Abstract
Previous studies have revealed significant differences in microbiome compositions between infants delivered via cesarean section (C-section) and natural vaginal birth. However, the importance of the delivery mode in the first days of life remains unclear. Importantly, this stage is minimally affected by infant feeding. Here, we used a metagenomic sequencing technique to characterize the meconium microbiome from the feces of a Chinese cohort of vaginally and C-section-delivered infants, including in vitro fertilization (IVF) newborns, during the first 24 h after birth. Meconium microbiome diversity was higher in vaginally delivered infants than that in C-section-delivered infants. Propionibacterium species were most abundant in the vaginally delivered infants, whereas the C-section group had high levels of Bacillus licheniformis. The two IVF newborns delivered by C-section harbored microbial communities similar to the vaginal microbiome in terms of taxonomic composition. Metabolic functions of the C-section group suffered more from the influence of the dominant group (B. licheniformis), whereas the vaginal group was more homogeneous, with a metabolism dominated by multi-microbes. Moreover, different modes of delivery affected the antibiotic resistance gene (ARG) prevalence. These findings provide novel information for the development of strategies to guide a healthy mode of delivery and promote the formation of healthy microbiota.
Similar content being viewed by others
Introduction
The complex community of the gut microbiota plays key roles at different stages of human life, including the newborn phase1. Early colonization by the microbiota is the basis of its assistance with later physiological growth, immunological maturity and neurological homeostasis for adequate infant development. However, it remains unclear at what stages of fertilization, pregnancy or birth this process of microbiota establishment begins.
Analyses of microorganisms in human follicular fluid and endometria early during the fertilization period in women undergoing in vitro fertilization (IVF) have revealed the presence of bacterial species correlated with IVF pregnancy outcomes and embryo transfer rates2,3. During pregnancy, a unique low-abundance microbiome has been detected in placental tissue4, amniotic fluid5, umbilical cord blood6, and fetal membranes7 from healthy newborns without any indication of infection or inflammation. However, major microbial colonization of the gastrointestinal tract begins at birth, when an infant comes into contact with microbes from the extra-uterine environment8,9. This initial exposure of the fetus during pregnancy and delivery is assumed to lead to gradual compositional development and finally to the establishment of a stable, individual-specific microbiota10. The mode of delivery and subsequent environment exposures greatly influence the composition of the microbiota in the infant. These early colonization events and the establishment of the intestinal microbiota have an effect on the development of a variety of diseases later in life11. Compared with vaginal delivery, cesarean section (C-section) was demonstrated to correlate with increase in the incidence of diseases such as asthma12, allergic rhinitis, obesity13, celiac disease14,15 and type I diabetes16. The microbiota of infants delivered by C-section can be partially restored by re-exposure to maternal vaginal fluids at birth17. Therefore, identifying the pioneer environment established by the colonization of pioneer microbes may provide a new perspective for the prevention and treatment of disease.
A recent systematic reviewer by Rutayisire et al.11 indicated that C-sections were associated with a lower abundance and diversity of the phyla Actinobacteria and Bacteroidetes and a higher abundance and diversity of the phylum Firmicutes from birth to 3 months of life based on analyses using culture-dependent or culture-independent techniques (16 S rRNA gene sequencing/amplification, molecular methods or fluorescence in situ hybridization). At the taxonomic level, the genera Bifidobacterium and Bacteroides appear significantly more frequently in vaginally delivered infants than in C-section delivered infants. Conversely, these latter infants are colonized more by bacteria from the genera Clostridium and Lactobacillus. A study by Dominguez Bello et al.8 using 16 S rRNA sequencing demonstrated that the guts microbiota of vaginally delivered Amerindian infants resembled their mothers’ vaginal microbiota, which were dominated by Lactobacillus, Prevotella, or Sneathia spp.; in contrast, infants delivered by C-section were colonized by common skin and environmental microbes, such as Staphylococcus, Corynebacterium, and Propionibacterium spp. Gosalbes et al.18 employed 16 S rRNA sequencing to examine the meconium microbiota of Spanish infants and found that these meconium microbiota were dominated by lactic acid or enteric bacteria. Dong Liu et al.19 investigated the intestinal microbiota of Chinese newborn infants on days 2 and 4 of postnatal life and demonstrated that the dominant bacteria found in vaginally delivered infants were Acinetobacter spp., Bifidobacterium spp., and Staphylococcus spp., whereas Citrobacter spp., E. coli, and Clostridium difficile were more common in C-section delivered infants.
Diversity analyses of the microbiota in vaginal and C-section delivered infants are partially dependent on race or geographical differences20. Additionally, the interval of meconium sample collection varies from within 24 h after delivery to 4 days, which might contribute to differences in the microbiota with different mode of delivery8,19,21.
Here, we assessed the gut microbiota composition in the first-pass meconium of 8 vaginally and 10 C-section (including two IVF newborn)-delivered Chinese infants. We assembled gut microbial genomes and demonstrated gut microbiome signatures that are characteristic of early life. In conclusion, we identified the characteristics of the pioneer environmental pattern according to different modes of delivery, including both the composition and metabolic functions of the microbiota.
Results
Characteristics of the meconium microbiomes associated with different modes of delivery
To characterize the infant meconium microbiome, we performed metagenomic sequencing analysis and compared the microbial community structures between vaginally and C-section-delivered infants. As shown in Table 1, 16 of 18 infants (8 from vaginal delivery and 8 from C-section delivery) were born at term after a normal pregnancy at a gestational age of 37–42 weeks, and 2 of 18 infants (C-section) were born at a gestational age of 36–37 weeks after an IVF pregnancy. There were no significant differences in the mother’s age, mother’s height, mother’s gestational weight gain, birth weight or infant length between the two groups. The vaginal delivery group had higher gestational age days than the C-section group (P = 0.044). Patients with a higher pregnancy weight were more common in the vaginal delivery group.
We obtained an average of 33.74 GB of raw data for each sample, with a range from 30.15 GB to 60.04 GB. After filtering out the low-quality reads and host contamination, we obtained 2.33 GB to 5.38 GB of clean data for each sample (average 3.62 GB).
The data yielded sequences belonging to 9 phyla and 141 species, including several DNA viruses (Fig. S1A,B) and a small amount of fungi (data not shown). Firmicutes was the most abundant phylum, accounting for 37.4% of the total reads, followed by Actinobacteria with 28.3%, Proteobacteria with 21.2%, unidentified viruses with 8.9%, Bacteroidetes with 3.7% and other phyla with 0.4% (Fig. S1C).
Mode of delivery significantly affected the first-pass meconium microbiome
To illustrate the relationship between the microbial composition and metadata, redundancy analysis (RDA), multivariate ANOVA based on dissimilarities (Adonis) and principal component analysis (PCA) was performed using species data at all sample levels. Figures 1A and S2 revealed that the delivery modes were the most significant contributors to the composition of the newborn neonate gut microbiomes in contrast to other factors (Figs 1A, S2A,B and Table S1). The Simpson and Shannon-Weiner indices, which measure the alpha-diversity of the species richness and the evenness of distinct microbes in a community, respectively, were plotted and compared. The results indicated that the microbiome of the vaginally delivered infants was somewhat more species-rich and diverse than the microbiome of the C-section-delivered infants (Fig. S3). The metagenomic sequence data were equivalent at the species (strain) level classification. A LEfSe analysis and cladogram were performed to investigate differences in the community composition between groups (Fig. 1B,C). The phyla Firmicutes and Deinococcus-Thermus were significantly enriched in the vaginally delivered group, whereas Actinobacteria was higher in abundance in the C-section delivered group (Fig. S4). There were six significantly different classes, with enrichment of Actinobacteria, Gammaproteobacteria, and Betaproteobacteria in the vaginal group and Deinococcus, Alphaproteobacteria and Bacilli in the C-section delivered group (Fig. 1B,C). The microbial composition also significantly differed at the species level, with 10 significantly different species between groups. Propionibacterium acnes, Parabacteroides unclassified, and Comamonas testosteroni exhibited relatively higher abundance in the vaginally delivered group. Bacillus licheniformis, Bacillus amyloliquefaciens, Eubacterium rectale, Bacteroides vulgatus, Aeromonas veronii, Faecalibacterium prausnitzii and Deinococcus unclassified were relatively more abundant in the C-section-delivered group (Fig. 1B,C).
Comparison of the microbiomes of vaginally delivered and C-section-delivered newborns. (A) Principal-component analysis (PCA) plot based on the relative taxa abundance in the fecal microbiomes of neonates with mode of delivery, gestation age, pregnancy weight or gender groups. Sample are marked by the group type. The number is the grouping boundaries. (B) LEfSe comparison of microbiomes from meconium samples from vaginally delivered and C-section-delivered neonates. Enriched taxa in samples from vaginally delivered infants with different level classifications with a positive linear discriminant analysis (LDA) score are shown in green; C-section samples with a negative LDA score are shown in red (>3.5 in both cases). (C) Cladogram derived from LEfSe analysis of metagenomic sequences from C-section delivered newborn and vaginally delivered neonate meconium samples. Green shaded areas indicate microbe orders that more consistently describe the fecal microbiome from vaginally delivered infants; red shaded areas indicate microbe orders that more consistently describe the meconium microbiome from C-section-delivered infants. The prefixes “p”, “c”, “o”, “f”, “g”, “s”, and “t” represent the annotated level of phylum, class, order, family, genus, species and strain.
Additionally, we identified a number of viruses in the meconium. The bacteriophage families were the most prevalent DNA virus families detected in early life specimens. Further study using an extended error bar plot analysis found that the Pseudomonas phage F116 was more enriched in the C-section delivered neonate specimens (Fig. S5).
The phylogenetic composition of the IVF newborn samples (CS4 and CS5) clustered similarly to the samples from the vaginally delivered fecal microbiomes (Fig. 2A). Additionally, the CS4 samples had an abundance of the human herpesvirus 6B, B. licheniformis and Cronobacter sakazakii microbes, whereas CS5 were richer in Comamonas unclassified, porcine type-C oncovirus and Escherichia unclassified (Fig. 2B). Other samples showed distinct phylogenetic compositions between the vaginal and C-section delivery groups using Bray Curtis cluster analysis.
Spearman correlation heat map analysis showed that the fecal microbiomes from IVF infant samples were clustered with those from the vaginally delivered infant samples. (A) CS4 and CS5 samples represent IVF neonates whose fecal microbiomes clustered with those from the vaginally delivered infants. (B) Distribution of the relative abundance of taxa between the CS4 and CS5 IVF samples. Horizontal bars indicate 95% confidence intervals from the mean proportion.
Correlation of the predominant microbiome with different modes of delivery
Extended error bar plots and associated confidence intervals with bootstrapping were constructed to determine the mean differences for each species between the vaginally and C-section-delivered fecal samples (Fig. 3A). Compared with the vaginal delivery samples, the C-section group contained significantly higher levels of B. licheniformis and B. amyloliquefaciens at the species level (P < 0.05) (Fig. 3A). The relative abundance of P. acnes was higher in the vaginal delivery group than in the C-section delivered samples. Both the C-section- and vaginally delivered infants showed a relatively low abundance of Lactobacillus. However, due to the relative lack of lactobacilli in the C-section infants, the diversity and abundance of lactobacilli in the vaginal abundance and delivery samples were still higher than the abundance in the C-section group (Fig. S6).
Correlation of the predominant microbiomes under different modes of delivery. (A) Heat map analysis generated with LEfSe showing the taxonomic strains with significantly different abundance in C-section and vaginally delivered neonates. The dendrogram of pathway abundance is colored red to indicate a negative LDA score and green to indicate a positive LDA score. Over/underrepresentation is depicted by a color gradient. (B,C) Correlations were expressed by Spearman’s correlation coefficients. Only the first 30 abundant species were analyzed. Network analysis of C-section-delivered infant meconium microbes (B) and vaginally delivered infant meconium microbes (C). Nodes indicate taxonomic affiliations at the species level. Red lines indicate negative correlations; gray lines indicate positive correlations. The line thickness represents the correlation strength. The size of a node is determined by the thickness of all lines on the node.
Recently, research has shown that the neonate intestinal microbiota is significantly affected by the mother’s microbiome. The differences in the mode of delivery may affect the fecal, vaginal, skin, mouth and environmental microbiota and, consequently, the establishment of the pioneer environment. The mode of delivery affected taxonomies and consistencies across individuals, demonstrating that a pioneer environmental pattern could directly affect the colonization of the gut microbiome during early life. We focused on the different predominant microbiomes based on the mode of delivery and explored the relationships among the microbes using a network analysis. Figure 3B,C depict a complex network-based analysis of the meconium microbiome obtained using the Cytoscape program. This analysis revealed that the microbial communities in the C-section group were dominated by inhibitory relationships. Among them, B. licheniformis displayed the strongest inhibitory relationship to the surrounding microbes in the cluster. In the vaginally delivered group, the interactions between microbial communities were higher, with larger correlations that were both positive and negative. Bacillus was associated with Pseudomonas and P. acnes in terms of connections with other microbe clusters and was not a major factor in the relationships between the entire microbiota of the C-section group and that of the vaginally delivered group.
Functions of metabolism in the meconium microbiome in vaginally and C-section delivered neonates
To detect changes in microbiome metabolites induced by changes in the microbiota abundance, the metagenomics data were annotated with metabolic pathways from the MetaCyc database using the HumanN2 program. According to the LEfSe analysis heatmap of the metabolic function pathways, most differences occurred in amino acid metabolism pathways. The meconium microbiome of the vaginally delivered infants included more relative amino acid biosynthesis and amino acid degradation than did the C-section microbiome. Nucleoside and nucleotide biosynthesis was more abundant in the vaginally delivered infants, whereas their degradation was elevated in the C-section infants. Moreover, carbohydrate biosynthesis, inorganic nutrient metabolism, acetyl-CoA biosynthesis, and amine and polyamine metabolism were more active in the C-section delivered newborns. Consistent with the taxonomic composition according to the mode of delivery, PCA also showed a significant difference in pathways between the C-section and vaginal groups (Fig. 4).
Extended error bar plots showing functional properties that differ between C-section and vaginally delivered neonate gut microbiomes. Species identified by LEfSe as different between the two groups are depicted. The proportion (left side) indicates the possible abundance of microbes possessing each functional feature and the difference between proportions for each feature. For this analysis, features were filtered by the q value (0.05) and effect size (0.01). Blue indicates the C-section group; orange indicates the vaginal group.
Influence of different modes of delivery on the ARG prevalence
We quantified the relative abundance of ARGs in the meconium DNA to characterize the pioneer gut microbiome environment. The top 15 main ARG classes in each sample are shown in Fig. 5A. The BLA_A showed significant abundance in the meconium microbiome and enrichment in the vaginally delivered neonates. MexHI, tet_RPP and MexEF were more represented in the vaginally delivered neonates than in the C-section-delivered counterparts. Neonates delivered by C-section showed a significantly higher prevalence of MLS_ABC, MacAB, VanA, TET_EFFLUX, Ykk, mdr, bcr, Blt and MLS_hdr than the vaginally born neonates (Fig. 5B).
Correlation between metabolism and microbiome in different modes of delivery
To understand the correlation between microbiota community structure differences and metabolic differences between the vaginally delivered group and the C-section delivered group, we conducted a correlation analysis between the two groups. The results revealed a closer relationship between microbes in the vaginal group.
As shown in Fig. 6, the abundance of B. licheniformis and B. amyloliquefaciens in the C-section group had a significant positive effect on the metabolic pathway abundance; almost all other species showed a negative correlation. The abundance of these metabolic pathways was affected by multiple microbiota in addition to B. licheniformis in the vaginal group. The diversity of the microbial composition in vaginal delivery involved in the newborn intestinal microbial metabolism was also higher, which helped avoid the emergence of certain types of metabolic deficiencies. Because a greater variety of microorganisms were present in the vaginal group, the metabolic pathways were more abundant and diversified (Fig. S7A). The diversity index is a quantitative measure that reflects the number of different types in a dataset. The Shannon index refers to the amount of the difference, whereas the Pielou index refers to the degree of closeness. Thus, the Shannon and Pielou indices of the metabolic pathways were also calculated. Although there was no difference in metabolic pathway diversity between the two groups in the Shannon index, the vaginal group exhibited more homogeneity in metabolic pathways in the Pielou index analysis (Fig. S7B). Together, these results indicate that the C-section group suffers more from the influence of the dominant group (B. licheniformis), whereas the vaginal group is more homogeneous in its metabolism, which is dominated by multi-microbes.
Microbial metabolic differences in the meconium microbiome between vaginally delivered and C-section-delivered neonate samples. Genera identified by LEfSe as different between the two biological samples are depicted. Metabolic pathway designations are delineated at the bottom of the figure. The shading intensity and size of the circles indicate the Kendall rank correlation coefficient between matrices. Negative correlations are shown in violet, and positive correlations are shown in blue. Red squares surrounding the circles are indicative of a P value ≤ 0.05.
Discussion
Colonization by pioneer microbes and gradual diversification in the infant intestinal tract play important roles in the establishment of a symbiotic system of interactions between the host and microbes22. Different modes of delivery may affect the colonization of pioneer microbes, which may be the first group of microbes that infants encounter as they pass through the birth canal or are delivered by C-section. We used metagenomic sequencing to detect differences in the neonatal meconium between vaginally and C-section delivered neonates. The results demonstrated that the composition of the meconium microbiome was significantly affected by the mode of delivery. The predominant microbes were Firmicutes, Actinobacteria and Proteobacteria at the phylum level in our study, which was consistent with previous studies23,24. We also found that the predominant species in the vaginal neonatal meconium was Propionibacterium, whereas the dominant species was B. licheniformis in the C-section group. Moreover, we report for the first time that the characteristic gut microbiota in two IVF neonates differed from that of other C-section neonates and more closely resembled that of the vaginally delivered infants.
The pioneer bacteria that colonize the postnatal intestinal tract contribute to the establishment of host-microbe interactions essential for optimal symbiosis, which is essential for health throughout life10,25. It has been reported that the infant microbiota undergoes substantial reorganization within the first 6 weeks of life in a manner that is driven not by mode of delivery but by body site26. That investigation found that there was no significant clustering in the neonatal meconium microbiota by mode of delivery26. Nevertheless, several studies have demonstrated that the neonatal intestine is rapidly colonized by maternal and environmental bacteria at birth and is strongly affected by the delivery mode8,21,24,27,28,29. It was recently reported finding that the fecal microbiota of vaginally delivered infants had significant enrichment of Bacteroides, Parabacteroides and Megamonas, whereas that of cesarean delivered infants had enrichment of Prevotella, Streptococcus and Trabulsiella in both neonates and 2-month-old Chinese infants30. Moreover, alterations in the microbiota composition and the effect of the delivery mode may be associated with many diseases, including metabolic diseases (i.e., obesity) and immune-related diseases (i.e., inflammatory bowel disease, asthma, and allergies)10. Therefore, clarifying the intestinal microbiome habitat of the pioneer environmental pattern during the early life of healthy neonates is very important for the prevention and treatment of related diseases.
Our metagenomic analyses showed that Propionibacterium was the predominant bacterium in vaginally delivered infants. Additionally, Propionibacterium was widely and closely cross-linked with other bacteria. A review by Sabrina Tamburini et al.31 suggested that Propionibacterium was the main group observed in C-sections; the authors reported that these bacteria were sourced from the common skin and environment microbiota and could even be detected at early stages in fetal umbilical cord blood. However, in a recent study by Fredrik Backhed et al.24, Propionibacterium was enriched in the meconium from the vaginal group based on a metagenomic analysis. Propionibacterium are dominant in skin flora and have also been reported to be present in the adult gut microbiota. Therefore, Propionibacterium in the meconium may be derived from skin or fecal microbes through contact during vaginal delivery.
We also observed that B. licheniformis was significantly abundant in the C-section infants. Because there is little contact with the mother during childbirth by C-section, the neonatal gut microbiota cluster is relatively isolated. These relatively isolated microbial clusters may have the same origin; for instance, P. acnes originates from the skin32, Enhydrobacter aerosaccus originates from the air, and Prevotella copri33 and Escherichia originate from the intestinal microbiota. However, the Lactobacillus cluster representing vaginal microbes is lacking, and the B. licheniformis-based cluster is not a common microbial component from the mother; instead, this cluster most likely originates from the environment. B. licheniformis is an aerobic, spore-forming, Gram-positive rod that can be isolated from a number of hospital and community environments, including water, air, soil, dust and wet surfaces34. Survival in these adverse conditions is characteristic of this bacillus species and is partially due to the formation of resistant endospores. These bacteria are usually regarded as nonpathogenic organisms or as simple contaminants. Although B. licheniformis was previously found in vaginal and adult feces, it has not been reported in the meconium. Because each area has its own unique environmental factors, the usefulness of the metagenome in in-depth studies of the meconium is small, especially for first day samples. The intestinal flora may also show significant changes with an increase in dietary factors. Therefore, our study found that environmental factors had a significant effect on the early microbiota in newborns delivered by C-section, which helped us better understand the correlational relationship with potential diseases.
Recently, viral microbiomes were also shown to play important roles in human health. Lim ES et al.35 demonstrated that the eukaryotic virome and the bacterial microbiome expanded in concert with a contraction of and shift in the bacteriophage virome composition from birth to 2 years of age. Our results also showed that bacteriophages were more abundant in early life. Additionally, there were significant differences in the pathogenic virus distribution, which was consistent with the process of delivery. The mode of delivery, which led to different bacteriophage compositions, might control the number of microbes operating under a “predator-prey” model, as described by Lotka-Volterra. Therefore, further research on the pioneer gut microbiological environment should also consider viral impacts, especially from bacteriophages.
The presence of microbes in the genitourinary system has an important effect on reproduction36, especially in vitro fertilization2,3. Some studies have investigated the presence of Lactobacillus spp. in ovarian follicular fluids, which are associated with embryo maturation and transfer. Other studies found that the Lactobacillus-dominated endometrial microbiota was associated with positive reproductive outcomes in IVF patients37. However, the effect of IVF on the microbiota of C-section newborns is not clear. In our study, we compared the microbiome composition of the two C-section-delivered IVF patients to that of the vaginally delivered group and found similarities.
The intestinal microbiomes can be altered by many factors, such as the mode of delivery, diet and drugs; its metabolic capabilities are altered accordingly and affect host health. Thus, it is critical to not only identify the types of bacteria that compose the intestinal microbiota but also understand their ability to participate in metabolic functions. Many studies have shown that intestinal microbial genes that do not include carbohydrate metabolism enzymes, such as glycoside hydrolase and polysaccharide lyase, cannot contribute to the degradation or metabolism of these materials by the body. These enzymes are particularly important for neonatal intestinal development. C-section deliveries are influenced by the dominant group (B. licheniformis). The lack of metabolic variety and the basal metabolism of a single type of Bacillus would more easily lead to dysbiosis once disturbed. In the vaginal group, the metabolic pathways were more abundant and diversified, and a variety of microorganisms were involved in each pathway. More aerobic metabolic pathways were enriched in the vaginal group, suggesting that environmental factors had a large influence during the vaginal delivery process.
The gut microbiota are currently considered reservoirs of ARGs with the potential for horizontal transfer to pathogenic species38. Additionally, several resistance genes, such as the bla genes, originate from environmental bacteria and pose major problems in clinical treatment39. The human gut microbiome harbors numerous functional ARGs that are influenced by many factors, such as the mode of delivery, duration of breastfeeding and number of siblings of the infant40. However, few studies have investigated the presence of ARGs in the meconium. Alicea-Serrano et al. tested for the presence of four Tcr genes [tet(M), tet(O), tet(Q) and tet(W)] in rectal swab samples from 10 infants taken after meconium passage and only detected tet(W) in one sample41. M. J. Gosalbes et al.38 revealed a high prevalence of BLr and Tcr in both meconium and early fecal samples. Here, we compared our sequencing data with the Antibiotic Resistance Genes Database (http://ardb.cbcb.umd.edu/) and found that the BLA_A genes were the most abundant ARGs in the meconium. Furthermore, we found that the C-section delivered meconium microbiome contained more ARGs than the vaginally delivered group in terms of both diversity and abundance, which might influence the infant’s health in later life through the dissemination of other potentially harmful bacteria in the gut.
Our results demonstrated that the microbiome and metabolic diversity of vaginally delivered infants were significantly higher than the corresponding factors in the C-section group. The predominant bacteria were Propionibacterium in the vaginally delivered infants and B. licheniformis in the C-section group. This dominant distribution of B. licheniformis in the gut microbiome of the C-section delivered newborns may inhibit the diversity of the overall microbiome. In contrast, long-term contact with the birth canal and access to the distribution of maternal microbes in the gut microbiomes of vaginally delivered neonates may inhibit the excessive growth of Bacillus bacteria. Therefore, vaginally delivered newborns will have more diversified species, resulting in more homogeneous metabolism.
It has been reported that significant differences in gestational age, maternal pregnancy weight gain and gender between groups can influence the microbiome21,29,42. In this study, with a relative limited sample size, the impact of these factors was not concerned. Despite this, we were able to unambiguously identify the delivery mode as the most significant factor contributing to the difference between the meconium samples. Of course, larger studies are warranted to validate our findings and further studies are needed to investigate the long-term influence of delivery mode on microbiome and infant health.
Methods
Sample collection and informed consent
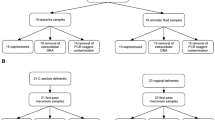
The initial meconium samples were collected by the Department of Gynecology and Obstetrics of the Chinese PLA General Hospital from April 2012 to November 2012. All mothers enrolled in the study gave their informed consent. The collected clinical data included the gestational age, gestational weight and weight gain of the mother, mode of delivery, birth weight, and body length of the infant. None of the mothers who delivered vaginally received antibiotics during delivery. Each mother who delivered by C-sectionreceived prophylactic antibiotics (metronidazole 0.5 g iv. bid, and cefmetazole 1 g iv. bid, or amoxicillin clavulanate potassium 3 g iv. bid) after the child was delivered. None of infants received antibiotic treatment. Ten infants in this study were born via C-section delivery, of which 8 infants were from healthy term pregnancies (gestational age 267 to 288 days) and 2 infants were fertilized using IVF (gestational age 254 days). Among the C-section delivered neonates, that mode of delivery was chosen because of breech/abnormal fetal position (n = 5), uterine scarring (n = 2), or the pregnant women’s request (n = 3). Eight infants were born with healthy term pregnancies via vaginal delivery (gestational age 273 to 287 days). The exclusion criteria were infants with a low birth weight (<2500 g) and infants who received postnatal antibiotics or whose mothers used antibiotics before C-section. All neonates were transferred to the postpartum room within 30 min after delivery and were breastfed with some colostrum after delivery. Internal portions of the first-pass meconium stools from within 24 h of delivery were collected from sterile single-use diapers into sterile single-use collection tubes, and then frozen at −80 °C within 30 min of collection.
Ethics statement
This study was undertaken with the approval of the Chinese PLA General Hospital Ethics Service Committee (2012–027). All experiments were carried out in accordance with the approved guidelines. Informed consent was obtained from all the parents of each child prior to sampling.
DNA extraction and metagenomic sequencing
In the laboratory, the meconium was divided into 5 aliquots (each 200-mg) and immediately stored at −80 °C. A frozen aliquot (200 mg) of each fecal sample was processed using the QIAamp DNA Stool Mini Kit (QIAGEN, Hilden, Germany) for DNA extraction as previously described43. The DNA concentration was measured with a NanoDrop (Thermo Scientific), and the molecular size was estimated by agarose gel electrophoresis.
Metagenomic DNA libraries were constructed with 0.2 μg of genomic DNA according to the Illumina TruSeq DNA Sample Prep v2 Guide, with an average insert size of 500 bp. The quality of all libraries was evaluated using an Agilent Bioanalyzer with a DNA LabChip 1000 kit. Negative controls (sterile water) were included for all the experimental process and showed no amplification in the final DNA library. Sequencing was performed using an Illumina Hiseq2500.
Quality control of Illumina HiSeq. 2500 reads
Illumina raw reads were subject to the following treatments: (1) reads with more than 3 ambiguous N bases were removed; (2) reads with less than 60% high-quality bases (Phred score ≥20) were deleted; and (3) the 3′ ends of the reads were trimmed to the first high-quality base. The subsequent high-quality reads were further mapped to some cereal genomes (Oryza sativa, Beijing Rice indica and Triticum aestivum) using SOAPaligner (version 2.21); any hit associated with the reads and their mated reads was removed. These filtered reads were admitted to the next steps of the analysis.
De novo assembly of the Illumina short reads
SOAPdenovo (version 2.04), which is based on DeBrujin graph construction, was employed to assemble short reads with the parameters ‘-M 3 -u -L 100 -d 1 -F’. The k-mers (varying from 39 to 59 by 4) from each sample were tested. The resulting scaffolds were cut into contigs at ambiguous Ns, and only contigs longer than 500 bp were retained. The N50 was calculated for contigs of different k-mers, and only contigs with the largest N50 assemblies were attributed to the sample. All contigs were applied for gene prediction.
Taxonomic and gene profiling
The microbial composition at each taxonomic level was defined using the MetaPhlAn2 program with default parameters. The program is available at http://huttenhower.sph.harvard.edu/metaphlan.
The relative gene abundance was determined with the procedure introduced by Qin N et al.44. When calculating the abundance of genes, the high-quality reads from each sample were aligned against the gene catalog using SOAPalign2.21 with the parameters ‘-r 2 -m 100 -x 1000′; only paired-end reads that could be mapped to the same gene were accepted. The antibiotic resistance genes (ARGs) were annotated against the ARG database using the same method.
Pathway analysis
MetaCyc is a curated database of experimentally elucidated metabolic pathways from all domains of life. The database contains 2453 pathways from 2788 different organisms and is currently the largest available curated collection of metabolic pathways. The HUMAnN2 program can efficiently and accurately determine the presence, absence, and abundance of metabolic pathways in a microbial community based on metagenomic sequencing data. The MetaCyc metabolic pathway modules for imputed genes were assigned using the HUMAnN2 program with default parameters (prescreen-threshold = 0.01, identity-threshold = 0.50). The methodology is available at http://huttenhower.sph.harvard.edu/humann, and the program can be installed from PyPI (https://pypi.python.org/pypi/humann2) as a source code from the HUMAnN2 Bitbucket repository (https://bitbucket.org/biobakery/humann2/wiki/Home).
Statistical analysis
The STAMP program is a statistical/econometric software system for time-series models with unobserved components, such as trends (available at http://stamp-software.com/). Using the program, we generated extended error bar (EEB) plots to show that some properties differed significantly between C-section and natural vaginal births with some filter parameters (ratio of effect proportion ≥2, difference between proportion ≥1, and P value = 0.05).
The LEfSe (linear discriminant analysis effect size) algorithm determines the features most likely to explain differences between classes by coupling standard tests for statistical significance with additional tests encoding biological consistency and effect relevance (available at http://huttenhower.sph.harvard.edu/galaxy)45.
R is a software environment for statistical computing and graphics, and Cytoscape is a software platform for visualizing complex networks and integrating them with any type of attribute data. Spearman’s correlation coefficient was calculated using R and visualized as a network using Cytoscape. These programs are available at https://www.r-project.org/ and http://www.cytoscape.org/, respectively. Redundancy analysis (RDA) and Adonis was performed to identify the relationship between the microbial composition and metadata with R program.
References
Cong, X. et al. Gut Microbiome Developmental Patterns in Early Life of Preterm Infants: Impacts of Feeding and Gender. PloS one 11, e0152751, https://doi.org/10.1371/journal.pone.0152751 (2016).
Hyman, R. W. et al. The dynamics of the vaginal microbiome during infertility therapy with in vitro fertilization-embryo transfer. Journal of assisted reproduction and genetics 29, 105–115, https://doi.org/10.1007/s10815-011-9694-6 (2012).
Pelzer, E. S. et al. Microorganisms within human follicular fluid: effects on IVF. PloS one 8, e59062, https://doi.org/10.1371/journal.pone.0059062 (2013).
Aagaard, K. et al. The placenta harbors a unique microbiome. Science translational medicine 6, 237ra265, https://doi.org/10.1126/scitranslmed.3008599 (2014).
Oh, K. J. et al. Detection of ureaplasmas by the polymerase chain reaction in the amniotic fluid of patients with cervical insufficiency. Journal of perinatal medicine 38, 261–268, https://doi.org/10.1515/jpm.2010.040 (2010).
Jimenez, E. et al. Isolation of commensal bacteria from umbilical cord blood of healthy neonates born by cesarean section. Current microbiology 51, 270–274, https://doi.org/10.1007/s00284-005-0020-3 (2005).
Steel, J. H. et al. Bacteria and inflammatory cells in fetal membranes do not always cause preterm labor. Pediatric research 57, 404–411, https://doi.org/10.1203/01.pdr.0000153869.96337.90 (2005).
Dominguez-Bello, M. G. et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proceedings of the National Academy of Sciences of the United States of America 107, 11971–11975, https://doi.org/10.1073/pnas.1002601107 (2010).
Palmer, C., Bik, E. M., DiGiulio, D. B., Relman, D. A. & Brown, P. O. Development of the human infant intestinal microbiota. PLoS biology 5, e177, https://doi.org/10.1371/journal.pbio.0050177 (2007).
Wopereis, H., Oozeer, R., Knipping, K., Belzer, C. & Knol, J. The first thousand days - intestinal microbiology of early life: establishing a symbiosis. Pediatric allergy and immunology: official publication of the European Society of Pediatric Allergy and Immunology 25, 428–438, https://doi.org/10.1111/pai.12232 (2014).
Rutayisire, E., Huang, K., Liu, Y. & Tao, F. The mode of delivery affects the diversity and colonization pattern of the gut microbiota during the first year of infants’ life: a systematic review. BMC gastroenterology 16, 86, https://doi.org/10.1186/s12876-016-0498-0 (2016).
Black, M., Bhattacharya, S., Philip, S., Norman, J. E. & McLernon, D. J. Planned Cesarean Delivery at Term and Adverse Outcomes in Childhood Health. Jama 314, 2271–2279, https://doi.org/10.1001/jama.2015.16176 (2015).
Kuhle, S., Tong, O. S. & Woolcott, C. G. Association between caesarean section and childhood obesity: a systematic review and meta-analysis. Obesity reviews: an official journal of the International Association for the Study of Obesity 16, 295–303, https://doi.org/10.1111/obr.12267 (2015).
Adlercreutz, E. H., Wingren, C. J., Vincente, R. P., Merlo, J. & Agardh, D. Perinatal risk factors increase the risk of being affected by both type 1 diabetes and coeliac disease. Acta paediatrica (Oslo, Norway: 1992) 104, 178–184, https://doi.org/10.1111/apa.12836 (2015).
Decker, E., Hornef, M. & Stockinger, S. Cesarean delivery is associated with celiac disease but not inflammatory bowel disease in children. Gut microbes 2, 91–98 (2011).
Algert, C. S., McElduff, A., Morris, J. M. & Roberts, C. L. Perinatal risk factors for early onset of Type 1 diabetes in a 2000-2005 birth cohort. Diabetic medicine: a journal of the British Diabetic Association 26, 1193–1197, https://doi.org/10.1111/j.1464-5491.2009.02878.x (2009).
Dominguez-Bello, M. G. et al. Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. 22, 250–253, https://doi.org/10.1038/nm.4039 (2016).
Gosalbes, M. J. et al. Meconium microbiota types dominated by lactic acid or enteric bacteria are differentially associated with maternal eczema and respiratory problems in infants. Clinical and experimental allergy: journal of the British Society for Allergy and Clinical Immunology 43, 198–211, https://doi.org/10.1111/cea.12063 (2013).
Liu, D. et al. Bacterial community structure associated with elective cesarean section versus vaginal delivery in Chinese newborns. Journal of pediatric gastroenterology and nutrition 60, 240–246, https://doi.org/10.1097/mpg.0000000000000606 (2015).
Fallani, M. et al. Intestinal microbiota of 6-week-old infants across Europe: geographic influence beyond delivery mode, breast-feeding, and antibiotics. Journal of pediatric gastroenterology and nutrition 51, 77–84, https://doi.org/10.1097/MPG.0b013e3181d1b11e (2010).
Dogra, S. et al. Dynamics of infant gut microbiota are influenced by delivery mode and gestational duration and are associated with subsequent adiposity. mBio 6, https://doi.org/10.1128/mBio.02419-14 (2015).
Neu, J. Developmental aspects of maternal-fetal, and infant gut microbiota and implications for long-term health. Maternal health, neonatology and perinatology 1, 6, https://doi.org/10.1186/s40748-015-0007-4 (2015).
Moles, L. et al. Bacterial diversity in meconium of preterm neonates and evolution of their fecal microbiota during the first month of life. PloS one 8, e66986, https://doi.org/10.1371/journal.pone.0066986 (2013).
Backhed, F. et al. Dynamics and Stabilization of the Human Gut Microbiome during the First Year of Life. Cell host & microbe 17, 690–703, https://doi.org/10.1016/j.chom.2015.04.004 (2015).
Secher, T., Brehin, C. & Oswald, E. Early settlers: which E. coli strains do you not want at birth? American journal of physiology. Gastrointestinal and liver physiology 311, G123–129, https://doi.org/10.1152/ajpgi.00091.2016 (2016).
Chu, D. M. et al. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nature medicine 23, 314–326, https://doi.org/10.1038/nm.4272 (2017).
Nagpal, R. et al. Sensitive Quantitative Analysis of the Meconium Bacterial Microbiota in Healthy Term Infants Born Vaginally or by Cesarean Section. Frontiers in microbiology 7, 1997, https://doi.org/10.3389/fmicb.2016.01997 (2016).
Bokulich, N. A. et al. Antibiotics, birth mode, and diet shape microbiome maturation during early life. Science translational medicine 8, 343ra382, https://doi.org/10.1126/scitranslmed.aad7121 (2016).
Martin, R. et al. Early-Life Events, Including Mode of Delivery and Type of Feeding, Siblings and Gender, Shape the Developing Gut Microbiota. PloS one 11, e0158498, https://doi.org/10.1371/journal.pone.0158498 (2016).
Kuang, Y. S. et al. Composition of gut microbiota in infants in China and global comparison. Scientific reports 6, 36666, https://doi.org/10.1038/srep36666 (2016).
Tamburini, S., Shen, N., Wu, H. C. & Clemente, J. C. The microbiome in early life: implications for health outcomes. Nature medicine 22, 713–722, https://doi.org/10.1038/nm.4142 (2016).
Bruggemann, H. et al. The complete genome sequence of Propionibacterium acnes, a commensal of human skin. Science (New York, N.Y.) 305, 671–673, https://doi.org/10.1126/science.1100330 (2004).
Hayashi, H., Shibata, K., Sakamoto, M., Tomita, S. & Benno, Y. Prevotella copri sp. nov. and Prevotella stercorea sp. nov., isolated from human faeces. International journal of systematic and evolutionary microbiology 57, 941–946, https://doi.org/10.1099/ijs.0.64778-0 (2007).
Blue, S. R., Singh, V. R. & Saubolle, M. A. Bacillus licheniformis bacteremia: five cases associated with indwelling central venous catheters. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America 20, 629–633 (1995).
Lim, E. S., Zhou, Y. & Zhao, G. Early life dynamics of the human gut virome and bacterial microbiome in infants. 21, 1228–1234, https://doi.org/10.1038/nm.3950 (2015).
Reid, G. et al. Microbes central to human reproduction. American journal of reproductive immunology (New York, N.Y.: 1989) 73, 1–11, https://doi.org/10.1111/aji.12319 (2015).
Moreno, I. et al. Evidence that the endometrial microbiota has an effect on implantation success or failure. American journal of obstetrics and gynecology 215, 684–703, https://doi.org/10.1016/j.ajog.2016.09.075 (2016).
Gosalbes, M. J. et al. High frequencies of antibiotic resistance genes in infants’ meconium and early fecal samples. Journal of developmental origins of health and disease 7, 35–44, https://doi.org/10.1017/s2040174415001506 (2016).
Potron, A., Poirel, L. & Nordmann, P. Origin of OXA-181, an emerging carbapenem-hydrolyzing oxacillinase, as a chromosomal gene in Shewanella xiamenensis. Antimicrobial agents and chemotherapy 55, 4405–4407, https://doi.org/10.1128/aac.00681-11 (2011).
von Wintersdorff, C. J. et al. The gut resistome is highly dynamic during the first months of life. Future microbiology 11, 501–510, https://doi.org/10.2217/fmb.15.154 (2016).
Alicea-Serrano, A. M., Contreras, M., Magris, M., Hidalgo, G. & Dominguez-Bello, M. G. Tetracycline resistance genes acquired at birth. Archives of microbiology 195, 447–451, https://doi.org/10.1007/s00203-012-0864-4 (2013).
Robinson, A. et al. Association of Maternal Gestational Weight Gain With the Infant Fecal Microbiota. Journal of pediatric gastroenterology and nutrition. https://doi.org/10.1097/mpg.0000000000001566 (2017).
Zhang, J. et al. Mongolians core gut microbiota and its correlation with seasonal dietary changes. Scientific reports 4, 5001, https://doi.org/10.1038/srep05001 (2014).
Qin, N. et al. Alterations of the human gut microbiome in liver cirrhosis. Nature 513, 59–64, https://doi.org/10.1038/nature13568 (2014).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome biology 12, R60, https://doi.org/10.1186/gb-2011-12-6-r60 (2011).
Acknowledgements
We are very grateful to all the volunteers for their participation. Helpful advice for Shu Cheng at BGI. Statistical analysis was supported by Yun Huang and Fang-Fang Fan in the Realbio Genomics Institute. This study was supported by the Beijing Natural Science Foundation (S150010) and the National High Technology Research and Development Program of China (863 Program) (2015AA020701).
Author information
Authors and Affiliations
Contributions
Yi-Chao Shi analyzed the data and drafted the manuscript; He Guo collected the samples; Jing Chen assisted analysis and statistics; Gang Sun revised the manuscript for important intellectual content; Rong-Rong Ren and Ming-Zhou Guo assisted analysis; Li-Hua Peng assisted with design and provided funding; and Yun-Sheng Yang designed, funded and supervised the study. All authors have read and approved the final version to be published.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shi, YC., Guo, H., Chen, J. et al. Initial meconium microbiome in Chinese neonates delivered naturally or by cesarean section. Sci Rep 8, 3255 (2018). https://doi.org/10.1038/s41598-018-21657-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21657-7
This article is cited by
-
Effect of different delivery modes on intestinal microbiota and immune function of neonates
Scientific Reports (2024)
-
Compositional and functional variability of the gut microbiome in children with infantile colic
Scientific Reports (2023)
-
Neonatal intensive care unit (NICU) exposures exert a sustained influence on the progression of gut microbiota and metabolome in the first year of life
Scientific Reports (2021)
-
Maternal diet during pregnancy and intestinal markers are associated with early gut microbiota
European Journal of Nutrition (2021)
-
The effects of perineal disinfection on infant’s oral microflora after transvaginal examination during delivery
BMC Pregnancy and Childbirth (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.