Abstract
Dilated cardiomyopathy (DCM) is a primary cause of heart failure, life-threatening arrhythmias, and cardiac death. Pathogenic mutations have been identified at the loci of more than 50 genes in approximately 50% of DCM cases, while the etiologies of the remainder have yet to be determined. In this study, we applied whole exome sequencing in combination with segregation analysis to one pedigree with familial DCM, and identified a read-through mutation (c.2459 A > C; p.*820Sext*19) in the myosin light chain kinase 3 gene (MYLK3). We then conducted MYLK3 gene screening of 15 DCM patients (7 familial and 8 sporadic) who were negative for mutation screening of the previously-reported cardiomyopathy-causing genes, and identified another case with a MYLK3 frameshift mutation (c.1879_1885del; p.L627fs*41). In vitro experiments and immunohistochemistry suggested that the MYLK3 mutations identified in this study result in markedly reduced levels of protein expression and myosin light chain 2 phosphorylation. This is the first report that MYLK3 mutations can cause DCM in humans. The clinical phenotypes of DCM patients were consistent with MYLK3 loss-of-function mouse and zebrafish models in which cardiac enlargement and heart failure are observed. Our findings highlight an essential role for cardiac myosin light chain kinase in the human heart.
Similar content being viewed by others
Introduction
Dilated cardiomyopathy (DCM) is a genetic disorder that causes heart failure, life- threatening arrhythmias, and cardiac death1. Familial DCM accounts for 20–35% of all cases1, and mutations in more than 50 genes are known to cause the development of DCM2; however, the etiological causes remain unidentified in approximately 50% of DCM cases. Recently, whole exome sequencing (WES) has been used successfully to identify genetic abnormalities in relatively rare hereditary diseases3.
Cardiac myosin light chain kinase (cMLCK) encoded by the myosin light chain kinase 3 (MYLK3) phosphorylates cardiac myosin light chain 2 (MLC2, encoded by MYL2) and facilitates actin−myosin interactions that enhance contractile force in the animal heart4,5,6. In neonatal rat cardiomyocytes, overexpression of MYLK3 promoted sarcomere assembly, while knockdown disrupted sarcomere formation5. Furthermore, MYLK3 loss-of-function in mice and zebrafish induced cardiac dysfunction and heart failure4,7, suggesting that this kinase is indispensable for cardiac homeostasis.
In this study, we report the first DCM cases harboring MYLK3 mutations identified using WES. Sequencing analyses of unrelated DCM patients revealed an additional MYLK3 frameshift mutation. Our human genetic analysis suggests that MYLK3 is a novel pathogenic gene for DCM.
Results
Study population
A 16-year-old male (family A-III-1; Fig. 1a) diagnosed with DCM at 10 months of age was started on β-blocker and angiotensin-converting enzyme (ACE) inhibitor therapy. An endomyocardial biopsy of the proband was consistent with DCM and showed mild fibrosis (Fig. 1b). He exhibited substantially improved left ventricular (LV) function after a few years of treatment. The sister of the proband (family A-III-2; Fig. 1a) was diagnosed with DCM at 9 months of age. She was also started on β-blocker and ACE inhibitor therapy, and exhibited improved LV function after the initiation of treatment. The mother of the proband (family A-II-5; Fig. 1a) was diagnosed with DCM at 40 years of age. She showed relatively mild symptoms and a later disease onset compared to her children. The father of the proband (family A-II-4; Fig. 1a) had no symptoms of heart failure and echocardiography showed normal cardiac contraction.
WES analysis identified a read-through mutation in the myosin light chain kinase 3 gene (MYLK3) in all three affected individuals. (a) Familial pedigree of DCM patients analyzed by whole exome sequencing. Squares and circles represent males and females, respectively. A slash represents a deceased individual. Black-filled symbols indicate patients with a clinical diagnosis of DCM. Open symbols represent unaffected family members. The proband is indicated by an arrow. MYLK3 mutation carriers are indicated by a plus sign. (b) Endomyocardial biopsy findings of the proband revealed myocardial changes and fibrosis (Masson trichrome stain; scale bar, 50 µm). (c) Summary of variant filtering. The number of variants is described in each category. MAF, minor allele frequency. (d) Sanger sequencing analysis verified the presence of a MYLK3 read-through mutation in family A. The stop codon affected by this mutation is well conserved across evolution.
WES of the three affected patients with familial DCM
The inheritance pattern was thought to be autosomal dominant, and we performed WES on the three affected patients in family A (Figs 1a and 2). The median read depth in the targeted exonic region was 91 × , and 96.3% of the targeted exonic regions had over 20 × read depth. Variant filtering was conducted as shown in Fig. 1c, and yielded 19 rare variants that potentially affected protein function (Supplementary Table 1). We found no shared mutations in genes that were previously reported to be associated with DCM. Then, we reviewed the variants according to the guidelines for implicating sequence variants8. Among the genes, MYLK3 was highly expressed and functionally established in the heart, and was considered to be the only candidate likely to cause DCM. The variant was a read-through mutation (c.2459 A > C; p.*820Sext*19) resulting in a protein product with a 19-amino acid C-terminal extension not present in any population database. The variant was predicted to be deleterious based on its combined annotation-dependent depletion (CADD) score (18.79). Sanger sequencing confirmed that the variant was present in the three affected patients (Fig. 1d, Supplementary Fig. 1), but not in the unaffected father of the proband (family A-II-4; Fig. 1a), indicating that the variant segregated with the disease phenotype.
MYLK3 genetic screening
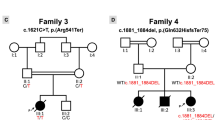
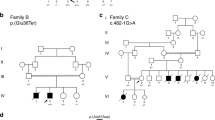
To examine whether other MYLK3 variants cause DCM, we first conducted targeted sequencing in our 52-patient unrelated DCM cohort and identified 15 patients (8 familial and 7 sporadic) with uncharacterized genetic etiology in known cardiomyopathy-related genes (Fig. 2, Supplementary Table 2). MYLK3 gene sequencing in these 15 patients identified an additional unique frameshift variant (c.1879_1885del; p.L627fs*41) in one familial DCM patient (family B-III-1; Fig. 3a and b). This frameshift was also predicted to be deleterious (CADD score of 35), resulting in a premature stop codon at E667 that likely disrupts both the catalytic and regulatory domains of the protein (Fig. 3c). Familial analysis revealed that the affected brother of the family B proband (family B-III-2; Fig. 3a) harbored the same variant. The proband and his brother were diagnosed with DCM at 56 and 52 years of age, respectively. Their cardiac function was improved after the initiation of treatment. One of their sisters (family B-III-3; Fig. 3a) had complained of dyspnea, but died in her twenties without an accurate diagnosis, so a DNA sample and detailed clinical information were not available. The mother of the proband (family B-II-2; Fig. 3a) also complained of dyspnea and palpitation, but her detailed clinical information and the presence or absence of DCM were also unclear. The father of the proband (family B-II-1; Fig. 3a) had no symptoms. We then performed WES on the two affected patients in family B and conducted variant filtering as described above to confirm the absence of additional pathogenic variants shared by these two affected patients (Supplementary Table 3).
MYLK3 gene sequencing analysis revealed a frameshift MYLK3 mutation in family B. (a) Familial pedigree of DCM patients harboring a MYLK3 frameshift variant. Squares and circles represent males and females, respectively. Slashes represent deceased individuals. Black-filled symbols indicate patients with a clinical diagnosis of DCM. Gray-filled symbols denote a suspicion of DCM. Open symbols represent unaffected family members. The proband is indicated by an arrow. MYLK3 mutation carriers are indicated by a plus sign. (b) Sanger sequencing analysis verified the presence of a MYLK3 frameshift mutation in family B. The amino acid residue altered by this mutation is well conserved across evolution. (c) MYLK3 domain structure. The stars represent the positions of the mutations identified in this study.
Overall, we identified two MYLK3 mutations in five DCM patients. The clinical characteristics of these patients are summarized in Table 1. Two patients (both in family A) developed severe DCM in infancy and the other three (one in family A and two in family B) developed adult-onset DCM. Life-threatening arrhythmia was not seen in any case. All of the patients with MYLK3 mutations identified in our study showed an improvement of cardiac function after the initiation of medical therapy.
Western blot analysis using MYLK3 constructs
We examined the effects of both MYLK3 mutations on protein expression. We transfected HEK293T cells with either wild-type or mutant MYLK3 (p.*820Sext*19 and p.L627fs*41) complementary DNA (cDNA) constructs. Western blot analysis using an antibody against residues 27–144 of human MYLK3 revealed significantly reduced mutant protein expression levels compared to the wild-type protein (p.*820Sext*19: 28.6% ± 7.8%, p.L627fs*41: 4.3% ± 1.7%; Fig. 4a and b). Cycloheximide (CHX) chase experiments revealed that mutant proteins were more rapidly degraded compared to the wild-type proteins, suggesting the instability of mutant proteins (Fig. 4c).
Western blot analysis suggested decreased expression levels of mutant cMLCK and MLC2 phosphorylation. (a) Expression of wild-type and mutant cMLCK protein in transfected HEK293T cells. Western blot analysis showed the reduced expression of mutant cMLCK protein. The blots were run under the same experimental conditions. The uncropped images are in Supplementary Fig. 2. (b) Densitometric quantification of western blots. Data are expressed as mean ± SEM (N = 3). *P < 0.05. (c) Wild-type or mutant MYLK3 transfected HEK293T cells were treated with cycloheximide (CHX) for indicated times. Western blot analysis showed rapid reduction of mutant proteins. Data are expressed as mean ± SEM (N = 3). *P < 0.05. The blots were run under the same experimental conditions. The uncropped images are in Supplementary Fig. 3. (d) Phos-tag SDS-PAGE followed by western blot analysis showed phosphorylated and non-phosphorylated forms of MLC2 using anti-FLAG antibody. Levels of phosphorylated MLC2 were remarkably reduced when co-transfected with mutant MYLK3 and MYL2 constructs. The ratio of phosphorylated MLC2 to total MLC2 (phosphorylated MLC2+ non-phosphorylated MLC2) was determined. Data are expressed as mean ± SEM (N = 3). *P < 0.05. The uncropped images are in Supplementary Fig. 4.
Analysis of MLC2 Phosphorylation
We performed Phos-tag9 SDS-PAGE followed by western blot analysis to determine the activity of mutant proteins on MLC2 phosphorylation. We co-trasnfected HEK293T cells with constructs of either wild-type or mutant MYLK3 and MYL2 with Myc-FLAG-Tag at C-terminal. Western blot analysis using an anti-FLAG antibody revealed that MLC2 phosphorylation levels were significantly reduced when co-transfected with mutant MYLK3 as compared with wild-type MYLK3 (Fig. 4d).
Immunohistochemistry of heart tissue
We performed immunohistochemistry of heart tissue from the patients to validate the effects of MYLK3 mutation on protein expression. The antibody against residues 27–144 of human cMLCK showed a strong and uniform staining pattern in the cardiomyocytes of a non-DCM patient or a DCM patient harboring an RBM20 mutation or a BAG3 truncation mutation in our cohort. However, a tissue from the proband in family A with the MYLK3 read-through mutation (p.*820Sext*19) showed significantly reduced cMLCK protein levels (Fig. 5a), although actin expression levels were comparable in all subjects (Fig. 5b).
Immunohistochemistry suggested decreased expression levels of cMLCK. (a) Expressions of cMLCK protein were examined in heart tissue by immunohistochemistry (scale bar, 50 µm.). Column 1 represents cardiac tissue from the proband of family A. Column 2 represents cardiac tissue from individual without cardiac disorders. Column 3 and 4 represents cardiac tissue from a DCM patient harboring an RBM20 mutation (without a MYLK3 mutation) and a BAG3 mutation (without a MYLK3 mutation), respectively. (b) Detection of actin protein in heart tissue. Expression levels of actin were comparable in all subjects.
Variants shared by early-onset DCM siblings in family A
Since the two siblings in family A showed an early-onset and severe DCM phenotype compared to the other three affected patients harboring MYLK3 mutations, we further analyzed whether the only two siblings in family A shared a second-hit variant. After variant filtering in the WES data, we found 14 rare variants that potentially affected protein function (Supplementary Table 4). Among these variants, a filamin C (FLNC) nonsense mutation (c.1444 C > T; p.R482*) was most likely to contribute to the severe phenotype because recent studies have suggested an association between FLNC truncation mutations and DCM10,11. Sanger sequencing confirmed that the variant was inherited from their unaffected father.
Discussion
In this study, we identified two mutations in MYLK3 that were associated with human DCM pathogenesis. WES analysis identified candidate pathogenic mutations in three affected individuals within one pedigree, and led to the discovery of a MYLK3 read-through mutation. Subsequent MYLK3 gene sequencing analysis revealed a MYLK3 frameshift mutation responsible for DCM in another family. Although the importance of cMLCK for proper cardiac function is strongly suggested by animal cardiomyopathy models in vivo and experimental data in vitro 4,5,7, no MYLK3 mutation has been identified as a pathogenic factor of human cardiomyopathy. Our present findings strongly suggest that MYLK3 mutations cause DCM, which emphasizes a pivotal role for cMLCK in human cardiac function.
The clinical characteristics of these five patients suggest that MYLK3-related DCM responds to pharmacological therapy and exhibits reverse remodeling. LV ejection fraction or end-diastolic diameter of all patients with MYLK3 mutations in this study improved significantly after the initiation of treatment. Responsiveness to medical therapy is associated with the prognosis of DCM12. None of the MYLK3-related DCM patients developed a life-threatening arrhythmia or required heart transplantation during follow-up. These findings suggest that the progression of MYLK3-related DCM could be slowed or prevented by appropriate medical treatment.
The two novel mutations in MYLK3 identified in this study were not present in any database and both were predicted to be deleterious in silico. One was a read-through mutation that causes the continued translation of the mRNA into the 3′-untranslated region, resulting in the production of an aberrant protein with a 19-amino acid extension. mRNAs lacking a stop codon and C-terminally extended proteins can be degraded more easily than wild-type molecules through a mechanism called non-stop decay or protein instability, leading to low protein expression13,14,15,16. In fact, more than 20 read-through mutations have been reported to cause inherited diseases15 and, for example, a case of heterozygous read-through mutation causing remarkable reduction of normal protein was reported17. These observations suggest that the read-through mutation identified in this study would not only create a simple extended protein but also would affect cMLCK protein expression levels. The other mutation was a frameshift predicted to create a premature stop codon, leading to nonsense-mediated decay or truncation with a disrupted catalytic domain (Fig. 3c). Western blot analysis revealed decreased expression levels of both mutant proteins compared to wild-type proteins (Fig. 4a and b) and CHX treatment showed instability of both mutant proteins (Fig. 4c). Phos-tag SDS-PAGE in combination with western blot analysis revealed the kinase activity of mutant proteins were lower than wild-type proteins (Fig. 4d). Furthermore, expression levels of cMLCK were significantly lower in the heart tissue from the proband of family A compared with that from the other DCM patients without a MYLK3 mutation or non-DCM patient (Fig. 5a). Therefore, we suggest that these mutations result in decreases of expression levels and activity of cMLCK.
MYLK3 hypomorphic mice exhibited reduced cMLCK expression and MLC2 phosphorylation as well as mild heart failure and fibrosis with a clinical course resembling human DCM, while cardiac development and sarcomeric maturation were normal7. Animal studies have shown that cardiac MLC2 phosphorylation increases actin−myosin interactions and cardiac contractile strength through the regulation of myosin head position, lever-arm stiffness, and myosin step-size18,19. MLC2 phosphorylation is reportedly reduced in end-stage heart failure20. These findings suggest that MLC2 phosphorylation by cMLCK is essential for normal cardiac function, and that the loss of cMLCK causes LV dysfunction and enlargement, consistent with our results.
Age of onset and disease severity varied substantially among our study patients, as in most previously reported cases of familial DCM21,22. Two siblings in family A developed DCM in infancy and the proband’s sister with suspected DCM in family B died in her twenties. Additionally, the two childhood-onset DCM patients in family A had severely impaired cardiac function at diagnosis. In this way, a wide spectrum of clinical features can be present21. This heterogeneity complicates the assessment of familial DCM. Previous studies have demonstrated that multiple variants in cardiomyopathy-related genes can be present in patients with DCM or other cardiomyopathies23,24. In these cases, patients harboring multiple variants are predisposed to early-onset and severe clinical phenotypes compared with those of other family members harboring only a single variant. In this study, we identified an additional FLNC nonsense mutation in two early-onset and severe DCM siblings of family A and this variant was inherited from their unaffected father. Reinstein et al. reported an early-onset DCM case with FLNC compound heterozygous mutations11. In their report, all relatives with only a nonsense mutation in the FLNC gene were completely normal as was the unaffected father in our family A, and an additional missense mutation was necessary to cause early-onset DCM. These findings exactly correspond to our observation that the coexistence of the MYLK3 and FLNC mutations could contribute to the early-onset and severe phenotype of DCM, highlighting the possibility of oligogenic effects on DCM. Furthermore, MYLK3 knockout mice have been reported to show a mild contractile abnormality under basal conditions and exhibit severe heart failure under cardiac stress25. Therefore, we consider that additional stresses including second-hit mutations could explain the heterogeneity of MYLK3-related DCM phenotypes, although further functional studies are required to prove this hypothesis.
Our results suggest that MYLK3 functions as the causal gene of DCM in two unrelated families, but there are some limitations in this study. First, the number of cases, especially of heart samples is limited. We assessed expression levels of cMLCK semi-quantitatively by immunohistochemical staining but not quantitatively by western blot analysis because there is not sufficient amount of heart tissues. Since the biopsy sample could not be obtained from any family members, immunohistochemical staining was performed only for the proband. It is also difficult to conclude that reduced expression levels of cMLCK are due to mutation or secondary to heart failure, since we did not examine expression levels of cMLCK in the heart of patients with mutation who do not show heart failure. Expression levels of mutated cMLCK were significantly lower than those of wild type cMLCK in our transfection experiments (Fig. 4a) and expression levels of cMLCK were well-preserved in DCM patients harboring an RBM20 mutation or a BAG3 mutation (Fig. 5). Although these results suggest that the reduced cMLCK expression in the patient with MYLK3 mutation is not secondary to heart failure, but due to a pathological consequence of mutated MYLK3, we need more heart samples to reach conclusion. To elucidate the mechanical linkage between gene mutation and heart phenotype, further studies using disease models such as zebrafish, mice, and induced pluripotent stem cells from patients are needed. Also, studies of larger multiethnic cohorts are necessary to validate the role of MYLK3 mutations in DCM pathogenesis and to examine specific genotype−phenotype associations.
In conclusion, we have first identified MYLK3 mutations associated with DCM in humans. This MYLK3-associated subtype appears relatively responsive to medical treatment, and our findings have important implications for the prognosis and treatment of DCM patients and their family members. Non-symptomatic MYLK3 mutation-carriers in a pedigree with familial DCM could profit from early genetic diagnosis, and preemptive pharmacological treatment from an early disease stage, thereby contributing to a better prognosis for such DCM patients. We also suggest that chemical modulation of cMLCK activity could be a potential therapeutic strategy for cardiomyopathy.
Materials and Methods
Subjects
The study protocol followed the ethical guidelines of the Declaration of Helsinki and was approved by the ethics committees of the Tokyo Women’s Medical University and the University of Tokyo. All subjects provided written informed consent and all experiments were performed in accordance with relevant guidelines and regulations. DCM diagnosis was based on the presence of LV dilatation and dysfunction in the absence of heart disorders such as hypertensive heart disease, primary valvular disease, and coronary artery disease. Genomic DNA was extracted from whole blood by standard procedures.
WES, targeted sequencing, and data analysis
Exome capture was performed with an Agilent SureSelectXT Human All Exon Kit V6 and a sequence library was prepared according to the manufacturer’s protocol for Illumina paired-end sequencing (Illumina, San Diego, CA, USA). Exome sequencing was performed on an Illumina HiSeq. 2500 using 100-bp paired-end reads. For target sequencing, we designed a panel of 95 genes (Supplementary Table 2) associated with DCM or other inherited cardiovascular diseases using SureDesign for Haloplex technology (Agilent Technologies, Santa Clara, CA, USA) that covered 99.4% of the exons of the target genes. Targeted sequencing was performed on an Illumina HiSeq. 2500 system in rapid-run mode, producing 150-bp paired-end reads.
For data analysis, filtered reads were mapped to the GRCh37/hg19 human reference genome with BWA-MEM26. The initial detection of variants was carried out using Surecall 3.0 (Agilent Technologies), which incorporates SAMtools27 and SNPPET (Agilent Technologies). We excluded intronic variants and synonymous mutations, as well as variants with minor allele frequency >0.01% in any freely accessible, ethnically-matched population database, such as the 1000 Genomes Project28, Exome Aggregation Consortium29, Human Genetic Variation Database (http://www.hgvd.genome.med.kyoto-u.ac.jp/index.html)30, Tommo database31, and our internal database of more than 800 exomes. Variants were also annotated with information from the Genome Aggregation database (gnomAD; http://gnomad.broadinstitute.org). Finally, mapped reads and called mutations were confirmed by inspection using the Integrative Genomics Viewer32. Deleterious variants were predicted in silico using a CADD score33, with a cutoff of 10. Function and tissue expression for each gene was investigated using database of pubmed (https://www.ncbi.nlm.nih.gov/pubmed), Mouse Genome Informatics (MGI; http://www.informatics.jax.org)34, GeneCards (http://www.genecards.org)35, and our internal database. The positions of MYLK3 mutations are based on NCBI RefSeq NM_182493.
MYLK3 gene sequencing
All exons and flanking introns of MYLK3 were amplified by PCR using the primer pairs listed in Supplementary Table 5. The products were then sequenced with a BigDye Terminator V3.1 kit (Applied Biosystems, Foster City, CA, USA).
DNA constructs
A full-length cDNA clone of human MYLK3 (NM_182493) with an N-terminal His Tag and an additional 57 C-terminal bases was purchased from Eurofins (Tokyo, Japan). Both p.*820Sext*19 and p.L627fs*41 mutations of MYLK3 were induced by site-directed mutagenesis using a QuikChange XL Site-Directed Mutagenesis Kit (Agilent Technologies). Wild-type and mutant constructs were subcloned into the pShuttle-CMV vector36 (a gift from B. Vogelstein). The specific primers for mutagenesis are shown in Supplementary Table 6. Introduction of the correct mutations was verified by DNA sequencing. A full-length cDNA clone of human MYL2 (NM_000432) with an C-terminal Myc-FLAG-Tag was obtained from Origene (Rockville, MD, USA).
Cell culture studies of HEK293T cells
HEK293T cells were grown in Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum and L-glutamine at 37 °C under a 5% CO2 atmosphere. The cells were transfected using Lipofectamine 3000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. In brief, the cells were seeded to reach 60−70% confluence at 24 h later, and then transfected with wild-type or mutated MYLK3 cDNA constructs. Total cell lysates were extracted at 36 h after transfection and analyzed by immunoblotting. For the protein stability experiments, medium containing 100 µg ml−1 of cycloheximide (Sigma) was added at 16 h after transfection and total cell lysates were extracted at indicated time points.
Protein extraction and immunoblot analysis
The cells were lysed in buffer containing 10 mM Tris-HCl, 150 mM NaCl, 5 mM EDTA, 1% Triton X-100, 1% sodium deoxycholate, 0.1% sodium dodecyl sulfate, complete protease inhibitor cocktail, and PhoStop EASYpack (Roche). After centrifugation at 15,000 rpm for 20 min at 4 °C, the supernatants were recovered and total protein concentrations were measured using a bicinchoninic acid assay (Pierce BCA Protein Assay Kit; Thermo Scientific). For immunoblot analysis, extracted proteins were separated on 4–15% Mini-PROTEAN TGX precast gradient gels (Bio-Rad) and transferred onto nitrocellulose membranes. The membranes were blocked with 2% nonfat dry milk powder in Tris-buffered saline plus 0.05% Tween and incubated overnight at 4 °C with an anti-MYLK3 antibody (1:500; Novus Biologicals, Littleton, CO, USA; cat# NBP1–86648) recognizing residues 27−144 of human cMLCK and an anti-β-actin antibody (1:5000; Sigma cat# A5441) as a loading control. Primary antibodies were detected with horseradish peroxidase (HRP)-conjugated species-specific secondary antibodies (GE Healthcare) and ECL plus (Thermo Scientific) using a LAS 3000 analyzer (GE Healthcare). Human cardiac tissue and non-transfected HEK293T cells were used as controls. Immunoblot band intensities were measured using ImageJ software (National Institutes of Health).
Analysis of the MLC2 phosphorylation
For the phosphorylation analysis, HEK293T cells were co-transfected with wild-type or mutated MYLK3 and MYL2 cDNA constructs. The cells were lysed at 16 h after transfection in buffer containing 50 mM Tris-HCl, 120 mM NaCl, 0.5% NP-40, complete EDTA free protease inhibitor cocktail, and PhoStop EASYpack and proteins were extracted as described above. The extracted proteins were separated by SDS-PAGE on 12.5% polyacrylamide gels containing 50 μmol/L Phos-tag (Wako Pure Chemical Industries, Japan) and 100 μmol/L ZnCl2. After electrophoresis, the gel was washed three times in transfer buffer containing 10 mM EDTA for 10 min. The proteins were then transferred onto nitrocellulose membranes. The membranes were blocked with 5% nonfat dry milk powder in Tris-buffered saline plus 0.05% Tween and incubated 1 h at room temperature with an anti-FLAG antibody (1:100; Sigma cat# A8592). Detection and measurement of band intensity were performed as described above. MLC2 phosphorylation ratio was calculated as follows: MLC2 phosphorylation = (phosphorylated MLC2)/(non-phosphorylated MLC2+ phosphorylated MLC2)
Immunohistochemistry
Immunohistochemistry was performed on paraffin-embedded 4 µm-thick tissue sections obtained from an endomyocardial biopsy of the proband in family A. Deparaffinization and rehydration were performed using a standard procedure with xylene and ethanol. To block non-specific binding, the sections were incubated for 5 min with normal horse serum (Vector Laboratories, La Jolla, CA, USA; cat# PK-7200). The samples were incubated for 1 h at room temperature with an anti-MYLK3 antibody (1:500; Novus Biologicals; cat# NBP1-86648) and an anti-muscle actin antibody (1:200; Dako; cat# M0635). The slides were incubated with biotinylated anti-mouse and anti-rabbit IgG secondary antibodies for 30 min and with the VECTASTAIN ABC reagent (Vector Laboratories; cat# PK-7200) for 30 min. Then, the slides were visualized with the ImmPACT Diaminobenzidine (DAB) Peroxidase (HRP) Substrate (Vector Laboratories; cat# SK-4105) followed by counterstaining with hematoxylin. For comparison, heart tissues of right ventricle from a 29-year-old male with no evidence of cardiac disorders and endomyocardial biopsy specimens from a 21-year-old male DCM patient with an RBM20 mutation (p.R634W), and a 50-year-old female DCM patient with a BAG3 truncation mutation (p.H166fs*6) in our cohort were used. They were confirmed not to have any MYLK3 mutations by Sanger sequencing. The positions of the RBM20 and BAG3 mutations are based on NCBI RefSeq NM_001134363 and NM_004281, respectively.
Statistical analysis
Data are shown as the mean ± standard error of the mean (SEM). Statistical analysis was performed with SAS software JMP version 11.0. Group means were compared by Student’s t-test, and P < 0.05 was considered statistically significant.
Data availability
Genetic data have been deposited at the Japanese Genotype−phenotype Archive (http://trace.ddbj.nig.ac.jp/jga), which is hosted by the DNA Data Bank of Japan, under accession number JGAS00000000110.
References
Yancy, C. W. et al. ACCF/AHA guideline for the management of heart failure: a report of the American College of Cardiology Foundation/American Heart Association Task Force on practice guidelines. Circulation. 128, e240–327 (2013).
McNally, E. M., Golbus, J. R. & Puckelwartz, M. J. Genetic mutations and mechanisms in dilated cardiomyopathy. J Clin Invest. 123, 19–26 (2013).
Boycott, K. M., Vanstone, M. R., Bulman, D. E. & MacKenzie, A. E. Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat. Rev. Genet. 14, 681–691 (2013).
Seguchi, O. et al. A cardiac myosin light chain kinase regulates sarcomere assembly in the vertebrate heart. J Clin Invest. 117, 2812–2824 (2007).
Chan, J. Y. et al. Identification of cardiac-specific myosin light chain kinase. Circ Res. 102, 571–580 (2008).
Davis, J. S. et al. The overall pattern of cardiac contraction depends on a spatial gradient of myosin regulatory light chain phosphorylation. Cell. 107, 631–641 (2001).
Ding, P. et al. Cardiac myosin light chain kinase is necessary for myosin regulatory light chain phosphorylation and cardiac performance in vivo. J Biol Chem. 285, 40819–40829 (2010).
MacArthur, D. G. et al. Guidelines for investigating causality of sequence variants in human disease. Nature. 508, 469–476 (2014).
Kinoshita, E., Kinoshita-Kikuta, E., Takiyama, K. & Koike, T. Phosphate-binding tag, a new tool to visualize phosphorylated proteins. Mol Cell Proteomics. 5, 749–757 (2006).
Ortiz-Genga, M. F. et al. Truncating FLNC Mutations Are Associated With High-Risk Dilated and Arrhythmogenic Cardiomyopathies. J Am Coll Cardiol. 68, 2440–2451 (2016).
Reinstein, E. et al. Congenital dilated cardiomyopathy caused by biallelic mutations in Filamin C. Eur J Hum Genet. 24, 1792–1796 (2016).
Merlo, M. et al. Prevalence and prognostic significance of left ventricular reverse remodeling in dilated cardiomyopathy receiving tailored medical treatment. J Am Coll Cardiol. 57, 1468–1476 (2011).
van Hoof, A., Frischmeyer, P. A., Dietz, H. C. & Parker, R. Exosome-mediated recognition and degradation of mRNAs lacking a termination codon. Science. 295, 2262–2264 (2002).
Tsuboi, T. et al. Dom34:hbs1 plays a general role in quality-control systems by dissociation of a stalled ribosome at the 3′ end of aberrant mRNA. Mol Cell. 46, 518–529 (2012).
Shibata, N. et al. Degradation of stop codon read-through mutant proteins via the ubiquitin-proteasome system causes hereditary disorders. J Biol Chem. 290, 28428–28437 (2015).
Arribere, J. A. et al. Translation readthrough mitigation. Nature. 534, 719–723 (2016).
Clegg, J. B., Weatherall, D. J. & Milner, P. F. Haemoglobin Constant Spring−a chain termination mutant? Nature 234, 337–340 (1971).
Wang, Y., Ajtai, K. & Burghardt, T. P. Ventricular myosin modifies in vitro step-size when phosphorylated. J Mol Cell Cardiol. 72, 231–237 (2014).
Sweeney, H. L., Bowman, B. F. & Stull, J. T. Myosin light chain phosphorylation in vertebrate striated muscle: regulation and function. Am J Physiol. 264, C1085–1095 (1993).
van der Velden, J. et al. Increased Ca2+ -sensitivity of the contractile apparatus in end-stage human heart failure results from altered phosphorylation of contractile proteins. Cardiovasc Res. 57, 37–47 (2003).
Hershberger, R. E. & Siegfried, J. D. Update 2011: clinical and genetic issues in familial dilated cardiomyopathy. J Am Coll Cardiol. 57, 1641–1649 (2011).
Crispell, K. A., Wray, A., Ni, H., Nauman, D. J. & Hershberger, R. E. Clinical profiles of four large pedigrees with familial dilated cardiomyopathy: preliminary recommendations for clinical practice. J Am Coll Cardiol. 34, 837–847 (1999).
Roncarati, R. et al. Doubly heterozygous LMNA and TTN mutations revealed by exome sequencing in a severe form of dilated cardiomyopathy. Eur J Hum Genet 21, 1105–1111 (2013).
Maron, B. J., Maron, M. S. & Semsarian, C. Genetics of hypertrophic cardiomyopathy after 20 years: clinical perspectives. J Am Coll Cardiol 60, 705–715 (2012).
Warren, S. A. et al. Myosin light chain phosphorylation is critical for adaptation to cardiac stress. Circulation. 126, 2575–2588 (2012).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv. 1303, 3997 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics. 25, 2078–2079 (2013).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature. 526, 68−74 (2015).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 536, 285–291 (2016).
Higasa, K. et al. Human genetic variation database, a reference database of genetic variations in the Japanese population. J Hum Genet. 61, 547–553 (2016).
Nagasaki, M. et al. Rare variant discovery by deep whole-genome sequencing of 1,070 Japanese individuals. Nat. Commun. 6, 8018, https://doi.org/10.1038/ncomms9018 (2015).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Bello, S. M. & Eppig, J. T. Inferring gene-to-phenotype and gene-to-disease relationships at Mouse Genome Informatics: challenges and solutions. Journal of Biomedical Semantics 7, 14 (2016).
Stelzer, G. et al. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr Protoc Bioinformatics 54, 1.30.1–1.30.33 (2016).
He, T. C. et al. A simplified system for generating recombinant adenoviruses. Proc. Natl. Acad. Sci. USA 95, 2509–2514 (1998).
Acknowledgements
This work was supported by grants from the Japan Foundation for Applied Enzymology (to S.N.), SENSHIN Medical Research Foundation (to S.N.), KANAE Foundation for the Promotion of Medical Science (to S.N.), MSD Life Science Foundation (to S.N.), The Tokyo Biomedical Research Foundation (to S.N.), a Grant-in-Aid for Young Scientists (B) (to S.N.), and the Japan Agency for Medical Research and Development 16ek0109069h0103 (to H.A.) and 16e0109009h0003 (to I.K.). We thank the patients and family members for participating in this study. We would also like to thank K. Shiina, S. Kawanabe, and A. Nonaka for assistance with the experiments; T. Suzuki, M. Shimizu, A. Suzuki, M. Matsuo, M. Ito, M. Satoh, Y. Furutani, T. Shiga, K. Saito, and T. Nakanishi for acquiring informed consent.
Author information
Authors and Affiliations
Contributions
T.T., S.N., H.A. and I.K. designed the study and interpreted the results. T.T. and T.F. analyzed the data. T.T., T.K., H.T. and K.U. performed the experiments. H.M., N.H., H.A. and I.K. provided overall guidance throughout the study. T.T., S.N. and I.K. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tobita, T., Nomura, S., Morita, H. et al. Identification of MYLK3 mutations in familial dilated cardiomyopathy. Sci Rep 7, 17495 (2017). https://doi.org/10.1038/s41598-017-17769-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-17769-1
This article is cited by
-
Pangenomics of the cichlid species (Oreochromis niloticus) reveals genetic admixture ancestry with potential for aquaculture improvement in Kenya
The Journal of Basic and Applied Zoology (2023)
-
Striated preferentially expressed gene deficiency leads to mitochondrial dysfunction in developing cardiomyocytes
Basic Research in Cardiology (2023)
-
Regulation of myosin light-chain phosphorylation and its roles in cardiovascular physiology and pathophysiology
Hypertension Research (2022)
-
Translating emerging molecular genetic insights into clinical practice in inherited cardiomyopathies
Journal of Molecular Medicine (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.