Abstract
In this paper, the enzymatic activity, substrate specificity and antibiofilm feature of bacteriophage dual-function tail proteins are presented. So far, tail tubular proteins A–TTPAgp31 and TTPAgp44-have been considered as structural proteins of Klebsiella pneumoniae bacteriophages KP32 and KP34, respectively. Our results show that TTPAgp31 is able to hydrolyze maltose as well as Red-starch. The activity of 1 µM of the protein was calculated as 47.6 milli-Units/assay relating to the α-amylase activity. It degrades capsular polysaccharides (cPS), slime polysaccharides (sPS) and lipopolysaccharide (LPS) of K. pneumoniae PCM 2713 and shows antibiofilm reactivity towards S. aureus PCM 519 and E. faecalis PCM 2673. TTPAgp44 hydrolyses trehalose and cPS of E. faecium PCM 1859. TTPAgp44′s activity was also observed in the antibiofilm test against P. aeruginosa PCM 2710 and B. subtilis PCM 2021. TTPAgp31 has been identified as α-1,4-glucosidase whereas, TTPAgp44 exhibits trehalase-like activity. Both proteins contain aspartate and glutamate residues in the β-stranded region which are essential for catalytic activity of glycoside hydrolases. The significant novelty of our results is that for the first time the bacteriophage tubular proteins are described as the unique enzymes displaying no similarity to any known phage hydrolases. They can be used as antibacterial agents directed against bacterial strains producing exopolysaccharides and forming a biofilm.
Similar content being viewed by others
Introduction
Biofilm forming pathogens are a widespread problem especially in medicine, food technologies, pharmaceutical and biohydrometallurgical industry. In the food industry, the biofilm-forming bacteria cause spoilage of the products due to the colonization of industrial equipment surface and food, leading to infections among consumers. Similarly, biofilm formed in the interior of water supply pipes results in the rise of human health risk and additionally contributes to metal corrosion and reduction of water flow. Biofilm formation on medical implant devices such as catheters and mechanical heart valves, is also a medical problem. All these aspects determine the widespread interest among researchers in the biofilm formation and functioning. In recent years attention has been paid to methods of recognizing and combating biofilm-forming microbial populations1,2,3.
Human pathogens associated with biofilm development include species of: Staphylococcus aureus, Staphylococcus epidermidis, Enterococcus faecalis, Enterococcus faecium, Escherichia coli, Proteus mirabilis, Klebsiella pneumoniae, Streptococcus viridans, Pseudomonas aeruginosa and Acinetobacter spp. These bacteria are the cause of chronic and recurrent infections (over 60% cases), particularly in the hospital environment4.
Klebsiella pneumoniae strains belonging to the Enterobacteriaceae family are widely distributed in the environment5. The genus Klebsiella comprises bacteria that cause different types of healthcare-associated infections, including pneumonia, bloodstream, urinary and intestinal tract infections, wound or surgical site infections, meningitis and neonatal sepsis6. An important virulence factor and a defense barrier of K. pneumoniae is polysaccharide layer, referred to as a capsule7. Inhibition of capsular polysaccharides (cPSs) synthesis enhances the phagocytosis of encapsulated bacteria by human neutrophils and consequently their elimination8. It has been shown that the larger the capsule produced, the more pathogenic the strain is7. Encapsulated K. pneumoniae also produces large quantities of cell-free PS (slime PS) which is secreted into the external environment and is another important virulence factor contributing to pathogenicity8. PSs are also a crucial structural component of bacterial biofilm, which promotes attachment to the surface and protects microorganisms from antimicrobials9. Therapeutic agents destroying bacterial capsular and/or slime PSs provide a useful adjunct to therapy against infections caused by encapsulated bacteria, especially if they are not chemically synthesized antibiotics7.
Bacteriophages are a promising alternative for this purpose. Their antibacterial efficiency is well documented10,11,12,13,14. However, they are still classified as high-risk medications due to the release of endotoxin into the bloodstream after bacterial cell lysis15. Therefore, the use of molecules such as hydrolyzing enzymes of phage origin provides better control over the therapeutic effects on the organism16,17. Some of the hydrolyzing enzymes are localized on bacteriophage tail. According to recent scientific data, 120 of the 160 different phage hydrolyses are coded in the same open reading frame as structural proteins (tail and fibers proteins). Therefore, they have been also considered as structural proteins, not as enzymes18.
Previously, we have reported the preliminary results of biological activity of the tail tubular protein A of the phage KP32 tail which is a product of a gene number 31 (TTPAgp31)19. This protein we called the dual-function protein due to its structural and hydrolytic features related to binding and hydrolysis of EPS obtained from K. pneumoniae PCM 2713.
In this paper, we show the results of the next biological tests of TTPAgp31 and for the first time we demonstrate the results of biological activity of the protein named TTPAgp44 which comes from KP34 bacteriophage tail and it is a product of a gene number 44.
Here we present our efforts to determine the substrate specificity of two TTPAs using versatile chromogenic polysaccharide substrates with well-defined chemical structures. We also demonstrate screening tests of TTPAs mentioned above displaying antibiofilm effect which is due to hydrolytic activity towards extracellular polysaccharides.
Results
Cloning, genes expression, proteins purification and analysis
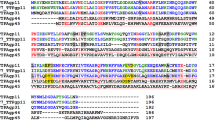
Tail tubular proteins A from K. pneumoniae bacteriophages were selected for study as potential secondary adhesive proteins (adhesins), which bind to receptors localized on the bacterial cell walls. These proteins belong to a group of homologous proteins present in a wide range of bacteriophages specific to several strains of Gram-negative and Gram-positive bacterial species. Both TTPAs are built of 192 amino acid residues however, their isoelectric points (Ip) are quite different, for TTPAgp31 the Ip is 4.26 and for TTPAgp44 is 8.13. BLAST amino-acid sequence analysis showed that the homology between them is about 21.5%, on the other hand, analysis using Phyre 2 server20 revealed that 88% of TTPAgp44 amino acids share the same structure as TTPAgp31. As we reported previously19, in the case of TTPAgp31 high amino acid sequence similarity (about 64%) to the gatekeeper protein (gp11) of T7 bacteriophage has been found, while TTPAgp44 is similar to gp11 only at 29%. Further analysis using a HHPred tool21 showed that both TTPAs share peptidoglycan hydrolase domain with probability about 40%.
The results of the protein expression and purification are presented in Fig. 1. As a result: 25 mg of TTPAgp31 and 20 mg of TTAgp44 from one liter of bacterial culture was obtained. The purified proteins were analyzed by SDS-PAGE. The electrophoretic analysis of TTPAgp31 (Fig. 1A) have been previously reported by Pyra et al.19 and the SDS-PAGE results for TTPAgp44 (Fig. 1B) are presented here for the first time.
Electrophoretic analysis of TTPAs proteins expressed in E. coli DE3plysS in 12.5% SDS-PAGE.TTPAgp31 before (line 2) and after (line 4) purification – The results have been previously reported by Pyra et al.19; 1) Low molecular mass standard (Bio-Rad), B) TTPAgp44 before (line 2) and after (line 3) purification on Ni2+-affinity chromatography, (1) molecular mass standard (Thermo Scientific).
Agar overlay test
Initial screening tests that assess the enzymatic activity of the phage proteins were agar spot assays on plates with mature bacterial lawns. Eighteen selected strains both from Gram-negative and positive group have been tested. TTPAgp31 showed its ability to create activity zone on K. pneumoniae PCM 2713, S. aureus PCM 519 and E. faecalis PCM 2673 lawn in the same test conditions. TTPAgp44 was active against thermoresistant E. faecium PCM 1859, P. aeruginosa PCM 2710 and B. subtilis PCM 2021. The activity zones were translucent halo (not clear plaques) and all sensitive strains have been able to produce slime as well as to form a biofilm. Moreover, strains: PCM 2673, 2710 and 2713 have been documented as clinical human isolates. Further analysis showed that TTPAs can also reduce bacterial biomasses of above strains. It was shown that 0.47 µM of TTPAgp31 can reduce about 80%, 60% and 50% of K. pneumoniae, E. faecalis and S. aureus strains’ biomass, respectively after one-day biofilm formation assay. While 0.47 µM of TTPAgp44 reduces about 80% of E. faecium biomass and about 40% of P. aeruginosa and B. subtilis strains; biomass in the same test conditions. The antibiofilm activity of the two phage proteins is shown in Fig. 2. To compare the spots created by both TTPAs, we have reused the figure of TTPAgp31 activity on agar plate (Fig. 2A) presented previously in19.
The activity of TTPAs towards bacterial strains. (A) overlay agar test towards Klebsiella pneumoniae PCM 2713 and (B) Enteroccocus faecium PCM 1859 lawn. In both cases 10 µl of 2 mM of the proteins were spotted on mature bacterial lawns and incubated at 37 °C overnight. (C) Antibiofilm activity of TTPAgp31 and TTPAgp44 (0.47 µM) (D) detected by TTC assay. The amount of biofilm formed after 20 h of incubation was estimated. Absorbance was measured at 540 nm following incubation with TTC (0.1%). All results are presented as averages of results from three independent replicates in three parallel trials. Error bars represent the means, standard deviations (GraphPad Prism version 5.00 for Windows, GraphPad Software, San Diego California USA, (www.graphpad.com)). The authors have reused the Fig. 2A previously reported in19.
Hydrolytic activity
The hydrolytic activity of the TTPAs was first performed towards disaccharides: α-lactose, β-lactose, trehalose, melibiose, cellobiose and maltose. Only maltose was hydrolyzed by TTPAgp31, while TTPAgp44 hydrolyzed trehalose. In the test conditions, TTPAgp31 released 35% of RSs with regards to the whole number of RSs obtained after acid hydrolysis (positive control). TTPAgp44 hydrolyzed trehalose with the efficiency of 87%. The hydrolyzing activity of the TTPAs towards disaccharides is shown in Fig. 3.
The hydrolytic activity of TTPAs toward disaccharide and PSs substrates. TTPAgp31 activity toward maltose and K. pneumoniae PCM 2713 PSs. *The given results are without the background of the negative control. TTPAgp44 activity toward trehalose and E. faecium PCM 1859 cPS. All results are presented as averages of results from three independent replicates in three parallel trials. Error bars represent the means, standard deviations (GraphPad Prism version 5.00 for Windows, GraphPad Software, San Diego California USA, (www.graphpad.com)).
Based on bioinformatics analysis, both proteins carry peptidoglycan hydrolase domain with the probability of 40%. To check their activity towards peptidoglycan pNP-GlcNAc as well as resorufin-labeled casein, chromogenic substrates were used. No activity of both TTPAs towards both substrates was observed.
Red-starch was tested as a next chromogenic substrate. Only TTPAgp31 was active against the polysaccharide showing almost as high activity as the α-amylase used in the assay as a positive control. The activity of the B. subtilis α-amylase (positive control) towards the Red-starch was calculated as 128.6 milli-Ceralpha U/ml, while TTPAgp31 as 112.8 milli-Ceralpha U/ml. It means that 1 µM of TTPAgp31 possess over 47 U of an amylase activity per 1 milliliter of the solution. The calculation was performed based on a protocol for assay of α-amylase using Red-starch from Megazyme22. Results of the experiment on Red-starch paper are presented in Fig. 4.
Hydrolytic activity of TTPAgp31 towards Red-starch. The activity of TTPAgp31 was compared to the B. subtilis α-amylase (positive control). The calculation was performed based on a protocol for assay of Alpha-amylase using Red-starch from Megazyme. All results are presented as averages of results from three independent replicates in three parallel trials. Error bars represent the means, standard deviations (GraphPad Prism version 5.00 for Windows, GraphPad Software, San Diego California USA, (www.graphpad.com)). Red-starch saturated filter paper treated with (3) TTPAgp31, (2) B. subtilis α-amylase, positive control, (1) phosphate buffer, the negative control.
The overlay test on the agar plates showed that TTPAgp44 was significantly active towards E. faecium (PCM 1859) strain. It means that TTPAgp44 possess antibacterial activity towards other bacterial strain than TTPAgp31 reported before19.
To check the hydrolyzing activity of both proteins against the bacterial polysaccharides (PS), cPS, sPS and LPS from the K. pneumoniae and cPS from the E. faecium strains, were isolated.
The acid hydrolysis has been performed to obtain the total amount of the RSs of the bacterial PSs. The RSs calculation was made using Nelson-Somogoi assay23. The RSs concentration for K. pneumoniae capsular and the slime PSs (100 μg of dry mass), as well as for LPS was calculated as 0.39 μM ml−1, 0.47 μM ml−1 and 0.29 μM ml−1, respectively. After the K. pneumoniae PS hydrolysis using TTPAgp31 the RSs in the samples were 0.12, 0.21 and 0.11 μM ml−1 for cPS, sPS and LPS, respectively. The calculation of the RSs release has been performed considering the RSs present in the control sample lacking TTPAgp31.
The hydrolytic activity of TTPAgp44 was tested towards cPS of E. faecium. The RSs amount released after incubation with the protein was 0.143 μM ml−1 and reached almost 50% of the total RSs obtained when the same amount of the PSs was subjected to acid hydrolysis (positive control). No such release was observed in the control sample lacking the phage protein.
Discussion
Bacteriophages are attractive and very effective antibacterial agents. It has been shown that phages degrade PSs and allow access to bacterial cell24,25,26,27. It is well-known that K. pneumoniae phages produce PS hydrolases which are associated with the tail spike and form characteristic halo zones in a spot agar plate assay28. In our studies, phage tail proteins named TTPAs belonging to KP32 and KP34 phages were tested in the screening assays on agar plates towards randomly selected bacterial strains, however, most of them have been isolated from human body and fluids. Their enzymatic activity displayed as the translucent halo zones creating on the bacteria lawns. The zones were not clear plaques meaning that these proteins didn’t lyse the bacterial cells, but destroyed the components localized outside the cells. In consequence, the translucent, but not clear zones were observed24. Interestingly, the proteins were active towards different bacterial strains. TTPAgp31 was active against K. pneumoniae (PCM 2713), S. aureus PCM 519 and E. faecalis PCM 2673, while TTPAgp44′s activity was observed towards E. faecium PCM 1859, P. aeruginosa PCM 2710 and B. subtilis PCM 2021. The antibacterial activity has been confirmed in antibiofilm assay. We used trimethyl tetrazolium chloride (TTC) assay that is versatile, high-throughput method of biofilm measure, applicable to a broad range of microorganisms29.
As we can see, although these proteins are classified to the same family of tail tubular protein A and have similar molecular masses they differ with respect to their physico-chemical features (such as Ip mentioned above) and the substrate specificity. That is in agreement with the previous observation that phages proteins have also different host specificities30. TTPAgp31 belongs to KP32 bacteriophage that is Kp32virus in the subfamily Autographivirinae (http://www.ictv.global/proposals-16/2016.023a-dB.A.v1.Kp32virus.pdf) and TTPAgp44 belongs to KP34 phage (Kp34likevirus in the subfamily Autographivirinae)30. Both phages come from Podoviridae family and are characterized as Klebsiella pneumoniae bacteriophages30,31. It must be also emphasized, that the substrate specificity of the proteins as separate molecules may differ to the specificity of the proteins which are combined together forming phage tail, by extension to the specificity of the bacteriophage particles.
In this paper, the hydrolytic activity of two TTPAs towards different saccharides substrates has been shown. In our previous paper, we have discussed the glucanase activity of TTPAgp31 that can be related to the presence of lectin-like domain and peptidoglycan hydrolase domain19. Despite low amino acid sequence homology between TTPAgp31 and TTPAgp44 (21%) the structural homology can be high as predicted by Phyre2 analysis (88% with 100.0% confidence) and both potential hydrolytic domains are present in TTPAgp44 as well. pNp-GlcNAc and resorufin-labeled casein as substrates were used to verify the N-acetyl-β-D-glucosaminidase and peptidase activity, respectively. These two enzymes are involved in peptidoglycan (PG) degradation, however, the PG hydrolysis is not limited only to those two enzymes (Moak and Molineux, 2004)32. Therefore, even if both TTPAs were not active against above substrates they should not be discriminated as PG hydrolases. PG consists of cross-linked amino acids and sugars which form alternating disaccharide of amino sugars: N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM). N-acetyl-β-D-glucosaminidase cleaves the N-acetylglucosaminyl-β-1,4-N-acetylmuramine bond, N-acetyl-β-D-muramidase (also termed lysozyme) cleaves N-acetylmuramyl-β-1,4-N-acetylglucosamine bond and N-acetylmuramyl-L-alanine amidase can hydrolyze the amide bond between the sugar and the peptide moieties. Endopeptidase hydrolyzes any of the peptide bonds between amino acids. Finally, the recently discovered γ-D-glutaminyl-L-lysine endopeptidase cleaves the gamma bond between D-glutamine and L-lysine residues33.
In the next experiments, disaccharides and Red-starch (according to Megazyme the substrate for α-amylase) have been used as substrates. TTPAgp31 hydrolyzed maltose and starch (better results were obtained at pH = 5.5–6.0 than at 8.0) suggesting the similar activity to α-glucosidase (EC.3.2.1.20) in terms of substrate specificity and pH range. α-Glucosidases are typical amylolytic hydrolases cutting the α-1,4 bonds releasing single glucose residue from the non-reducing end of substrates (oligosaccharides or polysaccharides). However, these enzymes are diverse in substrate specificity, optimum temperature reaction and transglucosylation activities34. α-Amylases belong to glycoside hydrolases (GH) number 13 and contain the catalytic site and subsides, which interact with single glucose residue in the substrate35. In general, differences in substrate specificity of the enzymes are explained by the difference in numbers of the subsites, a different affinity for glucose residues of each subsite, different amino acid composition and the differences in the catalytic35.
TTPAgp44 hydrolyzed trehalose breaking down the α,α-1,1-glycosidic bond that is characteristic for α,α-trehalase (glucohydrolase) (EC 3.2.1.28).
Previously19, for TTPAgp31 we have suggested the motif of D-X-D and/or D-X-E responsible for enzymatic activity of glycosyl hydrolases36. Our analysis let us suppose that the similar catalytic motif, which is D116-X-E118, is present in TTPAgp44.
The results of the hydrolytic activity tests towards K. pneumoniae PSs, TTPAgp31 hydrolyzed the capsular, slime PSs and the LPS. TTPAgp44 was active toward cPS of E. faecium. PSs of above bacteria usually contain mannose, galactose and glucose as predominant carbohydrate components which are linked via glycosidic bond and that are potential targets for TTPAs37,38. However, the substrate specificity depends on many factors, for example, a tertiary structure of the PSs39. In general, bacterial extracellular PSs can comprise homopolysaccharides and heteropolysaccharides synthesized at the cell membrane through processes involving nucleoside diphosphate sugars and isoprenoid lipid intermediates. The other group comprises levans and dextrans, formed essentially by extracellular processes without nucleoside diphosphate sugars or lipid intermediates, but dependent on the presence of specific substrates such as sucrose and closely related oligosaccharides39. Moreover, to understand the broad specificity of the proteins, we must emphasize that the TTPAs are the secondary phage adhesins and they come into action after a host recognition proses. Being a part of the phage tail, they are not naturally exposed to recognize the bacteria, unlike in the experiments described in this paper.
Conclusions
Bacteriophage tail tubular proteins A displaying enzymatic activity, are not well known as dual-function proteins. They have been considered only as structural proteins and they enzymatic features were not studied. In our previous paper19 and here we present the results of our research aimed to explain the biological function of TTPAs based on two proteins, TTPAgp31 and TTPAgp44.
Our research showed that both proteins have a high structural homology, but different substrate specificity. TTPAgp31 has been identified as α-1,4-glucosidase which hydrolyzes maltose as well as starch. TTPAgp44 possesses trehalase-like activity. As it is known, hydrolases are very diverse in terms of their molecular mass and specificity40 and almost 16 different combinations of the catalytic domains of these enzymes have been documented18.
But our studies let us suggest that both proteins contain D-X-E motif localized in the β-stranded region which is essential for catalytic activity of glycoside hydrolases.
Our results showed also that TTPAs might be useful in the treatment of infections caused by biofilm-forming bacteria. TTPAgp31 is very effective against clinical isolates of E. faecalis, P. aeruginosa and K. pneumoniae while TTPAgp44 is effective against P. aeruginosa, B. subtilis and thermoresistant E. faecium strain which is a very dangerous foodborne pathogen. It must be emphasiezed that both proteins possess antibacterial activity against different bacterial strains which means that they have different antibacterial specificity.
Our results show that the phage proteins being separate molecules in experimental conditions may possess not obvious enzymatic activity and for that reason, they should be litigious examined.
Materials and Methods
Bacterial strains
All of the bacterial strains used in the experiments were obtained from the Polish Collection of Microorganisms (PCM) of the Institute of Immunology and Experimental Therapy, Polish Academy of Sciences (Wroclaw, Poland), namely: Klebsiella pneumoniae (PCM 1; PCM 2713), Shigella flexneri (PCM 2336), Escherichia coli (PCM 172; PCM 185; PCM 195; PCM 2711), Proteus mirabilis (PCM 543), Citrobacter freundii (PCM 1562), Enterobacter aerogenes (PCM 532), Hafnia alvei (PCM 1223); Pseudomonas aeruginosa (PCM 499, PCM 2710), Enterococcus faecium (PCM 1858; PCM 1859), Enterococcus faecalis (PCM 2673), Bacillus subtilis (PCM 2021) and Staphylococcus aureus (PCM 502; PCM 2054, PCM 519). The strains were stored at −80 °C and cultivated in Luria-Bertani (LB) broth medium (Difco). Bacteria were cultured at 37 °C stationary or with shaking.
Cloning procedure
The genomes of phage KP32 and KP34 are deposited in the genomic database (GenBank): GQ413937 and GQ413938, respectively. TTPA encoding genes were selected for overexpression. Bacteriophage genes were obtained using polymerase chain reaction (PCR) with the following primers: TTPA from KP32 FW – GGATCCCATATGAACATGCAAGATGCTTAC, RV – GAATTCAAAGCTTACGACCGATGAGACCCT, TTPA from KP34 FW – GGATCCCATATGAGAGAACTTGATGCAATT, RV – GAATTCAAAGCTTAATACCATAAAACGAGCGCG.
The DNA of the bacteriophages was prepared as previously described41. PCR reactions were conducted using a two-phase program. The first phase consisted of seven and the second phase of 23 cycles. Taq polymerase (Fermentas) was used and the extension times for each gene were appropriate to gene length, min. 30 seconds to max. 2 minutes. Annealing temperature in the first phase was 48–52 °C and in the second phase 55–65 °C.
PCR products were cloned into the pGEM T-easy vector (T-vector, Promega) using T4 ligase. Constructs were transformed into E. coli DH5α bacteria using the heat-shock method and sequenced. Correct sequences were recloned into pET28a (Promega) expression vectors to obtain the phage tail proteins with an N-terminal six-histidine tag. Plasmid transformation into the competent E. coli BL21(DE3)plysS (Promega) cells was done using the heat shock method.
Expression and purification of phage tail proteins
Bacteriophage proteins were expressed in E. coli BL21(DE3)plysS strain (Promega). Bacterial clones were propagated in LB broth (37 °C with shaking) with kanamycin and chloramphenicol to reach OD600 = 0.8. Protein expression was performed using 0.05 mmol l−1 IPTG (Roche) as an inductor and followed by overnight incubation at 9 °C. Cells were pelleted and suspended in 50 mmol l−1 Tris/HCl pH = 8.0 lysis buffer containing 0.2 mol l−1 NaCl supplemented with protease inhibitor cocktail tablets (Roche). Cells were sonicated 8 times for 30 seconds with 1 minute breaks. After debris removal via centrifugation (14000 × g for 50 minutes) supernatant was mixed with Ni2+-agarose beads and incubated at 37 °C for 1 hour on a rotary shaker. After batch chromatography, the beads were washed using lysis buffer to remove unbound proteins. Bounded proteins were eluted using lysis buffer containing 250 mmol l−1 imidazole. Imidazole was removed from the protein’s solution via dialysis on centrifugal filters containing membrane (Millipore) with a cutoff of 3 kDa. The concentration of protein was determined using the BCA method described by Smith et al.42 and eluted fractions were analyzed by SDS-PAGE using 12.5% gels according to the method of Laemmli et al.43.
Agar overlay test
The hydrolytic activity of the phage proteins was determined using the spot assay described by Adams and Park (1956)24. The overnight cultures of bacteria were diluted to OD = 0.2 and pipetted onto agar plates. 10 µl of 47 µM of the proteins were spotted on the bacterial lawns and incubated at 37 °C overnight. The nutrient agar (pH = 7.2) composition was: beef extract (10 g) peptone (10 g) NaCl (5 g) and agar (20 g) suspended in 1000 ml and sterilized at 121 °C for 20 minutes.
Glycolytic hydrolase activity measurement towards saccharide substrates and bacterial PSs
The hydrolytic activity of the two proteins has been first demonstrated towards the disaccharide substrates such as: α-lactose, β-lactose, trehalose, melibiose, cellobiose, maltose and saccharose. 0.1 ml of disaccharide solution (5 mg/ml) was incubated with 0.1 ml of protein solution (47 µM dissolved in 50 mmol l−1acetate buffer, pH = 5.5) for 1 h at 37 °C on a rotary shaker.
Slime polysaccharide was precipitated using 3 volumes of 96% ethanol (4 °C, overnight) from the supernatant obtained after centrifugation of the one-day bacterial culture. The precipitated sPS was centrifuged (14000 × g for 30 minutes at 4 °C) then dissolved in water, dialyzed against water, frozen and lyophilized. The capsular polysaccharide was extracted from the freeze-dried bacterial mass using 10% trichloroacetic acid (TCA) according to the method described by Gorska-Frączek et al.44.
Lipopolysaccharide was extracted by hot phenol-water method45 with some modifications. 1 g of dry bacterial mass was suspended in the phenol-water mixture (25 ml 1:1 v/v). LPS was extracted at 65 °C for 15 minutes and next, after cooling to 5 °C it was pelleted by centrifugation at 2500 rpm for 15 minutes. The water fraction was dialyzed to water for three days, concentrated by ultrafiltration. In order to eliminate contaminating nucleic acids and proteins treatment with DNase, RNase and Proteinase K was performed. For this purpose, proteinase RNase (40 µg/mL) (Roche) and DNase (20 µg/mL) (Roche) in the presence of 1 µL/mL 20% MgSO4 were added to the mixture and kept at 37 °C overnight. Finally, the Proteinase K was added (100 µg/mL) (Roche) at 65 °C for 1 hour. To remove the contamination, the ultrafiltration was performed at 100 000 g for 6 hours. The pelleted LPS was suspended in miliQ water and the ultrafiltration was repeated 4 times. Final purified LPS product was lyophilized and stored at 4 °C.
The enzyme activity test was performed as follows: 60 μl of PS (0.1 mg ml−1) was mixed with 50 μl of the phage protein (47 µM dissolved in 50 mmol l−1acetate buffer, pH = 5.5). The reaction mixtures were incubated at 37 °C on a rotary shaker for 3 hours. The enzyme activity was evaluated using the reducing sugars (RSs) determination according to Nelson-Somogoi method (1952)23. The RSs amount was compared to the total amount of RSs released after PSs and the disaccharides acid hydrolysis (10 mol l−1 HCl, 85 °C, 25′). The negative control contained disaccharides/PS and buffer lacking the phage proteins. The acid hydrolysis was performed according to the method of Kubler-Kielb et al.46.
Enzymatic activity measurements toward chromogenic substrates
p-Nitrophenyl-2-acetamino-2-deoxy-β-D-glucopyranoside (pNP-GlcNAc) (Sigma-Aldrich), RedCL-Amylose (Megazyme) and Red-Starch (Megazyme) were used as substrates for TTPAs. 0.1 g of pNP-GlcNAc was dissolved in 10 ml of 50 mM sodium citrate pH 4.6 supplemented with 1% BSA. TTPAs (2 mM) were mixed with the substrate and incubated at 37 °C for 1 hour. The reaction was stopped using 0.4 M glycine buffer pH 10.4.
RedCL-Amylose (0.2 g) was suspended in 10 ml of 100 mM Bis-Tris Buffer pH 7.0. TTPAs (50 µl of 47 µM) were mixed with the substrate and incubated at 37 °C for 1 hour. The reaction was stopped using 96% ethanol.
0.6% Red-starch was made up in phosphate buffer pH 6.8. TTPAs (50 µl of 47 µM) were mixed with the substrate (0.95 ml) and incubated at 37 °C for 1 hour. The reaction was stopped using 96% ethanol. As a positive control 50 µl of α-amylase from Bacillus subtilis (Sigma-Aldrich) was used in the concentration of 50 µM.
The assay was also performed on Whatman filter paper grade 1 according to the procedure described by Martin et al.47. The filter paper was saturated with the starch solution, dried and then, 10 µl of 47 µM of TTPAs and Bacillus subtilis α-amylase (positive control) and phosphate buffer (negative control) were spotted on the Red-Starch paper and incubated at 37 °C for 15 minutes. Positive results were characterized by a marked color change from dark pink to white. For each test above, positive and negative controls were included.
Antibiofilm activity
One-day biofilm preparation: Bacteria were cultured overnight in 5 ml of LB broth at 37 °C with shaking. Overnight cultures were diluted to OD600 = 0.2 using fresh LB. 100 µl of bacteria suspension was inoculated into a 96-well plate (CytoOne) and incubated for 20 hours at 37 °C. After that time bacteria were removed and the plate was dried for 15 minutes up-side-down on a sterile paper towel. TTPAs were added (0.47 µM) and incubated at 37 °C for the next 24 hours. The next day, the OD600 was measured and after that 50 µl of 0.1% trimethyl tetrazolium chloride (TTC) was added to each well. After 1 hour of incubation at 37 °C the OD540 was measured on a Biotec microplate reader. Microtitre plate wells containing growth medium without any bacterial culture–sterility control; wells containing cell cultures but without TTPAs–control regarded as 100% cell mass. All trials were performed in triplicate and the mean value was calculated with the standard deviation range.
References
Vu, M. et al. Bacterial extracellular polysaccharides involved in biofilm formation. Molecules 14, 2535–2554 (2009).
Donlan, M. Preventing biofilms of clinically relevant organisms using bacteriophage. Trends Microbiol. 17, 66–72 (2009).
Sutherland, I. The biofilm matrix – an immobilized but dynamic microbial environment. Trends Microbiol. 9, 222–227 (2001).
Morin A. Screening of polysaccharide-producing microorganisms, factors influencing the production and recovery of microbial polysaccharides. Polysaccharides: structural diversity and functional versatility, Dumitriu, S. (Ed.), Marcel Dekker Inc. Publication, New York, 275–296, (1998).
Ahmad, T. A., El-Sayed, L. H., Haroun, M., Hussein, A. A. & El Ashry, E. S. Development of immunization trials against Klebsiella pneumoniae. Vaccine 30, 2411–2420 (2011).
Stock, I. & Wiedemann, B. Natural antibiotic susceptibility of Klebsiella pneumoniae, K. oxytoca, K. planticola, K. ornitinolithica and K. terrigena strains. J Med Microbiol 50, 396–406 (2001).
Domenico, P., Schwartz, S. & Cunha, B. A. Reduction of capsular polysaccharide production in Klebsiella pneumoniae by sodium salicylate. Infect Immun. 57, 3778–3782 (1989).
Domenico, P., Salo, R. J., Cross, A. S. & Cunha, B. A. Polysaccharide capsule-mediated resistance to opsonophagocytosis in Klebsiella pneumoniae. Infect. Immun 62, 4495–4499 (1994).
Karatan, E. & Watnick, P. Signals, regulatory networks, and materials that build and break bacterial biofilms. Mol Biol Rev. 73, 310–347 (2010).
Sulakvelidze, A., Alavidze, Z. & Morris, J. G. Bacteriophage Therapy. Antimicrob Agents Chemother. 45, 649–659 (2001).
Kucharewicz- Krukowska, A. & Slopek, S. Immunogenetic effect of bacteriophage in patients subjected to phage therapy. Arch. Immunol. et Therapie Experiment. 35, 553–561 (1987).
Cislo, M., Dabrowski, M., Weber-Dabrowska, B. & Woyton, A. 1987. Bacteriophage treatment of suppurative skin infections. Arch. Immunol. et Therapie Experiment. 35, 175–183 (1987).
Slopek, S., Weber-Dabrowska, B., Dabrowski, M. & Kucharewicz-Krukowska, A. Results of bacteriophage treatment of suppurative bacterial infections in the years 1981–1986. 1987. Ach. Immunol. et Therapie Experiment. 35, 569–583 (1987).
Weber-Dabrowska B., Mulczyk M., Gorski A. Bacteriophage Therapy of bacterial infections: an update of our institute’s experience. Arch Immunol. et Therapie Experiment. 48; 547–551,(200).
Peterson, J.W. Bacterial pathogenesis. Medical Microbiology, (4th edition, Baron, S. editor). (1996).
Steinbacher, S. et al. Crystal structure of phage P22 tailspike protein complexes with Salmonella sp. 0-antigen receptors. Biochem. 93, 10584–10588 (1996).
Brzozowska, E., Bazan, J. & Gamian, A. The functions of bacteriophage proteins. Adv. Hig. Exp. Med 65, 167–176 (2011).
Pires, D. P., Oliviera, H., Melo, L. D. R., Sillankorova, S. & Azeredo, J. Bacteriophage-encoded depolymerases: their diversity and biotechnological application. Appl. Microbiol Biotechnol 100, 2141–2151 (2016).
Pyra, A. et al. Tail tubular protein A: a dual – function tail protein of Klebsiella pneumoniae bacteriophage KP32. Scient. Rep. 7, 2223 (2017).
Kelly, L. A. & Sternberg, M. J. E. Subject Categories: Bioinformatics In silico modeling structural analysis Protein structure prediction on the Web: a case study using the Phyre server. Nature Protocols 4, 363–371 (2009).
Soding J., Biegerd A., Lupas A.N. The HHpred interactive server for protein homology detection and structure prediction. Nucleic Acids Res. 33, (2005).
Megazyme International Ireland Bray Business Park, Bray, Co. Wicklow, Ireland, www.megazyme.com.
Nelson, N. A photometric adaptation of the Somogyi method for the determination of glucose. J Biol Chem. 153, 375–380 (1944).
Adams, M. H. & Park, B. H. An enzyme produced by a phage host cell system. II. The properties of the polysaccharide depolymerase. Virol. 2, 719–736 (1956).
Hughes, K. A., Sutherland, I. W. & Jones, M. V. Biofilm susceptibility to bacteriophage attack: the role of phage-borne polysaccharide depolymerase. Microbiol. 144, 3039–3047 (1998).
Tait, K., Skillman, L. C. & Sutherland, I. W. The efficacy of bacteriophage as a method of biofilm eradication. Biofouling 18, 305–311 (2002).
Leiman, P. G. et al. The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities. J Mol Biol 371, 836–849 (2007).
Rieger-Hug, D. & Strim, S. Comparative study of host capsule depolymerases associated with Klebsiella bacteriophages. Virol. 113, 363–378 (1981).
Azeredo, J. et al. Critical review on biofilm methods. Critical Rev. Microbiol 43, 313–351 (2016).
Kęsik-Szeloch, A. et al. Characterising the biology of novel lytic bacteriophages infecting multidrug resistant Klebsiella pneumoniae. Virol. 10, 100–112 (2013).
Eriksson, H. et al. A suggested new bacteriophage genus, “Kp34likevirus”, within the Autographivirinae subfamily of Podoviridae. Viruses 7, 1804–22 (2015).
Moak, M. & Molineux, J. Peptidoglycan hydrolytic activities associated with bacteriophage virions. Molec. Microbiol. 51, 1169–1183 (2004).
Schmelcher, M., Donovan, D. M. & Loessner, M. J. Bacteriophage endolysis and novel antimicrobials. Future Microbiol. 7, 11-47–1171 (2012).
Nashiru, O., Koh, S., Lee, S. Y. & Lee, D. S. Novel a-Glucosidase from Extreme Thermophile Thermus caldophilus GK24. Biochem. Mol. Biol. 34, 347–354 (2001).
Bijttebier, A., Goesaert, H. & Delcour, J. A. Amylase action pattern on starch polymers. Biol. 63, 989–999 (2008).
Davies, G. & Henrssat, B. Structures and mechanism of glycosyl hydrolases. Curr. Biol. 3, 853–859 (1995).
Beles, P. M., Mijkovic, R., May, S. L., Shen, Y. & Nelson, D. C. Purification and characterization of biofilm-associated EPS exopolysaccharides from ESKAPE organisms and other pathogens. PlosOne. 8, 1–8 (2013).
Hancock, L.E., Murray, B.E. & Sillanpää, J. Enterococcal Cell Wall Components and Structures. (Gilmore, M.S., Clewell, D.B. & Ike, Y., et al. editors). Enterococci: From Commensals to Leading Causes of Drug Resistant Infection [Internet]. Boston: Massachusetts Eye and Ear Infirmary, (2014).
Sutherland I. Surface carbohydrates of the prokaryotic cell. Academic Press London, (1977).
Latka, A., Maciejewska, B., Majkowska-Skrobek, G., Briers, Y. & Drulis-Kawa, Z. Bacteriophage-encoded virion-associated enzymes to overcome the carbohydrate barriers during the infection process. Appl Microbiol Biotechnol. 101, 3103–3119 (2017).
Maciejewska, B. et al. Klebsiella phages representing a novel clade of viruses with an unknown DNA modification and biotechnologically interesting enzymes. Appl Microbiol Biotechnol. 101, 673–684 (2017).
Smith, P. K. et al. Measurement of protein using bicinchoninic acid. Anal. Biochem. 150, 76–85 (1985).
Laemmli Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685, (1970).
Górska-Fraczek, S. et al. The Structure and Immunoreactivity of Exopolysaccharide Isolated from Lactobacillus johnsonii Strain 151. Carbohyd. Res. 378, 148–153 (2013).
Westphal, O. & Jann K. Bacterial lipopolysaccharides. Extraction with phenol-water and further applications of the procedure. In: (Whistler, R. L. & Wolfan M. L. editors). Methods in Carbohydrate Chemistry. New York: Academic press; 83–91, (1965).
Kubler-Kielb, J., Coxon, B. & Chneerson, R. Chemical Structure, Conjugation, and Cross-Reactivity of Bacillus pumilus Sh18 Cell Wall Polysaccharide. J. Bacteriol. 186, 6891–6901 (2004).
Martin, N. C., Clayson, N. J. & Scrimger, D. G. The sensitivity and specificity of Red-Starch paper for the detection of saliva. Scient. Tech. 46, 97–105 (2006).
Acknowledgements
This research was supported by Wroclaw Centre of Biotechnology, programme The Leading National Research Centre (KNOW) for years 2014–2018 and by Wroclaw Research Center EIT+ under the project “Biotechnologies and advanced medical technologies – BioMed” (POIG 01.01.02–02–003/08–00) financed from the European Regional Development Fund (Operational Program Innovative Economy, 1.1.2).
Author information
Authors and Affiliations
Contributions
conceived and designed the experiments: E.B., A.P., Performed the experiments: E.B., M.J., N.U. Analyzed the data: E.B., A.P., and K.P. Contributed to the writing of the manuscript: E.B., A.P., S.G., Z.D.K. and A.G.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Brzozowska, E., Pyra, A., Pawlik, K. et al. Hydrolytic activity determination of Tail Tubular Protein A of Klebsiella pneumoniae bacteriophages towards saccharide substrates. Sci Rep 7, 18048 (2017). https://doi.org/10.1038/s41598-017-18096-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-18096-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.