (D). Malat1 expression in sorted Cd31+ cells of murine kidney after I/R injury (n = 5) (E). Malat1 expression in cultured HK-2 cells (F) and HUVEC (G) (n = 5 each). Malat1 expression in subcellular fractions (cytosolic, nuclear soluble, nuclear chromatin associated) of HK-2 cells (H) and HUVECs (I) (n = 4). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. CIT = cold ischemia time, CTL = control, I/R = ischmemia/reperfusion-injury.
Malat1 expression in HUVEC and HK-2 cells is transcriptionally activated by Hypoxia-inducible Factor-1α
To identify transcription factors activating Malat1 in response to hypoxia, we employed the Genomatix suite of sequence analysis tool (MatInspector). One putative hypoxia response elements (HRE) and 2 Hypoxia-inducible Factor 1 (HIF1) ancillary sequence were found within the promoter region of the Malat1 gene 5 kb upstream of the transcriptional start site in mice as well as one putative HRE and 1 Hypoxia-inducible Factor 1 (HIF1) ancillary sequence in humans (Fig. 2A). In an ELISA based transcriptional activation study, we found Hypoxia-inducible Factor 1 alpha (HIF-1α) highly induced in HUVECs (Fig. 2B) and HK-2 cells (Fig. 2E) upon exposure to hypoxia, as expected. Immunoblotting confirmed HIF-1α activation by hypoxia (Supplemental Fig. 1A) as well as transcription of established HIF-1 target genes CA9, Glut1, PHD2 in HK-2 cells (Supplemental Fig. 1B–D). We used a specific HIF-1α antibody to chromatin-immunoprecipitate HIF-1α bound to the promoter region of Malat1. Subsequently, we designed specific primers for quantitative PCR analysis detecting the HIF-1α binding region in the Malat1 promoter region and found a significantly increased expression level in HUVECs (Fig. 2C) and HK-2 cells (Fig. 2F). In order to stabilize HIF-1α under normoxic conditions, we treated HUVECs and HK-2 cells with Dimethyloxaloylglycine (DMOG), an inhibitor of the prolyl hydroxylase (PHD) and the asparaginyl hydroxylase factor inhibiting HIF (FIH). DMOG induced activation of HIF-1α increased Malat1 expression in HUVECs (Fig. 2D) and HK-2 cells (Fig. 2G). Altogether, we identified HIF-1α as transcriptional activator of Malat1 in HUVECs and HK-2 cells. HIF-1α is part of a heterodimer with HIF-β activating hypoxia-sensitive transcription11.
Malat1 gene expression in HUVECs and HK-2 cells is transcriptionally activated by HIF-1α (=HIF1a). Upstream promoter region of Malat1 spanning 5 kb from the transcriptional start site (TSS) identifies 1 putative Hypoxia Response Elements (HRE) and 2 HIF1 ancillary sequence in mice as well as 1 putative Hypoxia Response Elements (HRE) and 1 HIF1 ancillary sequence in humans (A). Transcriptional activation of HIF-1α in hypoxic HUVECs (B) and HK-2 cells (E) detected by transcriptional activation ELISA (n = 4). qPCR of Malat1 primer amplifying the HIF-1α– promoter region after immunoprecipitation of hypoxic HUVECs (C) and HK-2 cells (F) (n = 5). Malat1 expression after Dimethyloxaloylglycin (DMOG) treatment of HUVECs (D) and HK-2 cells (G) for 24 h. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. ChIP = Chromatin immunoprecipitation, HIF-1α = Hypoxia-inducible factor 1 alpha, TSS = Transcriptional Start Site.
Functional role of Malat1 in HUVEC and HK-2 cell biology
To investigate the functional role of Malat1 in vitro, we used antisense oligonucleotides (GapmeR) to silence Malat1 expression. First, we analysed the impact of Malat1 on cell cycle regulation under basal normoxic conditions and in response to hypoxia/reoxygenation. In this context, cellular DNA was labeled with propidium iodide to differentiate cells in all phases of the cell cycle. Under normoxic conditions, Malat1 antagonism decreased the G1- to S-phase transition in HUVECs (Fig. 3A–E). Likewise, cell cycle progression was diminished in response to hypoxia/reoxygenation in HUVECs demonstrated by a significantly reduced amount of HUVECs entering the S-phase of the cell cycle (Fig. 3F–J). Furthermore, we studied the impact of Malat1 on the proliferation rate of HUVECs. We observed, that silencing of Malat1 in HUVECs reduced the proliferation rate significantly in both normoxia and hypoxia/reoxygenation (Fig. 3K,L). In addition, we compared the migratory capacity of HUVECs depending on Malat1 expression. Intriguingly, antagonizing Malat1 in HUVECs significantly increased the capacity to migrate (Fig. 3M–O). To reveal an effect of Malat1 on apoptosis induction in HUVECs, we stained cells with Annexin and 7AAD and subsequently used fluorescent-activated cell sorting to detect cells undergoing early apoptosis (Annexin + /7AAD-). Here we found a protective effect of Malat1 antagonism on early apoptosis after hypoxia/reoxygenation (Supplemental Fig. 2A). Likewise, Malat1 inhibition decreased Caspase 3/7 activity in hypoxic/reoxygenated HUVECs (Supplemental Fig. 2B). In contrast, Malat1 silencing in HK-2 cells did not result in any functional alteration, concerning cell cycle regulation in normoxia (Supplemental Fig. 3A–C) and upon hypoxia/reoxygenation (Supplemental Fig. 3D–F), proliferation rate in normoxia and upon hypoxia/reoxygenation (Supplemental Fig. 3G,H) as well as apoptosis induction upon hypoxia/reoxygenation (Supplemental Fig. 3I–J). These findings suggest Malat1 to impact on endothelial cell cycle progression, proliferation, migration capacity and apoptosis induction, at least in vitro.
Functional role of Malat1 in HUVEC biology. DNA histograms showing HUVECs detected in G0/G1, S, or G2/M phase of the cell cycle after GapmeR Control (A) and GapmeR Malat1 (B) treatment under basal conditions. Quantification of HUVECs detected in G0/G1 phase (C), S phase (D), and G2/M phase (E) under basal conditions (n = 5). DNA histograms showing HUVECs detected in G0/G1, S, or G2/M phase of the cell cycle after GapmeR Control (F) and GapmeR Malat1 (G) treatment after hypoxia/reoxygenation. Quantification of HUVECs detected in G0/G1 phase (H), S phase (I), and G2/M phase (J) after hypoxia/reoxygenation (n = 5). BrdU incorporation in HUVECs treated with GapmeR Control and GapmeR Malat1 under normoxic conditions (K) and after hypoxia/reoxygenation (L) (n = 5). Scratch migration analysis comparing the migrated area of GapmeR Control- (M) and GapmeR Malat1 (N) treated HUVEC at 12 hours and quantification of the results (O). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. BrdU = Bromodeoxyuridine incorporation.
In vivo role of Malat1 in tubular epithelial cell injury, proliferation and capillary density
As we found Malat1 upregulated in patients and mice upon renal I/R Injury (Fig. 1), our aim was to elucidate the contribution of Malat1 to renal I/R injury in vivo. We used Malat1 knockout - and wildtype mice on a C57BL/6 background and induced unilateral renal I/R injury to analyze possible effects on epithelial- and endothelial injury. First, we detected increased renal expression levels of Kidney Injury Molecule – 1 (KIM-1) after I/R injury in wildtype mice, that were increased similarly in mice lacking the Malat1 gene (Fig. 4A). PAS staining was performed to score the degree of outer medullary tubular epithelial injury. After unilateral renal I/R injury, approximately 70–80% of tubules were injured in both Malat1 knockout and wildtype mice (Fig. 4B–F). To detect proliferating cells, we performed Ki67 staining and observed proliferating cells in Malat1 knockout and wildtype mice to be equally increased (Fig. 4G–K). Capillary integrity upon I/R injury was analyzed by Tomato-Lectin staining. Severe capillary rarefaction was found in Malat1 knockout and wildtype mice without any differences (Fig. 4L–P). These results indicate, that Malat1 is not associated with impaired proliferation or epithelial and capillary integrity in post-ischemic kidneys in vivo.
In vivo role of Malat1 in tubular epithelial cell injury, proliferation and capillary density. KIM-1 gene expression after 24 h of I/R injury (A). Epithelial injury analyzed by PAS staining in wild-type control mice (B), knockout control mice (C), wildtype mice with ischemia/reperfusion injury (D) and knockout mice with ischemia/reperfusion injury (E) and quantification of the results (F). Detection of proliferating cells by Ki67 staining after 24 h of reperfusion in wildtype control mice (G), knockout control mice (H), wildtype mice with ischemia/reperfusion injury (I) and knockout mice with ischemia/reperfusion injury (J) and quantification of the results (K). Evaluation of capillary density by Tomato-Lectin staining after 24 h of reperfusion in wildtype control mice (L), knockout control mice (M), wildtype mice with ischemia/reperfusion injury (N) and knockout mice with ischemia/reperfusion injury (O) and quantification of the results (P). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n = 6 mice in each group. KIM-1 = kidney injury molecule-1; KO = knockout, WT = wildtype, CTL = control, I/R = ischemia/reperfusion-injury, FC = foldchange.
Inflammatory cell influx upon unilateral renal I/R injury
Renal I/R injury is associated with inflammatory cell infiltration. Accordingly, we tested, whether Malat1 mediates inflammatory responses. Expression levels of Il-1beta (Interleukin-1beta), MIP2a (macrophage inflammatory protein 2 alpha), Il-6 (Interleukin-6) and MCP-1 (monocyte chemoattractant protein-1) were highly upregulated after renal I/R injury, but without any signs of amelioration in mice lacking the Malat1 gene (Fig. 5A–D). Moreover, F4/80 staining revealed that macrophages were infiltrating the kidney tissue of Malat1 knockout and wildtype mice upon unilateral renal I/R injury to the same degree (Fig. 5E–I). Likewise, T-cell infiltration (CD3 staining) upon unilateral renal I/R injury was not affected by Malat1 loss (Fig. 5J–N). Taken together, these data indicate that Malat1 does not affect renal I/R-injury induced inflammatory cell infiltration.
Inflammatory gene expression and cell influx upon unilateral renal I/Rinjury. Il-1b (A). MIP2a (B), Il-6 (C) and MCP-1 expression (D) in ischemic murine kidneys. Analysis of F4/80 (red) positive macrophage infiltration in wildtype control (E), knockout control (F), wildtype with ischemia/reperfusion injury (G) and knockout mice with ischemia/reperfusion injury (H) as well as quantification of the results (I) at 7 days of reperfusion. Analysis of CD3 (red) positive T-cell infiltration in wildtype control (J), knockout control (K), wildtype with ischemia/reperfusion injury (L) and knockout mice with ischemia/reperfusion injury (M) as well as quantification of results (N) at 7 days of reperfusion. *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001; n = 6 mice in each group. Il-1b = Interleukin-1beta, MIP2a = Macrophage inflammatory protein-2 alpha; Il-6 = Interleukin-6; MCP-1 = monocyte chemoattractant protein – 1, KO = knockout, WT = wildtype, CTL = control, I/R = ischemia/reperfusion-injury, FC = foldchange.
Accumulation of interstitial fibrosis after unilateral renal I/R injury
In the course of renal I/R injury, accumulation of tubulointerstitial fibrosis ultimately results in an impairment of kidney function. In order to investigate a potential contribution of Malat1, we analyzed mRNA levels of pro-fibrotic genes and additionally used Masson’s trichrome staining to detect interstitial fibrosis. Expectedly, Col1a2 (Collagen1-alpha2), Col III (Collagen 3) and TGF-beta (Transforming growth factor beta) expression levels were increased after unilateral renal I/R injury, but there was no significant difference between Malat1 knockout and wildtype mice (Fig. 6A–C). Interstitial fibrosis was increased after renal I/R injury, but did not show detectable differences between Malat1 knockout and wildtype mice (Fig. 6D–H).
Accumulation of interstitial fibrosis after unilateral renal I/R injury at 7 days of reperfusion. Col1a2 (A), Col III (B) and TGFb (C) expression in ischemic murine kidneys. Detection of interstitial fibrosis by Massons’s trichrome staining in wildtype control (D), knockout control (E), wildtype with ischemia/reperfusion injury (F) and knockout mice with ischemia/reperfusion injury (G) as well as quantification of the results (H). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. Col III = Collagen 3, TGFbeta = Transforming growth factor beta, KO = knockout, WT = wildtype, CTL = control, I/R = ischemia/reperfusion-injury, FC = foldchange. n = 6 mice in each group.
Impact of Malat1 on kidney function and survival after bilateral renal I/R injury
To evaluate the influence of Malat1 on kidney function, we collected blood samples of mice before, at day 1, day 3 and day 7 after bilateral renal I/R injury in Malat1 knockout and wildtype mice and measured creatinine and urea levels. Creatinine and urea levels in Malat1 knockout and wildtype mice were similarly increased at day 1 after bilateral renal I/R injury and decreased continuously without any significant difference (Fig. 7A,B). In addition, our survival study of Malat1 knockout and wildtype mice subjected to bilateral renal I/R injury showed no significant differences (Fig. 7C).
Impact of Malat1 on kidney function and survival after bilateral renal I/R injury. Creatinine levels in Malat1 knockout and wildtype mice before induction of ischemia/reperfusion injury, 1 day after ischemia/reperfusion injury, 3 days after ischemia/reperfusion injury and 7 days after ischemia/reperfusion injury (A). Urea levels in Malat1 knockout and wildtype mice before induction of ischemia/reperfusion injury, 1 day after ischemia/reperfusion injury, 3 days after ischemia/reperfusion injury and 7 days after ischemia/reperfusion injury (B). Survival study in Malat1 knockout- and wildtype mice after bilateral renal ischemia/reperfusion injury (C). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. KO = knockout, WT = wildtype. n = 10 mice in each group.
Whole genome RNA analysis in Malat1 knockout and wildtype mice
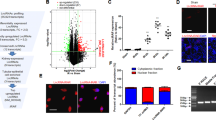
In order to find potentially dysregulated signalling pathways and small RNA expression levels in vivo we performed a whole genome mRNA array as well as a small RNA sequencing approach in ischemic kidneys of Malat1 knockout and wildytpe mice. The data was subsequently filtered for differentially expressed mRNA and small RNA. Passed candidates were finally hierarchically clustered and displayed as heatmap (Fig. 8A for mRNA and B for small RNA). Additionally, most highly regulated candidates are summarized in Supplemental Table 1 (mRNA) and Supplemental Table 2 (small RNA). Only minor differences in mRNA and small RNA expression levels were detected. Epidermal growth factor receptor kinasesubstrate 8-like protein 3 (Eps8l3; log2 fold change of 1.4), 2010003K11Rik (log2 fold change of 1.4) and aldehyde dehydrogenase family 1, subfamily A7 (Aldh1a7; log2 fold change of 1.2) were found to be the top upregulated candidates. Gm5878 (log2 fold change of 3.9), aldehyde dehydrogenase 1 familymember A3 (Aldh1a3; log2 fold change of FC −3.0) and solute carrier family 14 (urea transporter), member 2 (Slc14a2 log2 fold change of 2.7) were top downregulated candidates. These candidates were validated by qPCR (see Supplemental Fig. 4A–F). Small RNA sequencing revealed several transcripts to be slightly regulated: miR-15 (log2 fold change 1.5), miR-383–5p (log2 fold change of 1.3) and miR-146b-5p (log2 fold change of 1.1) were top upregulated candidates, while Gm24706 (log2 fold change of 1.4), miR-7046–3p (log2 fold change of 1.1) and miR-203–5p (log2 fold change of 0.9) were top downregulated transcripts. Supplemental Fig. 4G shows PANTHER gene ontology enrichment analysis of dysregulated gene expression. Gene ontology analysis revealed the 5-Hydroxytryptamin degradation as the strongest influenced pathway, biological adhesion processes as the mainly affected biological processes, antioxidant activity as the primarily controlled molecular function and cell adhesion molecules as the mostly affected protein class.
Heatmap of hierarchically clustered mRNA-array data in ischemic kidneys of Malat1 knockout and wildtype mice (A): Signal intensities were log2 transformed, centered and normalized. Data was clustered using Spearman rank correlation and average linkage. Heatmap of hierarchically clustered small RNA-sequencing data in ischemic kidneys of Malat1 knockout and wildtype mice (B): Signal intensities were log2 transformed, centered and normalized. Data was clustered using Spearman rank correlation and average linkage.
Discussion
In the present study we investigated the in vivo role of the lncRNA Malat1 on the induction and progression of renal I/R-injury. While we did detect increased expression levels of Malat1 in ischemic kidney biopsies and plasma of patients with AKI, ischemic mouse kidneys and sorted endothelial cells and tubular epithelial cells ex vivo, we were unable to report an overt phenotype of Malat1 knockout mice in contrast to previous studies investigating the in vivo role of Malat1 in other disease settings. More specifically, we found that in unilateral renal I/R-injury Malat1 knockout mice exhibited the same degree of outer medullary injury, capillary rarefaction, fibrosis, infiltration of inflammatory cells, inflammatory gene expression and proliferation as did Malat1 wildtype mice. In bilateral renal I/R-injury Malat1 knockout mice showed a similar level of kidney dysfunction and survival as Malat1 wildtype mice.
The role of microRNAs in the kidney has been extensively studied over the past 10 years. Very limited information is available on the role and functional importance of lncRNAs. We here provide the first study investigating the role of a lncRNA in experimental acute kidney injury. It is becoming evident that lncRNAs are important epigenetic regulators of tissue homeostasis during development and disease4. Although the function of most lncRNAs is unexplored to date, a mechanistic annotation has been suggested by Howard Chang’s group4,12. Here, lncRNAs were described to either function as (1) signal lncRNAs, which are spatiotemporally transcribed in response to developmental cues or cellular context; (2) decoy lncRNAs, which impact on transcriptional regulation by titrating away transcription factors and other proteins from chromatin; (3) guide lncRNAs, which sequester ribonucleoprotein complexes and direct them to chromatin; (4) scaffold lncRNAs, which interact with multiple partners to form a chromatin modifying complex.
A few lncRNAs have thus far been functionally characterized. For instance, the long intergenic RNA (lincRNA) Air and X-chromosome inactivation by Xist (X-inactive specific transcript) play a critical role in the regulation of imprinting13. The lncRNA H19 is imprinted with maternal expression14. It has been shown to have a tumour-suppressive effect both in vitro and in vivo15. A lncRNA termed cardiac hypertrophy associated transcript (Chast) was found to be increased in mice and humans with cardiac hypertrophy16. Gain- and loss-of function studies underlined the importance of this transcript for cardiac hypertrophy and fibrosis development. RNA-sequencing analysis in a UUO mouse model of kidney fibrosis recently revealed a number of lncRNAs to be highly dysregulated17.
Malat1 was first described in metastasizing non-small cell lung cancer, hence its name18. Owing to the high degree of conservation between different species and high expression levels Malat1 knockout mice have been generated to further elucidate the role of this lncRNA. Abnormalities during embryonic or post-natal development have not been detected19,20,21, indicating that Malat1 may only have a major role under pathological conditions. Malat1 has been demonstrated to sequester serine/arginine (SR) splicing factors in nuclear speckle domains, thereby regulating alternative splicing4,5. In endothelial cells hypoxic stress has been shown to induce the expression of Malat1, leading to enhanced proliferation and migration as well as a reduction of apoptosis8. In vivo, Malat1 knockout mice were characterized by impaired proliferation of endothelial cells and reduced vascularization in the developing retina8. In a hindlimb ischemia model, pharmacological silencing of Malat1 reduced blood flow recovery during reperfusion and capillary density8. High glucose treatment induced the expression of Malat1 in endothelial cells. This was paralleled by induction of inflammatory gene expression6. In another study Malat1 was found to be increased in the cortex of mice with diabetic nephropathy as well as in cultured podocytes under high glucose conditions22. Malat1 silencing restored podocyte function in vitro22. In diabetic rats Malat1 was highly upregulated in the kidney. Here, Malat1 influenced pyroptosis by sponging miR-23c23. Recently, the induction of Malat1 by inspiratory hypoxia was assessed in mice9. It was shown that the induction was most pronounced in kidney and testis. Hypoxic induction in the kidney was confined to proximal rather than distal tubular epithelial cells as shown by in situ hybridization9. Direct oxygen-dependent regulation of Malat1 was further validated in isolated primary kidney epithelial cells9. However, most of the available literature did not assess the in vivo role of Malat1 by using Malat1 knockout mice, which are available19,20,21.
In contrast to the aforementioned studies the absence of Malat1 does not seem to impact on the resolution of disease. In a study involving genotoxic stress using diethylnitrosamine (DEN) to induce liver cancer Malat1 knockout mice and littermate controls were investigated24. Here, Malat1 knockout mice did not differ with respect to tumor development after 1 year compared to controls. In thoracic aortic constriction (TAC), a model of cardiac pressure overload, Malat1 knockout mice were investigated compared to wildtype controls25. Absence of Malat1 did not affect cardiac hypertrophy upon pressure overload. There are numerous in vitro studies claiming a role for Malat1 in liver cancer progression26,27,28. In addition, in blood samples of patients with myocardial infarction Malat1 was shown to be regulated29. Malat1 was shown to impact on proliferation of cardiomyocytes in vitro30. However, in the in vivo studies referenced above changes between wildtype and Malat1 knockout animals were not detected24,25. Taking into account these in vitro and biomarkers studies as well as our results of increased expression levels in hypoxic cells and blood of patients, we believe, that Malat1 may have a role as a biomarker of different pathologies, including AKI. Most importantly, however, its role in vivo is either minimal or masked by redundant and/or overwhelming mechanisms. Whole genome expression analysis as well as small RNA sequencing revealed only minimal differences between the two groups. Most highly dysregulated genes included Epidermal growth factor receptor kinase substrate 8-like protein 3, 2010003K11Rik and aldehyde dehydrogenase family 1, subfamily A7. Intriguingly, ALDHs are described to be upregulated in response to oxidative stress to prevent further impairment and are associated with disease, in which oxidative stress plays an important role, e.g. diabetes and acute lung injury31. Most highly downregulated candidates were Gm5878, aldehyde dehydrogenase 1 family member A3 and solute carrier family 14 (urea transporter), member 2. Small RNA sequencing revealed miR-15, miR-383-5p and miR-146b-5p as top upregulated candidates. Interestingly, miR-15 is described to play a role in apoptosis by targeting Bcl2 in chronic lymphocytic leukemia (CLL)32 and mir-146b-5p was found to be induced in AKI and fibrosis33. Top downregulated small RNA candidates in our sequencing analysis were Gm24706, miR-7046-3p and miR-203-5p. Although we identified several mRNA and small RNA with potential impact on oxidative cellular stress and apoptosis induction to be slightly dysregulated and to be in association with AKI and fibrosis, the differences were only marginal and indicate Malat1 loss to only have a minor influence in vivo.
It has previously been shown, that Malat1 affects hind limb ischemia8. In contrast to this study, in which Malat1 was pharmacologically silenced by GapmeRs we used mice with a genetic deletion of Malat1. Compensatory mechanisms might be activated in Malat1 knockout mice during the embryonic developmental phase, which might cause possible adaptive response mechanisms, which could contribute to the absence of discernable effects in our study. Inducible knockout strategies could be used to bypass this effect and to identify possible compensatory mechanisms.
In conclusion, using two models of renal I/R-injury we did not detect a disease-specific phenotype of Malat1 knockout mice. Malat1 may be a viable biomarker of I/R-injury of the kidney, as evidenced by increased levels in plasma and kidney biopsies of patients in our study. However, its role in vivo on relevant signalling pathways is likely limited.
Concise Methods
Patients with AKI
Plasma samples of patients with AKI were collected as part of the HANDOUT trial34. In brief, in this study patients in seven ICUs of the tertiary care centre at the Hannover Medical School suffering from AKI were evaluated for inclusion. The study protocol was approved by the Hannover Medical School Ethics Committee and was conducted in accordance with the declaration of Helsinki and German Federal Guidelines. Written informed consent was obtained by the patient or his/her legal representative. The inclusion criteria were non-post-renal AKI with RRT dependence indicated by the loss of kidney function of >30% calculated estimated glomerular filtration rate (eGFR) with either the Modification of Diet in Renal Disease (MDRD) or Cockcroft–Gault equation and/or cystatin C-GFR within 48 hours prior to inclusion and oliguria/anuria (<30 mL/h >6 hours prior to inclusion) or hyperkalemia (>6.5 mmol/L) or severe metabolic acidosis (pH <7.15, bicarbonate <12). Exclusion criteria were pre-existing chronic kidney disease as defined by eGFR <60 mL/min or a serum creatinine concentration >1.7 mg/dL more than 10 days prior to initiation of the first RRT. Patient characteristics are shown in Table 1.
Kidney biopsies of patients
In kidney transplant recipients, who presented with prolonged (707 ± 73 minutes, n = 5) and short (208 ± 56 minutes, n = 5) cold ischemia time (CIT), renal biopsy specimens and clinical and demographic data were collected. Patient characteristics are shown in Table 2. The study protocol was approved by the Hannover Medical School Ethics Committee (“Ethikkommission der Medizinischen Hochschule Hannover”) and was conducted in accordance with the declaration of Helsinki and German Federal Guidelines. Written informed consent was obtained by the patient or his/her legal representative.
Animals
Male Malat1 knockout mice and littermate wildtype control mice (both on a C57BL/6 background) were housed under standard conditions20. These mice were previously generated and described20. These mice are viable and fertile and do not display a pathological phenotype under basal conditions. Mice that were 10 to 12 weeks old weighing between 20 and 30 g were used for all experiments. In unilateral I/R-injury experiments n = 6 animals per group were included, in bilateral I/R-injury experiments 10 animals per group were included. All animal experimental procedures were in agreement with institutional and legislational regulations and were approved by local authorities of lower Saxony (“Niedersächsisches Landesamt für Verbraucherschutz und Lebensmittelsicherheit”; approval number 13/1245).
Genotyping of mice
As described previously20 a three-primer-PCR strategy was used for genotyping of Malat1 knockout mice. Primer 1 (CACTCTGGGAATGTTTTTGG), Primer 2 (CAGGAAAACGCAAAAGGTGT) and Primer 3 (TGTCGAAAAGAGGTGGTGTG) produced a 120 bp fragment for the wild-type allele and a 204 bp fragment for the Malat1-deleted locus. PCR conditions were as follows: 4 min 95 °C, followed by 30 cycles of 30 sec. 95 °C, 30 sec. 56 °C, 30 sec. 72 °C, and a final elongation for 10 min at 72 °C. Genotyping analyses of Malat1 wildtype and knockout mice are shown in Supplemental Fig. 4E. Malat1 absence was also confirmed by qPCR (Supplemental Fig. 1F).
References
Kelly, K. J. Acute renal failure: Much more than a kidney disease. Semin. Nephrol. 26, 105–113 (2006).
Bon, D. et al. New strategies to optimize kidney recovery and preservation in transplantation. Nat. Rev. Nephrol 8, 339–347 (2012).
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Lorenzen, J. M. & Thum, T. Long noncoding RNAs in kidney and cardiovascular diseases. Nat. Rev. Nephrol. 12(6), 360–73 (2016).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell. 39, 925–938 (2010).
Puthanveetil, P., Chen, S., Feng, B., Gautam, A. & Chakrabarti, S. Long non-coding RNA MALAT1 regulates hyperglycaemia induced inflammatory process in the endothelial cells. J Cell Mol Med 19, 1418–1425 (2015).
Tano, K. et al. MALAT-1 enhances cell motility of lung adenocarcinoma cells by influencing the expression of motility-related genes. FEBS Lett. 584, 4575–4580 (2010).
Michalik, K. M. et al. Long noncoding RNA MALAT1 regulates endothelial cell function and vessel growth. Circ. Res 114, 1389–1397 (2014).
Lelli, A., et al. Induction of long noncoding RNA MALAT1 in hypoxic mice. Hypoxia (Auckl) 3, 45–52. eCollection (2015).
Choudhry, H. et al. Extensive regulation of the non-coding transcriptome by hypoxia: role of HIF in releasing paused RNApol2. EMBO Rep. 15(1), 70–6 (2014).
Mole, D. R. & Ratcliffe, P. J. Cellular oxygen sensing in health and disease. Pediatr. Nephrol. 23(5), 681–94 (2008).
Wang, K. C. & Chang, H. Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Lorenzen, J. M., Martino, F. & Thum, T. Epigenetic modifications in cardiovascular disease. Basic Res. Cardiol. 107, 245 (2012).
Bartolomei, M. S. Genomic imprinting: employing and avoiding epigenetic processes. Genes Dev. 23, 2124–2133 (2009).
Hao, Y., Crenshaw, T., Moulton, T., Newcomb, E. & Tycko, B. Tumour-suppressor activity of H19 RNA. Nature 365, 764–767 (1993).
Viereck, J. et al. Long noncoding RNA Chast promotes cardiac remodeling. Sci. Transl. Med. 8(326), 326ra22 (2016).
Arvaniti, E. et al. Whole-transcriptome analysis of UUO mouse model of renal fibrosis reveals new molecular players in kidney diseases. Sci. Rep. 6, 26235 (2016).
Ji, P. et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 22(39), 8031–41 (2003).
Zhang, B. et al. The lncRNA Malat1 is dispensable for mouse development but its transcription plays a cis-regulatory role in the adult. Cell.Rep. 2(1), 111–23 (2012).
Eissmann, M. et al. Loss of the abundant nuclear non-coding RNA MALAT1 is compatible with life and development. RNA Biol. 9(8), 1076–87 (2012).
Nakagawa, S. et al. Malat1 is not an essential component of nuclear speckles in mice. RNA. 18(8), 1487–99 (2012).
Hu, M. et al. LncRNA MALAT1 is dysregulated in diabetic nephropathy and involved in high glucose-induced podocyte injury via its interplay with β-catenin. J. Cell. Mol. Med. 21(11), 2732–2747 (2017).
Li, X. et al. Long noncoding RNA MALAT1 regulates renal tubular epithelial pyroptosis by modulated miR-23c targeting of ELAVL1 in diabetic nephropathy. Exp. Cell Res. 350(2), 327–335 (2017).
Miard, S. et al. Absence of Malat1 does not prevent DEN-induced hepatocarcinoma in mice. Oncol. Rep. 37(4), 2153–2160 (2017).
Peters, T. et al. Long Non-Coding RNA Malat-1 Is Dispensable during Pressure Overload-Induced Cardiac Remodeling and Failure in Mice. PLoS One 11(2), e0150236 (2016).
Chen, L., Yao, H., Wang, K. & Liu, X. Long Non-Coding RNA MALAT1 Regulates ZEB1 Expression by Sponging miR-143-3p and Promotes Hepatocellular Carcinoma Progression. J. Cell. Biochem. 118(12), 4836–4843 (2017).
Hou, Z. et al. The long non-coding RNA MALAT1 promotes the migration and invasion of hepatocellular carcinoma by sponging miR-204 and releasing SIRT1. Tumour Biol. 39(7), 1010428317718135 (2017).
Malakar, P. et al. Long Noncoding RNA MALAT1 Promotes Hepatocellular Carcinoma Development by SRSF1 Upregulation and mTOR Activation. Cancer Res. 77(5), 1155–1167 (2017).
Vausort, M., Wagner, D. R. & Devaux, Y. Long noncoding RNAs in patients with acute myocardial infarction. Circ. Res. 115(7), 668–77 (2014).
Zhao, J., Li, L. & Peng, L. MAPK1 up-regulates the expression of MALAT1 to promote the proliferation of cardiomyocytes through PI3K/AKT signaling pathway. Int. J. Clin. Exp. Pathol. 8(12), 15947–53 (2015).
Singh, S. et al. Aldehyde dehydrogenases in cellular responses to oxidative/electrophilic stress. Free. Radic. Biol. Med. 56, 89–101 (2013).
Cimmino, A. et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl. Acad. Sci. USA 102(39), 13944–9 (2005).
Pellegrini, K. L. et al. Application of small RNA sequencing to identify microRNAs in acute kidney injury and fibrosis. Toxicol. Appl. Pharmacol. 312, 42–52 (2016).
Faulhaber-Walter, R. et al. The Hannover Dialysis Outcome study: comparison of standard versus intensified extended dialysis for treatment of patients with acute kidney injury in the intensive care unit. Nephrol. Dial. Transplant. 24(7), 2179–86 (2009).
Acknowledgements
We acknowledge support of the Else Kröner Fresenius foundation in form of the memorial scholarship (to JML),the National Competence Center in Research Kidney.CH in form of a junior grant (to JML) and a nephro physician scientist grant (to MK). TT acknowledges support of the Integrated Treatment and Research Center Transplantation (IFB-Tx).
Author information
Authors and Affiliations
Contributions
J.M.L. conceived the study, planned the experiments and wrote the paper. M.K. performed in vitro experiments. G.H., H.S., A.K. and A.H. contributed to in vitro experiments. T.K. performed ex vivo cell sorting experiments. R.S., M.Z., T.T., S.R., T.T., H.H., I.S.Z., C.G. performed animal experiments. J.T.K., D.F., R.W. helped in obtaining plasma and tissue samples of patients.
Corresponding author
Ethics declarations
Competing Interests
TT is founder of Cardior Pharmaceuticals GmbH. The other authors have nothing to disclose.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kölling, M., Genschel, C., Kaucsar, T. et al. Hypoxia-induced long non-coding RNA Malat1 is dispensable for renal ischemia/reperfusion-injury. Sci Rep 8, 3438 (2018). https://doi.org/10.1038/s41598-018-21720-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21720-3
This article is cited by
-
Ischemic tubular injury: Oxygen-sensitive signals and metabolic reprogramming
Inflammopharmacology (2023)
-
The renoprotective effects of gallic acid on cisplatin-induced nephrotoxicity through anti-apoptosis, anti-inflammatory effects, and downregulation of lncRNA TUG1
Naunyn-Schmiedeberg's Archives of Pharmacology (2022)
-
Divergent regulation of lncRNA expression by ischemia in adult and aging mice
GeroScience (2022)
-
Arid2-IR promotes NF-κB-mediated renal inflammation by targeting NLRC5 transcription
Cellular and Molecular Life Sciences (2021)
-
Reverse-genetics studies of lncRNAs—what we have learnt and paths forward
Genome Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.