Abstract
In this report, we present an improved protocol for CRISPR/Cas9 genome editing in mice. The procedure consists in the electroporation of intact mouse zygotes with ribonucleoprotein complexes prepared in vitro from recombinant Cas9 nuclease and synthetic dual guide RNA. This simple cloning-free method proves to be extremely efficient for the generation of indels and small deletions by non-homologous end joining, and for the generation of specific point mutations by homology-directed repair. The procedure, which avoids DNA construction, in vitro transcription and oocyte microinjection, greatly simplifies genome editing in mice.
Similar content being viewed by others
Introduction
Since its adaptation to mammalian cells1,2, CRISPR/Cas9 genome editing has become a key element of the genetic toolbox in mice3,4. The method relies on Streptococcus pyogenes Cas9 (SpCas9), a RNA-guided DNA endonuclease. Annealing between genomic DNA and the 5′-end of the guide RNA defines the position of the cleavage site by SpCas9, provided that the homologous genomic DNA sequence is immediately followed by a so-called protospacer adjacent motif (PAM: 5′-NGG-3′). SpCas9 cleaves both DNA strands 3 nucleotides (nt) upstream to the PAM. Therefore, the sequence of the first 20 nt of the guide RNA defines the genomic location of the double strand break5. Cells can repair double strand breaks in two ways: (1) non-homologous end-joining (NHEJ), an error prone mechanism which often generates small insertions and deletions (indels) at the cleavage site, (2) homology-directed repair (HDR) which uses an intact DNA molecule as repair template. The repair template can be provided as a single-stranded oligodeoxynucleotide (ssODN) to generate point mutations or small insertions. In that case, HDR introduces predefined mutations, and therefore can be used to precisely modify coding sequences.
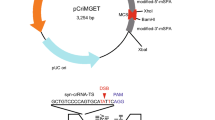
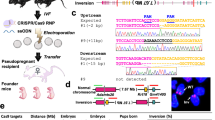
The methodology for CRISPR/Cas9 mediated genome editing is improving rapidly to increase in simplicity, efficiency and versatility, as exemplified by the recent development of cloning-free procedures. They combine the use of recombinant Cas9 protein with chemically synthetized guide RNA6, usually in the form of a dual guide RNA7 obtained by annealing two RNA molecules: a long component (72-mer) with a constant nucleotide sequence called tracrRNA and a short variable component (42-mer) called crRNA. The 5′ part (20 nt) of the crRNA is homologous to the targeted genomic DNA sequence, while its 3′ part is complementary to the tracrRNA (Fig. 1). Using a dual guide RNA, rather than a single guide RNA (sgRNA), circumvents the difficult chemical synthesis of long RNA molecules and is cost-effective since the same batch of tracrRNA can be aliquoted and used for several projects. Chemical modifications introduced at the extremities of the RNAs protect from exonucleases, and significantly improve efficiency6,8. When mixed in vitro, SpCas9 and the dual guide RNA assemble into ribonucleoprotein complexes (RNPs) which can be introduced into mouse zygotes by microinjection7 or in cultured cells by electroporation9,10.
General structure of the dual guide RNA. Two synthetic RNA molecules are annealed. The short variable 42mer called crRNA has its first 20 nt designed to match a genomic sequence located immediately upstream to a PAM consensus motif (5′NGG) and the sequence is identical to the bottom genomic DNA strand on this figure. The second RNA, called tracrRNA, is a constant 72mer with partial homology to the crRNA, which secondary structure permits the tight association to the Cas9 nuclease (grey). Black crosses indicate the modified nucleotides, to which 2′-O-methyl and 3′ phosphorothioate are added to protect from nuclease degradation. Arrows indicate the cutting sites of the Cas9 nuclease.
Electroporation can also be efficiently used to introduce RNP complexes made of Cas9 and sgRNA into mouse zygotes11,12,13. In the present manuscript, we show the clear advantage of combining electroporation, with the cloning-free procedure using dual guide RNA/SpCas9 RNP complexes to perform genome editing in mice. We found that these RNPs electroporation can generate mutations by NHEJ with 100% efficiency. The coelectroporation of two different RNPs can be used to generate deletions, or to simultaneously generate small indels at two different loci. If a ssODN is added to the RNPs preparation in order to provide template for HDR, the procedure also leads to the generation of specific point mutations. Overall, this new protocol is cost-effective and makes CRISPR/Cas9 in mice amenable for small facilities and laboratories without previous expertise in mouse transgenesis.
Results
The four following experiments of genome editing were recently performed in our facility by combining the cloning-free procedure and zygote electroporation as depicted in Table 1. Technical details are presented in the method section, while all RNA and DNA sequences are detailed in Supplementary Table S1.
Generation of small indels by NHEJ repair after electroporation of RNPs
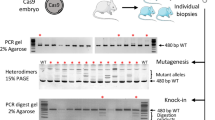
The aim of the experiment was to generate frameshift mutations in order to inactivate the function of Brcc3 gene (encoding subunit 3 of the BRCA1-BRCA2-containing complex), a gene located on the X chromosome. RNPs directed against Brcc3 gene were electroporated into zygotes. DNA samples were prepared from 27 pups born after embryo reimplantation. The DNA sequence surrounding the expected double strand break was amplified by PCR and analyzed by agarose gel electrophoresis to identify major changes in fragment size. Large deletions were detected in two DNA samples (close to 200 bp, Supplementary Fig. S1). Four DNA samples repeatedly failed to amplify, suggesting the presence of larger deletions, as DNA integrity was verified with primers targeting an independent locus. This hypothesis was confirmed for three samples by an additional PCR analysis targeting a larger sequence (Supplementary Fig. S1). Sanger sequencing and TIDE analysis were performed for the 21 remaining samples in which deletions were not obviously visible. We identified small indels in the genome of all the corresponding pups (Fig. 2A, Table 2). This analysis failed to reveal the persistence of the wild-type allele in the somatic tissues of the F0 pups. In 20 out of these 21 mice, up to two alleles were detected (Table 3), reflecting mosaicism, at least in male pups. This is caused by NHEJ-generated mutations occurring after the first mitotic cell division of the embryo. Males n°3 and 8 were mated with wild-type C57BL/6 mice. As anticipated, genotyping of the female offspring revealed the germline transmission of one mutant allele to each pup (n = 6 and n = 4). F0 mice transmitted only one of the two mutant alleles detected in somatic tissues. Surprisingly, the allele transmitted by mouse n°8 was the least frequent in somatic tissue (8.9%), according to TIDE analysis. Therefore, information from genotyping of F0 toe clips cannot always be extrapolated to predict the germline genetic content.
Mice genotyping (A) Use of a dual guide RNA to generate short indels on Brcc3 gene. DNA Sanger sequencing of control and mutated F0 n°4 mice are shown. Top chromatogram represents a part of wild-type Brcc3 gene: guide sequence is underlined in blue, green rectangle indicates PAM, red arrow indicates Cas9 cutting site. Bottom chromatogram represents mutant mouse n°4, in which NHEJ generated a 2 nt deletion (dotted red lines) without apparent mosaicism, as correlated by TIDE software analysis represented on the right side of chromatograms. (B) Simultaneous use of two dual guide RNAs to generate short indels on two vasohibin genes located on different chromosomes. DNA Sanger sequencing of control and mutated F0 mice n°2 are shown for Vash1 (top chromatograms) and Vash2 (bottom). The TIDE software is used to analyze the spectrum and frequency of indels, as represented on the right side of chromatograms. On control sequences, guide sequences are underlined in blue and green rectangles indicate PAM. Red arrows indicate Cas9 cutting site. On Vash1 mutant lane, mosaicism and heterozygosity generate a scrambled sequence. Vash2 mutant chromatogram represents a case without apparent mosaicism, in which NHEJ generated a 28 nt deletion (dotted red lines) on both Vash2 alleles. (C) Simultaneous use of two dual guide RNAs to generate a 83 bp deletion in Ctse gene. Agarose gel analysis (cropped picture) of PCR performed on F0 mice genomic DNA, showing the frequent presence of a shorter PCR product of the expected size: the intended deletion of 83 bp leaves 398 bp instead of the 481 bp for the wild-type allele. Although most mice seem to harbor the expected deletion, additional small indels were identified by Sanger sequencing for some of them. Samples 8, 9 and 12 have larger deletion. Samples 10 and 17 repeatedly failed to amplify, suggesting large deletion, as DNA integrity was verified with primers targeting an independent locus (Supplementary Fig. S3). C represents control mouse with only the 481 bp wild-type allele. Full-length gels are presented in Supplementary Fig. S3. (D) Generation of specific point mutations using ssODN as template for HDR. A fraction of the Mct8 genomic sequence and the corresponding reading frame are shown. The sequence underlined in blue is complementary to the 5′ end of the crRNA. The green rectangle indicates the complement of the PAM sequence. Red arrow indicates Cas9 cutting site. The ssODN has perfect homology for the Mct8 gene and is identical to the “non PAM” strand on 92 nt at the 5′ end and 36 nt at the 3′ end. Mismatches are with red letters. The first two mismatches change the proline codon into a leucine codon. The following ones eliminate the PAM sequence, preventing Cas9 cleavage after HDR, and create a Pac I restriction site facilitating mouse genotyping. Top chromatogram represents control mouse sequence and the bottom one represents mutated F0 male n°7. Asterisks indicate mutated nucleotides.
Generation of small indels on two loci by NHEJ repair after electroporation of RNPs
The aim of the experiment was to generate frameshift mutations in order to inactivate the function of both Vash1 and Vash2 genes (encoding vasohibin 1 and vasohibin 2) located on two different chromosomes. RNPs directed against each gene were electroporated together. Sequences surrounding the expected double strand breaks were amplified by PCR from 11 mice. Agarose gel electrophoresis did not identify major changes in fragment size, except for one mouse, where a large insertion was found (Supplementary Fig. S2). Sanger sequencing identified indels in both Vash1 and Vash2 genes for all analyzed newborns (Fig. 2B and Table 2). Deletion of up to 35 nt and 42 nt were respectively found for Vash1 and Vash2 (Table 4). By TIDE analysis, we identified the wild-type allele in only 4 pups. We identified several mutations in each newborn (2 to 5 different alleles identified for Vash1 gene and 1 to 3 alleles for Vash2 gene), revealing mosaicism. Two F0 mice were mated with wild-type C57BL/6 mice for germline transmission. Only 2 F1 pups from breeder n°7 were genotyped, and both inherited Vash1 and Vash2 mutations. Mice n°6 produced F1 offspring in which all pups (n = 15) inherited one mutant allele for Vash1 and one mutant allele for Vash2 genes, confirming the complete absence of wild-type alleles in the F0 mice.
Generation of a deletion by coelectroporation of two RNPs
Our goal was to generate a 83 bp deletion in the Ctse gene, encoding Cathepsin E. This was achieved by co-introducing two RNPs targeting neighboring sequences. Analysis of PCR products by agarose gel electrophoresis indicated the presence of a deletion with the approximate expected size in 15 out of 25 mice (Fig. 2C). Larger deletions, up to 230 bp, were identified for 3 mice (n°8, 9 and 12). PCR amplification failed repeatedly for 2 mice suggesting the presence of large deletions, as DNA integrity was verified with primers targeting an independent locus (Supplementary Fig. S3). Sanger sequencing of PCR products confirmed gel analysis results regarding deletions, and identified 5 mice with small indels on one or both targeted sites (Table 2). Only the 83 bp deletion, and no other allele, was identified in 9 out of 23 mice, suggesting the deletion of both alleles in all cells (Supplementary Fig. S3). Two of these 9 F0 mice were mated with wild-type mice for germline transmission of the deleted allele. The result of the first litter, in which 6 out of 11 pups inherited the mutation, was finally indicative of germline heterozygosity or mosaicism in the F0 mice. In the second litter, all individuals (n = 8) were heterozygous for the deletion, suggesting germline homozygosity in the F0 mice.
Introduction of in frame missense mutation by HDR from an exogenous single-stranded DNA template
We used HDR to introduce an amino-acid substitution (P321L) in the MCT8 transporter, encoded by Mct8 located on the X chromosome. The dual guide RNA was designed to cut the DNA sequence next to codon 321. Several mismatches were introduced in the ssODN template: (1) the first two mutated one codon introducing P321L, (2) a silent mutation eliminated the PAM sequence from the genome preventing further Cas9 cleavage after HDR, (3) silent mutations of 3 nucleotides created a PacI restriction site to facilitate genotyping (Fig. 2D). Allele-specific PCR and PCR/Sanger sequencing identified all the designed mutations in 69% of the pups (Fig. 2D, Table 2 and Supplementary Fig. S4). We also found 2 unusual cases in which a portion of the ssODN sequence was inserted at the Cas9 cleavage site by NHEJ. Of note, only the expected mutant allele, and no other allele, was detected in 5 males and 3 females by TIDE analysis, suggesting that there was no mosaicism in these F0 mice and that they were hemizygous males and homozygous females. Absence of mosaicism in the germline was confirmed for 3 of these F0 males. Indeed, all F1 female offspring (n = 5; n = 6; n = 8) from each of these mutated F0 males were heterozygous for the mutation. On the contrary, offspring analysis from one F0 female was rather suggestive of mosaicism or heterozygosity in this female. Indeed, only 5 out of 12 pups inherited the mutation. We also successfully verified the absence of a non-intended mutation on a predicted intronic off-target site on the X chromosome (Supplementary Fig. S4 and Table S2).
Altogether these data show the outstanding performance of this new procedure combining zygote electroporation with the use of dual guide/SpCas9 RNP complexes.
Discussion
A number of possible combinations have been used for CRISPR/Cas9 genome editing in mice. Cas9 can be encoded by a microinjected plasmid14, a microinjected mRNA15, or directly provided as a recombinant protein7,16,17. Single guide or dual guide10 RNA can be used, transcribed either in vivo from a microinjected plasmid or in vitro with a phage polymerase. Guide RNA can also be produced by chemical synthesis10. Finally, microinjection can be substituted by electroporation11,12,13,18,19,20, which can also be performed in situ in oviduct21. Here, the presented examples illustrate the efficiency of mouse zygotes electroporation with RNPs composed of recombinant Cas9 and synthetic dual guide RNA with chemical modifications. This strategy enables to achieve genome editing in mice with unprecedented simplicity. Although we did not perform side by side comparisons, it seems that this protocol outperforms the other methods that we have tested (Supplementary Fig. S5). Comparison of the mutation rates that we obtained with published results also supports the high performance of this new method12,13,22. In particular, the possibility to achieve HDR in the majority of the treated embryos is impressive. Although the cloning-free procedure does not eliminate mosaicism, which is a limitation common to all current methods, we observed the complete elimination of the wild-type allele in most experiments. In some cases, only one mutant allele was detected, suggesting that all cells were homozygous for the desired mutation. However, later analysis showed that the apparent homozygosity of somatic tissue biopsies does not necessarily imply homozygosity of the germline. If mosaicism has to be reduced, one possible improvement would be to perform electroporation at earlier developmental stage12. We also found that two different genes can be mutated independently in a single experiment, a non-negligible advantage on both ethical and financial points of view. Indeed, it reduces the number of animals needed and can significantly improve the throughput of genome editing facilities. Nevertheless, this strategy has limitation as multiplexing might increase the chances of generating chromosomal translocation23,24,25.
The other limitation of genome editing is the likely occurrence of off-target mutations, which we did not specifically addressed here. Although the occurrence of off-target mutations seems to be only partially predictable26, these can be kept infrequent by proper guide RNA design. Here we used the CRISPOR algorithm to optimize the design, without favoring efficiency over specificity. It should be stressed however that off-target mutations do not raise major problems for later analysis, provided that mouse phenotyping is performed only after generation F2. In this case, mutations localized far enough from on-targets will segregate during mouse mating. The current procedure can still be improved to limit the frequency of off-target mutations, since modified Cas9 nuclease with increased specificity27,28,29 become commercially available as recombinant proteins.
One attractive aspect of the cloning-free procedure is that it does not require any specific expertise in recombinant DNA manipulation and notably in in vitro transcription. Producing high quality guide RNA with bacteriophage RNA polymerases often appears as a limiting factor. RNA chemical synthesis also facilitates the introduction of chemical modifications which increase the genome editing efficiency by protecting dual guide RNA from degradation.
Dual guide RNPs electroporation of fertilized oocytes, with intact zona pellucida, is feasible with limited equipment, technical skills and financial resources. The technology is thus amenable to facilities which are familiar with embryo transfer, but not with molecular cloning or oocyte microinjection. Moreover, the remarkable efficiency of this method allows reducing the number of mice required to generate genetically modified models. It should also be easy to transpose to other mammalian species whenever embryo electroporation is possible18,30.
Methods
The research projects were approved by a local ethics committee (CECCAPP, registered as CEEA015 by the French ministry of research) and subsequently authorized by the French ministry of research. All procedures were in accordance with the European Community Council Directives of September 22, 2010 (2010/63/EU) regarding the protection of animals used for scientific purposes.
Embryo collection
Embryos from several genetic backgrounds were used: C57BL/6 J for Brcc3 gene edition, B6D1F1xFVB or FVBxFVB for Vash1 and Vash2 genes, B6.SJL-PtprcaPepcb/BoyCrl for Ctse gene and B6D2F1xFVB for Mct8 point mutations. Four or five weeks old female mice (Charles River, France) were superovulated by intraperitoneal (i.p.) administration of 5 IU of Pregnant Mare Serum Gonadotropin (PMSG, Alcyon, France), followed by an additional i.p. injection of 5 IU Human Chorion Gonadotropin 48 h later (hCG, Alcyon, France). Superovulated females were mated with adult males (1 male/2 females) and euthanatized at 0.5 d post coitum (usually between 10 and 11 am). Oviduct were dissected, and the ampulla nicked to release zygotes associated with surrounding cumulus cells into a 200 µL droplet of hyaluronidase (Sigma) in M2 solution (300 µg/ml, Sigma) under a stereomicroscope (Olympus SZX9). Zygotes were incubated for 1 min at room temperature and passed with a mouth pipette through 3 washes of M2 medium to remove cumulus cells. Zygotes were kept in M16 medium (Sigma) in a water jacketed CO2 incubator (5% CO2, 37 °C) until electroporation, which was usually performed between 2 to 3 pm.
Guide RNA design
Design was optimized to maximize cutting specificity and efficiency, using the online CRISPOR freeware (http://crispor.tefor.net/)31. We selected sequences (20 nt) which were unique in the genome, avoiding the presence of predicted off-target sites on the same chromosome as on-targets. Except for one case discussed below, all predicted off-targets were unlikely as they displayed at least 3 (one case) or 4 mismatches with on-target sequences, partly localized close to the PAM, which is more detrimental for off-target cleavage (Supplementary Table S2). The exception was for the achievement of MCT8 amino-acid substitution that left no latitude in choosing the guide RNA32. A predicted site for off-target mutation was located in an intronic sequence on the same X-chromosome as on-target, with 3 mismatches all residing distally to PAM (non-seed region). The likelihood of this unintended target to be cut was thus increased. Although this off-target site was localized 60 Mb away from on-target and thus could segregate during mice mating, we verified by PCR/Sanger sequencing that no off-target mutation was transmitted at this position (Supplementary Fig. S4).
Preparation of RNP components and ssODN
Recombinant SpCas9 (PNABio, reference CP02) was dissolved in 1 mM DTT (5 µg/µL; 30 µM). 2 µL aliquots were stored at −80 °C for up to one year. Custom RNA synthesis was ordered either from TriLink (tracrRNA; PAGE and HPLC purification) or from Eurogentec (crRNA; HPLC purification). 2′-O-methyl +3′phosphorothioate chemical modifications were introduced on nt 1 to 3 of crRNA and nt 69 to 71 of tracrRNA. All sequences are on Supplementary Table S1. crRNA and tracrRNA were dissolved in ultrapure water (2 µg/µL). 15 µL of a 1 µg/µL equimolar mix of tracrRNA+ crRNA (4.8 µL of tracrRNA + 2.7 µL of crRNA + 7.5 µL H2O) were prepared and filtrated on a Millex-GV 0.22 µm filter (Millipore). The absence of contaminating RNase in these dual guide RNA preparations was ascertained as follows: 2 µL of annealed RNA were diluted in H2O (1/10). Half of this sample was incubated at 37 °C for 1 h to favor possible contaminating RNase activity. The migration on a 2% agarose gel of this sample was then compared to the migration of the non-incubated control. Clear bands with equivalent intensity and without smearing should be visible for both samples. Dual-guide RNA preparations which passed the quality control test were aliquoted (4 µL) and stored at −80 °C. The design of the ssODN used as template for HDR was performed following published recommendations33: strand choice was such that ssODN sequence was complementary to crRNA 5′ end (copying the “non PAM strand”), and the position of the ssODN sequence was asymmetric with respect to the double strand break, displaced toward the 3′ region. Size of 5′ homology was 92 nt and 36 nt for 3′ homology. Phosphorothioate modifications were introduced at positions 1, 2, 3, 136, 137 and 138 to protect the ssODN from nucleases34. ssODN purified by PAGE (Eurogentec) was resuspended in ultrapure water (4 µg/µL) and filtrated on a Millex-GV 0.22 µm filter.
RNPs assembly
RNP assembly was performed just before electroporation. One aliquot of 4 µg RNA mix (4 µL) was incubated at 80 °C for 2 min for denaturation, and then at 37 °C for 10 min to favor annealing of the tracrRNA and crRNA into a dual guide RNA. 10 µg of Cas9 protein (2 µL) were added before additional 10 min incubation at room temperature for RNP complex formation. When two RNPs were electroporated, the amount of each dual-guide RNA was reduced by half. When needed, 4 µg (1 µL) of ssODN was finally included. The final volume was adjusted to 20 µL with Opti-MEM (Life Technologies). This provides enough RNPs solution for the electroporation of at least 3 batches of 20–30 zygotes each.
Electroporation of pronuclear-stage embryos with intact zona pellucida
We adapted a procedure consisting in repeated pulses of electroporation delivered by a NEPA 21 electroporator (NEPA GENE Co. Ltd., Chiba, Japan) to first damage the zona pellucida and then favor the intracellular entry of RNPs18. A glass chamber with 1 mm gap platinum plate electrodes (CUY5001P1–1.5, NEPA GENE Co. Ltd) was filled with 5 μL of RNPs-containing medium, with or without ssODN. Between 15 and 30 zygotes were aligned in the chamber, taking care to avoid contacts between zygotes and electrodes. Impedance was measured and maintained between 120 and 180 kΩ by liquid volume adjustment (reducing volume increases impedance). Four “poring pulses” were applied (40 V, 3.5 ms, interval 50 ms, 10% voltage decay, polarity+), followed by 5 “transfer pulses” (5 V, 50 ms, interval 50 ms, 40% voltage decay; alternating + and − polarity). Poring pulses worked equally well when voltage was between 30 V and 50 V (in the latter case, duration was reduced to 2.5 ms). Zygotes were then transferred into M16 medium and kept overnight in incubator. The embryos that reached the 2-cell stage (89 to 97%, Table 2) were transferred into the oviduct of B6CBAF1 (Charles River, France) pseudopregnant females (15–20 embryos per female). Embryo mortality, comparing the number of newborns to the number of transferred embryos, was similar to the one usually observed after microinjected embryo transfer, and was not significantly affected by the electroporation procedure (Table 2).
Genotyping
PCR amplifications were performed directly on alkaline lysates of toe clips. Briefly, each toe was incubated 30 min at 95 °C in 100 µL of alkaline solution (NaOH 25 mM, EDTA 0.2 mM, pH 12.0). Neutralization was performed by adding 100 µl Tris 40 mM, pH 5.0. Two microliters of lysates were analyzed by PCR (25 µL) with appropriate primers. Large deletions were identified by agarose gel electrophoresis. Small indels and point mutations were identified by Sanger sequencing of PCR products (Genewiz). For indels’ identification, Sanger electrophoretograms obtained from mosaic F0 mice were analyzed using the TIDE software35 which decomposes sequencing traces and estimates the respective contribution of each PCR fragment. Indel size that can be identified by TIDE is limited to 50 nt, therefore this software was not used for mice with bigger indels as detected by agarose gel electrophoresis. TIDE analysis becomes difficult when spacing between the nucleotides on the Sanger electrophoretogram is not constant. In this case, TIDE appends a “spacing not constant” message to the result.
Data availability
All data generated or analyzed during this study are included in this published article and its Supplementary Information file.
Change history
13 March 2018
A correction to this article has been published and is linked from the HTML and PDF versions of this paper. The error has not been fixed in the paper.
References
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826, https://doi.org/10.1126/science.1232033 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823, https://doi.org/10.1126/science.1231143 (2013).
Williams, A., Henao-Mejia, J. & Flavell, R. A. Editing the Mouse Genome Using the CRISPR-Cas9 System. Cold Spring Harbor protocols 2016, pdb top 087536, https://doi.org/10.1101/pdb.top087536 (2016).
Markossian, S. & Flamant, F. CRISPR/Cas9: a breakthrough in generating mouse models for endocrinologists. J Mol Endocrinol 57, R81–92, https://doi.org/10.1530/JME-15-0305 (2016).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nature protocols 8, 2281–2308, https://doi.org/10.1038/nprot.2013.143 (2013).
Hendel, A. et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat Biotechnol 33, 985–989, https://doi.org/10.1038/nbt.3290 (2015).
Aida, T. et al. Cloning-free CRISPR/Cas system facilitates functional cassette knock-in in mice. Genome Biol 16, 87, https://doi.org/10.1186/s13059-015-0653-x (2015).
Rahdar, M. et al. Synthetic CRISPR RNA-Cas9-guided genome editing in human cells. Proc Natl Acad Sci USA 112, E7110–7117, https://doi.org/10.1073/pnas.1520883112 (2015).
Schumann, K. et al. Generation of knock-in primary human T cells using Cas9 ribonucleoproteins. Proc Natl Acad Sci USA 112, 10437–10442, https://doi.org/10.1073/pnas.1512503112 (2015).
Jacobi, A. M. et al. Simplified CRISPR tools for efficient genome editing and streamlined protocols for their delivery into mammalian cells and mouse zygotes. Methods 121-122, 16–28, https://doi.org/10.1016/j.ymeth.2017.03.021 (2017).
Chen, S., Lee, B., Lee, A. Y., Modzelewski, A. J. & He, L. Highly Efficient Mouse Genome Editing by CRISPR Ribonucleoprotein Electroporation of Zygotes. J Biol Chem 291, 14457–14467, https://doi.org/10.1074/jbc.M116.733154 (2016).
Hashimoto, M., Yamashita, Y. & Takemoto, T. Electroporation of Cas9 protein/sgRNA into early pronuclear zygotes generates non-mosaic mutants in the mouse. Dev Biol 418, 1–9, https://doi.org/10.1016/j.ydbio.2016.07.017 (2016).
Wang, W. et al. Delivery of Cas9 Protein into Mouse Zygotes through a Series of Electroporation Dramatically Increases the Efficiency of Model Creation. Journal of genetics and genomics = Yi chuan xue bao 43, 319–327, https://doi.org/10.1016/j.jgg.2016.02.004 (2016).
Mashiko, D. et al. Generation of mutant mice by pronuclear injection of circular plasmid expressing Cas9 and single guidedRNA. Scientific reports 3, 3355, https://doi.org/10.1038/srep03355 (2013).
Horii, T. et al. Validation of microinjection methods for generating knockout mice by CRISPR/Cas-mediated genome engineering. Scientific reports 4, 4513, https://doi.org/10.1038/srep04513 (2014).
Menoret, S. et al. Knockin (KI) in rat zygotes with TALENs and CRISPR/Cas9 systems as protein forms. Transgenic Research 23, 883–883 (2014).
Jung, C. J. et al. Comparative Analysis of piggyBac, CRISPR/Cas9 and TALEN Mediated BAC Transgenesis in the Zygote for the Generation of Humanized SIRPA Rats. Scientific reports 6, 31455, https://doi.org/10.1038/srep31455 (2016).
Kaneko, T., Sakuma, T., Yamamoto, T. & Mashimo, T. Simple knockout by electroporation of engineered endonucleases into intact rat embryos. Scientific reports 4, 6382, https://doi.org/10.1038/srep06382 (2014).
Qin, W. et al. Efficient CRISPR/Cas9-Mediated Genome Editing in Mice by Zygote Electroporation of Nuclease. Genetics 200, 423–430, https://doi.org/10.1534/genetics.115.176594 (2015).
Hashimoto, M. & Takemoto, T. Electroporation enables the efficient mRNA delivery into the mouse zygotes and facilitates CRISPR/Cas9-based genome editing. Scientific reports 5, 11315, https://doi.org/10.1038/srep11315 (2015).
Gurumurthy, C. B. et al. GONAD: A Novel CRISPR/Cas9 Genome Editing Method that Does Not Require Ex Vivo Handling of Embryos. Current protocols in human genetics/editorial board, Jonathan L. Haines… [et al.] 88, Unit 15 18, https://doi.org/10.1002/0471142905.hg1508s88 (2016).
Kaneko, T. & Mashimo, T. Simple Genome Editing of Rodent Intact Embryos by Electroporation. PLoS One 10, e0142755, https://doi.org/10.1371/journal.pone.0142755 (2015).
Jiang, J. et al. Induction of site-specific chromosomal translocations in embryonic stem cells by CRISPR/Cas9. Scientific reports 6, 21918, https://doi.org/10.1038/srep21918 (2016).
Birling, M. C., Herault, Y. & Pavlovic, G. Modeling human disease in rodents by CRISPR/Cas9 genome editing. Mammalian genome: official journal of the International Mammalian Genome Society, https://doi.org/10.1007/s00335-017-9703-x (2017).
Choi, P. S. & Meyerson, M. Targeted genomic rearrangements using CRISPR/Cas technology. Nature communications 5, 3728, https://doi.org/10.1038/ncomms4728 (2014).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR-Cas nucleases. Nat Biotechnol 33, 187–197, https://doi.org/10.1038/nbt.3117 (2015).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science, https://doi.org/10.1126/science.aad5227 (2015).
Kleinstiver, B. P. et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495, https://doi.org/10.1038/nature16526 (2016).
Chen, J. S. et al. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Nature 550, 407–410, https://doi.org/10.1038/nature24268 (2017).
Tanihara, F. et al. Somatic cell reprogramming-free generation of genetically modified pigs. Science advances 2, e1600803, https://doi.org/10.1126/sciadv.1600803 (2016).
Haeussler, M. et al. Evaluation of off-target and on-target scoring algorithms and integration into the guide RNA selection tool CRISPOR. Genome Biol 17, 148, https://doi.org/10.1186/s13059-016-1012-2 (2016).
Paquet, D. et al. Efficient introduction of specific homozygous and heterozygous mutations using CRISPR/Cas9. Nature 533, 125–129, https://doi.org/10.1038/nature17664 (2016).
Richardson, C. D., Ray, G. J., DeWitt, M. A., Curie, G. L. & Corn, J. E. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat Biotechnol 34, 339–344, https://doi.org/10.1038/nbt.3481 (2016).
Renaud, J. B. et al. Improved Genome Editing Efficiency and Flexibility Using Modified Oligonucleotides with TALEN and CRISPR-Cas9 Nucleases. Cell reports 14, 2263–2272, https://doi.org/10.1016/j.celrep.2016.02.018 (2016).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res 42, e168, https://doi.org/10.1093/nar/gku936 (2014).
Acknowledgements
We thank Farida Henry and Agnès Couzon for technical help and animal care at mouse breeding and transgenesis facilities (AniRA-PBES, SFR Biosciences, Lyon, France). AniRA platforms belong to the national CELPHEDIA Infrastructure (http://www.celphedia.eu/). We thank Séverine Chautard (CIRI) and Frédérique Vossier (GIN) for technical assistance; Eric Denarier (GIN), Christophe Arnoult (Institute for Advance Biosciences, Grenoble), Annie Andrieux (GIN) and Sophia Djebali (CIRI) for helpful discussions. This work was supported by grants from SFR Biosciences (US8 INSERM, UMS3444CNRS, UCBL, ENS de Lyon) to SM; Agence Nationale de la Recherche (Thyromut2 program; ANR-15-CE14-0011-01) to FF; Ligue Contre leCancer Comités de l’Isère et de Savoie to MJM; Association pour la Recherche sur le Cancer (SFI20111204053) to MJM; European Research Council (ERC-2013-CoG_616986) to BP; Agence Nationale de la Recherche (ANR-13-JSV3-0002-01) to BP; Agence Nationale de la Recherche Agence Nationale de la Recherche (ANR-12-RPIB-0011) to JM; Ligue Nationale Contre leCancer Comité du Rhône to JM.
Author information
Authors and Affiliations
Contributions
M.T., C.B., D.L., B.P., S.M. organized and performed experiments. B.P., C.B., M.J.M., J.M., F.F. and S.M. designed experiments. F.F. and S.M. wrote the manuscript, which was reviewed and corrected by all authors.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A correction to this article is available online at https://doi.org/10.1038/s41598-018-22724-9.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Teixeira, M., Py, B.F., Bosc, C. et al. Electroporation of mice zygotes with dual guide RNA/Cas9 complexes for simple and efficient cloning-free genome editing. Sci Rep 8, 474 (2018). https://doi.org/10.1038/s41598-017-18826-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-18826-5
This article is cited by
-
Generation of a C57BL/6J mouse strain expressing the CD45.1 epitope to improve hematopoietic stem cell engraftment and adoptive cell transfer experiments
Lab Animal (2023)
-
Highly reliable creation of floxed alleles by electroporating single-cell embryos
BMC Biology (2022)
-
H3K27me3 at pericentromeric heterochromatin is a defining feature of the early mouse blastocyst
Scientific Reports (2022)
-
From genotype to phenotype: genetics of mammalian long non-coding RNAs in vivo
Nature Reviews Genetics (2022)
-
The cell cycle stage of bovine zygotes electroporated with CRISPR/Cas9-RNP affects frequency of Loss-of-heterozygosity editing events
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.