Abstract
Open-angle glaucoma (OAG) is a major cause of blindness worldwide. To identify new risk loci for OAG, we performed a genome-wide association study in 3,071 OAG cases and 6,750 unscreened controls, and meta-analysed the results with GWAS data for intraocular pressure (IOP) and optic disc parameters (the overall meta-analysis sample size varying between 32,000 to 48,000 participants), which are glaucoma-related traits. We identified and independently validated four novel genome-wide significant associations within or near MYOF and CYP26A1, LINC02052 and CRYGS, LMX1B, and LMO7 using single variant tests, one additional locus (C9) using gene-based tests, and two genetic pathways - “response to fluid shear stress” and “abnormal retina morphology” - in pathway-based tests. Interestingly, some of the new risk loci contribute to risk of other genetically-correlated eye diseases including myopia and age-related macular degeneration. To our knowledge, this study is the first integrative study to combine genetic data from OAG and its correlated traits to identify new risk variants and genetic pathways, highlighting the future potential of combining genetic data from genetically-correlated eye traits for the purpose of gene discovery and mapping.
Similar content being viewed by others
Introduction
OAG is characterized by optic nerve damage and progressive loss of peripheral vision, with many patients remaining undiagnosed until severe irreversible vision loss has occurred1,2. OAG has a significant genetic component with a relative risk of over 9 in first-degree relatives of affected individuals compared to relatives of unaffected people3. Our previous genome-wide association studies (GWAS) have reproducibly identified several risk loci for OAG including TMCO1, CDKN2B-AS1, SIX6, CAV1, CAV2, ABCA1, AFAP1, GMDS, ARHGEF12, TXNRD2, ATXN2, and FOXC14,5,6,7,8,9. However, the majority of the genetic variance contributing to OAG remains unexplained, emphasizing that further studies to identify additional risk loci for OAG are required in order to make genetic risk prediction more clinically useful.
Optic disk parameters including cup area (CA; the central area), disc area (DA; the total area of optic disc including cup area and the surrounding area containing axons of the retinal ganglion cells), and vertical cup-disc ratio (VCDR; the ratio of the vertical diameter of cup area to the vertical diameter of the optic disc) are key measurements used to assess OAG diagnosis and progression10. Elevated intraocular pressure (IOP) is the major known risk factor for OAG2. We refer to CA, DA, VCDR, and IOP as OAG endophenotypes or quantitative traits. There are high genetic correlations between these quantitative traits and OAG, with several of the already known risk loci for these traits overlapping with each other and with OAG loci, demonstrating their utility as endophenotypes11,12,13,14. These findings suggest that combining genetic data from OAG and its endophenotypes has the potential to increase the probability of identifying genetic variants that are common between traits, thus enabling the extraction of greater genetic power from valuable disease cohorts.
In this study, we sought to identify additional risk loci contributing to OAG susceptibility by (1) increasing the sample size for OAG, (2) combining GWAS data from OAG and its endophenotypes in order to increase our statistical power to identify new risk loci for OAG, and (3) applying gene and pathway based approaches.
Results
In total, 3,071 OAG cases from the Australian & New Zealand Registry of Advanced Glaucoma (ANZRAG) obtained in three phases of data collection, and 6,750 unscreened controls of European descent were used as the GWAS discovery dataset in this study (Supplementary Table 1). Five loci were associated with OAG at genome-wide significance level in the meta-analysis of GWAS results between the three phases of ANZRAG data (P < 5 × 10−8), including regions near or within CDKN2B-AS1, ABCA1, C14orf39 and SIX6, TMCO1, and ARHGEF12, all of which are now well established risk loci for OAG5,6,7,9. Manhattan and Q-Q plots are shown in Supplementary Figure 1. Genomic inflation factor lambda was 1.006 for this analysis.
Next, to increase the power of this study to identify new risk loci for OAG, we performed GWAS meta-analyses of ANZRAG OAG and each of the endophenotypes (CA, DA, VCDR, and IOP) that we obtained from our previous study11 (Supplementary Table 1). Before performing the meta-analyses, we confirmed validity of the endophenotypes for OAG by showing that there were significant genetic correlations between OAG and the endophenotypes (ranging between 20% and 47%, Supplementary Table 2; p ≤ 0.018) using the cross-trait bivariate LD score regression approach15. By design, the endophenotype studies did not include any of the OAG cases. This was further confirmed in the LD score bivariate analyses where the intercepts were close to zero with 95% confidence intervals (CI) overlapping zero, indicating that there was not significant sample overlap between our OAG and the endophenotypes studies. Moreover, intercepts of the univariate LD score regression analyses16 were close to 1 with 95% CIs overlapping 1 (Supplementary Table 3), indicating that there was no model misspecification and other sources of bias such as population stratification and cryptic relatedness in either study16.
Four genomic regions that were genome-wide significant in meta-analyses of ANZRAG OAG and one of the endophenotypes (Table 1), and were not previously known risk loci for OAG, and had at least P < 0.05 in the OAG separate analysis, were taken forward for validation. The best SNPs within these regions were rs72815193[G] (risk alleles are indicated within brackets) (P = 6.10 × 10−10) on chromosome 10 near MYOF, XRCC6P1, and CYP26A1 for combined OAG and VCDR (in European and Asian ancestries), rs56962872[G] (P = 2.81 × 10−8) on chromosome 3 within LINC02052 and near CRYGS for combined OAG and VCDR (in European ancestry), rs6478746[G] (P = 4.54 × 10−8) on chromosome 9 near LOC105376277 and LMX1B for combined OAG and CA (in European ancestry), and rs148639588[T] (P = 3.53 × 10−8) on chromosome 1 near COL11A1 for combined OAG and CA (in European ancestry).
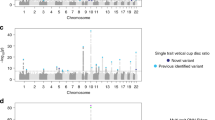
Of the above four loci that were genome-wide significant in our discovery meta-analyses, two loci (LOC105376277/LMX1B and LINC02052/CRYGS) were replicated (P < 0.0125, the Bonferroni-corrected threshold considering four independent tests) for OAG in an independent replication study, the National Eye Institute Glaucoma Human Genetics Collaboration Heritable Overall Operational Database (NEIGHBORHOOD) (Supplementary Notes), containing 3,853 OAG cases and 33,480 controls8 (Table 2). Furthermore, rs72815193 near MYOF and XRCC6P1 had a P = 0.06, and rs4918865 (in high LD with rs72815193; LD r2 = 0.93) had a P = 0.02 for OAG in NEIGHBORHOOD. Although the SNPs near MYOF and XRCC6P1 did not pass the Bonferroni-corrected threshold of P < 0.0125, rs4918865 was more strongly associated with the high-tension glaucoma (HTG) subset (P = 0.003 for HTG vs. P = 0.37 for normal-tension glaucoma (NTG)) in NEIGHBORHOOD. The COL11A1 association was not replicated in NEIGHBORHOOD (P = 0.41 for rs148639588). The statistics including effect sizes of the top SNPs within the three new replicated loci with OAG separately (the meta-analysed OAG data from ANZRAG and NEIGHBORHOOD studies, without including the endophenotype data) are summarized in Table 2. All of the three new replicated loci were associated with CA and VCDR at, at least, nominal significance (P < 0.05), while LMX1B was also nominally (P = 0.03) associated with IOP (Table 3). Manhattan and Q-Q plots are shown in Supplementary Figure 1, and regional association plots in Fig. 1. Genomic inflation factor lambda ranged between 1.03 and 1.05 for these analyses.
Regional plots for the new risk loci identified in single variant analyses in this study. The most significantly associated SNPs in each region are marked as solid purple diamonds. Pairwise correlations (LD r2) between the top SNP and the other SNPs in a 400 kb flanking region are illustrated by different colours. Blue spikes show estimated recombination rates. (a) rs72815193 on chromosome 10 near MYOF, CYP26A1, and CYP26C (the most significant results were obtained in combined OAG and VCDR analysis conducted in combined Asians and European ancestry). (b) rs56962872 on chromosome 3 within LOC253573 (LINC02052), near CRYGS and TBCCD1 (the most significant results were obtained in combined OAG and VCDR analysis conducted in European ancestry). (c) rs6478746 on chromosome nine near LMX1B (the most significant results were obtained in combined OAG and CA in European ancestry). (d) rs9530458 on chromosome 13 within LMO7 (combined OAG data from ANZRAG and NEIGHBORHOOD). cM = centimorgan.
Next, results for the SNPs that were not genome-wide significant, but approached this threshold (SNPs with P < 1 × 10−7) in the ANZRAG OAG meta-analysis or the meta-analysis of OAG and its endophenotypes, were combined with those from the NEIGHBORHOOD replication data, using a fixed-effects meta-analysis. A fourth locus on chromosome 13 within LMO7 that was nearly genome-wide significant in the OAG and CA (European ancestry) meta-analysis (rs9530458 [T], P = 2.71 × 10−7) became genome-wide significant (rs9530458 [T], OR = 1.148, P = 3.45 × 10−9) in the meta-analysis of the OAG data (ANZRAG discovery and NEIGHBORHOOD replication studies), without including the endophenotypes. This SNP was nominally associated with CA, VCDR and IOP (Table 3). Regional association for this locus is plotted in Fig. 1.
We also investigated association of the new loci with NTG and HTG subsets within the ANZRAG and NEIGHBORHOOD data (overall 1,546 NTG cases, 3,412 HTG cases, and 40,230 controls). The results summarized in Supplementary Table 4 show that the 95% CIs overlap between the NTG and HTG analyses, suggesting that these loci may affect both NTG and HTG. However, larger sample sizes are required to further investigate this, as especially for NTG, the 95% CIs are quite wide, and for the LMO7 SNP (rs9530458) overlaps 1.
We performed a series of sensitivity analyses by excluding the ANZRAG cases in which visual field data was unavailable (585 people) as well as people with mixed-mechanism glaucoma (277 people with OAG as well as a secondary glaucoma) to ensure that the results were not driven by uncertainty in phenotype assignment. The results from the sensitivity analyses in ANZRAG were meta-analysed with the endophenotype or NEIGHBORHOOD results as for the main analysis (Supplementary Tables 5 and 6). Overall, effect sizes obtained from the original and sensitivity analyses were similar, suggesting that our results were not biased by presence of any phenotype uncertainties.
Interestingly, rs72815193 and rs4918865 within the MYOF and XRCC6P1 locus are in high LD (r2 = 0.840 and r2 = 0.9, respectively) with rs10882165, a SNP that has been shown to be associated with refractive error (P = 1 × 10−11)17, indicating that this locus may affect glaucoma and its endophenotypes as well as myopia. In addition, SNPs within LMO7 have been suggestively associated with corneal astigmatism (P = 4 × 10−6 for rs11841001)18. However, rs9530458 is in low LD (r2 = 0.14) with rs11841001 (P = 0.06 in the OAG and CA analysis in Europeans), suggesting that even if the LMO7 gene affects glaucoma as well as corneal astigmatism, this effect may come from independent risk variants within LMO7. On the other hand, the MYOF and XRCC6P1 locus is ~1 Mb away from PLCE1, a known risk locus for VCDR. However, rs72815193 (MYOF) is not in LD with rs7072574 (PLCE1) (LD r2 = 0.001) (P = 3.86 × 10−6 in the OAG and VCDR analysis in Europeans), suggesting that these are independent loci.
Gene-based results
We used the approaches implemented in MetaXcan19, fastBAT20, and EUGENE21, to identify genes whose genetic variants or expression levels were significantly associated with development of OAG and its endophenotypes. These gene-based tests are complementary since they make different assumptions and use different approaches and input data to identify associated genes. After Bonferroni correction for multiple testing (refer to the Methods section), nine genes (one from fastBAT, five from MetaXcan, and three from Eugene approaches, Table 4) that were genome-wide significant in the gene-based methods, and were not overlapping with the known risk loci for OAG and its endophenotypes, were taken forward for validation in NEIGHBORHOOD. For MetaXcan and EUGENE approaches, we investigated replication of the significant genes in the same tissues that showed significance in the discovery set since the results from these approaches are tissue-specific.
One previously unreported gene became gene-wide significant (P < 7 × 10−7; see the Methods section) in the fastBAT approach (Table 4). This gene, complement factor 9 (C9) (P = 4.93 × 10−7 in the combined OAG and IOP analysis in European ancestry) was replicated in the NEIGHBORHOOD data (P = 0.04). The best result in the single variant tests for rs56345442, the top SNP within C9, was observed in the combined OAG and IOP analysis (P = 4.43 × 10−6 for combined European ancestry and Asians, and P = 9.998 × 10−6 in European ancestry), suggesting that a larger sample size would be required to detect this association at genome-wide significance threshold in the single variant analysis.
Five previously unreported genes were gene-wide significant (P < 5 × 10−8; see the Methods section) in the MetaXcan approach (Table 4), all of which were located within 1 Mb of a previously known locus. Of those genes, association of two genes, RNF26 (P = 2.66 × 10−9 in the discovery set) and NPAS4 (P = 3.97 × 10−9 in the discovery set), were replicated in the NEIGHBORHOOD data (P = 0.0004 and P = 0.05 for RNF26 and NPAS4, respectively). However, four of the eQTL SNPs (rs1893261, rs11823300, rs61898351, and rs11217821) used by MetaXcan to impute the gene expression levels for RNF26 are in LD r2 = 0.3 with rs11827818, located within a previously known locus (near ARHGEF12) for IOP and OAG. Repeating the analysis without SNPs in LD r2 > 0.2 with rs11827818 (56 SNPs remained out of the original 60 SNPs) led to a non-significant result for this gene (P = 0.85). Similarly, excluding the eQTL SNPs in LD r2 > 0.2 with rs7931311, an already known locus near SCYL1 showed a non-significant association for NPAS4 (P = 0.82; twenty SNPs remained out of the original 25 SNPs for this analysis). Although these results are valuable for the purpose of fine-mapping of the previously known associations, they suggest that RNF26 and NPAS4 are not new risk loci for OAG, but are driven by the eQTL SNPs within the previously known loci.
Three previously unreported genes were gene-wide significant (P < 9 × 10−6; see the Methods section) in brain using the EUGENE approach. However, none were replicated in brain using the EUGENE approach applied to the NEIGHBORHOOD data. Despite this, while DHRS7 was associated at P = 0.07 in brain in NEIGHBORHOOD, there was a stronger association at P = 0.0006 in blood. These data suggestively support that DHRS7 may also be a risk locus for OAG.
Accordingly, in addition to the risk loci identified in the single variant analyses, our gene-based approaches identified and validated C9 as an additional new risk locus for OAG. The previously reported OAG loci that also passed the gene-wide significance threshold in the gene-based tests included TMEM136 in the MetaXcan approach, AFAP1, AFAP1-AS, ARHGEF12, and TXNRD2 in the EUGENE approach, and TMCO1, ABCA1, C9orf53, CDKN2A, CDKN2B, CDKN2B-AS1, ARHGEF12, TMEM136, SIX1, SIX4, SIX6, AFAP1, GMDS, CAV1, and CAV2 in the fastBAT approach (Supplementary Table 7). These data provide further support for these genes being the target genes within the previously reported risk loci for OAG.
Pathway-based results
Two genetic pathways survived the significance threshold of P < 1 × 10−6 and false discovery rate < 0.05 in the pathway-based analysis in DEPICT22. One pathway was the “response to fluid shear stress” (GO: 0034405, P = 2.09 × 10−7, FDR < 0.01) in the combined OAG and CA analysis, and the other was “abnormal retina morphology” (MP: 0001325, P = 2.50 × 10−7, FDR < 0.01) in the combined OAG and VCDR analysis. The “abnormal retina morphology” pathway is interesting because it emphasizes that common risk loci between OAG and VCDR could be functioning through mechanisms related to retinal formation.
Gene expression
We also investigated the expression of the nearest genes to the best associated SNPs within the new OAG risk loci using RNA sequencing data from relevant human tissues including optic nerve, optic nerve head, retina, ciliary body pars plicata, trabecular meshwork, corneal endothelium, corneal stroma, and corneal epithelium (see Methods section). We observed a higher expression of LMX1B in trabecular meshwork, corneal endothelium, and corneal stroma, MYOF in trabecular meshwork and corneal epithelium, and LMO7 in corneal epithelium (Supplementary Figure 2). A relatively higher expression of LMX1B and MYOF in the trabecular meshwork is interesting because it is consistent with the previous observations for other known OAG genes such as MYOC showing high expression profile in the trabecular meshwork23. In addition, the LMX1B results are also consistent with the results from the GTEx eQTL studies where rs4837100, the SNP in high LD with the top variant in the LMX1B locus is an eQTL (P = 7 × 10−5) for LMX1B in sub-cutaneous adipose tissues24, suggesting that risk variants within this locus may alter expression levels of LMX1B.
Discussion
Our study identified four new OAG risk loci in single variant analyses as well as an additional locus using a gene-based approach. Interestingly, some of these new risk loci contribute to risk of other partially correlated eye diseases including age-related macular degeneration (AMD) and myopia (more details below). We also highlighted two genetic pathways associated with the development of OAG, one of which is gene-sets contributing to morphology of retina.
This study highlights the potential of combining genetic data from correlated eye traits for the purpose of gene discovery and mapping. We showed that meta-analysis of GWAS summary statistics from OAG and its correlated traits (VCDR, CA, DA, and IOP) is capable of identifying new risk loci by increasing statistical power. To our knowledge, this is the first study to use an integrative approach for OAG and its endophenotypes to identify new risk loci for OAG. This approach identifies risk variants common between OAG and its correlated traits, while increasing statistical power to detect variants with small effect sizes at the genome-wide significance threshold, which otherwise requires a much larger OAG sample for successful detection.
While all the new risk loci were at least nominally associated with CA and VCDR, none were associated with DA. This suggests that the new loci identified in this study are more likely to influence the size of the central area of the optic disk, rather than the total disc size. There have been some debates on whether the total size of the optic disc is a suitable trait to predict OAG risk and progression25. In this study we estimated a much smaller genetic correlation between DA and OAG as compared with genetic correlation between OAG and CA, VCDR, and IOP. In addition, the majority of the genome-wide significant loci in our meta-analyses of OAG and DA showed significant heterogeneity (P < 0.05) between the GWAS results from OAG and DA (data not shown), further suggesting that DA may not be as suitable as CA and VCDR to be used as an endophenotype for POAG.
Bioinformatics functional features of the newly identified risk loci or variants in high LD (r2 > 0.8) with them are discussed in Supplementary Materials. These loci are either quantitative trait loci that regulate the expression of genes within the regions, change sequence motifs for protein binding sites, or are located within DNAase hypersensitivity regions and within regions with enhancer or promoter motifs (Supplementary discussion).
rs72815193 is an intergenic SNP on chromosome 10 and is located near several genes including XRCC6P1, MYOF, CYP26A1, CYP26C1, and EXOC6. MYOF encodes a calcium/phospholipid-binding protein that plays a role in membrane repair of endothelial cells damaged by mechanical stress (http://www.genecards.org/cgi-bin/carddisp.pl?gene=MYOF). CYP26A1 and CYP26C1 are involved in regulation of cellular retinoic acid metabolism, eye development, and maturation of vision function by their effect on retina and retinal ganglion cells during the later stages of eye development26,27,28. Interestingly, microdeletion of approximately 363 kb within this region of chromosome 10, which included CYP26A1, CYP26C1, and EXOC6, was reported in three patients affected by non-syndromic bilateral and unilateral optic nerve aplasia in a Belgian pedigree29. Moreover, this locus is also a risk locus for refractive error, where rs10882165 (P = 1 × 10−11 for refractive error) is in high LD with rs72815193 (r2 = 0.84), the top risk SNP within this locus17. XRCC6P1 is a pseudogene with limited data available on function of this gene or its relevance to diseases.
rs56962872 on chromosome 3 is an intronic variant within the LOC253573 (LINC02052) gene, near CRYGS and TBCCD1. LINC02052 is highly expressed in retina and vitreous humor CRYGS is a member of the crystallin gene families, which are expressed in human lens, retina, and cornea30. Mutations in CRYGS are associated with autosomal dominant paediatric cortical cataract in humans31. TBCCD1 is a centrosomal protein that plays a role in the regulation of centrosome and Golgi apparatus positioning, with consequences on cell shape and cell migration32. Interestingly, human XRP2 is a TBCC-domain containing protein mutated in certain forms of retinitis pigmentosa, a retinal degenerative disease33,34. Thus, TBCC-domain containing proteins including TBCCD1 may play a role in OAG through their effect on retinal formation or mechanisms such as cell shape and function.
rs6478746 on chromosome 9 is located near LMX1B and LOC105376277. LMX1B is mutated in Nail-Patella Syndrome, characterized by nail, patella and elbow dysplasia, in which some patients develop OAG35. In support of this, a mouse model study showed that a dominant-negative mutation of Lmx1b causes glaucoma36. This gene is required for murine trabecular meshwork formation and thus has an important role in controlling IOP37, suggesting that this gene may influence OAG through the mechanisms related to increased eye pressure. In support of this, rs6478746 was nominally associated with IOP (P = 0.03), and associated at P = 6.38 × 10−7 in the combined OAG and IOP analysis in European ancestry.
rs9530458 is an intronic variant within LMO7, a protein-coding gene that may be involved in protein-protein interaction (http://www.genecards.org/cgi-bin/carddisp.pl?gene=LMO7&keywords=LMO7). An engineered 800 kilobase deletion of Uchl3 and Lmo7 caused defects in viability, postnatal growth and degeneration of muscle and retina in mice38. In addition, LMO7 has been suggestively (P = 4 × 10−6 for rs11841001) associated with corneal astigmatism. However, rs9530458 is not in high LD with the top corneal astigmatism SNP (rs11841001, LD r2 = 0.14, P = 0.06 in the OAG and CA analysis in Europeans), suggesting that independent variants within this gene may be involved in the development of OAG and corneal astigmatism.
C9, the gene identified in the gene-based approaches in this study also has interesting implications for OAG. This gene is one component of the complement system, a part of the innate immune response whose deregulation is considered to have a major role in pathogenesis of AMD39. Common and rare variants in multiple complement genes including C9 have been associated with AMD40,41,42, consistent with studies showing significant genetic correlation between AMD and glaucoma43. Moreover, there is some evidence that the complement system including C9 is activated in glaucomatous optic nerve head astrocytes44, suggesting a possible role of C9 in the development of OAG.
This study has several limitations. First, we did not use the recently proposed approaches for meta-analysis of correlated traits using GWAS summary statistics45 which adjust for overlapping or related subjects, population stratification, and heterogeneity of effect between studies. In our study, as confirmed with the LD score regression analyses, we did not have biases such as population stratification and sample overlap between the OAG and endophenotype studies. Thus, we did not use the proposed approaches that adjust for such biases in this study. In addition, approaches such as that proposed by Zhu and colleagues are susceptible to detecting association for a trait that is mainly contributed to via only a subset of the traits. Although our approach has a similar limitation, we investigated the heterogeneity of association between studies to ensure that the results were not biased towards one study. Another limitation of this study is that we performed the combined analysis of OAG and each endophenotype separately, rather than including all the endophenotypes in the same analysis. This was justified due to two reasons: (1) IOP and VCDR loci act through two distinct pathways (intraocular pressure vs optic disc morphology), and (2) the GWAS results for the endophenotypes included in this study were obtained from the same consortia study11, and thus the subjects overlap substantially between these phenotypes.
In conclusion, this study highlighted several novel genes and cellular pathways likely to be involved in the development of OAG. Fine-mapping and functional validation of the new risk loci will help to better understand disease pathophysiology. Identification of additional risk loci using larger sample sizes in the future may lead to more accurate genetic risk prediction algorithms for OAG as well as identification of new molecular targets for prevention and intervention strategies.
Methods
Study design and participants
In total 3,071 OAG cases from the Australian & New Zealand Registry of Advanced Glaucoma (ANZRAG)46, and 6,750 unscreened controls of European descent were included in this study. This dataset involves three phases of OAG data collection, and hence, quality control (QC), imputation, and association analysis were conducted separately for each phase before combining the results in a meta-analysis. The first phase was previously published and comprises 1,155 advanced OAG cases and 1,992 controls genotyped on Illumina Omni1M or OmniExpress arrays (Illumina, San Diego, California, USA)7. The second phase includes a further 579 advanced OAG cases genotyped on Illumina HumanCoreExome array and 946 controls selected from parents of twins previously genotyped on the same array. The third phase comprises 1,337 OAG cases (11 advanced, 741 non-advanced, and 585 cases with visual field data unavailable) genotyped on Illumina HumanCoreExome array and 3,812 controls selected from a study of endometriosis previously genotyped on the same array. The diagnostic criteria have been described previously7. The Approval was obtained from the Human Research Ethics Committees of Southern Adelaide Health Service/Flinders University, University of Tasmania, QIMR Berghofer Medical Research Institute and the Royal Victorian Eye and Ear Hospital. Written informed consent was obtained from all participants. All the methods were carried out in accordance with relevant guidelines and regulations for human subject research, in accordance with the Declaration of Helsinki.
For the endophenotypes we used GWAS results from our previously published data, which includes varying numbers of participants for each trait; between 22,000 and 24,000 Europeans, and between 7,000 and 9,000 Asians11 (Supplementary Table 1). By design, the endophenotype studies did not include any of the OAG cases. We combined the OAG GWAS results with the results obtained from the endophenotype GWASs in Europeans in the primary analysis as well as those obtained from combined European and Asian endophenotype GWASs in a secondary analysis. The QC, imputation, and association testing has been previously described for these studies11 as well as for the first phase of the ANZRAG OAG study7– specifically imputation was done using the 1000 Genomes Phase 1 Europeans reference panel. The following paragraphs provide this information for the second and third phases of the ANZRAG OAG dataset.
Quality Control (QC)
We used the same QC protocol as was used for the first phase of the ANZRAG OAG GWAS. Briefly, we performed QC using PLINK 1.947,48 by removing individuals with more than 3% missing genotypes, and SNPs with call rate less than 97%, minor allele frequency (MAF) < 0.01, and Hardy-Weinberg equilibrium P < 0.0001 in controls and P < 5 × 10−10 in cases. The same QC protocol was used for case and control datasets before merging to avoid mismatches between the merged datasets. We used PLINK1.9 to compute identity by descent based on autosomal markers, with one of each pair of individuals with relatedness of greater than 0.2 removed within each phase of the ANZRAG data as well as between the three phases. PLINK 1.9 was used to compute principal components for all participants and reference samples of known northern European ancestry (1000 Genomes British, CEU, Finland participants). Participants with PC1 or PC2 values > 6 standard deviations from the mean of known northern European ancestry group were excluded.
Imputation
Phasing of the genotyped SNPs was conducted using ShapeIT49 and imputation was performed using Minimac3 through the Michigan Imputation Server50, with the Haplotype Reference Consortium (HRC)51 r1.1 as the reference panel. SNPs with imputation quality (r2) > 0.3 and MAF > 0.01 were carried forward for analysis.
Association testing
We assessed associations between SNPs and OAG status adjusted for sex and the first six principal components under an additive genetic model using the dosage scores obtained from imputation. Association analysis was performed either using SNPTEST v2.552,53 or PLINK 1.9. Genomic inflation factor lambda was calculated to investigate the presence of inflation due to model miss-specification or population stratification. We also performed a sensitivity analysis by excluding the OAG cases in which visual field data was unavailable to ensure that the association results were not driven by including those people in the analysis. Similarly, people with mixed-mechanism glaucoma (277 people with OAG as well as a secondary glaucoma) were excluded in a sensitivity analysis as a further robustness check. Association of the top loci were also investigated in NTG and HTG subsets within the ANZRAG dataset (821 NTG cases, 1,544 HTG cases, and 6,750 controls).
To increase the power of this study to identify new risk loci for OAG, we meta-analysed the OAG GWAS results with those obtained from the endophenotype GWASs. To confirm the validity of the endophenotypes for OAG, we estimated genetic correlation between OAG and the endophenotypes using the cross-trait bivariate LD score regression approach15. This approach estimates genetic correlation between traits from regression of the combined Z scores of each SNP for two traits obtained from GWAS summary statistics on LD scores calculated from a reference panel. LD scores are incorporated in estimation of genetic correlation based on the fact that SNPs with high LD have, on average, higher chi-square statistics for association with a trait as compared with SNPs with low LD. In addition, an intercept close to zero in these analyses indicates that there is not a significant sample overlap between studies. Moreover, we used the univariate LD score regression approach16 to investigate presence of model or structural bias in the OAG and endophenotype GWAS data. An intercept close to 1 in a univariate analysis indicates that there is no model misspecification and other sources of bias such as population stratification and cryptic relatedness16.
Meta-analysis of the ANZRAG OAG results between the three phases was performed in METAL54 using the fixed-effects inverse-variance weighting approach using SNP effect sizes and their standard errors. In addition, the quantitative trait GWAS results were meta-analysed with the ANZRAG OAG GWAS using the P-value approach in METAL. In this approach, Z scores are created for each SNP from P-values and direction of effect for tested alleles, and combined as weighted sum of the individual statistics where the weights are proportional to the square root of the number of individuals examined in each study. Genomic control correction was applied to each GWAS dataset prior to the meta-analysis to ensure that inflation was not driving our results. We also investigated the heterogeneity of Z scores between studies using the approach implemented in METAL. Q-Q and Manhattan plots were created in R. For the purpose of creating these plots, we excluded genome-wide significant SNPs that showed heterogeneity of effect (Cochran’s Q Test P < 0.05) between OAG and the quantitative traits that included Asians. Regional association plots were created using LocusZoom55.
SNPs with P < 1 × 10−7 from the overall meta-analysed results that were previously unreported for OAG, and were at least nominally associated (P < 0.05) with OAG in the combined OAG and the quantitative trait analyses, were taken forward for validation in an independent US dataset, the National Eye Institute Glaucoma Human Genetics Collaboration Heritable Overall Operational Database (NEIGHBORHOOD), containing 3,853 OAG cases and 33,480 controls8. More details on the NEIGHBORHOOD study has been provided in the Supplementary Notes.
Gene-based tests
Gene-based tests were conducted using the approaches implemented in MetaXcan19 fastBAT20, and EUGENE21. We used the GWAS results from OAG as well as combined OAG and its endophenotypes for the gene-based tests. MetaXcan is an extension of PredixCan56, a gene-based approach that uses GWAS summary results to impute the genetic component of gene expression in different tissues (thus eliminating the need to directly measure gene expression levels), and correlates the imputed gene expressions with phenotypes of interest. The Bonferroni-corrected threshold for multiple testing was set to 5 × 10−8, considering the maximum number of 7,230 genes tested in 44 tissues for three traits, OAG, IOP, and VCDR (note that VCDR is the ratio of CA to DA, so highly correlated with these traits). The MetaXcan method is developed based on the publically available European reference data; however, this method is quite robust to ethnicity differences19. Thus, we ran the MetaXcan analyses using European ancestry as well as combined Asians and European ancestry data. The combined ethnicity dataset was >80% European.
fastBAT (fast and flexible set-Based Association Test) is a gene-based approach that calculates the association p-values for a set of SNPs (within ±50 Kb of a gene for this study) using GWAS summary data while accounting for LD between SNPs. The Bonferroni-corrected significance threshold was set to 7 × 10−7, considering the maximum number of 24,654 genes tested for three traits. We ran the fastBAT analyses using European ancestry and Asians data separately, and combined P-values using the sum of Z scores method. In addition, we also used the combined Asians and European ancestry meta-analysis results as input for this analysis.
EUGENE is a gene-based approach that captures the aggregate effects of independent eQTL SNPs (both cis-acting and trans-acting) for each gene using GWAS summary statistics. The most suitable tissue for OAG that is available to use with the EUGENE approach is the brain tissue. Considering the maximum number of 5,487 genes tested in brain for three traits, the Bonferroni-corrected threshold was set to 9 × 10−6. Since the current version of EUGENE is developed based on publically available European reference data, we ran the EUGENE analyses using the meta-analysis results from subjects with European ancestry only. However, since the combined ethnicity analyses comprised mainly (at least 80%) Europeans, we also ran these analyses using combined Asian and European ancestry data.
Pathway-based tests
We used the results from the ANZRAG OAG meta-analysis as well as the meta-analysis of ANZRAG OAG and its endophenotypes to do a pathway analysis using the approach implemented in DEPICT22. Although it is preferable to use genome-wide significant loci for DEPICT provided there are at least 10 independent risk loci available for a trait, because we did not have this many independent genome-wide significant loci for each of the meta-analyses we used SNPs with P < 1 × 10−7 for the pathway analyses. Due to the polygenic nature of the studied traits, as well as our relatively low statistical power to detect SNPs with small effect sizes, including more associated SNPs in the analysis may result in improved power to detect associated pathways. Assuming that all the 14,463 pathways used by DEPICT are independent, and considering testing those pathways for three traits, we set the Bonferroni-corrected significance threshold to P < 1 × 10−6 and false discovery rate <0.05.
Gene expression
Ocular tissues of interest (corneal epithelium, corneal stroma, corneal endothelium, trabecular meshwork, pars plicata of the ciliary body, retina, optic nerve head and optic nerve) were collected from donor human eyes within 24 hours post-mortem (mean = 9.7 ± 5.3 hours) and fixed in RNAlater. RNA quality was assessed using Agilent Bioanalyzer 2100 RNA 6000 Nano Assay (Catalog #G2938C, Santa Clara, USA) (mean RNA integrity number = 6.5 ± 1.8) and concentrations were quantified on the Qubit® 2.0 Fluorometer (Catalog #Q32866, Carlsbad, USA) using Qubit™ RNA Assay Kits (Catalog #Q32852, Carlsbad, USA). 250 nanograms of total RNA from each tissue sample was indexed using Bioo Scientific® NEXTflex™ Rapid Directional mRNA-Seq Kit Bundle with RNA-Seq Barcodes and poly(A) beads (Catalog #5138-10, Austin, Texas) and sequenced on the Illumina NextSeq® 500 using High Output v2 Kit (75 cycles) (Catalog #FC-404-2005, San Diego, USA). All raw sequences were quality-control filtered and trimmed with Trimgalore v0.4.0 (http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/), aligned to the human genome (GRCh38 assembly) using TopHat v2.1.157,58 and normalized using the trimmed mean of M-values (TMM) normalisation method59 in Bioconductor R package EdgeR v3.10.260. Gene differential expression was analysed using EdgeR software with Benjamini Hochberg false-positive adjustment61.
In silico functional analyses
Bioinformatics functional analyses were performed for the novel genome-wide significant loci using HaploReg62, RegulomeDB63, ENCODE Project Consortium64, and eQTL-browsers including Blood eQTL-Browser65 and GTEx-Browser24. The top SNP in each locus as well as those with LD r2 > 0.8 with the top SNPs were used for these analyses.
Data availability
The datasets generated during and/or analysed during the current study are not publicly available due to ethical issues.
References
Quigley, H. A. & Broman, A. T. The number of people with glaucoma worldwide in 2010 and 2020. Br J Ophthalmol 90, 262–267, https://doi.org/10.1136/bjo.2005.081224 (2006).
Casson, R. J., Chidlow, G., Wood, J. P., Crowston, J. G. & Goldberg, I. Definition of glaucoma: clinical and experimental concepts. Clin Exp Ophthalmol 40, 341–349, https://doi.org/10.1111/j.1442-9071.2012.02773.x (2012).
Wolfs, R. C. et al. Genetic risk of primary open-angle glaucoma. Population-based familial aggregation study. Arch Ophthalmol 116, 1640–1645 (1998).
Thorleifsson, G. et al. Common variants near CAV1 and CAV2 are associated with primary open-angle glaucoma. Nat Genet 42, 906–909, https://doi.org/10.1038/ng.661 (2010).
Burdon, K. P. et al. Genome-wide association study identifies susceptibility loci for open angle glaucoma at TMCO1 and CDKN2B-AS1. Nat Genet 43, 574–578, https://doi.org/10.1038/ng.824 (2011).
Wiggs, J. L. et al. Common variants at 9p21 and 8q22 are associated with increased susceptibility to optic nerve degeneration in glaucoma. PLoS Genet 8, e1002654, https://doi.org/10.1371/journal.pgen.1002654 (2012).
Gharahkhani, P. et al. Common variants near ABCA1, AFAP1 and GMDS confer risk of primary open-angle glaucoma. Nat Genet 46, 1120–1125, https://doi.org/10.1038/ng.3079 (2014).
Bailey, J. N. et al. Genome-wide association analysis identifies TXNRD2, ATXN2 and FOXC1 as susceptibility loci for primary open-angle glaucoma. Nat Genet 48, 189–194, https://doi.org/10.1038/ng.3482 (2016).
Springelkamp, H. et al. ARHGEF12 influences the risk of glaucoma by increasing intraocular pressure. Hum Mol Genet 24, 2689–2699, https://doi.org/10.1093/hmg/ddv027 (2015).
Ramdas, W. D. et al. Defining glaucomatous optic neuropathy from a continuous measure of optic nerve damage - the optimal cut-off point for risk-factor analysis in population-based epidemiology. Ophthalmic Epidemiol 18, 211–216, https://doi.org/10.3109/09286586.2011.595038 (2011).
Springelkamp, H. et al. New insights into the genetics of primary open-angle glaucoma based on meta-analyses of intraocular pressure and optic disc characteristics. Hum Mol Genet 26, 438–453, https://doi.org/10.1093/hmg/ddw399 (2017).
Charlesworth, J. et al. The path to open-angle glaucoma gene discovery: endophenotypic status of intraocular pressure, cup-to-disc ratio, and central corneal thickness. Invest Ophthalmol Vis Sci 51, 3509–3514, https://doi.org/10.1167/iovs.09-4786 (2010).
van Koolwijk, L. M. et al. Common genetic determinants of intraocular pressure and primary open-angle glaucoma. PLoS Genet 8, e1002611, https://doi.org/10.1371/journal.pgen.1002611 (2012).
Fan, B. J., Wang, D. Y., Pasquale, L. R., Haines, J. L. & Wiggs, J. L. Genetic variants associated with optic nerve vertical cup-to-disc ratio are risk factors for primary open angle glaucoma in a US Caucasian population. Invest Ophthalmol Vis Sci 52, 1788–1792, https://doi.org/10.1167/iovs.10-6339 (2011).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat Genet 47, 1236–1241, https://doi.org/10.1038/ng.3406 (2015).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 47, 291–295, https://doi.org/10.1038/ng.3211 (2015).
Verhoeven, V. J. et al. Genome-wide meta-analyses of multiancestry cohorts identify multiple new susceptibility loci for refractive error and myopia. Nat Genet 45, 314–318, https://doi.org/10.1038/ng.2554 (2013).
Yazar, S. et al. Interrogation of the platelet-derived growth factor receptor alpha locus and corneal astigmatism in Australians of Northern European ancestry: results of a genome-wide association study. Mol Vis 19, 1238–1246 (2013).
Barbeira, A. et al. Integrating tissue specific mechanisms into GWAS summary results. bioRxiv, https://doi.org/10.1101/045260 (2017).
Bakshi, A. et al. Fast set-based association analysis using summary data from GWAS identifies novel gene loci for human complex traits. Sci Rep 6, 32894, https://doi.org/10.1038/srep32894 (2016).
Ferreira, M. A. et al. Gene-based analysis of regulatory variants identifies 4 putative novel asthma risk genes related to nucleotide synthesis and signaling. J Allergy Clin Immunol 139, 1148–1157, https://doi.org/10.1016/j.jaci.2016.07.017 (2017).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat Commun 6, 5890, https://doi.org/10.1038/ncomms6890 (2015).
Tomarev, S. I., Wistow, G., Raymond, V., Dubois, S. & Malyukova, I. Gene expression profile of the human trabecular meshwork: NEIBank sequence tag analysis. Invest Ophthalmol Vis Sci 44, 2588–2596 (2003).
Lonsdale, J. et al. The Genotype-Tissue Expression (GTEx) project. Nat Genet 45, 580–585, https://doi.org/10.1038/ng.2653 (2013).
Jonas, J. B. et al. Predictive factors of the optic nerve head for development or progression of glaucomatous visual field loss. Invest Ophthalmol Vis Sci 45, 2613–2618, https://doi.org/10.1167/iovs.03-1274 (2004).
Ross, A. C. & Zolfaghari, R. Cytochrome P450s in the regulation of cellular retinoic acid metabolism. Annu Rev Nutr 31, 65–87, https://doi.org/10.1146/annurev-nutr-072610-145127 (2011).
Luo, T., Sakai, Y., Wagner, E. & Drager, U. C. Retinoids, eye development, and maturation of visual function. J Neurobiol 66, 677–686, https://doi.org/10.1002/neu.20239 (2006).
Sakai, Y., Luo, T., McCaffery, P., Hamada, H. & Drager, U. C. CYP26A1 and CYP26C1 cooperate in degrading retinoic acid within the equatorial retina during later eye development. Dev Biol 276, 143–157, https://doi.org/10.1016/j.ydbio.2004.08.032 (2004).
Meire, F. et al. Nonsyndromic bilateral and unilateral optic nerve aplasia: first familial occurrence and potential implication of CYP26A1 and CYP26C1 genes. Mol Vis 17, 2072–2079 (2011).
Wistow, G. The human crystallin gene families. Hum Genomics 6, 26, https://doi.org/10.1186/1479-7364-6-26 (2012).
Sun, H. et al. Gamma-S crystallin gene (CRYGS) mutation causes dominant progressive cortical cataract in humans. J Med Genet 42, 706–710, https://doi.org/10.1136/jmg.2004.028274 (2005).
Goncalves, J. et al. TBCCD1, a new centrosomal protein, is required for centrosome and Golgi apparatus positioning. EMBO Rep 11, 194–200, https://doi.org/10.1038/embor.2010.5 (2010).
Veltel, S., Gasper, R., Eisenacher, E. & Wittinghofer, A. The retinitis pigmentosa 2 gene product is a GTPase-activating protein for Arf-like 3. Nat Struct Mol Biol 15, 373–380, https://doi.org/10.1038/nsmb.1396 (2008).
Schwarz, N., Novoselova, T. V., Wait, R., Hardcastle, A. J. & Cheetham, M. E. The X-linked retinitis pigmentosa protein RP2 facilitates G protein traffic. Hum Mol Genet 21, 863–873, https://doi.org/10.1093/hmg/ddr520 (2012).
Mimiwati, Z. et al. Nail-patella syndrome and its association with glaucoma: a review of eight families. Br J Ophthalmol 90, 1505–1509, https://doi.org/10.1136/bjo.2006.092619 (2006).
Cross, S. H. et al. A dominant-negative mutation of mouse Lmx1b causes glaucoma and is semi-lethal via LDB1-mediated dimerization [corrected]. PLoS Genet 10, e1004359, https://doi.org/10.1371/journal.pgen.1004359 (2014).
Liu, P. & Johnson, R. L. Lmx1b is required for murine trabecular meshwork formation and for maintenance of corneal transparency. Dev Dyn 239, 2161–2171, https://doi.org/10.1002/dvdy.22347 (2010).
Semenova, E., Wang, X., Jablonski, M. M., Levorse, J. & Tilghman, S. M. An engineered 800 kilobase deletion of Uchl3 and Lmo7 on mouse chromosome 14 causes defects in viability, postnatal growth and degeneration of muscle and retina. Hum Mol Genet 12, 1301–1312 (2003).
Cashman, S. M., Desai, A., Ramo, K. & Kumar-Singh, R. Expression of complement component 3 (C3) from an adenovirus leads to pathology in the murine retina. Invest Ophthalmol Vis Sci 52, 3436–3445, https://doi.org/10.1167/iovs.10-6002 (2011).
Klein, R. J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389, https://doi.org/10.1126/science.1109557 (2005).
Seddon, J. M. et al. Rare variants in CFI, C3 and C9 are associated with high risk of advanced age-related macular degeneration. Nat Genet 45, 1366–1370, https://doi.org/10.1038/ng.2741 (2013).
Fritsche, L. G. et al. A large genome-wide association study of age-related macular degeneration highlights contributions of rare and common variants. Nat Genet 48, 134–143, https://doi.org/10.1038/ng.3448 (2016).
Cuellar-Partida, G. et al. Assessment of polygenic effects links primary open-angle glaucoma and age-related macular degeneration. Sci Rep 6, 26885, https://doi.org/10.1038/srep26885 (2016).
Nikolskaya, T. et al. Network analysis of human glaucomatous optic nerve head astrocytes. BMC Med Genomics 2, 24, https://doi.org/10.1186/1755-8794-2-24 (2009).
Zhu, X. et al. Meta-analysis of correlated traits via summary statistics from GWASs with an application in hypertension. American journal of human genetics 96, 21–36, https://doi.org/10.1016/j.ajhg.2014.11.011 (2015).
Souzeau, E. et al. Australian and New Zealand Registry of Advanced Glaucoma: methodology and recruitment. Clin Exp Ophthalmol 40, 569–575, https://doi.org/10.1111/j.1442-9071.2011.02742.x (2012).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics 81, 559–575, https://doi.org/10.1086/519795 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7, https://doi.org/10.1186/s13742-015-0047-8 (2015).
Delaneau, O., Marchini, J. & Zagury, J. F. A linear complexity phasing method for thousands of genomes. Nat Methods 9, 179–181, https://doi.org/10.1038/nmeth.1785 (2011).
Das, S. et al. Next-generation genotype imputation service and methods. Nat Genet 48, 1284–1287, https://doi.org/10.1038/ng.3656 (2016).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet 48, 1279–1283, https://doi.org/10.1038/ng.3643 (2016).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat Rev Genet 11, 499–511, https://doi.org/10.1038/nrg2796 (2010).
Burton, P. et al. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678, https://doi.org/10.1038/nature05911 (2007).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191, https://doi.org/10.1093/bioinformatics/btq340 (2010).
Pruim, R. J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337, https://doi.org/10.1093/bioinformatics/btq419 (2010).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat Genet 47, 1091–1098, https://doi.org/10.1038/ng.3367 (2015).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111, https://doi.org/10.1093/bioinformatics/btp120 (2009).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol 14, R36, https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Bullard, J. H., Purdom, E., Hansen, K. D. & Dudoit, S. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics 11, 94, https://doi.org/10.1186/1471-2105-11-94 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140, https://doi.org/10.1093/bioinformatics/btp616 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society Series B (Methodological) 57, 289–300 (1995).
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res 40, D930–934, https://doi.org/10.1093/nar/gkr917 (2012).
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res 22, 1790–1797, https://doi.org/10.1101/gr.137323.112 (2012).
Dunham, I. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74, https://doi.org/10.1038/nature11247 (2012).
Westra, H.-J. et al. Systematic identification of trans eQTLs as putative drivers of known disease associations. Nat Genet 45, 1238–1243, https://doi.org/10.1038/ng.2756 http://www.nature.com/ng/journal/v45/n10/abs/ng.2756.html#supplementary-information (2013).
Acknowledgements
ANZRAG: Support for recruitment of ANZRAG was provided by the Royal Australian and New Zealand College of Ophthalmology (RANZCO) Eye Foundation. Genotyping was funded by the National Health and Medical Research Council of Australia (#535074 and #1023911). This work was also supported by funding from NHMRC #1031362 awarded to J.E.C., NHMRC #1037838 awarded to A.W.H., NHMRC #1048037 awarded to S.L.G., NHMRC #1009844 awarded to R.J.C. and I.G., NHMRC #1031920 and Alcon Research Institute grant awarded to D.A.M., Allergan Unrestricted grant awarded to A.J.W., and the BrightFocus Foundation and a Ramaciotti Establishment Grant. The authors acknowledge the support of Ms Bronwyn Usher-Ridge in patient recruitment and data collection, and Dr Patrick Danoy and Dr Johanna Hadler for genotyping. Controls for the ANZRAG cohort were drawn from the Australian Cancer Study, the Study of Digestive Health, a study of inflammatory bowel diseases, a study of endometriosis, and QIMR Berghofer twin study. The Australian Cancer Study was supported by the Queensland Cancer Fund and the National Health and Medical Research Council (NHMRC) of Australia (Program no. 199600, awarded to David C. Whiteman, Adele C. Green, Nicholas K. Hayward, Peter G. Parsons, David M. Purdie, and Penelope M. Webb; and program no. 552429 awarded to David C. Whiteman). The Study of Digestive Health was supported by grant number 5 R01 CA 001833 from the US National Cancer Institute (awarded to David C. Whiteman). The Barrett’s and Esophageal Adenocarcinoma Genetic Susceptibility Study (BEAGESS) sponsored the genotyping of oesophageal cancer and Barrett’s oesophagus cases, which were used as unscreened controls in the ANZRAG cohort. BEAGESS was funded by grant R01 CA136725 from the US National Cancer Institute. Genotyping for part of the Australian twin control samples included in the ANZRAG cohort was funded by an NHMRC Medical Genomics Grant. Genotyping for the remainder of twin controls was performed by the National Institutes of Health (NIH) Center for Inherited Research (CIDR) as part of an NIH/National Eye Institute (NEI) grant 1RO1EY018246, and we are grateful to Dr Camilla Day and staff. We acknowledge with appreciation all women who participated in the QIMR Berghofer endometriosis study. We thank Endometriosis Associations for supporting study recruitment. We thank Sullivan Nicolaides and Queensland Medical Laboratory for pro bono collection and delivery of blood samples and other pathology services for assistance with blood collection. The QIMR twin and endometriosis studies were supported by grants from the National Health and Medical Research Council (NHMRC) of Australia (241944, 339462, 389927,389875, 389891, 389892, 389938, 443036, 442915, 442981, 496610, 496739, 552485, 552498, 1049472 and 1050208), the Cooperative Research Centre for Discovery of Genes for Common Human Diseases (CRC), Cerylid Biosciences (Melbourne), and donations from Neville and Shirley Hawkins. We thank Matthew A. Brown, Margaret J. Wright, Megan J. Campbell, Anthony Caracella, Scott Gordon, Dale R Nyholt, Anjali K Henders, B. Haddon, D. Smyth, H. Beeby, O. Zheng, B. Chapman for their input into project management, databases, sample processing, and genotyping. We are grateful to the many research assistants and interviewers for assistance with the studies contributing to the QIMR Berghofer twin collection. NEIGHBORHOOD: The NEIGHBORHOOD data collection and analysis is supported by NIH/NEI R01EY022305 (JL Wiggs) and NIH/NEI P30 EY014104 (JL Wiggs). Support for collection of cases, controls and analysis for individual datasets is as follows. Genotyping services for the NEIGHBOR study were provided by the Center for Inherited Disease Research (CIDR) and were supported by the National Eye Institute through grant HG005259-01 (JL Wiggs). Genotyping for the MEEI dataset and some NHS and HPFS cases (GLAUGEN) was completed at the Broad Institute and supported by GENEVA project grant HG004728 (LR Pasquale) and U01-HG004424 (Broad Institute). Genotype data cleaning and analysis for the GLAUGEN study was supported by U01HG004446 (C Laurie). Collecting and processing samples for the NEIGHBOR dataset was supported by the National Eye Institute through ARRA grants 3R01EY015872-05S1 (JL Wiggs) and 3R01EY019126-02S1 (MA Hauser). Funding for the collection of NEIGHBOR cases and controls was provided by NIH grants: EY015543 (RR Allingham), EY006827 (D Gaasterland); HL73042, HL073389, EY13315 (MA Hauser); CA87969, CA49449, UM1 CA186107, UM1 CA167552, EY009149 (PR Lichter), HG004608 (C McCarty), EY008208 (FA Medeiros), EY015473 (LR Pasquale), EY012118 (M Pericak-Vance), EY015682 (A Realini), EY011671 (JE Richards), EY09580 (JE Richards), EY013178 (JS Schuman), RR015574, EY015872 (JL Wiggs), EY010886 (JL Wiggs), EY009847 (JL Wiggs), EY011008,EY144428 (K Zhang), EY144448 (K Zhang), EY18660 (K Zhang). The collection of Marshfield clinic cases and controls was supported by 1U02HG004608-01, 5U01HG006389-02 and NCATS/NIH grant UL1TR000427. In addition some NHS/HPFS cases and controls and analysis of GWAS data was supported by R01 CA131332. The WGHS is supported by HL043851 and HL080467 from the National Heart, Lung, and Blood Institute and CA047988 from the National Cancer Institute, the Donald W. Reynolds Foundation and the Fondation Leducq, with collaborative scientific support and funding for genotyping provided by Amgen. POAG case identification in WGHS was supported by 3R01 EY15473-5S1 (LR Pasquale). JL Wiggs and LR Pasquale are supported by the Harvard Glaucoma Center for Excellence and an unrestricted grant from Research to Prevent Blindness. Dr. Pasquale is also supported by a Harvard Medical School Distinguished Scholar award.
Author information
Authors and Affiliations
Consortia
Contributions
P.G., K.P.B., S. MacGregor, and J.E.C. were involved in designing the study. A.W.H., L.R.P., G.W.M., N.G.M., G.R.S., D.C.W., J.L.W., D.A.M., P.M. and J.E.C. were involved in participant recruitment, sample collection or genotyping. Analysis was performed by P.G., K.P.B., S.S., E.S., T.Z., O.M.S., D.L., M.H.L., J.N.C.B., L.R.P., J.H.K., J.L.H., J.L.W., and S. MacGregor. Clinician assessments were performed by R.A.M., D.A.M. J.L., M.A., B.R., R.C., S.L.G., I.G., A.W., P.R.H., J.G., J.Gale, M.L., P.M., J.R., M.C., M.W., S.B., A.V., J.Gale, and J.E.C. The initial draft was written by P.G., S. MacGregor, and J.E.C.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A comprehensive list of consortium members appears at the end of the paper
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gharahkhani, P., Burdon, K.P., Cooke Bailey, J.N. et al. Analysis combining correlated glaucoma traits identifies five new risk loci for open-angle glaucoma. Sci Rep 8, 3124 (2018). https://doi.org/10.1038/s41598-018-20435-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20435-9
This article is cited by
-
Genetic variants associated with glaucomatous visual field loss in primary open-angle glaucoma
Scientific Reports (2022)
-
MLIP genotype as a predictor of pharmacological response in primary open-angle glaucoma and ocular hypertension
Scientific Reports (2021)
-
MicroRNAs in the aqueous humor of patients with different types of glaucoma
Graefe's Archive for Clinical and Experimental Ophthalmology (2021)
-
A Comparison of Genomic Advances in Exfoliation Syndrome and Primary Open-Angle Glaucoma
Current Ophthalmology Reports (2021)
-
Genome-wide meta-analysis identifies 127 open-angle glaucoma loci with consistent effect across ancestries
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.