Abstract
Molecular studies sometimes reveal evolutionary divergence within accepted species. Such findings can initiate taxonomic revision, as exemplified in the formerly recognized species Ciona intestinalis. While an increasing number of studies have examined the ecology, reproductive barriers and genetics of C. intestinalis and C. robusta, there are still much uncertainties regarding other species of this genus. Using experimental crosses and mitochondrial data, we investigated the evolutionary relationships among four native and introduced Ciona spp., found in sympatry in the Mediterranean Sea or English Channel. Outcome of 62 bi-parental reciprocal crosses between C. intestinalis, C. robusta, C. roulei and C. edwardsi showed that C. edwardsi is reproductively isolated from the other taxa, which is in agreement with its distinct location in the phylogenetic tree. Conversely, hybrids are easily obtained in both direction when crossing C. intestinalis and C. roulei, reinforcing the hypothesis of two genetically differentiated lineages but likely being from a same species. Altogether, this study sheds light on the evolutionary relationship in this complex genus. It also calls for further investigation notably based on genome-wide investigation to better describe the evolutionary history within the genus Ciona, a challenging task in a changing world where biological introductions are shuffling species distribution.
Similar content being viewed by others
Introduction
Species can be defined as a group of lineages evolving separately from all others lineages, beginning with a speciation event and ending with its extinction or new speciation events1,2. Several species concept have been proposed, each of them relying on specific criteria such as reproductive isolation, biogeographic patterns, molecular divergence or phenotypes (including morphology)1. These various concepts reflect that evolutionary and ecological processes impact the divergence among groups of individuals with various effects and speed (e.g. accumulation of genetic changes, niche segregation, increased reproductive isolation). During the period of accumulation of differences, the ‘gray zone’ as coined by De Queiroz1, different groups of individuals undergoing a speciation process may be distinguished by some criteria, for instance deep genetic divergence, but not necessarily with other, for instance they may lack complete reproductive isolation. Speciation is a dynamic process which results from step-by-step accumulation of these differences of various nature. All the species concepts, and associated criteria, mentioned above are thus of interest, and jointly used in integrative taxonomy studies2,3,4,5.
In marine taxa, the joint use of biological species and phylogenetic concepts are particularly important regarding the difficulty to use other, for instance those based on biogeography and morphology. The natural distribution range of many marine species has indeed been strongly modified by human activities through biological introduction processes, leading Carlton6 to define cryptogenic species, i.e. species for which native versus non-native status is undetermined. Biological introductions not only shuffle species distribution and break biogeographic boundaries but also promote hybridization between previously isolated taxa, and thus changes in species evolutionary dynamics, (7; e.g. in Mytilus spp.,8). Molecular tools have revealed numerous cryptic species, i.e. species morphologically indistinguishable, across all biogeographical regions9. In the marine realm, the proportion of undiscovered molecular cryptic species had been estimated to vary from 11 to 43% of species described across 49 infra-ordo, with the highest proportion found in taxa with few diagnostic external morphological characters10. Cryptic taxa are often revealed by population genetic studies: unexpected genetic divergence and low gene flow within accepted species have often been a first step towards an investigation of the species status4. Whenever possible, taxonomic revision also benefits from experimental crosses testing for reproductive isolation (in support of the biological species concept). The biological species concept, as compared to the other ones, interestingly does not rely on a priori definition of biological units: reproductive isolation observed when using experimental crosses is a sufficient proof that two lineages have evolved separately.
The species complex Ciona intestinalis exemplifies the need to merge diverse approaches to achieve species delineation. In the early 2000’s, several molecular studies11,12,13,14,15 reveal that the formerly accepted species C. intestinalis, first described by Linnaeus16 was made of four cryptic distinct lineages that were named type A, B, C and D by Zhan, et al.13. Further studies focused on C. intestinalis type A and C. intestinalis type B. The two taxa have been introduced in different part of the world, in particular type A, putatively native to Asia, displayed a worldwide distribution17. Genome-wide studies showed a clear genetic distinctiveness between the two taxa, even in their range of sympatry in Europe, with a divergence time about 4 Myr17,18,19,20,21,22,23. Based on these molecular evidences and on morphological features19,20, type B and type A were finally reclassified as C. intestinalis sensu Millar and C. robusta (described by Hoshino and Tokioka24), respectively25. It is noteworthy that despite their clear distinctiveness, the two species lack reproductive isolation under laboratory conditions26,27 and hybridize (without marked introgression) in the wild in their area of sympatry in NE Atlantic26.
Interestingly, two accepted species, Ciona roulei28 and C. edwardsi29, are living in sympatry with the introduced and newly recognized species, C. robusta, in the Mediterranean Sea. Ciona roulei and C. edwardsi are putatively native to this region where they were first described. However, little is known about the relationships (i.e. evolutionary divergence, reproductive isolation) among the three taxa, and with C. intestinalis. For instance, Nydam and Harrison11,18 made detailed investigation of the phylogenetic relationships within the Ciona genus but did not include C. edwardsi in their studies.
Also it is not always straightforward to use previous published studies regarding these four species, because of the unknown date of introduction of C. robusta in the NE Atlantic and Mediterranean Sea17 and the recent taxonomic revision: until 2015, only C. intestinalis was accepted and thus reported in publications. For example, Lambert, et al.30 studied the reproductive isolation between C. edwardsi, C. roulei and “C. intestinalis” (Table 1). However, the latest is more likely to be C. robusta with the new nomenclature, as the individuals were collected in the Mediterranean Sea and along the Californian coasts, where only C. robusta has been reported so far. Based on this assumption, it can be hypothesized that Lambert, et al.30 showed that C. edwardsi is reproductively isolated from C. robusta whereas C. robusta and C. roulei can reproduce with each other, with asymmetric success. This has nevertheless to be confirmed in light of the new nomenclature, and the reproductive isolation between C. roulei and the newly reclassified C. intestinalis has still to be investigated. Reproductive success between taxa is a major criteria for determining if different taxa could potentially be merged or split into different species. This is particularly important to examine for C. roulei and C. intestinalis. C. roulei is an accepted species described for a long time but molecular phylogenetic studies failed to recognize this taxa as a distinct species from C. intestinalis11,18. It is likely that the two species are actually two lineages of the same species. If this is true, absence of reproductive isolation between C. roulei and C. robusta should be observed as for crosses between C. intestinalis and C. robusta with the same asymmetry (Table 1).
Finally, reproductive incompatibilities can also vary with the geographic origin (and the associated eco-evolutionary history). For instance, Caputi, et al.15 failed to produce juveniles from homospecific crosses of C. robusta collected in Plymouth (UK, English Channel) and Naples (Italy, Mediterranean Sea) (Table 1), raising the question of the variability in reproductive compatibility between populations of C. robusta and other species of the same genus. The extent of reproductive isolation between C. robusta collected in the Mediterranean Sea versus NE Atlantic with C. roulei and C. edwardsi remains to be ascertained.
To resolve the uncertainties listed above and clarify the evolutionary relationships between European Ciona spp., we carried out 1) a phylogenetic study using mitochondrial data with specimens collected over the same area; our study provides the first molecular data for C. edwardsi as compared to other Ciona species (native or introduced) found in Europe and 2) crossing experiments to examine in details the extent of reproductive isolation between populations or/and species within the C. intestinalis/C. robusta species complex and other Ciona spp. found in sympatry. We examined success rate at different stages of the life-cycle, focusing on early-stages, which are the most sensitive for marine invertebrates31, using reciprocal bi-parental crosses. We then discussed the relationships between reproductive isolation and mitochondrial genetic divergence.
Results
Mitochondrial Molecular inferences
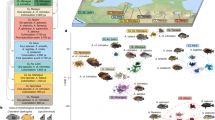
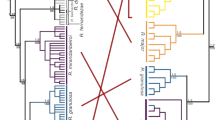
Altogether, we obtained 37 mitochondrial sequences over 717 base pairs, including the 33 individuals used in crossing experiments and the four additional C. edwardsi individuals, which led to the definition of 27 different haplotypes for the four targeted species (see details in Table S2). The six and nine individuals of C. edwardsi and C. roulei, sampled for this study were described by three and seven haplotypes, respectively. For C. roulei, six of the haplotypes found in this study were new as compared to those described by Nydam and Harrison11 and one (CrB4) was found in the two studies. Two and one haplotypes were new for C. robusta and C. intestinalis, respectively, as compared to the haplotypes described in Bouchemousse, et al.17. The phylogenetic tree was finally build-up with 46 unique sequences (Table S1). The phylogenetic tree displayed in Fig. 1 shows four well-supported (99–100%) monophyletic groups made of all sequences associated to (1) C. savignyi (used an outgroup and rooting the tree), (2) C. robusta, without differences between specimens collected in the English Channel or in the Mediterranean Sea, (3) C. edwardsi, and (4) C. roulei and C.intestinalis, the sequences of the two latter mixed with each other in the tree, with one haplotype (Hb2) shared by both C. roulei and C. intestinalis specimens (Fig. 1). It is noteworthy that the relative clustering of C. edwardsi as compared to C. robusta and C. intestinalis is poorly supported by bootstrap values. All the results above were supported using a distance method (Neighbor-joining; 1000 bootstraps) showing the same topology (Fig. S1) and by simple pairwise comparison based on the number of substitutions between sequences (Table S3).
Molecular phylogenetic tree constructed using the Maximum Likelihood method based on the T92 model with 67% of sites evolutionarily invariant. The tree was built with 42 unique sequences obtained (1) from 33 individuals used in this study for experimental crosses and (2) 15 additional haplotype17 (see Material & Methods). The tree is drawn to scale, with branch lengths measured in the number of substitutions per site.
Reproductive success. Early stages (fertilization to hatching)
For the crosses made with C. edwardsi, no fertilization could be observed before two hours. Egg cleavage took more time to be observed in this species as compared to the other ones (not shown). Because of this delay, the first measure of reproductive success (i.e. fertilization success) in C. edwardsi could not be compared with the other types of crosses. Note however that this result does not question the quality of the sperm and eggs of the two individuals used as hatching rates and survival was high (see below).
Excluding C. edwardsi all homospecific crosses worked well, as expected, with more than 80% of fertilization success (Fig. 2A). The geographic origin of C. robusta also did not affect the fertilization success within this species (see ‘CirobuMed x CirobuAtl’ in Fig. 2A).
Mean and standard deviation of fertilization rate (%) obtained with (A) homospecific crosses with different colors per species. (B) Heterospecific crosses (labelled with maternal lineage first and corresponding color). The number of bi-parental crosses per category is indicated above each bar. Abbreviations: CirobuMed, C. robusta from Mediterranean Sea; CirobuAtl, C. robusta from English Channel; Ciinte, C. intestinalis; Ciroul, C. roulei.
Interestingly, many of the heterospecific crosses also showed very good fertilization success. In particular, crosses between C. intestinalis and C. roulei led on average to 78% of fertilization success, as compared to 85% of success for each of homospecific crosses for these two species (Fig. 2B). The same holds when using oocytes of one of these two species with sperm of C. robusta (on average, 88.4% of fertilization success). The reciprocal crosses (i.e. C. robusta used as a mother) were however not successful (7% on average with either C. intestinalis or C. roulei). The geographical origin seems however to play a role (Fig. 2B): the mean fertilization rate of oocytes of C. robusta collected in Mediterranean Sea by sperm of C. intestinalis was 15% (n = 5 crosses, and one cross reached 50%), whereas fertilization success was null when fertilizing oocytes of C. robusta collected in the English Channel with sperm of C. intestinalis. Such an effect was not so clearly observed with the sperm of C. roulei although a slight reverse asymmetry was observed (3.2% and 1.0% for fertilization of eggs of C. robusta collected in the English Channel and the Mediterranean Sea, respectively).
Hatching rates were similar for all homospecific crosses, including for C. edwardsi (Fig. 3A), with variations mostly observed within categories (i.e. putative parental effects). Heterospecific crosses showed contrasted results across categories (Fig. 3B): very high hatching rates (>75%) were observed for crosses made with eggs of either C. intestinalis and C. roulei with sperm of C. robusta as well as between C. intestinalis and C. roulei whatever the direction of the crosses. Conversely, poor hatching rates were observed in crosses involving eggs of C. robusta (16.6% ± 15.1% with C. intestinalis and 5% ± 5.7% with C. roulei). And no hatching was observed when C. edwardsi was involved except for one cross made with oocytes of C. edwardsi and sperm of C. intestinalis but with low success (8.5% of hatching).
Mean and standard deviation of hatching rate (%) obtained with (A) homospecific crosses. (B) Heterospecific crosses (labelled with maternal lineage first). The number of bi-parental crosses per category is indicated above each bar. Abbreviations: Cirobu, C. robusta Ciinte, C. intestinalis; Ciroul, C. roulei; Ciedwa, C. edwardsi.
Survival and growth rate
For subsequent development, the geographic origin of C. robusta was not distinguished as results were identical (data not shown). Survival was first examined at D + 10 by comparing the number of juveniles with the number of post-larvae settled at D + 3. At D + 10, the three series were composed of 12, 10 and 17 crosses respectively, with variable categories of crosses within each. A non-parametric Kruskal-Walis test showed significant differences among series (p = 0.024) that may be due to change in food (see Material and Methods). Results are thus presented in Fig. 4 distinguishing both crosses and series. Several crosses were lost (i.e. all post-settled larvae died) between D + 3 and D + 10 in categories involving C. robusta as a mother and C. roulei as a father (Fig. 4A). For the other crosses, survival rates was very high for homospecific crosses involving C. intestinalis or C. robusta (90% on average; Fig. 4.A), whatever the series. For C. roulei one showed high mortality (15% of survival rate) but the three others (from two series) displayed high survival rate (86%). The same hold with heterospecific crosses which showed high survival rates at D + 10, often at level similar to homospecific crosses (Fig. 4A). At the end of the experiment, at D + 28, none of the crosses kept after D + 10 were lost (Fig. 4B) but survival rates were variable across crosses, the lowest values (ca. 30%) being observed for F1-C. intestinalis and hybrids made with eggs of C. intestinalis (Ciinte x Ciroul, Ciinte x Cirobu in Fig. 4B).
Survival rate per bi-parental crosses at a) D + 10 with survival estimated by comparison with the number of larvae settled at D + 3 and b) D + 28 with survival compared with the number of juveniles alive at D + 10. The biparental crosses are labelled with maternal lineage first; numbers in bracket indicate the number of crosses with live organisms as compared to the total number of crosses surveyed per category over the period. Abbreviations: Ciinte, C. intestinalis; Ciroul, C. roulei; Cirobu, C. robusta; Ciedwa, C. edwardsi.
Growth rates, shown in Fig. 5, were calculated at the end of the experiment (D + 28) using all juveniles alive in each category. Each series was however distinguished for the reasons explained above. An ANOVA confirmed that series as well as the interaction between series and categories have significant effects (p < 0.001 for both). For each of the three series, juvenile’s growth rates were significantly different among categories (Kruskal-Wallis, p-value < 10−5 each) but with variable ranking among categories according to series, suggesting either parental effects or/and food influence (series effect). Nevertheless, interestingly, all series show that the growth rates of either F1-C. intestinalis or F1-C. roulei were not significantly different from the growth rates of the F1-hybrids between these two species. In series 2, these hybrids even show the highest growth rate.
Boxplot showing the first quartile, the median, the third quartile, lower and upper values (coef = 1.5) and extreme values (dots) of growth rate (µm/day) of 28 day-old juveniles (number of juveniles measured is indicated at the top). Results are displayed separately for the three series (see Material and Methods). Within each series, letter on the top of each boxplot indicates different groups distinguished at 5% level by the Ryan-Einot and Gabriel-Welsch test. The biparental crosses are labelled with maternal lineage first. Abbreviations: Ciinte, C. intestinalis; Ciroul, C. roulei; Cirobu, C. robusta.
Discussion
This study aimed to investigate the evolutionary relationships between four species of the genus Ciona living in sympatry in NE Atlantic, namely C. roulei, C. intestinalis, C. robusta and C. edwardsi, by using and comparing the outcome of a mitochondrial phylogenetic tree and the extent of reproductive isolation among these taxa.
Experimental reciprocal bi-parental crosses and subsequent monitoring of survival and growth during 28 days first showed that homospecific crosses developed well, as expected, including when different geographic origins were used. For instance, individuals from two spatially disjoint populations of C. robusta have been used in crosses: some were sampled in the English Channel and some in the Mediterranean Sea. We did not see any evidence in support of variations in gamete compatibility between populations, conversely to what has been observed by Caputi, et al.15 who used individuals from Plymouth (English Channel) and Naples (Mediterranean Sea). The low success of crosses made with individuals from Plymouth in the study by Caputi, et al.15 is more likely due to stressed individuals (long-distance transport or poor acclimatization to local conditions) as already discussed by Sato, et al.27.
The first main outcome of our experimental study is that heterospecific crosses often showed results similar to homospecific crosses, except when involving C. edwardsi. Success was particularly high when considering crosses between C. intestinalis and C. roulei. Our results show that the two taxa behave like two non-reproductively isolated lineages: they can hybridize with high rate of success in both directions and display survival and growth rates similar to those observed in homospecific crosses of the two taxa. Concerning growth rates, no significant statistical differences were observed with the non-parametric test used. Regarding the low power of such tests, this is noteworthy that F1-C. roulei showed a higher growth rate than F1-C. intestinalis. This may be due to our raising condition (e.g. local filtered seawater) which could be more favorable to the local Mediterranean specimens (C. roulei) as compared to the Atlantic ones (C. intestinalis). Similar experiments should be repeated on a much larger number of bi-parental crosses in other places and using different seawaters.
The two taxa also behave similarly regarding the other tested species, in particular C. robusta. Both C. roulei and C. intestinalis hybridize well with C. robusta but in one direction only (oocytes of C. roulei and C. intestinalis could be fertilized by sperm from C. robusta). Such an asymmetry has been previously described for C. intestinalis by Bouchemousse, et al.26 and for C. roulei by Lambert et al.30 (note that in the later study, C. robusta was named C. intestinalis, following the accepted taxonomy and nomenclature at this time). Bouchemousse, et al.26 showed that despite (1) high rate of hybridization between C. intestinalis and C. robusta in laboratory conditions of and (2) a similar reproductive period in syntopic populations, hybridization between the two taxa is extremely rare, and introgression seems to be absent, in the wild. Extrinsic post-zygotic barriers (i.e. environmental barriers) do not seem neither to be effective against gene flow between the two species in their sympatry area. Hybrids are indeed performing as well as their parental species under various temperature and salinity conditions53. It is thus likely that intrinsic barriers, notably due to genomic incompatibilities in second generation hybrids (i.e. Dobzhansky-Muller incompatibilities (DMI)35,36,37), may be at play. Reproductive isolation and DMIs have often been shown to be asymmetrical in plants and animals56, in relation with differential selection on male and female functions. The genes involved in such asymmetrical and putative genomic incompatibilities between C. robusta and C. intestinalis are unknown. However, it is noteworthy that a self-sterility mechanism has been described in C. robusta by Harada, et al.54. These authors have identified two loci, Themis-A and Themis-B, interacting to reject self-fertilization. The Themis-B encodes for two genes, one expressed in eggs (v-ThemisB) and the other in sperm (s-ThemisB). A similar system could be involved in homospecific recognition between the vitelline coat of C. robusta oocytes and sperm, which do not recognize sperm of another species. Yamada et al.55 identified 800 proteins in the vitelline coat (VC) of C. robusta, including v-ThemisA and B. Further researches are needed to investigate the mechanisms of the genetic incompatibilities between C. robusta and C. intestinalis. A first step could be to compare VC proteins of C. robusta with VC proteins of C. intestinalis. In addition, taking into account our experimental results, it would be particularly interesting to test for the existence of first-generation hybrids between C. robusta and C. roulei in their sympatry area, the Western Mediterranean Sea. If results are similar to those observed between C. robusta and C. roulei (i.e. absence of introgression in the wild), it would be an ideal situation to look for conservation of the same DMIs across spatial replicates.
Altogether, in light of the biological species concept, the results of experimental crosses clearly support the hypothesis that C. roulei and C. intestinalis are actually the same species. The taxonomic hypothesis is strongly supported by the results of the phylogenetic analysis, and thus by a phylogenetic criteria. The mitochondrial tree indeed showed that individuals of the two species are mixed with each other. Lahille28 distinguished C. roulei from C. intestinalis with morphological criteria such as a longer atrial siphon than the oral one (the reverse being described for C. intestinalis), a different structure of the branchial wall and a reddish color due to specific pigments in C. roulei. Surprisingly, Brunetti et al.19 showed similar branchial structure between C. intestinalis and C. robusta. While we did not examine this latter characteristic on our specimens, we measured the relative length of the siphons but did not observe differences between C. roulei and C. intestinalis. However, consistent differences in body color were noticed between the C. intestinalis and C. roulei specimens, as described by Lahille28 (Fig. S2). By contrast, we observed a similar pigmentation of the gonoducts openings (orange pigmentation of the oviduct and absence of pigmentation of the spermiduct), a trait that serves to discriminate C. intestinalis from C. robusta (absence of pigmentation of the oviduct and dark red pigmentation of the spermiduct end)22. However, color is known to vary across specimens, notably according to the environment, in several species of Ciona19,22. The contrasted color we observed could thus be due to different regions of sampling for the two taxa (English Channel and Mediterranean Sea for C. intestinalis and C. roulei, respectively). Altogether based on Lahille28, Brunetti et al.19 and our study, further investigations by taxonomists are required to determine the robustness of the morphological criteria distinguishing the two taxa. Our phylogenetic tree is also in agreement with results obtained in previous studies that examined at once C. intestinalis and C. roulei. In particular, Nydam and Harrison11,18 and Zhan et al.13 clearly showed that C. roulei and C. intestinalis do not display reciprocal monophylly, a criteria required to be fitted in support of the phylogenetic species concept for species delineation. Our study confirmed these previous findings, expanding the origin of the individuals studied (including sequences from Bouchemousse, et al.17). Some marine species described as inhabiting both the Mediterranean Sea and the NE Atlantic were shown to be actually composed of cryptic species or cryptic lineages, such as Dicentrarchus labrax also distinct between Atlantic and Mediterranean Sea, when each group are homogenous38. However, the reverse situation, similar to the one described with C. roulei and C. intestinalis in this study, has also been reported. For instance, the species status of Pecten maximus and P. jacobaeus (L. 1758) is unclear: they have been recognized as two distinct species diverged 5My ago, but recent genetic studies showed low differentiation between the two species based on microsatellites39 confirming previous findings based on allozymes40 and mitochondrial data41. Populations of P. maximus show more differentiation along the Atlantic coast than with P. jacobeus in the Western Mediterranean Sea. The Almeria-Oran front has most often been shown to be associated with a strong genetic barrier between populations from Atlantic Ocean and Mediterranean Sea42,43 but there are exceptions. Patarnello, et al.42 hypothesize that populations with no genetic differentiation were separated during the late Pleistocene recolonization of the Mediterranean Sea. To better assess the historical processes that led to the faint genetic distinctiveness of C. roulei and C. intestinalis, a population-based study is needed. C. roulei has, so far, been described and sampled along the Western Mediterranean coast from southern France to northern Spain but the species distribution range is unknown. We thus first need to complement the survey (and sampling) along the Mediterranean coasts down to the Oran-Almeria front and along the NE Atlantic coasts on the other side of this front. Then we need to use nuclear markers to make fine-assessment of the demographic history of the two lineages, using coalescence-based methods to test for alternative divergence scenarios between them. Demographic inferences (e.g. estimate of the time of divergence) could help to understand why populations are now geographically isolated and morphologically distinct but not genetically and reproductively isolated. The two taxa could be at the onset of a speciation process, i.e. in the grey zone as defined by De Queiroz1. Roux, et al.44 shows that pairs of lineages/taxa are in the gray zone when their divergence is between 0.5–2%, which is congruent with our result (based on mitochondrial data) of a divergence of 0.84% between C. roulei and C. intestinalis.
Our study also brought new results regarding C. edwardsi. We showed that C. edwardsi is strongly divergent from C. intestinalis and C. robusta with on average 13.4% and 10.2% of mitochondrial divergence, respectively. Based on these molecular data, and assuming a relationship between evolutionary divergence and reproductive isolation44,45, C. edwardsi is expected to be either partially reproductively isolated with C. robusta, such as observed between C. robusta and C. intestinalis, or completely isolated with C. robusta, such as observed between C. robusta and C. savignyi46. Although with a limited number of individuals used (i.e. sperm from one individual and eggs from another one) because of sampling limitations, our experimental crosses clearly showed a complete reproductive isolation between C. edwardsi and C. intestinalis, C. roulei, C. robusta. A few larvae were produced with one cross made with C. intestinalis but it might be the result of enforced fertilization in petri dishes, as observed sometimes with crosses between eggs of C. robusta with sperm of C. intestinalis26. In addition, juveniles produced with this cross poorly survived after 10 days post-fertilization (16% of survival rate). To explore the evolutionary relationships between those four species, further studies are nevertheless required, notably to resolve the phylogenetic position of C. edwardsi as compared to C. robusta and C. intestinalis. A phylogenomic approach could be particularly interesting to carry out as they have been shown efficient to better understand relationship in species complex and in taxonomic groups with complex histories of migration and divergence, such as flowering plants (Pedicularis), American oak (Quercus) and deep-sea octocorals (Chrysogorgia, Anthomastus-Corallium)47,48,49,50. Phylogenies are indeed often complex to resolve when species hybridize along their evolutionary history, which is likely to have occurred among the Ciona spp. occurring in sympatry in European waters. Past introgression can indeed explain discordance between nuclear or mitochondrial gene trees as shown on marine vs freshwater fish phylogenies51; such gene flow can be maintained between highly divergent species as already shown for C. robusta and C. intestinalis23.
Our results also call for further population genetics investigation of the study species: two haplotypes for C. robusta and one for C. intestinalis were new as compared to those published by Bouchemousse, et al.17. The haplotypes associated to adult specimens of C. robusta collected in the Mediterranean Sea were not distinguished from the haplotypes of specimens sampled in the English Channel, confirming previous findings by Bouchemousse, et al.17 of a likely common origin of the populations introduced in the English Channel and along the Western Mediterranean coasts of France. As for C. intestinalis, we observed a high level of polymorphism for C. roulei and C. edwardsi: with only nine and six individuals respectively, no less than five and three haplotypes were found over ca. 700 base pairs, suggesting large effective size of the populations. The challenge here will be to identify the habitats where C. edwardsi can be sampled as unfortunately little is known about the ecology of this species, although it may be an interesting outgroup for further phylogenetic and genomic studies of the two model species C. intestinalis and C. robusta.
Conclusion
Despite a complex speciation history (divergence with gene flow, e.g. C. robusta and C. intestinalis) and background noise due to biological introductions (e.g. C. robusta introduced in the respective and disjoint native ranges of C. intestinalis and C. roulei), our study shows that coupling experimental studies and molecular analyses can help elucidating evolutionary relationships. Our results also give support to the classical assumption of a positive relationship between time of divergence and reproductive isolation: prezygotic barriers are indeed expected to increase with parental divergence even if the degree of divergence needed to the reproductive isolation varies along taxa45. In addition, this study confirmed our hypothesis regarding C. roulei and C. intestinalis: based on both biological and phylogenetic species concepts and criteria, these two accepted species are more likely to be two isolated populations, genetically poorly distinct, of a same species. At the opposite side of the population-species continuum, C. edwardsi is clearly a distinct species showing reciprocal monophylly and reproductive isolation with the three other study taxa. To our knowledge, this study was the first study attempting to resolve the phylogenetic relationship of C. edwardsi with species presumably living in sympatry (or parapatry). Other markers are however required to resolve in details the evolutionary history within this group of taxa, as the use of a single mitochondrial marker, besides some potential limitations due to its nature (maternally inherited, non-recombinant), does not allow fine investigation of historical gene flow and demography. Altogether, our results call for further integrative taxonomy studies and raise questions regarding the extent and barriers of inter-specific gene flow between these species living in sympatry in the wild.
Material and Methods
Adult sampling
Details about the samples used are provided in Table S1. Briefly, all adult specimens used in this study were collected by scuba diving in the western Mediterranean Sea (in Banyuls-sur-Mer and Thau Lagoon), except for C. intestinalis sampled in the English Channel and sent alive to the Banyuls Observatory where the experiments were made. Sampling C. roulei and C. intestinalis in different regions where the two taxa are in allopatry help to ensure their identification, in addition to morphological criteria (see discussion and Fig. S2). Specimens of C. robusta were also collected from the English Channel for sake of comparison between the two European regions (NE Atlantic and Mediterranean Sea) where this species is known to have been introduced. Only a small number of C. roulei and C. edwardsi specimens could be sampled (nine and six, respectively) despite repeated surveys, suggesting that the two species are either rare or in unknown (and/or difficult to access to) habitats. C. intestinalis and C. robusta were easily distinguished as described by Brunetti et al.19 C. roulei and C. edwardsi individuals were identified using morphological criteria following Lahille28 and Copello52 respectively. The color of the tunic (orange-reddish for C. roulei and bright yellow for C. edwardsi) clearly distinguished the two species. After the use of the adults to produce offspring, a piece of tissue was preserved for further DNA studies (see below).
Fertilization and juveniles raising
After sampling, individuals were kept in closed tanks with constant light to induce gametogenesis. Not all individuals reached maturity, for instance only two C. edwardsi specimens out of 6 collected produced enough gametes to be used in bi-parental crosses. A total of 33 individuals were used to make 62 bi-parental homo- (N = 22) and heterospecific (N = 40) crosses using the four species C. roulei, C. intestinalis, C. robusta and C. edwardsi. For logistic constraints, the 62 crosses were split across three series carried out between September 10 and September 25. Table S2 provides details about the number of crosses that were carried out per series and for each categories of crosses.
Experiments were made following protocols provided in Malfant, et al.53. Briefly, the number of oocytes of one individual was estimated in three drops of 150 µl. Oocytes were then split in equal number into four petri dishes to be fertilized by the sperm of four individuals: one from the same species, to be used as a control, and three belonging to each of the three other tested species, whenever possible. Fertilization was made in a controlled room at 15 °C, with 500 µl of sperm diluted in seawater (5 µl of dry sperm in 2 ml seawater). One hour after fertilization, the fertilization rate was estimated for each cross by counting cleaved eggs. Hatching rates were estimated 24 hours after fertilization by counting larvae. For each bi-parental cross, the larvae were then randomly distributed in petri dishes stored afterwards in a controlled room at 17 °C. Following larval metamorphosis, three days after fertilization, settlement rate was measured. The experiment was pursued for plates with more than 10 settlers at D + 3. Survival rate of juveniles was measured 10 days (D + 10) and 28 days (D + 28) after fertilization. For logistic constraints not all crosses were kept after D + 10 to be surveyed until D + 28 (see results). The experiment was stopped after 28 days with pictures of the individuals taken with a dissecting microscope, for subsequent size analysis. After the opening of the siphons and until the end of the experiment, the juveniles were fed every day with microalgae provided ad libidum. Because of technical problems with production of microalgae, food had to be modified over the course of the experiment. The first series of crosses were fed during 10 days with T-Isochrisis galbana and Tetraselmis sueccisa in proportion 50:50 (produced by local facilities) and then with a mixture of microalgae from Greensea (mainly composed of Nannocloropsis gaditana). The second serie of crosses were fed during 4 days with the microalgae produced locally and then with the Greensea product. The third series of crosses were fed exclusively with Greensea product. Changing food in the course of the experiment obviously did not influence fertilization, settlement and hatching rates but could have had an influence on survival and growth rates measured. Survival and growth rates were thus examined separately for the three series, each included different categories allowing for comparisons (Table S2). Using pictures taken at the end of the experiment (D + 28), the size of each juvenile was measured with Image J and transformed into growth rate (µm/day). Assumptions required for ANOVA not being fulfilled, non-parametric tests were carried out using R32 for examining growth rates between F1 categories. Post hoc tests were done using the Steel-Dwass-Critchlow-Fligner test procedure implemented in XLSTAT (Version 2015.4.01, Addinsoft).
Mitochondrial molecular data analysis
All adults (N = 33) used in this experiment were sequenced on the cytochrome c oxidase subunit I (COI) mitochondrial gene. The COI fragment was amplified using primer sequences and protocols obtained from Nydam and Harrison11. We also sequenced four additional individuals of C. edwardsi collected at the same time but which never reach sexual maturity. Purification of PCR products and sequencing reactions were performed by Eurofins. All PCR products were sequenced in both directions. Sequences obtained were edited using CodonCode Aligner v.4.0.2 (CodonCode Corporation, MA). In addition, we included in the dataset seven sequences (all different) of C. savignyi that were obtained as part of a side study. This species was used as outgroup of the targeted Ciona spp. Finally, we included six and 11 sequences (all different) obtained by Bouchemousse, et al.17 to be representative of the mitochondrial diversity and known distribution of C. robusta and C. intestinalis, respectively, and four sequences of C. roulei from Nydam and Harrison11. All the individuals and/or sequences used in this study are detailed in Table S1 which also provides Genbank accession numbers.
The sequences were aligned using BioEdit v.7.1.98133 leading to a 717 base pairs final alignment (all sequences were 737 bp except for the sequences of C. roulei from Nydam and Harrison11 shorter by 20 base pairs at the 5’ start of the sequence). A tree was built with MEGA734 with complete deletion of missing data. Goodness of fit with various evolution model was first tested: based on BIC criteria, the best model explaining the data is the Tamura 3-parameter (T92) model with 67% of the sites evolutionarily invariable. Using AIC criteria, the Best model (ΔAIC > 10) is the General Time Reversible (GTR) model with 63% of the sites evolutionarily invariable (I). The two models were used in subsequent analyses. Results were identical; only results based on T92 are shown. A rooted phylogenetic tree was constructed using a maximum likelihood method with heuristic search (Subtree-Prunning-Regrafting method). Initial tree(s) for the heuristic search were obtained by applying the Neighbor-Joining method to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach. To assess the reliability of the inferred tree, a boostrap test was carried out (1000 bootstraps). Note that similar topology was obtained using other tree construction method, in particular simple distance-based method (i.e. Neighbour-joining).
References
De Queiroz, K. Species concepts and species delimitation. SYST BIOL 56, 879–886, https://doi.org/10.1080/10635150701701083 (2007).
Samadi, S. & Barberousse, A. In Handbook of Evolutionary Thinking in the Sciences, (ed T. Heams et al.) Ch. 8, 141–157 (2015).
Kantor, Y. I., Stahlschmidt, P., Aznar-Cormano, L., Bouchet, P. & Puillandre, N. Too familiar to be questioned? Revisiting the Crassispira cerithina species complex (Gastropoda: Conoidea: Pseudomelatomidae). J Mollus Stud 83, 43–55, https://doi.org/10.1093/mollus/eyw036 (2017).
Pante, E. et al. Species are hypotheses: avoid connectivity assessments based on pillars of sand. Mol Ecol 24, 525–544, https://doi.org/10.1111/mec.13048 (2015).
Padial, J. M., Miralles, A., De la Riva, I. & Vences, M. The integrative future of taxonomy. Front Zool 7, 16, https://doi.org/10.1186/1742-9994-7-16 (2010).
Carlton, J. T. Molluscan invasions in marine and estuarine communities. Malacologia 41, 439–454 (1999).
Abbott, R. et al. Hybridization and speciation. J Evolution Biol 26, 229–246, https://doi.org/10.1111/j.1420-9101.2012.02599.x (2013).
Saarman, N. P. & Pogson, G. H. Introgression between invasive and native blue mussels (genus Mytilus) in the central California hybrid zone. Mol Ecol 24, 4723–4738, https://doi.org/10.1111/mec.13340 (2015).
Pfenninger, M. & Schwenk, K. Cryptic animal species are homogeneously distributed among taxa and biogeographical regions. Bmc Evol Biol 7, 121, https://doi.org/10.1186/1471-2148-7-121 (2007).
Appeltans, W. et al. The magnitude of global marine species diversity. Curr Biol 22, 2189–2202, https://doi.org/10.1016/j.cub.2012.09.036 (2012).
Nydam, M. L. & Harrison, R. G. Genealogical relationships within and among shallow-water Ciona species (Ascidiacea). Mar Biol 151, 1839–1847, https://doi.org/10.1007/s00227-007-0617-0 (2007).
Iannelli, F., Pesole, G., Sordino, P. & Gissi, C. Mitogenomics reveals two cryptic species in Ciona intestinalis. Trends Genet 23, 419–422, https://doi.org/10.1016/j.tig.2007.07.001 (2007).
Zhan, A., Macisaac, H. J. & Cristescu, M. E. Invasion genetics of the Ciona intestinalis species complex: from regional endemism to global homogeneity. Mol Ecol 19, 4678–4694, https://doi.org/10.1111/j.1365-294X.2010.04837.x (2010).
Suzuki, M. M., Nishikawa, T. & Bird, A. Genomic approaches reveal unexpected genetic divergence within Ciona intestinalis. J Mol Evol 61, 627–635, https://doi.org/10.1007/s00239-005-0009-3 (2005).
Caputi, L. et al. Cryptic speciation in a model invertebrate chordate. P NATL ACAD Sci Usa 104, 9364–9369, https://doi.org/10.1073/pnas.0610158104 (2007).
Linnaeus, C. Caroli Linnae. Systema naturae per regna tria naturae: secundum classes, ordines, genera, species, cum characteribus, differentiis, synonymis, locis. Editio duodecima. 1. Regnum Animale. 1 & 2 Holmiae, Laurentii Salvii. Holmiae [Stockholm], Laurentii Salvii (1767).
Bouchemousse, S., Bishop, J. D. & Viard, F. Contrasting global genetic patterns in two biologically similar, widespread and invasive Ciona species (Tunicata, Ascidiacea). SCI REP-UK 6, 24875, https://doi.org/10.1038/srep24875 (2016).
Nydam, M. L. & Harrison, R. G. Polymorphism and divergence within the ascidian genus. Ciona. Mol Phylogenet Evol 56, 718–726, https://doi.org/10.1016/j.ympev.2010.03.042 (2010).
Brunetti, R. et al. Morphological evidence that the molecularly determined Ciona intestinalis type A and type B are different species: Ciona robusta and Ciona intestinalis. J Zool Syst Evol Res. https://doi.org/10.1111/jzs.12101 (2015).
Pennati, R. et al. Morphological differences between larvae of the Ciona intestinalis species complex: hints for a valid taxonomic definition of distinct species. Plos One 10, e0122879, https://doi.org/10.1371/journal.pone.0122879 (2015).
Caputi, L., Crocetta, F., Toscano, F., Sordino, P. & Cirino, P. Long-term demographic and reproductive trends in Ciona intestinalis sp. A. Mar Ecol 36, 118–128, https://doi.org/10.1111/maec.12125 (2015).
Sato, A., Satoh, N. & Bishop, J. D. D. Field identification of ‘types’ A and B of the ascidian Ciona intestinalis in a region of sympatry. Mar Biol 159, 1611–1619, https://doi.org/10.1007/s00227-012-1898-5 (2012).
Roux, C., Tsagkogeorga, G., Bierne, N. & Galtier, N. Crossing the species barrier: genomic hotspots of introgression between two highly divergent Ciona intestinalis species. Mol Biol Evol 30, 1574–1587, https://doi.org/10.1093/molbev/mst066 (2013).
Hoshino, Z. I. & Tokioka, T. An unusually robust Ciona from the northeastern coast of Honsyu Island, Japan. (1967).
Gissi, C. et al. An unprecedented taxonomic revision of a model organism: the paradigmatic case of Ciona robusta and Ciona intestinalis. Zool Scr. https://doi.org/10.1111/zsc.12233 (2017).
Bouchemousse, S., Lévêque, L., Dubois, G. & Viard, F. Co-occurrence and reproductive synchrony do not ensure hybridization between an alien tunicate and its interfertile native congener. Evol Ecol 30, 69–87, https://doi.org/10.1007/s10682-015-9788-1 (2016).
Sato, A., Shimeld, S. M. & Bishop, J. D. Symmetrical reproductive compatibility of two species in the Ciona intestinalis (Ascidiacea) species complex, a model for marine genomics and developmental biology. Zool Sci 31, 369–374, https://doi.org/10.2108/zs130249 (2014).
Lahille, F. Recherches sur les Tuniciers Docteur des sciences naturelles (1890).
Roule, L. Recherches sur les ascidies simples des cotes de Provence. 1 - Phallusiadees. Ann. Mus. Hist. natur. Marseille 2, 7–270 (1984).
Lambert, C., Lafargue, F. & Lambert, G. Preliminary note on the genetic isolation of Ciona species (Ascidiacea, Urochordata). Vie milieu 40, 293–295 (1990).
Gosselin, L. A. & Qian, P.-Y. Juvenile mortality in benthic marine invertebrates. Mar Ecol Prog Ser 146, 265–282 (1997).
R: A language and environment for statistical computing. (R Foundation for statistical computing, Vienna, Austria, 2014).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acid S 41, 95–98 (1999).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol 33, 1870–1874, https://doi.org/10.1093/molbev/msw054 (2016).
Dobzhansky, T. Genetics and the origin of species. (New York: Columbia University Press., 1937).
Muller, H. J. Isolating mechanisms, evolution and temperature. Biol. Symp 6, 71–125 (1942).
Orr, A. H. D. Bateson, and the genetics of speciation. Genetics 144, 1331–1335 (1996).
Naciri, M., Lemaire, P., Borsa, P. & Bonhomme, F. Genetic study of theAtlantic/Mediterranean transition in sea bass (Dicentrarchus labrax). The American Genetic Association 90, 591–596 (1999).
Morvezen, R. et al. Genetic structure of a commercially exploited bivalve, the great scallop Pecten maximus, along the European coasts. Conserv Genet 17, 57–67, https://doi.org/10.1007/s10592-015-0760-y (2015).
Rı́os, C., Sanz, S., Saavedra, C. & Peña, J. B. Allozyme variation in populations of scallops, Pecten jacobaeus (L.) and P. maximus (L.) (Bivalvia: Pectinidae), across the Almeria–Oran front. J Exp Mar Biol Ecol 267, 223–244, https://doi.org/10.1016/S0022-0981(01)00371-9 (2002).
Saavedra, C. & Peña, J. B. Nucleotide diversity and Pleistocene population expansion in Atlantic and Mediterranean scallops (Pecten maximus and P. jacobaeus) as revealed by the mitochondrial 16S ribosomal RNA gene. J Exp Mar Biol Ecol 323, 138–150, https://doi.org/10.1016/j.jembe.2005.03.006 (2005).
Patarnello, T., Volckaert, F. A. & Castilho, R. Pillars of Hercules: is the Atlantic-Mediterranean transition a phylogeographical break? Mol Ecol 16, 4426–4444, https://doi.org/10.1111/j.1365-294X.2007.03477.x (2007).
Borsa, P. et al. Infraspecific zoogeography of the Mediterranean: population genetic analysis on sixteen Atlanto-Mediterranean species (fishes and invertebrates). VIE MILIEU (1997).
Roux, C. et al. Shedding light on the grey zone of speciation along a continuum of genomic divergence. PLOS BIOL 14, e2000234, https://doi.org/10.1371/journal.pbio.2000234 (2016).
Edmands, S. Does parental divergence predict reproductive compatibility? Trends Ecol Evol 17, 520–527 (2002).
Byrd, J. & Lambert, C. C. Mechanism of the block to hybridization and selfing between the sympatric ascidians Ciona intestinalis and Ciona savignyi. Mol reprod Dev 55, 109–116, https://doi.org/10.1002/(SICI)1098–2795(200001)55:1<109::AID-MRD15>3.0.CO;2-B (2000).
Eaton, D. A. & Ree, R. H. Inferring phylogeny and introgression using RADseq data: an example from flowering plants (Pedicularis: Orobanchaceae). Syst Biol 0, 1–18, https://doi.org/10.5061/dryad.bn281 (2013).
Hipp, A. L. et al. A framework phylogeny of the American oak clade based on sequenced RAD data. Plos One 9, e93975, https://doi.org/10.1371/journal.pone.0093975 (2014).
Pante, E. et al. Use of RAD sequencing for delimiting species. Heredity (2015).
Herrera, S. & Shank, T. M. RAD sequencing enables unprecedented phylogenetic resolution and objective species delimitation in recalcitrant divergent taxa. Mol Phylogenet Evol. https://doi.org/10.1101/019745 (2015).
Wallis, G. P. et al. Interspecific hybridization causes long-term phylogenetic discordance between nuclear and mitochondrial genomes in freshwater fishes. Mol Ecol 26, 3116–3127, https://doi.org/10.1111/mec.14096 (2017).
Copello, M. Ciona edwardsi (Roule, 1886) espece littorale de Mediterranee distinct de Ciona intestinalis Linne, 1767. Vie Milieu 31, 243–253 (1981).
Malfant, M., Coudret, J., Le Merdy, R. & Viard, F. Effects of temperature and salinity on juveniles of two ascidians, one native and one invasive, and their hybrids. J EXP Mar Biol Ecol 497, 180–187, https://doi.org/10.1016/j.jembe.2017.09.019 (2017).
Harada, Y. & Takagaki, Y. Sunagawa, m., Saito, T. & Yamada, L. Mechanism of self-sterility in a hermaphroditic chordate. Science 320, 548–550 (2008).
Yamada, L., Saito, T., Taniguchi, H., Sawada, H. & Harada, Y. Comprehensive egg coat proteome of the ascidian Ciona intestinalis reveals gamete recognition molecules involved in self-sterility. J Biol Chem 284, 9402–9410, https://doi.org/10.1074/jbc.M809672200 (2009).
Orr, A. H. & Presgraves, D. C. Speciation by postzygotic isolation: forces, genes and molecules. Bio Essays 22, 1085–1094 (2000).
Acknowledgements
We are very grateful to the divers of the Marine Operations Departments of the Station Biologique of Roscoff and of the laboratory Arago in Banyuls-Sur-Mer, for sampling in the field. We are also grateful to Pascal Romans and Michael Fuentes for additional sampling and tank installation and to Nancy Trouillard for algae production at the laboratory Arago in Banyuls-Sur-Mer. The project was supported by the ANR project HySea (no. ANR-12-BSV7-0011) and EMBRC-France. MM acknowledges a PhD grant from the University Pierre et Marie Curie (Sorbonne Universités).
Author information
Authors and Affiliations
Contributions
M.M., SD. and F.V. conceived the experiment design and contributed to sampling collection. M.M. and S.D. ran the experimental crosses, and M.M. the molecular labwork. M.M. and F.V. analyzed the data and prepared tables and figures. M.M., S.D. and F.V. wrote the mansucript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Malfant, M., Darras, S. & Viard, F. Coupling molecular data and experimental crosses sheds light about species delineation: a case study with the genus Ciona. Sci Rep 8, 1480 (2018). https://doi.org/10.1038/s41598-018-19811-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-19811-2
This article is cited by
-
Looks can be deceiving: Didemnum pseudovexillum sp. nov. (Ascidiacea) in European harbours
Marine Biodiversity (2020)
-
Assessing Cryptic Invasion State: Fine-Scale Genetic Analysis of Ciona savignyi Population in Putative Native Habitat of the Korean Coast
Ocean Science Journal (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.