Abstract
Olfactory function is a key sense for human well-being and health, with olfactory dysfunction having been linked to serious diseases. As the microbiome is involved in normal olfactory epithelium development, we explored the relationship between olfactory function (odor threshold, discrimination, identification) and nasal microbiome in 67 healthy volunteers. Twenty-eight subjects were found to have normal olfactory function, 29 had a particularly good sense of smell (“good normosmics”) and 10 were hyposmic. Microbial community composition differed significantly between the three olfactory groups. In particular, butyric acid-producing microorganisms were found to be associated with impaired olfactory function. We describe the first insights of the potential interplay between the olfactory epithelium microbial community and olfactory function, and suggest that the microbiome composition is able to mirror and potentially shape olfactory function by producing strong odor compounds.
Similar content being viewed by others
Introduction
The human olfactory system is able to discriminate a vast number of odors1. This function is mediated by olfactory receptors situated within the olfactory epithelium. The sense of smell shapes our perception of our external environment and is essential in decision-making and in guiding behavior, including eating behavior, and detection of danger, as well as contributing significantly to the hedonic component of everyday life (e.g. flavor and fragrance perception2,3).
Anosmia (complete loss of olfactory function) and hyposmia (decreased olfactory function) together affect approximately 20% of the population3. The risk of olfactory dysfunction increases with age, and the condition may also result from chronic sinonasal diseases, head trauma, upper respiratory infection or neurodegenerative diseases.
Negative effects on mood, enjoyment of food, safety, personal hygiene, social interactions, and sexual life have been reported in individuals with olfactory impairment4,5,6,7 (for review see8). Even mild olfactory impairment can negatively influence interpersonal relationships as shown for patients with traumatic brain injury9. The effect of olfactory dysfunction on bodyweight is unclear, with some studies reporting weight gain10,11,12 and others weight loss13,14. A study in 176 individuals with olfactory dysfunction found that 21% reported weight gain, 11% reported weight loss, and the rest reported no change15. Weight gain was more common in those with a better ability to identify odors and in younger subjects15.
Importantly, olfactory deficits have also been identified as indicators of several neurologic diseases, including multiple sclerosis, epilepsy, and possibly, Alzheimer’s and Parkinson’s disease16,17. They also represent ultra-high risk indicators for psychosis18. Very recently, olfactory dysfunction has been identified as the best predictor for 5-year mortality in older adults19; outperforming such factors as heart failure, stroke, diabetes, hypertension, liver damage and even cancer20.
The human microbiota, consisting of 3.8 × 1013 microorganisms21 mainly associated with the colon (3.8 × 1013 22) and skin (1011 23), fulfils numerous important tasks, ranging from food degradation to playing a key role in host immunity24. These microorganisms are known to interact with the body mostly via their secondary metabolites, which include hormone-like molecules25, short-chain fatty acids - the end products of gut bacterial fermentation - and serotonin precursors26. Dysbiosis (imbalance of the human microbial community), has been associated with a number of diseases, including cancer, metabolic disorders, inflammatory bowel disease, depression and many others. Thus, the microbial community has the capability to reflect health status and functionality, and may have potential as a diagnostic tool27,28.
While potentially important with respect to disorders such as chronic rhinosinusitis29, the nasal microbial community remains largely understudied. However, a recent report indicates that microorganisms contribute to normal development of the olfactory epithelium, as germ-free mice had a thinner olfactory cilia layer and reduced levels of olfactory epithelium cell apoptosis and proliferation30. In a recent study, focusing on the oral and nasal microbiome of Parkinson’s disease patients and a control group, no notable differences in nasal bacterial abundance or diversity were found, but overall olfactory function was associated with the abundance of certain bacterial taxa, including Moraxella and Staphylococcus31.
Based on these preliminary insights, we became interested in the relationship of nasal microbiome and olfactory function. We propose that the bacterial composition of the nasal olfactory epithelium per se can mirror olfactory function. To this end, the present study was implemented to explore correlations between the nasal microbial community composition and olfactory performance measures in healthy individuals. Briefly, we assessed olfactory function (odor threshold, discrimination, identification) in 67 healthy volunteers using the Sniffin’ Sticks test battery (Burghart Instruments, Wedel, Germany) and explored the relationship between their olfactory performance and nasal microbiome. As additional parameters, we as well included sex and BMI as potential factors for olfactory function and microbiome composition.
Results
Study cohort parameters
Overall, our cohort comprised 50 women and 17 men (mean weight 66.7 kg [SD = 13.4], and mean BMI 22.7 [SD = 3.8]): 53 were normal weight (BMI 18.5–25), 10 overweight (BMI > 25), and 4 underweight (BMI < 18.5). As shown in Supplementary Table 1, 57 were found to be normosmic and 10 hyposmic. Based on total (odor threshold: T, discrimination: D, identification: I) TDI scores, 29 of the normosmic subjects were categorized as “good normosmics” (see Fig. 1 for the olfactory scores and their distribution for each subgroup). We also classified subjects according to subscores for T, D, and I; see Supplementary Table 1). Across the whole dataset, there was no correlation (Spearman’s rho) between olfactory scores and age or BMI (Table 1, see Supplementary Figures 1 and 2 for distribution of BMI and age for each subgroup). In average normosmics, there was a marginal negative correlation between total TDI scores and age (rs = −0.37, p = 0.052). In good normosmics, there was a nonsignificant correlation between olfactory discrimination scores and age (rs = 0.33, p = 0.08). The only significant sex difference we observed was a marginal difference in BMI (t(65) = 1.7, p = 0.096), between men (mean 24.0, SD = 4.2) and women (mean 22.3, SD = 3.6). Olfactory scores did not differ significantly between men and women.
Sense of smell of the study cohort. (a) Boxplots showing odor threshold (T), discrimination (D) and identification (I) scores for each of the three groups (good normosmics, hyposmics and normosmics based on total TDI scores). (b) Boxplots showing the distribution of total TDI scores in each of the three groups (good normosmics, hyposmics and normosmics based on total TDI). Outliers are marked with circles, far outliers with stars.
Typical nasal microbiome was found
We characterized the nasal microbiome of the study subjects using Illumina MiSeq next generation sequencing (Supplementary Table 2). We identified 27 phyla, four belonging to the phylum Archaea (Thaumarchaeota [Nitrososphaera], Euryarchaeota [Methanosphaera, Methanobrevibacter, and Halogranum], Pacearchaeota and unclassified Archaea) and 23 to Bacteria. The most abundant phyla were Actinobacteria (50% of all sequence reads), Firmicutes (28%), and Proteobacteria (14%). The fourth most abundant bacterial phylum, Bacteroidetes, was represented by only 1.5% of all sequence reads. The most abundant bacteria at genus level were Corynebacterium (43%), a typical human skin bacterium32,33 also frequently found in the nose34,35, Staphylococcus spp. (15%), characteristic nasal residents34,36, and Dolosigranulum (4%), commensal inhabitants of the upper respiratory tract that have been implicated in causing infections37 but have also been identified as health-associated commensals38. Additionally we detected Peptoniphilus (4%), which is known to be part of the vaginal and gut microbiota39,40 and is among the most abundant bacteria in nasal samples, based on 16 S rRNA amplicon sequencing41, and culture42. Hyposmics had a higher proportion of Peptoniphilus (10%); this, however, was mainly driven by one subject (no. 49), who showed a predominant presence of signatures from this genus (see Supplementary Figures 3, 4 and 5, and Supplementary Table 2). An overview of the 300 most commonly detected bacterial genera that we identified is given in Fig. 2.
Hierarchical display of the human nasal bacteriome. Krona diagram visualization shows that the nasal microbiome of study subjects was dominated by typical skin and respiratory tract associated bacteria such as Corynebacterium, Staphylococcus and Dolosigranulum (normalized data from all study participants).
As BMI has previously been linked to olfactory function, we expected that subjects with differing BMI might also carry dissimilar nasal microbial communities. Indeed, the nasal microbiome did differ between those with normal, high and low BMI (Fig. 3), as the microbiome samples grouped separately. However, the differences were not found to be statistically significant (probably because of the small sample size), but a trend was obvious from a p-value of 0.084. The small sample size particularly for the underweight study subjects could potentially also explain why the association between BMI and olfactory function was not detected in this study.
Microbial community composition differs between normosmics and hyposmics
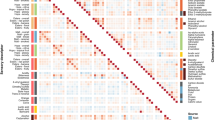
We compared the microbial communities between normosmics and hyposmics using alpha and beta diversity measures to assess the relationship between olfactory function and nasal microbiome. An overview of relationship between nasal microbiome in hyposmic, normosmic and good normosmic study subjects and the collected metadata is presented in Fig. 4.
Normosmic and hyposmic subjects were found to have significantly different nasal microbial community composition in the redundancy analysis (RDA) plot, both at operational taxonomic unit (OTU) (p = 0.007) (Fig. 5a) and genus (p = 0.037) levels. Furthermore, when the normosmic group was divided into average and good normosmics, there were significant differences between the 3 groups (hyposmics, average normosmics and good normosmics) at OTU level (p = 0.017) and near-significant differences at genus level (Fig. 5b). However, diversity (richness and evenness) of the nasal microbiome was not associated with overall olfactory function (based on total TDI score).
Microbial community composition and diversity reflects odor discrimination and threshold
We also evaluated whether subcategories of total olfactory function, i.e odor identification, threshold, and discrimination, are particularly associated with the dissimilarities that we detected in nasal microbial community structure.
The results show that odor identification was not associated with nasal microbial community structure or diversity, but some associations were found for odor threshold and discrimination. Subjects with hyposmia regarding odor threshold had higher microbial diversity (inverse simpson index) compared to normosmics (p = 0.003) (Fig. 6). Additionally, when the normosmic group was further divided into average and good normosmics (for threshold) the groups differed significantly (p = 0.01) in diversity. However, no significant difference was detected in diversity between average and good normosmics regarding odor threshold (p = 0.493).
Odor discrimination was related to nasal microbiome community composition (Fig. 7), with significant differences between hyposmic versus normosmic subjects (p = 0.006) (Fig. 6a), as well as between hyposmics, average and good normosmics (p = 0.004) (Fig. 7b), and nearly significant difference also between the average versus good normosmics (p = 0.075).
Decreased olfaction is associated with specific bacterial taxa
To investigate individual OTUs were correlated with total TDI scores in the dataset, we applied regression analysis (Pearson correlation), but no significant result was obtained. When we looked specifically at odor threshold score, the relative abundance of one OTU belonging to genus Campylobacter correlated negatively and significantly with odor threshold (Pearson correlation, p = 0.029). However, the relative abundance of any described OTU in this study did not correlate significantly with odor discrimination and identification scores. As the odor threshold, identification and discrimination seem to associate differently to the nasal microbiome (Fig. 4), we analysed the connections separately (Table 2, Supplementary Table 3) using LEfSe (Linear discriminant analysis Effect Size43) algorithm. LEfSe allows identification of features (e.g. taxons or functions) that most likely explain differences between groups. Our dataset was subjected to LEfSe analysis for determination of specific microbial markers for the subcategories odor threshold, identification and discrimination. Overall, for all subcategories, a number of specific microbial signatures were identified. I.e. low odor identification capability was associated with signatures of Proteobacteria, Actinobacteria, Firmicutes, and Bacteroidetes, as shown in Table 2 and Supplementary Table 3, which summarize the microbial genera significantly associated with olfactory function subcategories.
A number of signatures from bacterial taxa were associated with hyposmic threshold, low discrimination and low identification performance; these included mainly certain Actinobacteria, Bacteroidia, Bacilli, Clostridia and Proteobacteria. In particular signatures of Corynebacterium and Faecalibacterium appeared frequently as a biomarker for reduced odor discrimination and threshold, while Comamonadaceae and Enterobacteriaceae were significantly more abundant in nasal microbiome of study subjects with reduced odor threshold and identification. Signatures of genera associated with low performance in all three olfactory subcategories (T, D, I) belonged to Porphyromonas and unclassified Lachnospiraceae (Table 2 and Supplementary Table 3)44.
Discussion
In this study, we explored the relationship between olfactory function and nasal microbiome in 67 healthy volunteers. The aim was to determine whether: (i) olfactory epithelium microbiome community composition and diversity differ between normosmic and hyposmic individuals, (ii) specific bacteria therein are associated with above average or impaired olfactory function, and (iii) other factors correlate with a specific microbiome composition. Our results show that olfactory function was potentially connected to nasal microbiome community composition, as the microbiomes of normosmic and hyposmic study subjects differed significantly in RDA analyses and regarding their alpha diversity. Moreover, we could also detect nearly significant differences in the microbiome composition of subjects with normal, and those with particularly good olfactory function, based on odor discrimination scores.
Subjects with lower olfactory threshold scores had a more diverse nasal microbiome compared to those with normal odor thresholds. In general, a more diverse microbial community is accompanied by a wider range of microbial functions; this is usually acknowledged as a positive feature. Butyrate-producing Faecalibacterium or Porphyromonas in particular were strongly associated with reduced olfactory function in our study. Porphyromonas species are well known as butyrate-producing components of the human oral microbiome, whereas Corynebacterium representatives, which were correlated with reduced odor discrimination and threshold, are abundant on the skin, but, like Comamonadaceae, are also widely distributed in other human niches45. Futhermore, Lachnospiraceae, Faecalibacterium and Enterobacteriaceae, all associated with hyposmia, are typical gut microorganisms, also capable of butyric acid formation45.
Butyric acid, a typical product of anaerobic fermentation in the colon, plays a major role in energy balance and is involved in numerous body processes46,47,48. Characterized by a very strong and unpleasant odor, it can be detected in concentrations as low as 10 parts per million49. Commonly used as an odorant in chemosensory testing it has been investigated extensively as a stimulus. It has been reported that certain dietary restrictions can lead to butyric acid avoidance in mice50.
The microbiome has been shown to modulate the physiology of the olfactory epithelium, potentially changing the response to odorant stimulation30. Furthermore, olfactory receptors have been reported to detect microbial signals such as short-chain fatty acids51 hypothetically leading to changed olfactory perception, and it seems possible that by producing strong-smelling compounds such as butyric acid, microorganisms within the nasal microbiome could impact on olfactory perception and thus also on appetite. The sense of smell plays a major role in eating behavior, including anticipation and stimulation of appetite52, as well as flavor perception8,53,54. In our study, reflecting a healthy population we did not see a correlation of BMI with olfactory function, but a tendency towards a differing nasal microbiome between those with normal, high and low BMI. However, this relationship of BMI, body weight and olfactory function is also reflected in the literature: On one hand, individuals with an altered olfactory sense report alterations of dietary behavior and their BMI and olfactory function are negatively correlated15,55. On the other hand nutritional states are known to affect odor detection abilities due to hormonal and enzyme-related appetite regulation, and for patients with extreme weight/eating disorders, there is a negative impact on olfactory function for extremely high BMI and a positive impact for extremely low BMI (see for example56,57,58,59,60,61,62,63). Olfactory performance measures are known to be influenced by various molecules that modulate food intake64. The appetite-stimulating molecule ghrelin has been hypothesized to act as a moderator in emotional eating behavior, thereby influencing BMI60. Our finding that microbiome distribution differed with BMI level, although not quite statistically significant, might therefore be a direct result of this interaction.
Our study provides first insights into a potential correlation between olfactory function and olfactory epithelium microbiome composition. Future investigations will need to clarify whether the higher abundance of butyric acid-producing microorganisms in hyposmic individuals are a result or a cause of reduced olfactory function. Manipulation of microbiome content by using probiotics or fecal transplants has been shown to be beneficial for treating several gastrointestinal diseases associated with gut dysbiosis65,66. Whether altering the nasal microbiome in diseases in which the microbiota of the nasal cavity is affected, such as chronic rhinosinusitis, is feasible or has any therapeutic potential remains to be investigated49.
Methods
Subjects
Sixty-seven volunteers were recruited by means of public bulletins and announcements from the University of Graz and the Medical University of Graz, as well as announcements in local and upper regional newspapers. The following exclusion criteria were applied: consumption of antibiotics/probiotics within the last month before testing, acute hay fever, pollen allergies or common cold at the time of testing, nasal polyps, chronic neurologic, psychiatric or other severe disease, use of nasal spray on the day of testing, and age outside the specified range of 18–45 years. The study was evaluated and approved by the local ethics committee of the University of Graz (reference number GZ. 39/80/63 ex 2014/15) according to the Declaration of Helsinki. All study subjects provided an informed consent before participation.
Olfactory function testing
For all subjects, olfactory function was assessed using the Sniffin’ Sticks battery (Burghart Instruments, Wedel, Germany) following the standardized TDI (threshold, discrimination, identification) procedure67, see following section for details. An ENT physician examined the nasal cavity of each participant and obtained nasal swabs (Ultra minitip flocked swab, Copan, USA) from the olfactory mucosa, located at the ceiling of the nasal cavity (Supplementary Figure 7.). Samples were stored immediately on dry ice and transferred to Medical University of Graz for further analyses. The sequence of olfactory measurements and ENT examination was counterbalanced across participants.
The Sniffin’ Sticks battery consists of pen-like devices filled with odorants67,68,69. Three subtests designed to assess orthonasal olfactory function were carried out: detection threshold (the lowest concentration at which an odor can be perceived), odor discrimination (ability to distinguish between odors) and odor identification (ability to assign names to odors). The odor detection threshold was identified for each participant using a three-alternative, single-staircase, forced-choice procedure with the odorant n-butanol. Thus, participants were presented with triplets of odorant pens and asked to identify the pen containing n-butanol amongst two blank distractor pens. In the second subtest, odor discrimination ability was assessed using 16 triplets of odorants: within each triplet, two pens contained the same, while the third pen contained a different, odorant. In a forced-choice procedure, the participants were asked to detect the odd pen for each triplet. During the odor identification task, participants were presented with 16 common odors. Using a multiple choice answering format, they were asked to select which of four odor labels matched each presented odor.
Possible scores range from 1–16 for the detection threshold subtest and 0–16 for the other two subtests. The total TDI score is obtained by summing the scores for all three subtests. Normosmia is defined by a TDI score of ≥31, hyposmia by a TDI score 17 to 30.75 and anosmia a TDI score of <1770. The normosmic group was subdivided into average normosmics and good normosmics, using a cutoff TDI score ≥35.5 (median split). We also evaluated the association between olfactory function and microbiome separately for odor threshold, discrimination, and identification scores. The cutoff used to differentiate normosmics from hyposmics was ≥6.25 for threshold, ≥10 for discrimination, and ≥11 for identification70. Average normosmics and good normosmics were differentiated using a dividing value of ≥8.9 for threshold, ≥13 for discrimination, and ≥14 for identification (score higher than 50% of the normosmic group).
Statistical analysis
To evaluate whether olfactory scores were related to age or BMI, we calculated non-parametric Spearman’s rho correlations between these measures, as some parameters were not normally distributed. We also assessed whether olfactory scores, age or BMI differed between male and female subjects using two-sample t-tests. Statistical analyses were performed using the Statistical Package for the Social Sciences, Version 24.0 (SPSS, Chicago, Illinois). An alpha value of 0.05 was considered as statistically significant.
Microbiome analysis
The sample tubes were placed on dry ice immediately after sampling, then samples were stored at −80 °C until further processing.
Genomic DNA was extracted from the swabs using FastDNA SPIN Kit (MP Biomedicals, USA) according to manufacturer’s instructions. Concentration of the isolated DNA was quantified with Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific, USA). Variable region V4 of 16S rRNA gene was amplified with universal PCR primers 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 806 R (5′-GGACTACHVGGGTWTCTAAT-3′)71 using TaKaRa Ex Taq polymerase (Clontech, Japan) and 10–20 ng template in each reaction. Cycling conditions consisted of an initial denaturation at 94 °C for 3 min, followed by 35 cycles of denaturation (94 °C 45 sec), annealing (50 °C 60 sec), extension (72 °C 90 sec), and a final extension at 72 °C for 10 min. The resulting fragments were visualized in 3% agarose gel, and sequenced at ZMF Core Facility Molecular Biology in Graz, Austria, using the MiSeq platform.
Microbiome data processing
To analyze the microbial community structure and taxonomic diversity, the obtained raw reads were processed using mothur version 1.36.172 following the Standard Operation Procedure (SOP). In short, the paired end reads were joined together, the produced sequences were quality checked (minimum length 200, maximum length 300, maximum number of homopolymers 8) and aligned against the SILVA 123 database73. Good-quality sequences were then pre-clustered and chimeric sequences removed. Taxonomic assignment was carried out by querying the sequence reads against a trainset14_032015 reference database, after which the sequences were clustered into OTUs (threshold 0.03 dissimilarity) using average neighbor algorithm. A biom table was constructed for downstream analyses. OTUs represented by ≤5 sequences were removed from the analysis.
To evaluate alpha and beta diversities and differences in community composition, and visualize the results, we applied Calypso (Version 5.8), an online platform for mining, visualizing and comparing multiple microbial community composition data (cgenome.net/calypso). Total-sum normalization was applied for 16 S rRNA gene data. RDA and PCA plots were created with Calypso Multivariate tool, and inverse simpson indices were calculated and visualized with Calypso Diversity tool. The differences in alpha diversity were analyzed using anova. Individual OTUs were correlated with total TDI scores using regression analysis (Pearson correlation).
Declarations
Ethics approval and concent to participate
The study was evaluated and approved by the local ethics committee of the University of Graz (reference number GZ. 39/80/63 ex 2014/15) according to the Declaration of Helsinki. All study subjects provided an informed consent before participation.
Availability of data and materials
The dataset supporting the conclusions of this article is available in The European Nucleotide Archive (ENA) under study accession number PRJEB20526.
References
Bushdid, C., Magnasco, M. O., Vosshall, L. B. & Keller, A. Humans Can Discriminate More than 1 Trillion Olfactory Stimuli. Science 343, 1370–1372 (2014).
Croy, I. et al. Retronasal testing of olfactory function: An investigation and comparison in seven countries. Eur. Arch. Oto-Rhino-Laryngology 271, 1087–1095 (2014).
Boesveldt, S. et al. Anosmia - a clinical update. Chem. Senses in press, (2017).
Frasnelli, J. & Hummel, T. Olfactory dysfunction and daily life. Eur. Arch. oto-rhino-laryngology 262, 231–5 (2005).
Blomqvist, E. H., Brämerson, A., Stjärne, P. & Nordin, S. Consequences of olfactory loss and adopted coping strategies. Rhinology 42, 189–194 (2004).
Temmel, A. F. P. et al. Characteristics of Olfactory Disorders in Relation to Major Causes of Olfactory Loss. Arch. Otolaryngol. Neck Surg. 128, 635 (2002).
Kollndorfer, K., Reichert, J., Braunsteiner, J. & Schöpf, V. Assessment of olfactory memory in olfactory dysfunction. Perception 46, 516–529 (2017).
Croy, I., Nordin, S. & Hummel, T. Olfactory Disorders and Quality of Life - An Updated Review. Chem. Senses 39, 185–94 (2014).
Osborne-Crowley, K. & McDonald, S. Hyposmia, not emotion perception, is associated with psychosocial outcome after severe traumatic brain injury. Neuropsychology 30, 820–829 (2016).
Mattes, R. D. et al. Dietary evaluation of patients with smell and/or taste disorders. Am. J. Clin. Nutr. 51, 233–40 (1990).
Mattes, R. D. & Cowart, B. J. Dietary assessment of patients with chemosensory disorders. J. Am. Diet. Assoc. 94, 50–6 (1994).
Ferris, A. M. & Duffy, V. B. Effect of olfactory deficits on nutritional status. Does age predict persons at risk? Ann. N. Y. Acad. Sci. 561, 113–23 (1989).
Deems, D. A. et al. Smell and taste disorders, a study of 750 patients from the University of Pennsylvania Smell and Taste Center. Arch. Otolaryngol. Head. Neck Surg. 117, 519–28 (1991).
Schechter, P. J. & Henkin, R. I. Abnormalities of taste and smell after head trauma. J. Neurol. Neurosurg. Psychiatry 37, 802–10 (1974).
Aschenbrenner, K. et al. The influence of olfactory loss on dietary behaviors. Laryngoscope 118, 135–44 (2008).
Barresi, M. et al. Evaluation of olfactory dysfunction in neurodegenerative diseases. J. Neurol. Sci. 323, 16–24 (2012).
Rahayel, S., Frasnelli, J. & Joubert, S. The effect of Alzheimer’s disease and Parkinson’s disease on olfaction: A meta-analysis. Behav. Brain Res. 231, 60–74 (2012).
Lin, A. et al. Olfactory identification deficits at identification as ultra-high risk for psychosis are associated with poor functional outcome. Schizophrenia Research 161(2-3), 156–162 (2015).
Ekström, I. et al. Smell Loss Predicts Mortality Risk Regardless of Dementia Conversion. J. Am. Geriatr. Soc. https://doi.org/10.1111/jgs.14770 (2017).
Pinto, J. M., Wroblewski, K. E., Kern, D. W., Schumm, L. P. & McClintock, M. K. Olfactory Dysfunction Predicts 5-Year Mortality in Older Adults. PLoS One 9, e107541 (2014).
Sender, R., Fuchs, S. & Milo, R. Revised estimates for the number of human and bacteria cells in the body. Biorxiv 14, 1–21 (2016).
Savage, D. S. Microbial ecology of the gastrointestinal tract. Annu. Rev. Microbiol. 31, 107–133 (1977).
Leyden J, McGinley K, Nordstrom K, Webster G. Skin microflora. J Invest Dermatol 65–72 (1987).
Maynard, C. L., Elson, C. O., Hatton, R. D. & Weaver, C. T. Reciprocal interactions of the intestinal microbiota and immune system. Nature 489, 231–41 (2012).
Hughes, D. T. & Sperandio, V. Inter-kingdom signalling: communication between bacteria and their hosts. Nat. Rev. Microbiol. 6, 111–120 (2008).
Grenham, S., Clarke, G., Cryan, J. F. & Dinan, T. G. Brain-Gut-Microbe Communication in Health and Disease. Front. Physiol. 2, 94 (2011).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn’s disease. Cell Host Microbe 15, 382–392 (2014).
Hartstra, A. V., Bouter, K. E. C., Bäckhed, F. & Nieuwdorp, M. Insights into the role of the microbiome in obesity and type 2 diabetes. Diabetes Care 38, 159–165 (2015).
Wilson, M. T. & Hamilos, D. L. The nasal and sinus microbiome in health and disease. Curr. Allergy Asthma Rep. 14, 485 (2014).
François, A. et al. Olfactory epithelium changes in germfree mice. Sci. Rep. 6, 24687 (2016).
Pereira, P. A. B. et al. Oral and nasal microbiota in Parkinson’s disease. Parkinsonism Relat. Disord, https://doi.org/10.1016/j.parkreldis.2017.02.026 (2017).
Costello, E. K. et al. Bacterial community variation in human body habitats across space and time. Science 326, 1694–7 (2009).
Grice, E. A. et al. Topographical and Temporal Diversity of the Human Skin Microbiome. Science 324, 1190–1192 (2009).
Frank, D. N. et al. The Human Nasal Microbiota and Staphylococcus aureus Carriage. PLoS One 5, e10598 (2010).
Liu, C. M. et al. Staphylococcus aureus and the ecology of the nasal microbiome. Sci. Adv. 1, e1400216 (2015).
Ramakrishnan, V. R., Hauser, L. J. & Frank, D. N. The sinonasal bacterial microbiome in health and disease. Curr. Opin. Otolaryngol. Head Neck Surg. 24, 20–25 (2016).
Lecuyer, H. et al. Dolosigranulum pigrum Causing Nosocomial Pneumonia and Septicemia. J. Clin. Microbiol. 45, 3474–3475 (2007).
Bosch, A. A. T. M. et al. Development of Upper Respiratory Tract Microbiota in Infancy is Affected by Mode of Delivery. EBioMedicine 9, 336–45 (2016).
Sharma, H., Tal, R., Clark, N. A. & Segars, J. H. Microbiota and pelvic inflammatory disease. Semin. Reprod. Med. 32, 43–9 (2014).
Lagier, J.-C. et al. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin. Microbiol. Infect. 18, 1185–1193 (2012).
Biswas, K., Hoggard, M., Jain, R., Taylor, M. W. & Douglas, R. G. The nasal microbiota in health and disease: Variation within and between subjects. Front. Microbiol. 6, 1–9 (2015).
Kaspar, U. et al. The culturome of the human nose habitats reveals individual bacterial fingerprint patterns. Environ. Microbiol. 18, 2130–2142 (2016).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Summanen, P., Finegold, S. M., Summanen, P. & Finegold, S. M. in Bergey’s Manual of Systematics of Archaea and Bacteria 1–14 (John Wiley & Sons, Ltd, 2015), https://doi.org/10.1002/9781118960608.gbm00246 (2015).
Bergey’s Manual of Systematics of Archaea and Bacteria. (John Wiley & Sons, Ltd, 2015) https://doi.org/10.1002/9781118960608 (2015).
Hamer, H. M. et al. Review article: the role of butyrate on colonic function. Aliment. Pharmacol. Ther. 27, 104–119 (2007).
Bourassa, M. W., Alim, I., Bultman, S. J. & Ratan, R. R. Butyrate, neuroepigenetics and the gut microbiome: Can a high fiber diet improve brain health? Neurosci. Lett. 625, 56–63 (2016).
Plöger, S. et al. Microbial butyrate and its role for barrier function in the gastrointestinal tract. Ann. N. Y. Acad. Sci. 1258, 52–59 (2012).
Mahdavinia, M., Keshavarzian, A., Tobin, M. C., Landay, A. L. & Schleimer, R. P. A comprehensive review of the nasal microbiome in chronic rhinosinusitis (CRS). Clin. Exp. Allergy 46, 21–41 (2015).
Utsugi, C. et al. Hard-diet feeding recovers neurogenesis in the subventricular zone and olfactory functions of mice impaired by soft-diet feeding. PLoS One 9, e97309 (2014).
Pluznick, J. L. et al. Olfactory receptor responding to gut microbiota- derived signals plays a role in renin secretion and blood pressure regulation. Proc. Natl. Acad. Sci. 110, 4410–4415 (2013).
Boesveldt, S. & de Graaf, K. The Differential Role of Smell and Taste For Eating Behavior. Perception 30100661668557, https://doi.org/10.1177/0301006616685576 (2017).
Hummel, T. & Nordin, S. Olfactory disorders and their consequences for quality of life. Acta Otolaryngol. 125, 116–121 (2005).
Stevenson, R. J. The role of attention in flavour perception. Flavour 1, 2 (2012).
Schöpf, V., Kollndorfer, K., Pollak, M., Mueller, C. A. & Freiherr, J. Intranasal insulin influences the olfactory performance of patients with smell loss, dependent on the body mass index: A pilot study. Rhinology 53, 371–378 (2015).
Aschenbrenner, K., Scholze, N., Joraschky, P. & Hummel, T. Gustatory and olfactory sensitivity in patients with anorexia and bulimia in the course of treatment. J. Psychiatr. Res. 43, 129–137 (2008).
Bentz, M. et al. Heightened Olfactory Sensitivity in Young Females with Recent-Onset Anorexia Nervosa and Recovered Individuals. PLoS One 12, e0169183 (2017).
Claxton, A. et al. Sex and ApoE genotype differences in treatment response to two doses of intranasal insulin in adults with mild cognitive impairment or Alzheimer’s disease. J. Alzheimers. Dis. 35, 789–797 (2013).
Dazzi, F. et al. Alterations of the olfactory-gustatory functions in patients with eating disorders. Eur. Eat. Disord. Rev. J. Eat. Disord. Assoc. 21, 382–5 (2013).
Fernández-Aranda, F. et al. Smell-taste dysfunctions in extreme weight/eating conditions: analysis of hormonal and psychological interactions. Endocrine 51, 256–267 (2016).
Islam, M. A. et al. Olfaction in eating disorders and abnormal eating behavior: a systematic review. Front. Psychol. 6, 1431 (2015).
Rapps, N. et al. Olfactory deficits in patients with anorexia nervosa. Eur. Eat. Disord. Rev. 18, 385–389 (2010).
Schecklmann, M. et al. Olfaction in child and adolescent anorexia nervosa. J. Neural Transm. 119, 721–728 (2012).
Palouzier-Paulignan, B. et al. Olfaction under metabolic influences. Chem. Senses 37, 769–797 (2012).
Cryan, J. F. & O’Mahony, S. M. The microbiome-gut-brain axis: from bowel to behavior. Neurogastroenterol. Motil. 23, 187–92 (2011).
Duke, P. S. & Fardy, J. Recurrent Clostridium difficile infection treated with home fecal transplantation: a case report. J. Med. Case Rep. 8, 393 (2014).
Hummel, T., Sekinger, B., Wolf, S. R., Pauli, E. & Kobal, G. ‘Sniffin’ sticks’: olfactory performance assessed by the combined testing of odor identification, odor discrimination and olfactory threshold. Chem. Senses 22, 39–52 (1997).
Kobal, G. et al. Multicenter investigation of 1,036 subjects using a standardized method for the assessment of olfactory function combining tests of odor identification, odor discrimination, and olfactory thresholds. Eur. Arch. Oto-Rhino-Laryngology 257, 205–211 (2000).
Kobal, G. et al. ‘Sniffin’ Sticks’: Screening of olfactory performance. Rhinology 34, 222–226 (1996).
Hummel, T., Kobal, G., Gudziol, H. & Mackay-Sim, A. Normative data for the ‘Sniffin’ Sticks’ including tests of odor identification, odor discrimination, and olfactory thresholds: An upgrade based on a group of more than 3,000 subjects. Eur. Arch. Oto-Rhino-Laryngology 264, 237–243 (2007).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J. 6, 1621–1624 (2012).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microbiol. 75, 7537–41 (2009).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–6 (2013).
Acknowledgements
The authors thank Deepika Bagga, Katharina Gruber, Vanessa Hinterleitner, Christina Hirschmann, Sigrid Haider, Jennifer Kern, Margit List-Schleich, and Lisa Maria Rabl for their support with testing and inviting the participants. We also thank Lisa Wink for skillful laboratory assistance. Furthermore, we would like to thank Henkel Central Eastern Europe GmbH (Austria) for providing us with products for subject compensation. We also thank Sanne Boesveldt, Joel Mainland, and Leslie Vosshall for discussion of our results and helpful comments. Julia Balfour of Northstar Medical Writing and Editing Services provided English-language editing support.
Author information
Authors and Affiliations
Contributions
V.S. and C.M.E. conceived and designed the study, J.R., S.H., J.S., V.S. collected samples and acquired data, K.K. and S.D. processed the samples, K.K., J.R., C.M.E. and V.S. analysed and interpreted the data, K.K., J.R., C.M.E. and V.S. drafted and revised the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koskinen, K., Reichert, J.L., Hoier, S. et al. The nasal microbiome mirrors and potentially shapes olfactory function. Sci Rep 8, 1296 (2018). https://doi.org/10.1038/s41598-018-19438-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-19438-3
This article is cited by
-
Nasal Microbiome in COVID-19: A Potential Role of Corynebacterium in Anosmia
Current Microbiology (2023)
-
The Upper Respiratory Tract Microbiome Network Impacted by SARS-CoV-2
Microbial Ecology (2023)
-
A Review on the Nasal Microbiome and Various Disease Conditions for Newer Approaches to Treatments
Indian Journal of Otolaryngology and Head & Neck Surgery (2023)
-
Nasopharyngeal microbiota profiling of pregnant women with SARS-CoV-2 infection
Scientific Reports (2022)
-
Psychophysical chemosensory dysfunction in eating disorders: a qualitative systematic review
Eating and Weight Disorders - Studies on Anorexia, Bulimia and Obesity (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.