Abstract
Dysregulated autophagy is central to the pathogenesis and therapeutic development of cancer. However, how autophagy is regulated in cancer is not well understood and genes that modulate cancer autophagy are not fully defined. To gain more insights into autophagy regulation in cancer, we performed a large-scale RNA interference screen in K562 human chronic myeloid leukemia cells using monodansylcadaverine staining, an autophagy-detecting approach equivalent to immunoblotting of the autophagy marker LC3B or fluorescence microscopy of GFP-LC3B. By coupling monodansylcadaverine staining with fluorescence-activated cell sorting, we successfully isolated autophagic K562 cells where we identified 336 short hairpin RNAs. After candidate validation using Cyto-ID fluorescence spectrophotometry, LC3B immunoblotting, and quantitative RT-PCR, 82 genes were identified as autophagy-regulating genes. 20 genes have been reported previously and the remaining 62 candidates are novel autophagy mediators. Bioinformatic analyses revealed that most candidate genes were involved in molecular pathways regulating autophagy, rather than directly participating in the autophagy process. Further autophagy flux assays revealed that 57 autophagy-regulating genes suppressed autophagy initiation, whereas 21 candidates promoted autophagy maturation. Our RNA interference screen identified genes that regulate autophagy at different stages, which helps decode autophagy regulation in cancer and offers novel avenues to develop autophagy-related therapies for cancer.
Similar content being viewed by others
Introduction
Autophagy (also known as macroautophagy) is a subcellular catabolic process where cytoplasm and organelles are digested, and metabolic products are recycled to prolong cell survival under conditions of stress. Autophagy is characterized by the formation of autophagosomes (autophagy initiation) followed by fusion with lysosomes. The resulting autolysosomes degrade and recycle engulfed materials (autophagy maturation). This entire dynamic process is termed as autophagy flux. While autophagy often protects cells, excessive digestion triggers autophagic cell death. Hence, autophagy is critical for maintaining homeostasis and is central to the pathogenesis of many human diseases including cancer1,2,3,4.
Mounting evidence has established a fundamental role of autophagy in tumor initiation and progression5. As such, targeting autophagy as a means for cancer therapeutic development has been widely explored6,7,8,9. However, the relationship between autophagy and cancer therapeutic responses is complicated. On one hand, autophagy activators are appealing cancer drugs because excessive autophagy induces autophagic cell death. Many cancer signaling pathways such as PI3K/MTOR (phosphatidylinositol-4,5-bisphosphate 3-kinase/mechanistic target of rapamycin)10,11,12,13 or RAS (rat sarcoma viral oncogene homolog)14,15 suppress autophagy. However, targeting these kinases or small GTPase to specifically activate autophagic cell death has been proven difficult because of their important roles in other cell death pathways such as apoptosis6,16,17. On the other hand, established cancer often maintains a proficient level of autophagy to sustain cancer cell survival18, making autophagy blockade an attractive strategy to break this protection. Inhibition of autophagy can be achieved either by blocking autophagy initiation characterized by formation of autophagosomes or by impairing autophagy maturation to halt autophagy flux. Targeting autophagy maturation to block autophagy flux has been used in treating cancer. For example, chloroquine blocks the activity of autolysosomes to impede autophagy flux19. In combination with conventional therapies, this drug has prolonged the survival of glioblastoma patients; however, the statistical significance was low20. In addition, chloroquine exhibits severe side effects in the clinic limiting its application as an effective cancer treatment21,22. Many efforts have been invested to find new strategies to tackle autophagy with limited success, largely because autophagy regulation in cancer is not well understood and genes that regulate this dynamic process are not fully defined.
Chronic myeloid leukemia (CML) is characterized by the production of a single oncogenic protein BCR-ABL, which results from the translocation between chromosome 9 and 2223. Imatinib and its derivatives, the specific inhibitors of BCR-ABL, have been used in the clinic to treat CML24; however, resistance that is either BCR-ABL-dependent (i.e. BCR-ABL mutations) or -independent still develops in patients25. One possible reason for BCR-ABL-independent imatinib resistance is that CML cells activate autophagy to sustain their survival during imatinib treatment26,27,28,29. In our previous study, we show that BCR-ABL suppresses autophagy through activating a transcription factor called activating transcription factor 5 (ATF5), which in turn transcriptionally activates the expression of MTOR. Blocking ATF5 sensitizes CML cells to imatinib26,28. Based upon these results, it is imperative to identify novel autophagy regulators in CML so as to not only gain deeper insights into autophagy regulation but also provide novel therapeutic targets to develop effective autophagy-related therapies for cancer.
In this report, we isolated autophagic K562 cells using the fluorescent dye, monodansylcadaverine (MDC), coupled with fluorescence-activated cell sorting (FACS). The sensitivity of this approach in detecting autophagy was similar to other autophagy-detecting approaches such as LC3B immunoblotting and Cyto-ID fluorescence spectrophotometry. We carried out a large-scale RNA interference (RNAi) screen followed by rigorous multistep validations. We identified 82 genes that regulate autophagy in K562 cells. Among these genes, 62 candidates are novel autophagy regulators. Furthermore, we revealed that these genes either suppress autophagy initiation or promote autophagy maturation. Our results have important implications in understanding fundamental mechanisms of autophagy regulation in cancer and in the development of effective cancer therapies.
Results
MDC stains autophagic K562 cells
Labeling and isolating live autophagic cells is difficult30. To resolve this issue, we employed a fluorescent dye MDC that labels acidic compartments including autolysosomes31. To stain live autophagic cells with MDC, we treated K562 cells with 1 μM imatinib (IM) to induce autophagy in K562 cells as described in our previous report26. We then stained K562 cells with MDC and monitored MDC fluorescence in live cells using an inverted fluorescence microscope. MDC fluorescence was in low levels and barely detected in live K562 cells treated with DMSO (top panel), whereas strong MDC fluorescence was detected in IM-treated cells (Fig. 1A, bottom panel). To corroborate these results and quantitatively measure MDC fluorescence, we carried out a spectrophotometric assay using a microplate reader. IM significantly enhanced MDC fluorescence intensities by approximately 2-fold (Fig. 1B). LC3B (microtubule associated protein 1 light chain 3 β, MAP1LC3B) is an autophagy marker. Immuno-detection of the lipidated form of LC3B (LC3B-II) located on the membrane of autophagosomes and GFP-LC3B fluorescence microscopy that monitors the formation of autophagosomes (GFP-LC3B puncta) are well-established autophagy assays32,33. We determined levels of LC3B-II in K562 cells treated with DMSO or IM. IM induced a 1.4-fold increase of LC3B-II (Fig. 1C), similar to the fold changes of MDC fluorescence. Hence, the sensitivity of MDC in labeling autophagic cells is equivalent to that of LC3B. To determine whether MDC staining can be used to measure autophagy flux, we treated K562 cells with IM and bafilomycin A1 (BFA1), a selective inhibitor of vacuolar H+ ATPase that prevents the fusion of autophagosomes and lysosomes. Cells treated with both IM and BFA1 showed much stronger MDC fluorescence (Fig. 1D). Quantification revealed that the co-treatment of IM and BFA1 significantly enhanced MDC intensities compared to each treatment alone (Fig. 1E). To compare MDC staining with GFP-LC3B fluorescence microscopy, we transfected K562 cells with GFP-LC3B followed by the treatment of IM and/or BFA1. IM and BFA1 significantly increased percentages of cells with GFP-LC3B puncta (Fig. 1F,G), congruent with results of MDC staining (Fig. 1D,E). We did notice that the background levels of MDC staining were much higher than those of GFP-LC3B puncta. However, GFP-LC3B fluorescence microscopy is based on the morphological changes of GFP-LC3B rather than fluorescence intensity. As such, GFP-LC3B-based assays are not suitable for isolating autophagic cells using FACS. While MDC may stain cells with enhanced levels of both lysosomes and autolysosomes (see discussion for details), this approach is compatible with FACS and suitable for high-content studies. In addition, our results shown above demonstrate that MDC staining can be used to determine autophagy and autophagy flux at least in K562 cells upon IM treatment. We therefore used MDC staining coupled with FACS to isolate autophagic K562 cells in the following experiments.
Monodansylcadaverine (MDC) stains autophagic K562 cells with sensitivity equivalent to LC3B immunoblotting and GFP-LC3B fluorescence microscopy. (A) Fluorescence microscopy. K562 cells were treated with dimethyl sulfoxide (DMSO) or 1 μM imatinib (IM) overnight followed by MDC staining. Images of live cells were taken at different exposure times (milliseconds, ms) using an inverted fluorescence microscope. Scale bar: 25 μm. (B) MDC Fluorescence spectrophotometry. K562 cells treated with DMSO or IM were stained with MDC. MDC fluorescence of live K562 cells was quantified using a micro-plate reader at an excitation wavelength of 335 nm and an emission wavelength of 525 nm. The relative MDC levels were obtained by dividing MDC fluorescence of IM-treated cells to that of DMSO-treated cells. Error bars represent three independent experiments. (C) LC3B immunoblotting. Cropped images are shown and full images are included in supplemental materials. ACTB (β actin) is the loading control. The intensities of LC3B-II or ACTB were quantified using Image J. The fold changes of LC3B-II/ACTB were obtained by dividing the ratio of LC3B-II/ACTB in IM-treated cells with that of DMSO-treated cells. (D) MDC staining of K562 cells treated with IM and/or bafilomycin A1 (BFA1). K562 cells were treated with a combination of 1 μM IM (10 hours) and 5 nM of BFA1 (48 hours). Cells were imaged using a fluorescence confocal microscope. (E) Quantification of MDC intensities. Fluorescence intensities of MDC in 10 different cells from three images were quantified using Image J. (F) GFP-LC3B fluorescence microscopy. K562 cells were stably transfected with a construct encoding GFP-LC3B. Cells were treated with a combination of IM and BFA1 and then imaged. (G) Quantification of cells with GFP-LC3B puncta. Four to five images of more than thirty K562 cells from each treatment were randomly selected for the following quantification analyses. GFP-expressing K562 cells or K562 cells with GFP puncta (indicating autophagy) were counted by three persons. Percentages of K562 cells with puncta were obtained by dividing numbers of K562 cells with GFP puncta with those of GFP-expressing cells. Scale bar: 25 nm.
A genome-wide RNAi screen identifies novel autophagy-regulating genes
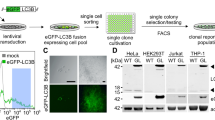
Next, we employed MDC staining followed by FACS in a genome-wide RNAi screen designed for identifying novel autophagy regulators. As illustrated in Fig. 2A, K562 cells were transduced with a library of 75,000 shRNAs that target nearly 15,000 human genes. After isolation of autophagic cells using MDC and FACS, genomic DNAs that harbor shRNAs were extracted. shRNAs were PCR-amplified using pre-designed primers, and subsequently identified using Sanger DNA sequencing. In more than 500 individual PCR products, we identified 336 shRNAs (Fig. 2A and Table S1) and more than 50 shRNAs were repeatedly sequenced.
A large-scale RNA interference screen identifies autophagy-regulating genes. (A) Schematic diagram illustrating a large-scale RNA interference screen. 336 shRNAs were identified in MDC positive cells. (B) Candidate validation using the Cyto-ID spectrophotometric assay. K562 cells were treated with non-silencing (NS) shRNA or individual 336 candidate shRNAs. The 1.4-fold increase of Cyto-ID fluorescence was empirically set as the cut-off line. (C) Candidate validation using the LC3B immunoblotting assay. K562 cells were treated with NS or 152 individual shRNAs of Cyto-ID positive candidates. Autophagy was monitored using LC3B immunoblotting. A cropped representative blot of LC3B immunoblotting is shown. Full images are included in supplemental materials. (D) Quantification of LC3B-II intensities. The intensities of protein bands were quantified using Image J. The ratios of the protein intensities of LC3B-II and ACTB are shown. The 1.4-fold increase of LC3B-II was empirically set as the cut-off line.
To further validate these candidates we employed another fluorescent dye, Cyto-ID, which specifically labels autophagosomes, amphisomes, and autolysosomes with minimal staining of lysosomes34,35. We transduced K562 cells with viruses containing individual shRNAs or a control non-silencing (NS) shRNA. Autophagy was measured using the Cyto-ID fluorescence spectrophotometric assay we have recently developed34,35. Among 336 candidates, 152 shRNAs induced a ≥ 1.4-fold increase of Cyto-ID fluorescence, compared to NS shRNA (Fig. 2B and Table S1). We next utilized LC3B immunoblotting to analyze 152 Cyto-ID positive candidates. A representative blot was shown in Fig. 2C. We found that 124 shRNAs increased LC3B-II intensities by ≥ 1.4-fold based on quantification of protein blots (Fig. 2D and Table S2). We empirically chose 1.4-fold increase of Cyto-ID fluorescence or LC3B-II as the cutoff lines because this approach allows us to include autophagy regulators with differential capabilities to activate autophagy (see below for details). The strength of autophagy responses often determines cell fate. For instance, autophagy often protects cells from cell death; however, a strong response often induces excessive autophagy leading to cell death.
To determine whether a specific shRNA is on or off target, we measured knockdown efficiency of 124 shRNAs on 122 target genes using quantitative RT-PCR. We found that 83 shRNAs induced a ≥2-fold reduction in mRNA levels of 82 genes (Fig. 3A and Table S3). We then selected four candidates and validated knockdown at protein levels using commercially available antibodies. shRNAs of ETS2, HCLS1, KRAS and LYN remarkably decreased protein expression at a factor of 2 to 5 folds (Fig. 3B–E), congruent with RT-PCR results.
Knockdown efficiency of candidate autophagy-regulating genes (ARGs). (A) Quantitative RT-PCR. K562 cells were treated NS or 124 individual shRNAs of Cyto-ID and LC3B-II positive candidates. mRNA levels of shRNA-targeting genes were measured using quantitative RT-PCR. The cut-off line was set as 0.5. Error bars represent standard deviations from three independent experiments. Protein levels of ETS2 (B), HCLS1 (G), KRAS (D), and LYN (E) in K562 cells treated with their shRNAs were determined using immunoblotting. Cropped images are shown and full images are included in supplemental materials. Protein levels were quantified using Image J. ACTB (β actin) is the loading control. Fold changes of ARG protein levels were obtained by dividing the ratios of ARG/ACTB in ARG shRNA-treated cells to those in NS shRNA-treated cells.
82 candidate genes were hereafter dubbed autophagy-regulating genes (ARGs, Table 1). As mentioned in our previous report, ATF5 regulates IM-induced autophagy in BCR-ABL positive CML cells26. The identification of ATF5 as an ARG in K562 CML cells indicates that the RNAi screen described above is unbiased. In addition to ATF5, 19 ARGs (ACAT1, CAMKV, CNOT2, ERGIC1, ETS2, HSF2, IGF2R, JAK1, KRAS, LPCAT2, LYN, PAH, PEX2, PLK1, PRKCD, S100A4, SCD, VAMP7, and YWHAZ) were also reported previously as autophagy modulators26,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54. Hence, the remaining 62 ARGs are novel autophagy mediators. The fact that approximately one-fourth of ARGs has been reported previously verifies the feasibility of MDC in isolating autophagic cells and the rigor of multistep validations in this large-scale RNAi screen. It is also notable that none of the 82 ARGs are autophagy-related genes, which are structural proteins or enzymes that directly participate in the formation of autophagic compartments such as autophagosomes and autolysosome55. This is perhaps because knockdown of autophagy-related genes impairs the formation of autophagic compartments and, therefore, is expected to decrease–rather than increase–the levels of MDC, Cyto-ID, or LC3B-II. Hence, a different approach needs to be taken to measure activities of autophagy-related genes during autophagy activation or inhibition in future RNA interference screens.
Cyto-ID labels most autophagic compartments such as autophagosomes, amphisomes, and autolysosomes34, whereas LC3B-II mostly highlights autophagosomes and autolysosomes32,33. Based on the changes in Cyto-ID fluorescence (Fig. 2B and Table S1) and LC3B-II intensities (Fig. 2D and Table S2), we defined the ability of ARG shRNAs to induce the formation of autophagic compartments using the following criteria: (1) High, if there was a ≥ 2-fold increase in both Cyto-ID and LC3B-II; (2) Medium, if there was a ≥ 2-fold increase of either Cyto-ID or LC3B-II; (3) Low, if there was a 1.4- to 1.9-fold increase of Cyto-ID and/or LC3B-II. More than half of ARG shRNAs was at medium levels, whereas 14 or 22 ARG shRNAs were at high or low levels, respectively (Table S4). Hence, ARGs have differential capabilities to modulate autophagy.
Taken together, our genome-wide RNAi screen followed by comprehensive and rigorous validations, identifies 82 ARGs with different capabilities in regulating autophagy in K562 cells. Of note, 62 candidate ARGs are novel autophagy mediators.
Bioinformatic analyses of ARGs
To further characterize ARGs, we performed several bioinformatic analyses. We queried the STRING v9.1 database56, which predicts protein-protein interactions. By using 16 core autophagy proteins (e.g. ATG3, ATG4, and ATG5, etc.) and LC3B (MAP1LC3B) as the bait (Fig. 4A, in red), we found that only two ARG proteins VAMP7 and AGPS (Fig. 4A, in black) showed potential interactions with the bait proteins. In line with the result that no autophagy-related genes were found in our RNAi screen, these results suggest that ARGs identified herein do not directly participate in the process of autophagy. We then investigated whether ARG proteins interact with autophagy signaling molecules. We set 22 known autophagy signaling molecules (Fig. 4B, in red) as the bait and detected potential protein-protein interactions in 13 candidate ARG proteins (Fig. 4B, in black). Because 62 ARGs are novel autophagy mediators, there is perhaps a lack of information of protein-protein interactions in the database. Hence, these results suggest that ARGs are upstream mediators of autophagy process.
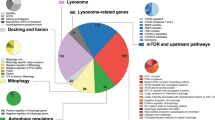
Bioinformatic analyses. (A) Predicted protein-protein interactions between candidate genes and core autophagy-related genes using the STRING program. 16 core autophagy-related genes (in red) were used as bait in the STRING database. ARGs (black) that show potential interactions with the bait are shown. Each line represents a potential interaction from one assay. The size of nodes represents the protein size. (B) Potential interactions between candidate genes and kinases in autophagy signaling. (C) Gene term enrichment analysis using the DAVID program. The biological processes that were significantly enriched were listed and color-coded. Each node represents genes involved in these biological processes. The larger the node, the more genes it represents. Two nodes are connected if they share genes or have possible interactions. (D) Biological processes in which candidate genes are involved. By using the data from the GeneCards database, candidate genes were categorized into different biological processes. The percentages of genes involved in these processes are shown.
By using the DAVID program57, we performed gene ontology analyses. Most ARGs were directly or indirectly involved in the signaling pathways emanating from interleukin-3, interleukin-5, or granulocyte-macrophage colony-stimulating factor (Fig. 4C). These hematopoietic cytokines play vital roles in leukemogenesis, and are important for autophagy regulation in normal and malignant blood cells58,59,60. Only a few ARGs are implicated in mitosis, vesicle biogenesis, diabetes, and lipid metabolism. By querying the GeneCards database, we grouped ARGs based on the molecular pathways to which they possibly target. Percentages of ARGs that exhibit potential activities in metabolism, transcription, signaling, and molecule transportation were 26%, 18%, 12%, and 10%, respectively (Fig. 4D). These molecular pathways have crucial assignments in autophagy regulation55,61,62. These results have implications in understanding the relationship between autophagy and ARGs-targeting biological processes.
ARGs target different stages of autophagy
Autophagy is a dynamic process (referred to as autophagy flux) including initiation (autophagosome formation) and maturation (autolysosome formation and cargo degradation). It is therefore crucial to distinguish whether the changes of autophagic compartments (autophagosomes, amphisomes, and autolysosomes) are due to autophagy activation or autophagy flux blockade because levels of autophagic compartments increase in both conditions32,33. In our RNAi screen and subsequent analyses, it is unclear whether the increase of MDC, Cyto-ID, and LC3B-II levels upon depletion of ARGs is due to the activation of autophagy initiation or the blockade of autophagy maturation.
To address this question, we employed the Cyto-ID fluorescence spectrophotometric assay. We have successfully monitored the changes of autophagic compartments in cells treated with autophagy activators and/or autophagy maturation blockers using this approach34. We treated ARG-deficient cells with chloroquine (an inhibitor of autophagy maturation)29,63,64. In principle, activation of autophagy in combination with chloroquine results in a significant increase of autophagy compartments compared to each treatment alone. For example, knockdown of IGSF1 significantly increased Cyto-ID levels together with chloroquine (P < 0.05; Fig. 5A). Consistently, LC3B-II levels also substantially increased in cells treated with IGSF1 shRNA and chloroquine (Fig. 5B). These results suggest that IGSF1 suppresses autophagy initiation and its depletion activates autophagy. In contrast, shRNAs of PTDSS1 failed to induce a significant increase of Cyto-ID (P = 0.08; Fig. 5C) or LC3B-II (Fig. 5D) in combination with chloroquine, compared to PTDSS1 shRNA alone. These results suggest that PTDSS1 targets the autophagy maturation stage. We did notice that the Cyto-ID assay detected similar changes in cells treated with either ARG shRNAs or chloroquine (Fig. 5A,C), whereas LC3B-II protein levels were much higher in chloroquine-treated cells than in ARG shRNAs-treated cells (Fig. 5B,D). This discrepancy may be due to the fact that Cyto-ID measures levels of most autophagic compartments and LC3B-II only labels autophagosomes. Based on P values that determine the statistical significance of difference between means of the combinational treatment (chloroquine and shRNA) and those of chloroquine or shRNA (Tables S5 and 2), we found that 57 ARGs significantly enhanced the levels of autophagic compartments together with chloroquine, indicating that these ARGs suppress autophagy initiation. The remaining 25 ARG shRNAs failed to do so (Tables S5 and 2, in bold).
Determination of autophagy stages to which ARGs target. (A,B) Combination of IGSF1 shRNA and chloroquine (CQ). K562 cells were treated with NS or IGSF1 shRNA followed by chloroquine treatment. Autophagy was assessed using the Cyto-ID spectrophotometric assay (A) or LC3B immunoblotting (B). (C,D) Combination of PTDSS1 shRNA and CQ. K562 cells were treated with NS or PTDSS1 shRNA. Autophagy was assessed using the Cyto-ID spectrophotometric assay (C) or LC3B immunoblotting (D). (E,F) p62 immunoblotting. K562 cells treated with NS shRNA or ARG shRNAs were subject to p62 immunoblotting. (G) Co-treatment of BFA1 and shRNAs of ARGs that suppress autophagy initiation. K562 cells were transduced with viruses harboring NS shRNA or shRNAs of TCERG1, ZNF197, or POLR3D. Cells were then treated with 5 nM of BFA1 for 48 h. Autophagy was monitored using LC3B immunoblotting. (H) Co-treatment of BFA1 and shRNAs of ARGs that promote autophagy maturation. (I). Autophagy flux in K562 cells treated with IM and/or shRNAs of ARGs that promote autophagy maturation. K562 cells were transduced with viruses containing NS shRNA or shRNAs of ARGs that promote autophagy maturation. Cells were then treated with DMSO (IM−) or 1 μM of IM (IM+). Autophagy was monitored using the Cyto-ID fluorescence spectrophotometric assay. ACTB is the loading control. Images cropped from same blot are shown and full images are included in supplemental materials. Error bars represent standard deviations of three independent experiments. *P < 0.05. P values determine the difference between combination of IM and ARG shRNA and each treatment alone.
To further determine whether these 25 ARGs target autophagy maturation stage, we performed a combinational treatment of individual ARG shRNAs and PP242. PP242 is a compound that inhibits mechanistic target of rapamycin and launches autophagy65. In principle, knockdown of an ARG that promotes autophagy maturation will significantly increase the quantities of autophagic compartments together with PP242. We found that 21 ARG shRNAs exhibited statistical significance in enhancing the Cyto-ID fluorescence together with PP242 (highlighted in bold, Tables S6 and 3). As such, these ARGs facilitate the completion of the autophagy process through promoting autophagy maturation. To corroborate these results, we also monitored the changes of another autophagy marker p62 (sequestosome 1, SQSTM1), which is degraded in autolysosomes. Activation of autophagy often leads to decreased levels of p62, whereas blockade of autophagy maturation causes accumulation of this protein. We found that shRNAs of TNNI3, PPP3R2, or MBTPS1 (ARGs that suppress autophagy initiation; Table 2) significantly reduced levels of p62 proteins (Fig. 5E). In contrast, p62 protein levels increased in cells treated with shRNAs of SLC2A8, RNASEH1, or CBWD1 (Fig. 5F), which impaired autophagy maturation (Tables 2 and 3). To further validate our autophagy flux assay presented above, we used BFA1 to block autophagy flux (Fig. 1D–G) followed by LC3B immunoblotting. We found that depleting ARGs that target autophagy initiation (e.g. TCERG1, ZNF197, or POLR3D) led to a significant increase of LC3B-II levels upon BFA1 treatment (Fig. 5G), whereas knockdown of ARGs that target autophagy maturation (e.g. CCDC77, SASH3, or GLO1) failed to do so (Fig. 5H). Furthermore, an autophagy flux assay using a combination of imatinib (activating autophagy) and shRNAs of ARGs that promote autophagy maturation revealed that depletion of these ARGs significantly increased levels of autophagic compartments in combination with IM (Fig. 5I), consistent with PP242 treatment (Table 3).
Of note, shRNAs of four ARGs (ERGIC1, PHF14, TBX15, or UBA6) did not significantly change the levels of autophagic compartments together with either chloroquine (Table 2) or PP242 (Tables S6 and 3). One possible explanation is that these ARGs may inhibit PP242-induced autophagy, thereby diminishing a further increase of Cyto-ID fluorescence in cells upon combinational treatments. ARGs that target autophagy at different stages were summarized in Fig. S1. To determine the functional relevance of ARGs in CML, we measured the cytotoxicity of IM in K562 cells upon depletion of ARGs. We found that depletion of ARGs that promote autophagy maturation sensitized K562 cells to IM. In contrast, knockdown of ARGs that suppress autophagy initiation showed no additive or synergistic inhibition of cell viability (Fig. S2). These results not only provide a further validation of candidate ARGs in autophagy regulation, but also highlight the importance of ARGs as potential therapeutic targets in CML and other cancers.
Discussion
In this study, we conducted a large-scale RNAi screen in K562 cells and identified 82 ARGs with 62 candidates as novel autophagy mediators. Our further analyses revealed two subclasses of ARGs: ARGs that suppress autophagy initiation and ARGs that promote autophagy maturation (Fig. S1). Identification of new autophagy regulators in cancer cells not only helps understand how autophagy is activated or inhibited, but also provides potential drug targets for future therapeutic intervention.
RNAi screens have been used previously in mammalian cells to identify genes that regulate autophagy. For instance, Chan et al., transiently transfected GFP-LC3B-positive HEK293 cells with small interfering (si) RNAs targeting human kinases and searched for kinases that block starvation-induced autophagy66. Following this study, Lipinsky et al.,44 and McKnight et al.,67 performed a genome-wide siRNA-based RNAi screen and identified genes important for autophagy activation during starvation. Several other research groups studied autophagy flux or autophagy signaling in unstressed conditions using siRNA or shRNA-based RNAi screening68,69,70,71. In these screens, LC3B immunoblotting or microscopic imaging of GFP-LC3B or GFP-p62 was used to detect autophagy. However, the workload of LC3B immunoblotting of individual shRNA-treated cells in a large-scale RNAi screen is overwhelming and time-consuming. Fluorescence imaging of GFP-LC3B or GFP-p62 depends on the availability of a high content and expensive imaging system. To date, only a few laboratories could afford to use this high-content imaging system in large-scale RNAi screens44,67,69,70,72,73,74.
In this report, we coupled MDC staining with FACS and successfully isolated autophagic K562 cells. Past research demonstrates that the specificity of MDC in labeling autophagic cells is relatively low compared to LC3B or p62 because this fluorescence dye non-specifically stains lysosomes32,33. However, our results shown in Figs 1, 2 indicate that MDC staining is at least feasible and efficient in labeling autophagic K562 cells. The sensitivity of this approach is equivalent to LC3B immunoblotting and GFP-LC3B fluorescence microscopy. While we could not rule out the possibility of non-specific staining of non-autophagic K562 cells by MDC in our primary screen, our rigorous validations using the Cyto-ID spectrophotometric assay and LC3B immunoblotting assay effectively eliminated the false positive candidates. This is supported by the fact that one-fourth of candidate ARGs have been reported previously. Hence, using MDC-based FACS to isolate autophagic cells is suitable for performing high-content screens (e.g. loss-of-function screens, gain-of-function screens, and small molecule screens, etc.) in normal or malignant cells. More importantly, this approach is inexpensive and affordable.
As described earlier, autophagy has multifaceted activities in cancer. On one hand, autophagy protects cancer cells from therapy-induced cell death. On the other hand excessive autophagy, by itself, induces cell death in cancer cells18. Hence, the determination of autophagy stages to which 82 ARGs target has important clinical implications in treating cancer. For example, 21 ARGs that target the autophagy maturation stage are of interest in developing novel combinational therapies in that effectively blocking autophagy maturation inhibits autophagy-promoted survival of CML cells, thereby enhancing the sensitivity of CML cells to BCR-ABL inhibitors such as IM29,75. Indeed, our results shown in Fig. S2 demonstrate the functional relevance of ARGs with IM sensitivity in K562 cells. Given the possibility of excessive autophagy in inducing cell death, 57 ARGs that suppress autophagy initiation are potential new therapeutic targets for CML. Our future research will uncover the therapeutic potential of 82 ARGs reported herein in CML or other cancers that benefit from autophagy-related therapies.
Methods
Materials and reagents
K562 cells were maintained in RPMI-1640 medium supplemented with 10% fetal bovine serum, 100 μg/ml streptomycin, and 100 μg/ml penicillin. The Cyto-ID kit was purchased from Enzo Life Sciences. PP242, chloroquine, bafilomycin A1 and monodansylcadaverine (MDC) were purchased from Sigma-Aldrich. Imatinib was purchased from LC Laboratories. Stock solutions were prepared as follows: imatinib (10 mM) was dissolved in either sterile water or DMSO; PP242 (20 mM) was dissolved in DMSO; chloroquine (10 mM) was dissolved in sterile water; bafilomycin A1 (5 mM) was dissolved in DMSO.
Fluorescence microscopy
Fluorescence microscopy was used to monitor MDC fluorescence and GFP-LC3B. For MDC fluorescence microscopy, K562 cells were treated with a low dose of imatinib (1 μM) to induce autophagy26 or 5 nM of bafilomycin A1 to block autophagy flux. Cells were then stained with 0.1 mM MDC for 10 minutes in the dark at 37 °C as described previously31. Live MDC positive cells were imaged using an inverted fluorescence microscope (Carl Zeiss Microscopy, LLC, Thornwood, NY). Photos were taken using a 20× or 40× lens at different exposure times (from 100 to 200 milliseconds). For GFP-LC3B microscopy, K562 cells were stably transfected with a pEGFP-LC3B plasmid that encodes a fusion protein GFP-LC3B. Cells were then treated with imatinib (1 μM) for 18 hours and/or BFA1 (5 nM) for 48 hours. GFP fluorescence images were taken using a TCS SP8 laser scanning confocal microscope equipped with a Plan Apochromat 63×/1.4 numerical aperture oil immersion objective, and a Leica HyD hybrid detector (Leica, Buffalo Grove, IL).
Quantification of cells with GFP-LC3B puncta
Four to five images of more than 30 K562 cells treated with DMSO, 1 μM imatinib, and/or 5 nM bafilomycin A1 were randomly selected. GFP-LC3B-expressing K562 cells or K562 cells with GFP-LC3B puncta (indicating the formation of autophagosomes) in these images were counted by three persons. Percentages of cells with GFP-LC3B puncta were obtained by dividing numbers of cells with GFP-LC3B puncta with those of GFP-LC3B-expressing cells. The statistical difference was obtained using the Student’s t test.
Fluorescence spectrophotometric assay
The MDC or Cyto-ID fluorescence spectrophotometric assay was previously described34,35. In brief, autophagy was induced or inhibited in K562 cells as described in each figure. To minimize spontaneous autophagy due to lack of nutrients, cells were seeded in freshly prepared media at 5 × 104 or 105 cells per ml overnight. Live autophagic cells were stained with MDC as described above or with Cyto-ID based on the manufacturer’s instructions (Enzo Life Sciences, Inc.). Cells were then seeded in a 96-well plate and fluorescence intensity was quantified using a microplate reader (Molecular Devices, Sunnyvale, CA). The wavelengths used for detecting MDC fluorescence were 335 nm (excitation) and 525 nm (emission), respectively. The Cyto-ID fluorescence was measured at excitation 485 nm and emission 535 nm. Fluorescence levels were shown as fold changes by normalizing fluorescence intensities in cells treated with bafilomycin A1, imatinib, or shRNAs to those in untreated or non-silencing (NS) shRNA-treated cells. There were three biological replicates in each treatment group. The statistical difference was obtained using the Student’s t test.
Lentivirus production
shRNAs used in this study were built on the pLKO.1 lentiviral vector. To produce lentivirus that harbors a library of about 75,000 shRNAs targeting 15,000 human genes (OpenBiosystem, GE Dharmacon), shRNA plasmids were divided into 16 pools and used to transfect HEK293T cells using effectene (QIAGEN). 2 μg of plasmid DNA was used to transfect 5 × 106 cells to yield 3 ml media containing lentivirus. A series of dilution of viral supernatant was then used to transduce HEK293T cells and puromycin (0.25 μg/ml) was used to select infected cells. Based on cell colonies stained by crystal violet (Sigma-Aldrich), the titer of lentivirus was determined.
RNA interference screening for autophagy-regulating genes
2 × 105 K562 cells were transduced with above 16 pools of lentiviruses at a multiplicity of infection (MOI) of 2.5 using the standard spinning infection approach. The volume of viruses was calculated using the formula: (MOI × cell number)/titer. Infected cells were then selected with 0.5 μg/ml puromycin for 1 week to facilitate the insertion of shRNAs into the host cell genome. Cells were stained with 0.1 mM MDC (Sigma-Aldrich) for 10 minutes in dark at 37 °C. Cells with strong MDC fluorescence (top 1%) were collected using FACS (wavelength: excitation 335 nm and emission 525 nm). Genomic DNA was isolated immediately after FACS using the QIAmp DNA mini kit (QIAGEN). DNA fragments that contained shRNA sequences were amplified using PCR as previously described76. The primer set used for this reaction was (Forward: 5′-acgatacaaggctgttagagag-3′; Reverse: 5′-cgaaccgcaaggaaccttc-3′). The PCR products were the cloned into a TA vector (Thermo Fisher Scientific) and submitted for Sanger DNA sequencing. The sequencing primer was 5′-aaacccagggctgccttggaaaag-3′. shRNAs were identified by searching the database RNAi codex. To help identify as many candidates as possible, 500 PCR-amplified DNA fragments were sent for sequencing. Among 500 sequences, 336 different shRNAs were identified and more than 50 shRNAs were repeatedly sequenced at least one more time, suggesting that nearly all PCR-amplified shRNAs in autophagic K562 cells have been identified in our screen. Primary candidates were further validated using the Cyto-ID fluorescence spectrophotometric assay and LC3B immunoblotting. For experiments involving individual shRNAs, lentivirus was prepared as described above. K562 cells were transduced with viruses with NS shRNA or candidate shRNAs followed by 1-week puromycin (0.1–0.5 μg/ml) selection.
Immunoblotting
Immunoblotting was performed as previously reported34,77. Antibodies used in this study include: anti-LC3B (Cell Signaling Technology®, 1:1000), anti-ACTB (β-actin) (Sigma-Aldrich, 1:5000) anti-KRAS, anti-LYN (Cell Signaling Technology®, 1:1000), anti-HCLS1 (Cell Signaling Technology®, 1:1000), anti-PLK1 (Cell Signaling Technology®, 1:1000), and anti-ETS2 (Santa Cruz Biotechnology, Inc., 1:200). Quantification of protein brand intensities was performed using Image J as described previously34. The band intensities of proteins of interest were first normalized to that of actin (ACTB), yielding relative intensities. The relative intensities of proteins of interest in candidate shRNA-treated cells were further normalized with that in NS shRNA-treated cells.
Quantitative RT-PCR
mRNA levels of genes of interest were measured using qRT-PCR as described previously76,78. In brief, K562 cells treated with NS shRNA or individual candidate shRNAs were subject to total RNA isolation using Trizol (Thermo Fisher Scientific). 1 to 5 μg total RNA was used to generate cDNA using SuperScript III reverse transcriptase (Thermo Fisher Scientific). The mRNA levels of each candidate gene were determined using quantitative real-time PCR.
Bioinformatic analyses
Potential protein-protein interaction was analyzed using the online program STRING v9.1 (http://string-db.org). Bait proteins included (1) proteins that directly participate in the autophagy process and (2) kinases that regulate autophagy. Results were exported as image files and modified using adobe illustrator. Lines between nodes represent potential interactions. Colors denote different assays. Gene ontology analyses were performed using the online program DAVID (https://david.ncifcrf.gov) or GeneCards database.
Autophagy stage determination assay
Autophagy stage determination assay was previously described34,35. K562 cells that stably express NS shRNA or individual candidate shRNAs were treated with either vehicle (water or DMSO), chloroquine (2.5 μM for 6 hours), or bafilomycin A1 (5 nM for 48 hours). In some cases, these cells were treated with DMSO, PP242 (5 μM), or imatinib (1 μM) for 8–10 hours. The amount of autophagic compartments was measured using the Cyto-ID fluorescence spectrophotometric assay or LC3B immunoblotting as described above. The Cyto-ID level in cells treated with the combinations was compared to that of each treatment alone. The statistical difference was obtained using the Student’s t test.
Data Availability
All data are available for sharing upon request.
Change history
10 June 2019
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has been fixed in the paper.
References
Kroemer, G. Autophagy: a druggable process that is deregulated in aging and human disease. J Clin Invest 125, 1–4, https://doi.org/10.1172/JCI78652 (2015).
White, E. Deconvoluting the context-dependent role for autophagy in cancer. Nat Rev Cancer 12, 401–410, https://doi.org/10.1038/nrc3262 (2012).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 8, 445–544 (2012).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42, https://doi.org/10.1016/j.cell.2007.12.018 (2008).
Galluzzi, L. et al. Autophagy in malignant transformation and cancer progression. Embo J 34, 856–880, https://doi.org/10.15252/embj.201490784 (2015).
Schmukler, E., Kloog, Y. & Pinkas-Kramarski, R. Ras and autophagy in cancer development and therapy. Oncotarget 5, 577–586 (2014).
Gewirtz, D. A. The four faces of autophagy: implications for cancer therapy. Cancer Res 74, 647–651, https://doi.org/10.1158/0008-5472.CAN-13-2966 (2014).
Cheong, H., Lu, C., Lindsten, T. & Thompson, C. B. Therapeutic targets in cancer cell metabolism and autophagy. Nat Biotechnol 30, 671–678, https://doi.org/10.1038/nbt.2285 (2012).
Kimmelman, A. C. The dynamic nature of autophagy in cancer. Genes Dev 25, 1999–2010, https://doi.org/10.1101/gad.17558811 (2011).
Xia, Y. et al. A combination therapy for KRAS-driven lung adenocarcinomas using lipophilic bisphosphonates and rapamycin. Science translational medicine 6, 263ra161, https://doi.org/10.1126/scitranslmed.3010382 (2014).
Carayol, N. et al. Critical roles for mTORC2- and rapamycin-insensitive mTORC1-complexes in growth and survival of BCR-ABL-expressing leukemic cells. Proc Natl Acad Sci U S A 107, 12469–12474, https://doi.org/10.1073/pnas.1005114107 (2010).
Iwamaru, A. et al. Silencing mammalian target of rapamycin signaling by small interfering RNA enhances rapamycin-induced autophagy in malignant glioma cells. Oncogene 26, 1840–1851, https://doi.org/10.1038/sj.onc.1209992 (2007).
Schuurman, H. J. et al. SDZ RAD, a new rapamycin derivative: synergism with cyclosporine. Transplantation 64, 32–35 (1997).
Karnoub, A. E. & Weinberg, R. A. Ras oncogenes: split personalities. Nature reviews. Molecular cell biology 9, 517–531, https://doi.org/10.1038/nrm2438 (2008).
Inoki, K., Li, Y., Zhu, T., Wu, J. & Guan, K. L. TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat Cell Biol 4, 648–657, https://doi.org/10.1038/ncb839 (2002).
Denton, D., Xu, T. & Kumar, S. Autophagy as a pro-death pathway. Immunol Cell Biol 93, 35–42, https://doi.org/10.1038/icb.2014.85 (2015).
Shanware, N. P., Bray, K. & Abraham, R. T. The PI3K, metabolic, and autophagy networks: interactive partners in cellular health and disease. Annual review of pharmacology and toxicology 53, 89–106, https://doi.org/10.1146/annurev-pharmtox-010611-134717 (2013).
Levine, B. Cell biology: autophagy and cancer. Nature 446, 745–747, https://doi.org/10.1038/446745a (2007).
Luiken, J. J., Aerts, J. M. & Meijer, A. J. The role of the intralysosomal pH in the control of autophagic proteolytic flux in rat hepatocytes. Eur J Biochem 235, 564–573 (1996).
Sotelo, J., Briceno, E. & Lopez-Gonzalez, M. A. Adding chloroquine to conventional treatment for glioblastoma multiforme: a randomized, double-blind, placebo-controlled trial. Ann Intern Med 144, 337–343 (2006).
Poklepovic, A. & Gewirtz, D. A. Outcome of early clinical trials of the combination of hydroxychloroquine with chemotherapy in cancer. Autophagy 10, 1478–1480, https://doi.org/10.4161/auto.29428 (2014).
Manic, G., Obrist, F., Kroemer, G., Vitale, I. & Galluzzi, L. Chloroquine and hydroxychloroquine for cancer therapy. Mol Cell Oncol 1, e29911, https://doi.org/10.4161/mco.29911 (2014).
Bartram, C. R. et al. Translocation of c-ab1 oncogene correlates with the presence of a Philadelphia chromosome in chronic myelocytic leukaemia. Nature 306, 277–280 (1983).
Druker, B. J. et al. Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nature medicine 2, 561–566 (1996).
Volpe, G., Panuzzo, C., Ulisciani, S. & Cilloni, D. Imatinib resistance in CML. Cancer Lett 274, 1–9, https://doi.org/10.1016/j.canlet.2008.06.003 (2009).
Sheng, Z., Ma, L., Sun, J. E., Zhu, L. J. & Green, M. R. BCR-ABL suppresses autophagy through ATF5-mediated regulation of mTOR transcription. Blood 118, 2840–2848, https://doi.org/10.1182/blood-2010-12-322537 (2011).
Helgason, G. V., Karvela, M. & Holyoake, T. L. Kill one bird with two stones: potential efficacy of BCR-ABL and autophagy inhibition in CML. Blood 118, 2035–2043, https://doi.org/10.1182/blood-2011-01-330621 (2011).
Donato, N. J. Bcr-Abl adds another twist to cell fate. Blood 118, 2646–2647, https://doi.org/10.1182/blood-2011-07-366153 (2011).
Bellodi, C. et al. Targeting autophagy potentiates tyrosine kinase inhibitor-induced cell death in Philadelphia chromosome-positive cells, including primary CML stem cells. J Clin Invest 119, 1109–1123, https://doi.org/10.1172/JCI35660 (2009).
Joachim, J., Jiang, M., McKnight, N. C., Howell, M. & Tooze, S. A. High-throughput screening approaches to identify regulators of mammalian autophagy. Methods, https://doi.org/10.1016/j.ymeth.2015.02.002 (2015).
Biederbick, A., Kern, H. F. & Elsasser, H. P. Monodansylcadaverine (MDC) is a specific in vivo marker for autophagic vacuoles. Eur J Cell Biol 66, 3–14 (1995).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222, https://doi.org/10.1080/15548627.2015.1100356 (2016).
Mizushima, N., Yoshimori, T. & Levine, B. Methods in mammalian autophagy research. Cell 140, 313–326, https://doi.org/10.1016/j.cell.2010.01.028 (2010).
Guo, S. et al. A rapid and high content assay that measures cyto-ID-stained autophagic compartments and estimates autophagy flux with potential clinical applications. Autophagy 11, 560–572, https://doi.org/10.1080/15548627.2015.1017181 (2015).
Guo, S., Pridham, K. J. & Sheng, Z. Detecting Autophagy and Autophagy Flux in Chronic Myeloid Leukemia Cells Using a Cyto-ID Fluorescence Spectrophotometric Assay. Methods Mol Biol 1465, 95–109, https://doi.org/10.1007/978-1-4939-4011-0_9 (2016).
Aoyagi, K. et al. VAMP7 Regulates Autophagy to Maintain Mitochondrial Homeostasis and to Control Insulin Secretion in Pancreatic beta-Cells. Diabetes 65, 1648–1659, https://doi.org/10.2337/db15-1207 (2016).
Dong, G. et al. STS-1 promotes IFN-alpha induced autophagy by activating the JAK1-STAT1 signaling pathway in B cells. Eur J Immunol 45, 2377–2388, https://doi.org/10.1002/eji.201445349 (2015).
Dupont, N. et al. Neutral lipid stores and lipase PNPLA5 contribute to autophagosome biogenesis. Current biology: CB 24, 609–620, https://doi.org/10.1016/j.cub.2014.02.008 (2014).
Ge, L., Melville, D., Zhang, M. & Schekman, R. The ER-Golgi intermediate compartment is a key membrane source for the LC3 lipidation step of autophagosome biogenesis. eLife 2, e00947, https://doi.org/10.7554/eLife.00947 (2013).
Jeong, K., Kwon, H. Y., Jeong, M. S., Sohn, E. J. & Kim, S. H. CNOT2 promotes degradation of p62/SQSTM1 as a negative regulator in ATG5 dependent autophagy. Oncotarget. https://doi.org/10.18632/oncotarget.17682 (2017).
Kim, M. J. et al. Involvement of autophagy in oncogenic K-Ras-induced malignant cell transformation. J Biol Chem 286, 12924–12932, https://doi.org/10.1074/jbc.M110.138958 (2011).
Kwak, S. S. et al. Autophagy induction by tetrahydrobiopterin deficiency. Autophagy 7, 1323–1334, https://doi.org/10.4161/auto.7.11.16627 (2011).
Li, X. et al. Lyn Delivers Bacteria to Lysosomes for Eradication through TLR2-Initiated Autophagy Related Phagocytosis. PLoS Pathog 12, e1005363, https://doi.org/10.1371/journal.ppat.1005363 (2016).
Lipinski, M. M. et al. A genome-wide siRNA screen reveals multiple mTORC1 independent signaling pathways regulating autophagy under normal nutritional conditions. Developmental cell 18, 1041–1052, https://doi.org/10.1016/j.devcel.2010.05.005 (2010).
Ma, S. et al. The C-ETS2-TFEB Axis Promotes Neuron Survival under Oxidative Stress by Regulating Lysosome Activity. Oxid Med Cell Longev 2016, 4693703, https://doi.org/10.1155/2016/4693703 (2016).
Ogasawara, Y. et al. Stearoyl-CoA desaturase 1 activity is required for autophagosome formation. J Biol Chem 289, 23938–23950, https://doi.org/10.1074/jbc.M114.591065 (2014).
Prasad, K. V., Taiyab, A., Jyothi, D., Srinivas, U. K. & Sreedhar, A. S. Heat shock transcription factors regulate heat induced cell death in a rat histiocytoma. Journal of biosciences 32, 585–593 (2007).
Ramakrishnan, R. & Gabrilovich, D. I. The role of mannose-6-phosphate receptor and autophagy in influencing the outcome of combination therapy. Autophagy 9, 615–616, https://doi.org/10.4161/auto.23485 (2013).
Sargent, G. et al. PEX2 is the E3 ubiquitin ligase required for pexophagy during starvation. J Cell Biol 214, 677–690, https://doi.org/10.1083/jcb.201511034 (2016).
Shen, W. et al. S100A4 interacts with mutant p53 and affects gastric cancer MKN1 cell autophagy and differentiation. Int J Oncol 47, 2123–2130, https://doi.org/10.3892/ijo.2015.3209 (2015).
Shibuya, Y., Chang, C. C., Huang, L. H., Bryleva, E. Y. & Chang, T. Y. Inhibiting ACAT1/SOAT1 in microglia stimulates autophagy-mediated lysosomal proteolysis and increases Abeta1-42 clearance. J Neurosci 34, 14484–14501, https://doi.org/10.1523/JNEUROSCI.2567-14.2014 (2014).
Valianou, M. et al. Pharmacological inhibition of Polo-like kinase 1 (PLK1) by BI-2536 decreases the viability and survival of hamartin and tuberin deficient cells via induction of apoptosis and attenuation of autophagy. Cell Cycle 14, 399–407, https://doi.org/10.4161/15384101.2014.986394 (2015).
Zhang, D., Xu, X. & Dong, Z. PRKCD/PKCdelta contributes to nephrotoxicity during cisplatin chemotherapy by suppressing autophagy. Autophagy 13, 631–632, https://doi.org/10.1080/15548627.2016.1269990 (2017).
Pozuelo-Rubio, M. Regulation of autophagic activity by 14-3-3zeta proteins associated with class III phosphatidylinositol-3-kinase. Cell Death Differ 18, 479–492, https://doi.org/10.1038/cdd.2010.118 (2011).
Parzych, K. R. & Klionsky, D. J. An overview of autophagy: morphology, mechanism, and regulation. Antioxid Redox Signal 20, 460–473, https://doi.org/10.1089/ars.2013.5371 (2014).
Franceschini, A. et al. STRINGv9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res 41, D808–815, https://doi.org/10.1093/nar/gks1094 (2013).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature protocols 4, 44–57, https://doi.org/10.1038/nprot.2008.211 (2009).
Lum, J. J. et al. Growth factor regulation of autophagy and cell survival in the absence of apoptosis. Cell 120, 237–248, https://doi.org/10.1016/j.cell.2004.11.046 (2005).
Baker, S. J., Rane, S. G. & Reddy, E. P. Hematopoietic cytokine receptor signaling. Oncogene 26, 6724–6737, https://doi.org/10.1038/sj.onc.1210757 (2007).
von Gunten, S. & Simon, H. U. Autophagic-like cell death in neutrophils induced by autoantibodies. Autophagy 3, 67–68 (2007).
Ryter, S. W., Koo, J. K. & Choi, A. M. Molecular regulation of autophagy and its implications for metabolic diseases. Current opinion in clinical nutrition and metabolic care 17, 329–337, https://doi.org/10.1097/MCO.0000000000000068 (2014).
Russell, R. C., Yuan, H. X. & Guan, K. L. Autophagy regulation by nutrient signaling. Cell Res 24, 42–57, https://doi.org/10.1038/cr.2013.166 (2014).
Sinclair, A., Latif, A. L. & Holyoake, T. L. Targeting Survival Pathways in Chronic Myeloid Leukaemia Stem Cells. Br J Pharmacol. https://doi.org/10.1111/bph.12183 (2013).
Crowley, L. C., Elzinga, B. M., O’Sullivan, G. C. & McKenna, S. L. Autophagy induction by Bcr-Abl-expressing cells facilitates their recovery from a targeted or nontargeted treatment. Am J Hematol 86, 38–47, https://doi.org/10.1002/ajh.21914 (2011).
Apsel, B. et al. Targeted polypharmacology: discovery of dual inhibitors of tyrosine and phosphoinositide kinases. Nat Chem Biol 4, 691–699, https://doi.org/10.1038/nchembio.117 (2008).
Chan, E. Y., Kir, S. & Tooze, S. A. siRNA screening of the kinome identifies ULK1 as a multidomain modulator of autophagy. J Biol Chem 282, 25464–25474, https://doi.org/10.1074/jbc.M703663200 (2007).
McKnight, N. C. et al. Genome-wide siRNA screen reveals amino acid starvation-induced autophagy requires SCOC and WAC. Embo J 31, 1931–1946, https://doi.org/10.1038/emboj.2012.36 (2012).
Puustinen, P. et al. CIP2A oncoprotein controls cell growth and autophagy through mTORC1 activation. J Cell Biol 204, 713–727, https://doi.org/10.1083/jcb.201304012 (2014).
Strohecker, A. M. et al. Identification of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase as a novel autophagy regulator by high content shRNA screening. Oncogene 34, 5662–5676, https://doi.org/10.1038/onc.2015.23 (2015).
Hale, C. M. et al. Identification of modulators of autophagic flux in an image-based high content siRNA screen. Autophagy 12, 713–726, https://doi.org/10.1080/15548627.2016.1147669 (2016).
Jung, J. et al. Multiplex image-based autophagy RNAi screening identifies SMCR8 as ULK1 kinase activity and gene expression regulator. eLife 6, https://doi.org/10.7554/eLife.23063 (2017).
Orvedahl, A. et al. Image-based genome-wide siRNA screen identifies selective autophagy factors. Nature 480, 113–117, https://doi.org/10.1038/nature10546 (2011).
Lefebvre, V. et al. Genome-wide RNAi screen identifies ATPase inhibitory factor 1 (ATPIF1) as essential for PARK2 recruitment and mitophagy. Autophagy 9, 1770–1779, https://doi.org/10.4161/auto.25413 (2013).
Ivatt, R. M. et al. Genome-wide RNAi screen identifies the Parkinson disease GWAS risk locus SREBF1 as a regulator of mitophagy. Proc Natl Acad Sci USA 111, 8494–8499, https://doi.org/10.1073/pnas.1321207111 (2014).
Calabretta, B. & Salomoni, P. Inhibition of autophagy: a new strategy to enhance sensitivity of chronic myeloid leukemia stem cells to tyrosine kinase inhibitors. Leuk Lymphoma 52(Suppl 1), 54–59, https://doi.org/10.3109/10428194.2010.546913 (2011).
Sheng, Z., Murphy, S. F., Guo, S. & Green, M. R. A diphtheria toxin negative selection in RNA interference screening. Methods in molecular biology 1176, 59–72, https://doi.org/10.1007/978-1-4939-0992-6_6 (2014).
Murphy, S. F. et al. Connexin 43 Inhibition Sensitizes Chemoresistant Glioblastoma Cells to Temozolomide. Cancer Res 76, 139–149, https://doi.org/10.1158/0008-5472.CAN-15-1286 (2016).
Sheng, Z. et al. A genome-wide RNA interference screen reveals an essential CREB3L2-ATF5-MCL1 survival pathway in malignant glioma with therapeutic implications. Nature medicine 16, 671–677, https://doi.org/10.1038/nm.2158 (2010).
Acknowledgements
We thank Sara Deibler for text editing and all members in the Sheng and Green lab for critical discussion of this study. This work was supported by the Startup Funds provided to Z.S. by the Virginia Tech Carilion Research Institute and US National Institutes of Health grant R01CA163926 to M.R.G. M.R.G is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Z.S. and M.R.G. conceived the project. Z.S. wrote the manuscript. S.G., K.J.P. and Z.S. performed most of the experiments. C.M.V. prepared viruses of shRNAs. L.Z. and B.H. performed some bioinformatic analyses. H.V. assisted candidate validation and manuscript editing.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, S., Pridham, K.J., Virbasius, CM. et al. A large-scale RNA interference screen identifies genes that regulate autophagy at different stages. Sci Rep 8, 2822 (2018). https://doi.org/10.1038/s41598-018-21106-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-21106-5
This article is cited by
-
Connexin 43 confers chemoresistance through activating PI3K
Oncogenesis (2022)
-
A high-throughput screen identifies the long non-coding RNA DRAIC as a regulator of autophagy
Oncogene (2019)
-
Casein Kinase 1 Epsilon Regulates Glioblastoma Cell Survival
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.