Abstract
The Asian honeybee Apis cerana is one of two bee species that have been commercially kept with immense economic value. Here we present the analysis of genomic sequence and transcriptomic exploration for A. cerana as well as the comparative genomic analysis of the Asian honeybee and the European honeybee A. mellifera. The genome and RNA-seq data yield new insights into the behavioral and physiological resistance to the parasitic mite Varroa the evolution of antimicrobial peptides, and the genetic basis for labor division in A. cerana. Comparison of genes between the two sister species revealed genes specific to A. cerana, 54.5% of which have no homology to any known proteins. The observation that A. cerana displayed significantly more vigilant grooming behaviors to the presence of Varroa than A. mellifera in conjunction with gene expression analysis suggests that parasite-defensive grooming in A. cerana is likely triggered not only by exogenous stimuli through visual and olfactory detection of the parasite, but also by genetically endogenous processes that periodically activates a bout of grooming to remove the ectoparasite. This information provides a valuable platform to facilitate the traits unique to A. cerana as well as those shared with other social bees for health improvement.
Similar content being viewed by others
Introduction
Of the ten honeybee species in the genus Apis, only the multiple-comb bee species, the European honeybee Apis mellifera and the Asian honeybee Apis cerana, have been commercially kept for crop pollination with immense economic and ecological value1,2,3,4. Nevertheless, in recent decades, there have been escalating concerns that populations of honeybees are in sharp decline as a result of the loss of habitat, introduction of invasive species, emergence of new pathogens and parasites, persistence of chemical residues and other environmental threats5,6,7,8,9. One of the most serious threats to honeybees is Colony Collapse Disorder (CCD), a mysterious plague that decimated entire bee colonies during the winter of 2006–2007 in the US and yet remains unsolved10. Hence, a genome-based approach to understanding the molecular mechanisms that regulate population dynamics of honeybees has been an integral part of pollinator conservation efforts. The complete genome sequence of A. mellifera has provided the first insights into honeybee biology and evolution11. To deepen our understanding of the complex social behavioral traits of honeybees, genome sequences of related species will enhance the understanding of the genetic basis associated with adaptive changes within and between honeybee species.
A. cerana is endemic to Asia with six morphologically distinct subspecies3,12,13 distributed throughout a series of climatic zones on the Asian landscape and has been used for pollination and commercial beekeeping over thousands of years in Asia. As close relatives, A. cerana and A. mellifera are very similar in morphology and behavior. Despite this, A. cerana has several distinct biological characteristics when compared with A. mellifera. For example, workers of A. cerana ventilate their hive with their heads pointing toward the outside which is the opposite of A. mellifera workers which fan with their head toward the entrance; foragers of A. cerana are good at collecting nectar from scattered floral resources that are often neglected by foragers of A. mellifera; A. cerana does not collect propolis, a resinous material used by A. mellifera to seal apertures in the hive and defend against pathogens. Moreover, as an indigenous species of Asia, A. cerana has evolved a series of striking biological characteristics to combat the adverse climatic conditions of their habitats. Foragers of the Chinese black honeybee (A. c. cerana) visit the flowers of Eurya spp under cloudy conditions when the air temperature is as low as 7 °C, a temperature at which the A. mellifera enters into a torpid state. A. cerana also has developed a diverse set of defense mechanisms to combat the invasion of predators, parasites and pathogens14,15. Compared to the steady and clumsy flight of A. mellifera, the flight of A. cerana is rapid, hasty and unpredictably zigzagging, which helps in escaping from hornets and bee-eating birds3. A. cerana is the native host to the ectoparasitic mite Varroa destructor16 which is the single most detrimental pest of A. mellifera, and has evolved high resistance to this pest over a long period of mutual adaptation17. Its workers can effectively remove mites from both adults and brood by performing a series of cleaning behaviors18,19.
Now many regions of Asia are suffering from a shortage of pollinators for plants and reduction of biodiversity3,20,21. The parasites and pathogens that have caused serious disease problems in A. mellifera were also found in A. cerana22. Therefore, it is urgent to take measures to protect A. cerana. Even so, compared to the wealth of genetic information available from A. mellifera which is used as a key model animal for social behavior11,23, the knowledge of A. cerana is limited.
Here we present our complementary efforts of a high-quality assembly and annotation of the southern strain of A. cerana genome and detailed analyses of its evolutionary history of domestication and selection. In particular, we identify and describe the genome variations that regulate different aspects of worker behavior and physiology between A. cerana and A. mellifera. Most importantly, we annotate the genome by coupling RNA-Seq analysis and laboratory assays to test gene functions and provide compelling experimental evidence of the molecular mechanisms underlying the Varroa mite resistance characteristics of A. cerana. These data provide a novel insight into honeybee biology and might be utilized to improve bee health globally.
Results
Genome sequencing and assembly
We sequenced 23.7 giga-bases (Gb) of Illumina pair-end reads, 8.7 Gb of Illumina mate-pair reads, and 636 mega-bases (Mb) of 454 GS FLX shotgun sequences using genomic DNA extracted from two haploid drone pupae from one A. cerana colony (Supplementary Table 1). A total of 21,784 contigs of 209.2 Mb, ranging from 500 bp to 174,956 bp with N50 of 21,160 bp were assembled. All contigs were constructed into 879 scaffolds with a total length of 228.8 Mb and N50 of 1.39 Mb (Supplementary Table 2), coinciding with the estimated genome size of 226 Mb calculated by k-mer (Supplementary Fig. 1).
A comparison with the A. mellifera genome (V4.5) demonstrated consistency between two genomes (Supplementary Fig. 2). Transcriptome sequencing of a sample of mixed brains of A. cerana workers produced 469,162 ESTs with an average length of 390 bp and 99% of the ESTs could be mapped on the A. cerana genome, suggesting the A. cerana genome covered most genes. The assessment of A. cerana genome coverage using 469,162 ESTs showed that the accumulative numbers of mapped EST aligned over 100%, 90%, 80%, 70%, 50%, and 20% of their length was 308128, 451855, 462797, 465583, 467197, and 467952, respectively. The accumulative mapped ratio aligned over 100%, 90%, 80%, 70%, 50%, and 20% of their length was 65.68%, 96.31%, 98.64%, 99.24%, 99.58%, and 99.74%, respectively.
Depending on a SNP-based A. cerana linkage map24 and A. mellifera DH4 linkage group, we linked 370 A. cerana scaffolds into 16 chromosomes, with a total size of 214.19 Mb (Fig. 1, Supplementary Table 3, Supplementary Fig. 3).
The atlas of A. cerana chromosomes and comparison to A. mellifera. The 16 chromosomes of A. cerana and A. mellifera genomes were drawn in the upper and bottom part, respectively. From outer side to inside, each circle represents the chromosome, genome homology, gene density, GC content and the translocation between the two species.
We found only 4.2% of the genome could be defined as repeat region, mainly composed of microsatellites (2.63%) (Supplementary Fig. 4). A total of 104,983 microsatellites were revealed in the genome, with an average distance of 1,948 bp and 10 repeat units in each microsatellite. Among them, (AT)n and (AG)n occupied 25% and 19%, respectively.
Genome characterization and annotation
By combining a gene prediction program based on EST alignments (Gnomon) and three ab initio prediction programs (GeneMark.hmm, Augustus and Snap) based on the A. mellifera model, we identified 10,182 protein-encoding genes in the A. cerana genome, with an average gene size of 7,577 bp and an average CDS size of 1,695 bp. The largest CDS was 27,474 bp long (ACC_02761), comprising 67 exons and encoding a myosin light chain kinase. The average exon and intron sizes were 237 bp and 988 bp, respectively (Table 1). The intron size of A. cerana is 250 bp smaller than that of A. mellifera, explaining the contradiction of bigger CDS length but smaller gene size for A. cerana (Supplementary Fig. 5). A total of 571 genes were predicted without introns (Supplementary Table 4).
BLASTP searches revealed that 9,911 (97.4%) of A. cerana proteins have homology with other known proteins. InterproScan assigned IPR domains to 8,767 genes, and 7,343 proteins were assigned GO terms. 3,849 proteins have Kyoto Encyclopedia of Genes and Genomes (KEGG) orthologs with 2,237 of them involved in at least one pathway. Eukaryotic Orthologous Groups (KOG) analysis assigned 7,713 A. cerana genes into different KOG categories, and 13.97% of these genes were involved in diverse signaling pathways, a similar ratio as in other insects like ants and bees, coinciding with the complex behavior of A. cerana. To study the gene roles of A. cerana genes in different stages, we performed RNA-seq for 16 different A. cerana samples (Supplementary Table 5).
CpG and DNA methylation
The A. cerana genome had an overall GC content of 32.97%. Based on observed CpG/expected CpG ratio, CpG dinucleotide distribution in A. cerana genes showed a di-peak pattern (Supplementary Fig. 6), with 2,577 genes under-represented (0.53-fold, higher methylated genes) and 2,297 genes over-represented (1.15 fold, rarely methylated genes) compared to the expectation from mononucleotide frequencies (Supplementary Information). It is known that the CpG deficiency in mammalian genomes represents a mutational hot spot through deamination of methylcytidine to thymidine.
There were a total of 793 lower CpG content genes and 248 higher CpG content genes that were involved in different pathways and showed significantly different enrichment (Supplementary Table 6). Lower CpG content genes showed significant enrichment in N-Glycan biosynthesis and protein processing in the endoplasmic reticulum (Supplementary Fig. 7) suggesting that genes of the two pathways were subject to germline methylation and subsequent deamination from a genomic history. N-glycans in invertebrate allergens (like bee venom) are a common cause of immunoglobulin (Ig) E cross-reactivity in vitro and are commonly known as cross-reactive carbohydrate determinants (CCDs). Higher CpG contentgenes showed significant enrichment in neuro-active ligand-receptor interactions, Cell Adhesion Molecules (CAMs) and insect hormone biosynthesis (Supplementary Fig. 8).
Comparison with other insects and unique genes
We compared six social insects including A. cerana, A. mellifera, A. florea, Bombus impatiens, Camponotus floridanus and Harpegnathos saltator with five non-social insects Drosophila melanogaster, Anopheles gambiae, Megachile rotundata, Bombyx mori and Nasonia vitripennis at the proteome level. By searching the Conserved Domain Database (CDD) with RPS-BLAST at E-value of 1e-5, we uncovered 2,010 domains conserved across all of the insects, and three domains uniquely to A. cerana (Table 2, Supplementary Table 7).
A total of 4,844 genes of A. cerana are confirmed as single copy, with less than 30% coverage and 20% identity with other genes in the whole proteome. The gene distribution in pathways was studied in detail (Supplementary Table 8). Non-social insects had more KEGG Orthologies (KOs) (144) than social insects (130) for the immune system, consistent with previous suggestions that the reduction of immune related genes at the individual level in honey bees may be a result of strengthened collective colony-level immunity (social immunity)11,25,26. However, the new evidence indicated that xenobiotic detoxification and immune genes are similarly depauperate in both honeybees and bumblebees which have colony lifestyles lying between completely solitary bees and highly eusocial species such as the honey bees27. The number of immune genes remains consistent regardless of bees’ degree of sociality suggests that a depauperate immune repertoire precedes evolution of sociality in bees and the differences in selection on immune genes likely reflect divergent pressures exerted by parasites across social contexts27,28.
Among the social insects, we only identified one unique KO (K12796) relative to non-social insects, which encoded the erbb2-interacting protein and was involved in the NOD-like receptor signaling pathway. Domain comparison and KOG analysis showed that there is no significant statistical difference between A. cerana and A. mellifera or A. florea.
To better understand the evolutionary position of A. cerana, 155 single copy genes with the best hit to another 10 species were chosen to create a phylogenetic tree. It is clear that A. cerana and A. mellifera were located together, and showed a close relationship with the other three species of Apoidae. Relative to ants (C. floridanus and H. saltator), the parasitoid wasp (N. vitripennis) showed a more distant relationship with A. cerana (Fig. 2, Supplementary Table 9).
8,202 orthologous gene pairs were identified between A. cerana and A. mellifera using Bidirectional Best Hit (BBH). By measuring the synonymous substitution rates (Ks) of orthologous gene pairs, we obtained the mean Ks of 0.149 and the peak Ks of 0.095. The mean sequence identity between A. cerana and A. mellifera homologous genes are 96.6%, with intron regions showing 92.5% identity. In addition, only 19 genes out of these 8,202 orthologs were under positive selection (Supplementary Table 10).
A comparison of A. cerana genes against genes of A. mellifera and A. florea revealed that 433 genes are specific to A. cerana, with 408 of them validated by RNA-seq data (Supplementary Table 5), though 236 genes (54.5%) have have no homology to any known proteins in the database like NR, KEGG and KOG (Supplementary Table 11). These A. cerana unique genes might contribute to the specific characteristics of the A. cerana. For example, ACC_04782 and ACC_08219 encode proteins involved in sensory perception of smell and sensory perception of taste, respectively; three NADH-ubiquinone oxidoreductase genes, which participate in oxidative phosphorylation, are unique in A. cerana compared with A. mellifera.
Three unique genes (ACC_00190, ACC_01436, and ACC_04893) encode MLE (maleless) which is one component of the dosage compensation complex, and the encoded MLE proteins have no homology at the amino acid level with A. mellifera or A. florea. Dosage compensation is a genetic regulatory mechanism, which balances systems of higher eukaryotes’ genome by equalizing the phenotypic expression of an unequal copy of sex chromosomes in the two sexes29. Five components are the core subunit of the dosage compensation complex (DCC) in Drosophila30, including MLE, MSL1, 2, 3 (male-specific lethal 1, 2 and 3) and MOF (males absent on the first). Unlike A. mellifera, lacking some component of X-specific dosage compensation11, the A. cerana genome has a full set of genes encoding the components of DCC. Genes, msl1, 2 and 3 (ACC_01543, ACC_01587 and ACC_04118), are all detected in A. cerana. We identified five genes (ACC_04472, ACC_00332, ACC_05236, ACC_01279 and ACC_02931) as Mof genes with histone acetyltransferase activity according to previous reports by Gu et al. (2000) and Hilfiker et al. (1997)31,32. According to KOG analysis, a total of 18 genes encode for MLE, of which three of them are unique for A. cerana. There are three NADH-ubiquinone oxidoreductase genes in A. cerana, and all of them are unique genes when compared with A. mellifera. The NADH-ubiquinone oxidoreductase gene participates in the process of oxidative phosphorylation.
Homeobox proteins
A total of 114 homeobox protein genes were revealed in the genome (Supplementary Table 12), including six genes encoding Paired box (Pax) proteins which are involved in the visual system development of the metazoan33. Among them, two Pax-6 genes, eye gone (eyg) and eyeless (ey) were known to act cooperatively in promoting eye development34, with eyg controlling eye growth and ey controlling eye specification35. Their duplicate genes, twin of eye gone (toe) plays only a subtle role in the Drosophila eye imaginal disc36, and twin of eyeless (toy) is required for initiation of ey expression in the embryo and acts through ey to activate eye development37. Phylogenetic analysis (Supplementary Fig. 9) revealed two A. cerana proteins (ACC_02837 and ACC_09296) located within the same clade with Eyg and Toe, indicating a possible role in eye growth by promoting cell proliferation. This is supported by the gene expression profile analysis which revealed that ACC_02837 only expressed in the pupal stage. Another A. cerana protein, ACC_01021, is located in the same branch as Ey and Toy, indicating its main role might be involved in retinal specification36.
Apitoxin
Apitoxin, or honeybee venom, is a complex mixture of proteins/peptides that cause an allergic reaction in humans. The main components of apitoxin include melittin, phospholipase A2 (PLA2), hyaluronidase, apamin, histamine, dopamine, noradrenaline38, acid phosphatase39,40, CUB serine protease, and Api m 541. We revealed 73 apitoxin candidate genes in the A. cerana genome (Supplementary Table 13), including the above-mentioned compounds and homologues of allergens from other species, like snake venom vascular endothelial growth factor toxin (ACC_00031). Among these components, phospholipase A2 (ACC_09897) is the most lethal42, together with hyaluronidase (ACC_10171), acid phosphatase (ACC_02702) and Api m 5 (ACC_09285), mainly contributing to IgE-mediated allergic reactions39,41. Melittin (ACC_10176), considered as a minor allergen, is the most abundant, occupying 50% of the venom38.
Though phylogenetic analysis had shown that A. cerana is most closely related with A. mellifera, the melittin gene (Api m 4) of A. cerana shows 99% identity with that of Vespa magnifica while only 93% identity with A. mellifera, which in turn, showed 100% identity with Vespula maculifrons prepromelittin (Fig. 3). A test for positive selection was performed on the five genes (Api m 1-Api m 5) encoding the major allergens and melittin (Supplementary Table 14). Noteworthy, the Api m 5 gene experienced position selection (dn/ds = 2.25) during the sequence divergence between A. cerana and other bees. We did not identify the complete coding sequence of another minor allergen Api m 643 in the A. cerana genome. We compared the genomic structure of Api m 6 locus in A. mellifera with the corresponding region of A. cerana genome (Supplementary Fig. 10) and found that the 2nd exon of Api m 6 was disrupted in the A. cerana genome, perhaps resulting in a degenerate gene. The residues 37–91 of Api m 6 encoded a trypsin inhibitor-like cysteine rich (TIL) domain, whose coding region was not affected by the disruption.
Immune System
Honeybees, like all insects, lack an adaptive immune system and have evolved an “innate immunity” to fight against foreign invaders. The immune system of A. cerana consists of 144 genes and 12 pathways (Supplementary Table 15), with six more genes and three more KOs than A. mellifera (Supplementary Table 8). Among them, 46 genes constitute the chemokine signaling pathway, which regulates inflammatory immune response by transducing chemokine signals through G-protein-coupled receptors (GPCR) expressed on the immune cells. There are 43 GPCR genes in the A. cerana genome (Supplementary Table 16) associated with neuropeptides, vision, odor, etc.
Adenylate Cyclase (AC) → cyclic adenosine monophosphate (cAMP) → Protein Kinase A (PKA) cascade is associated with many intracellular signaling pathways, such as calcium signaling, insulin secretion, circadian entrainment, and chemokine signaling pathway. A. mellifera has seven AC genes44, while A. cerana has eight AC genes, and seven of them are membrane-bound and one soluble (ACC_10007). AC8 is prominently expressed in the mushroom bodies45 which play important roles in some forms of associative learning and memory formation, such as olfactory memory45. Besides PKA-C3 (ACC_04702), AC8 (ACC_01531, membrane-bound) was highly expressed in worker pupae (p < 1e-4), suggesting that AC8 might be highly expressed in mushroom bodies in the pupal stage of workers for promoting olfactory system formation.
Fc gamma R-mediated phagocytosis plays an essential role in host-defense mechanisms through the uptake and destruction of infectious pathogens. The 36 candidate genes which involve in recognition of infectious pathogens through Fc gamma receptors and gamma R-mediated phagocytosis were identified in A. cerana, suggesting phagocytosis could be another important immune defense mechanism of A. cerana. Arp2/3 complex is highly conserved in almost all eukaryotes and has the capability to initiate actin filament formation46,47. In the A. cerana genome, the actin related protein 2/3 complex was intact and included two actin-related proteins, Arp2 (ACC_00048) and Arp3 (ACC_01435), and five subunits (ARPC1–5, ACC_01746, 00828, 08843, 09274, 07187). ARPC3, ARPC5, and Arp2, were all highly expressed in the dance stage (p < 1e-15) and ARPC5 was also highly expressed in the forage stage (p < 1e-25). Knockout of genes encoding ARPC3 leads to a defect of sustained migration directionality and a delay in wound closure in mouse fibroblast48. Perhaps the Arp2/3 complex is crucial for damage healing.
Honeybees are affected by a wide array of xenobiotic compounds. Three enzyme families, the cytochrome P450 - (P450), glutathione transferases (GST) and carboxy/cholinesterases (CCE) catalyze a wide range of detoxification reactions. A. cerana has 27, 52 and 12 genes encoding CCE, P450 and GST enzymes (Supplementary Table 17), respectively, with a similar number of genes found in A. mellifera44.
We found seven A. cerana genes in A. cerana encoding apidaecin, hymenoptaecin, defensin, apisimin, and abaecin, respectively (Supplementary Table 18). While all these antimicrobial peptide (AMPs) genes were also found in A. mellifera, it had been reported that A. cerana expresses many more transcripts of AMPs than A. mellifera49, suggesting that A.cerana might exhibit a stronger protecting ability against pathogens through encoding variable AMPs.
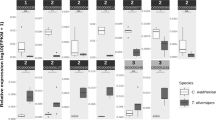
RNA-seq was performed to determine the AMP gene expression profiling in the brain of A. cerana nurses, guard bees, and foraging bees. The newly-emerged workers were set as a control. There were differences in the patterns of the AMP gene expression among A. cerana workers specializing in different tasks (Supplementary Fig. 11). For both abaecin and defensin2, the expression levels increased with age. The remarkable increase happened from nursing to guarding for abaecin while that from guarding to foraging for defensin2 (Supplementary Fig. 11). The mRNA levels of hymenoptaecin, apidaecin and defensin1 fluctuated with caste transition. Of them, nearly the same pattern was seen in hymenoptaecin and apidaecin gene expression and their transcriptions both peaked at the guarding caste then decreased sharply to a low level. However, for defensin1, its highest level of expression was seen in the nursing stage (Supplementary Fig. 11).
Genes underlying complex grooming and body shaking behaviors
Hygienic and grooming behaviors are important defensive mechanisms of honeybees against the parasitic mite V. destructor. The grooming behavior involves the ability of bees to shake, swipe, and bite Varroa mites from their bodies while the hygienic behavior involves the ability of bees to detect, uncap, and remove mite-infested brood17,50,51. Compared with A. mellifera, A. cerana possesses desirable behavioral traits including body shaking, rolling and tomb uncapping when they are infested by Varroa mites17,52, in addition to the behavioral strategies of grooming and hygienic behavior15. The results of our bioassay for grooming effectiveness towards Varroa mites confirmed that A. cerana and A. mellifera exhibited significant differences in their behavior responses to Varroa mites (Table 3). A. cerana workers responded immediately and vigorously to the presence of Varroa mites and displayed an average of 122 times of intensive body shaking, 228 times of light to moderate body shaking, 36 times of self-grooming, and 9 times of allogrooming within five minutes. In most of the cases, mites were finally removed from the body of A. cerana workers. Meanwhile, A. mellifera workers showed insignificant responses to the presence of Varroa mites in comparison to A. cerana. Their responses were more limited to the light to moderate body shaking and self-grooming, an average of 22.6 times and 5.93 times, respectively. The same, effective behaviors were rarely observed in A. mellifera workers.
From the transcriptomic study of A. cerana workers exhibiting Varroa-induced grooming and body shaking behaviors, 502 genes were significantly (FDR < 0.001) up-regulated and 11 were down-regulated in workers displaying grooming behaviors in response to the presence of Varroa mites relative to the control bees without Varroa (Supplementary Table 19). Meanwhile, 48 genes were up-regulated and 60 were down-regulated in the heads of body-shaking bees (Supplementary Table 20).
GO and pathway enrichment analysis suggested that up-regulated genes in grooming bees were enriched in DNA replication and repair, while down-regulated genes were enriched in response to biotic stimulus (GO:0009607) in both grooming and body-shaking bees (Supplementary Table 21). Six genes were significantly down-regulated in both grooming and body-shaking bees, including genes encoding antimicrobial peptides (apidaecin and hymenoptaecin), one pheromone-binding protein-related protein, one major royal jelly protein, and protein mab-21. Protein mab-21 is a homeotic regulator found in eukaryotes, which appears to be involved in the determination of cell fate. It is cell-autonomously required for specifying the identity of sensory ray 6 in the Caenorhabditis elegans male tail, and also for backward locomotion, normal body morphology, fecundity, and embryonic morphogenesis53. Of the 27 candidate genes identified influencing A. mellifera grooming behavior54, Ataxin-10 and a gene encoding gamma-tubulin complex component were up-regulated in A. cerana allogrooming workers. However, none of these candidate genes were found to be up-regulated in body-shaking bees.
Of 513 genes with altered expression in response to Varroa infestation, (Supplementary Table 19), the expression of 40 genes involved in signal recognition, modulation, and transduction as well as immune responses were confirmed by qRT-PCR in Varroa -challenged A. cerana that exhibited grooming and body shaking behavior and in A. cerana without exposure to Varroa. The qRT-PCR corroborated with the RNA-seq data (Table 4).
Chemoreceptor genes
Odorant binding proteins (OBPs) and chemosensory proteins (CSPs) are two gene families that transport odorant molecules through the sensillum lymph to the olfactory receptors on the membrane of chemosensory neurons. The A. cerana genome encodes only 14 OBPs (Supplementary Table 22), less than the 21 OBPs of A. mellifera. Correspondingly, 81 olfactory receptor (Or) genes were revealed in A. cerana, less than half the number of A. mellifera (163) but similar to the number of A. gambiae (79). Twenty-one of these tandemly arrayed genes were located in a 109 kb region, and showed 28%-79% identity to each other on the protein level, indicating a possible gene duplication. We also determined two CSP genes (ACC_03947 and ACC_09381) and 10 gustatory receptor (Gr) genes in the A. cerana genome.
Learning and Memory
The learning and memory performance of A. cerana is significantly better on both color and grating patterns than that of A. mellifera55. Protein sequences of learning and memory genes in mice, Drosophila, and other species were used to search homologous genes in the honeybees. Sixty-five and 50 homologous genes involved in learning and memory were identified in A. cerana and A. mellifera, respectively (Supplementary Table 23). Moreover, most of those learning and memory genes in higher mammals have a corresponding homolog in A. cerana and A. mellifera, indicating that the molecular regulation mechanisms of learning and memory are highly conserved from the lower insects to higher mammals56,57. The learning and memory process needs to convert extracellular stimuli into intracellular signals. The formation of long-term memory and synaptic plasticity needs to activate many signal pathways of neurons, such as cAMP-PKA, MAPK, and CaMK IV pathways58. Those signal transduction pathways converge on the cyclic AMP response element-binding protein (CREB), which is a transcription factor affected by cyclic AMP in the cell (Supplementary Fig. 12).
Division of labor
Analysis of RNA-seq data revealed that 66% of the A. cerana genes (6,722 of 10,182 genes) had significant expression differences (fold change > 2, FDR > 0.001) in the brain of newly emerged workers, nurses, guards, foragers, and dancers (Supplementary Fig. 13, Supplementary Tables 24 and 25). Enrichment analysis revealed that GO terms namely “energy metabolism”, “carbohydrate metabolism” and “amino acid metabolism” were significantly enriched (FDR < 0.05) in up-regulated genes in the brain of foragers and dancers compared to that of younger nest workers (Supplementary Table 26).
Intriguingly, foraging bees had a higher brain energy metabolism rate than in-hive workers (Fig. 4, Supplementary Information, Supplementary Tables 27 and 28). Of 147 genes belonging to “energy metabolism”, 109 (74%) and 87 (59%) genes were up-regulated in scouts and foragers relative to nurses; whereas 89 (61%) and 58 (39%) of genes were up-regulated relative to guard bees. Pathway analysis of the same data showed that the oxidative phosphorylation pathway, whereby a cell generates most adenosine triphosphate (ATP) during respiration, was significantly enriched (FDR < 0.001) in up-regulated genes in foragers and dancers relative to nurses and guard bees (Supplementary Table 27). The overall brain energy metabolic rate in different phenotypes could be arranged as: dancer > forager > newly-emerged > guard > nurse. Newly-emerged workers had a higher capacity for ATP production than the other in-hive workers, which not only helps to facilitate the bee to chew open the cell cap during emergence but also is required for nerve cell growth.
Changes in brain energy metabolism during worker maturation elucidated by pathway analysis of data from brain RNA-seq experiments. Gene expression data from nurse, forager and scout brains were mapped to citrate cycle pathways compiled by the Kyoto Encyclopedia of Genes and Genomes. Genes in the tricarboxylic acid pathway (shown) and other energy metabolism pathways were predominantly upregulated in foragers and scouts brains relative to nurses. Solid lines indicate the predicted enzymatic reactions catalyzed by the product of each gene and dotted lines indicate indirect links to other metabolic pathways. ACLY: ATP citrate (pro-S)-lyase; ACO: aconitate hydratase; CS: citrate synthase; DLAT: Dihydrolipoyllysine-residue acetyltransferase;DLD: dihydrolipoamide dehydrogenase;DLST: dihydrolipoamide succinyltransferase; FH: fumarate hydratase; IDH: isocitrate dehydrogenase; MDH: malate dehydrogenase; OGDH: 2-oxoglutarate dehydrogenase E1 component; PC: pyruvate carboxylase; PDH: Pyruvate carboxylase; PEPCK:phosphoenolpyruvate carboxykinase; SDH: succinate dehydrogenase; SUCLA: succinate–CoA ligase(ADP-forming); SUCLG: succinate–CoA ligase (GDP-forming).
Pathway analysis of the RNA-seq data revealed that 59% of the insulin/TOR network genes (Supplementary Fig. 14) were up-regulated in the brain during the transition from nurses to foraging workers (18 of the 44 genes were up in foragers, 19 up in dancers, and 11 up in both). Furthermore, the brain expression of the insulin/TOR/PI3K network was positively correlated with main energy metabolism pathways in addition to several transcription and cell cycle related pathways (Supplementary Information). This result indicates that the insulin/TOR signaling played a central role in linking cellular energy status to changes in behavior and in controlling the process of behavioral maturation, perhaps via regulation of cellular energy status and protein synthesis, which consequently affects nerve cell growth. The genome of A. cerana contains the same gene set for insulin-related neuropeptides and receptors as in A. mellifera, which includes two insulin-like peptides (ILPs), two ILP receptors (InRs), an adipokinetic hormone (AKH), and an AKH receptor (AKHR). The level of mRNA of these genes displayed an age- dependent manner, with foraging bees showing much higher brain expression than nurses (Supplementary Table 28, Supplementary Fig. 14). Brain mRNA levels for ILPs, InRs, and AKH increased ~2.3-fold as nurses turned into guard bees, and then stayed at a higher level thereafter (Supplementary Table 28). The brain expression level of AKH was much higher than its receptor gene, and the mRNA concentration of AKHR was just above detection limits. Because AKH is synthesized and stored in the corpora cardiaca, a neurohemal organ adjacent to the brain59, we speculated that this neuropeptide hormone might execute biological functions mainly in other tissues after being released into the hemolymph and transported to its target cells. An increase in AKH activity might correlate with loss of internal lipid stores during behavioral maturation from nurses to foraging workers, indicating AKH signaling causally influence lipid mobilization and metabolism (Supplementary Information).
Comparison with Acc genome of Korea
In the preparation of this paper, a Northern strain of A. cerana genome with 2,430 scaffolds and 10,631 predicted genes was published by scientists in Korea60. The comparison of Northern strain of A. cerana genome and South strain of strain of A. cerana genome from present study showed that the identity between two strains was 98.8%, with the Korea A. cerana genome only 475 kb shorter (Supplementary Table 50). The N50 of the Korea A. cerana genome is 1,421,626 bp, while the Chinese A. cerana genome is 1,393,515 bp. Though the predicted CDS size of Chinese A. cerana (1,696 bp) is 200 bp larger than that of Korea A. cerana (1,472 bp), the identity between the genes of two genomes was 99.71%. Meanwhile, we identified 1,618 unique genes (with an average length of 900 bp) in Chinese Southern strain of A. cerana genome relative to Korea strain of A. cerana, and 2,011 unique genes (with an average length of 438 bp) in the Korea strain of A. cerana. For the 2,011 unique genes in the Korea strain, 2,009 had homologous regions in the Chinese A. cerana genome, but might be excluded by our gene prediction criteria, since the average size of these genes is only 438 bp and 88.8% (1,786) of them encoded hypothetical proteins. For the 1,618 unique genes in the Chinese strain, 1,564 had homologous regions in the Korea A. cerana genome while only 14.9% (241) of them encoded hypothetical proteins. The 54 genes left might be taken as unique Chinese A. cerana genes, including odorant binding protein 21 precursor, cytochrome c oxidase subunit, etc. (Supplementary Table 51), suggesting that the Chinese A. cerana have evolved unique physiological adaptations in response to different climate and other environmental conditions in China.
Conclusions
We present a high-quality genome sequence for the Asian honeybee A. cerana which constitutes an important resource for further molecular studies of honeybees and other social insects. Our comparative genomic analysis of A. cerana and A. mellifera deepens our understanding of the relationship between genes, behavior, and genetic adaptations of honeybees and reveals many species-specific genes that are potentially related to specialized biological functions and life history of A. cerana.
Using whole-genome sequencing and transcriptome sequencing (RNA-seq) followed by laboratory experiments measuring and comparing behavior response and in A. cerana and A. mellifera induced by Varroa infestation, we identified and functionally validated genes underlying the A. cerana grooming behavior in cleaning and removing Varroa mites. Over the years, the unique biological features of Asian honeybees especially their high degree of resistance to Varroa infestation have attracted a great deal of interest from researchers around the world who are committed to understanding the Varroa mite and searching for solutions for its control. Previous studies on the mechanisms of honeybee grooming behaviors focused on the induction of the olfactory, visual, and immune responses to the exogenous stimuli from the parasitic mites. Our genome analysis reveals, while A. cerana has more immune system genes and encodes many more antimicrobial peptides than A. mellifera, A. cerana contains less than 34% and 50% of genes encoding odorant binding proteins (OBPs) and olfactory receptors (Ors), respectively, compared to A. mellifera. Our observation that A. cerana displayed significantly more rapid and vigilant grooming behaviors to the presence of Varroa mites than A. mellifera suggests that parasite-defensive grooming in A. cerana is likely triggered not only by exogenous stimuli through visual and olfactory detection of the parasite but also by genetically endogenous processes, the programmed grooming model that postulates an existence of central programming that periodically activates a bout of grooming to remove ectoparasites before they begin to feed. The difference in levels of grooming behaviors in response to Varroa infestation between A. cerana and A. mellifera may also reflect their genetic difference in DNA methylation-mediated learning and memory. It has been reported previously that the learning and memory performance of A. cerana is significantly better on both color and grating patterns than that of A. mellifera. The formation of long-term memory and synaptic plasticity needs to activate many signal pathways of neurons, such as cAMP-PKA, MAPK, and CaMK IV pathways. Our transcriptome analysis by RNA-seq following qRT-PCR confirmation showed that the expression level of genes involved in these genetic pathways was significantly upregulated in A. cerana when challenged by Varroa mites, suggesting that Asian honeybees have evolved species-specific behavior adaptation that helped them survive and thrive in their particular environments. We, therefore, conclude that the Asian honeybee is an excellent model system for studying the cellular and molecular mechanisms of hygienic and grooming behaviors to combat the Varroa mites, the greatest pest threat to the European honeybees.
Materials and Methods
Bees and Genomic DNA Extraction
A wild colony of A. cerana cerana was collected at the Chinese Honeybee Natural Protection Area of the Wufu Mountain, Jiangxi Province, China. It was believed that this place (28°09′, 118°03′) was where Fabricius collected bee samples for his original descriptions of A. cerana in 1793. The bee colony was maintained at the Chinese National Human Genome Center at Shanghai. Genomic DNA was extracted from two drone pupae from the colony of A. cerana after getting rid of its intestine using the AxyPrep Multisource Genomic DNA Kit (AxyGen, Cat.No. AP-MN- MS-GDNA-50, CA, USA).
Genome Sequencing and Assembly
A 300 bp paired-end library was constructed using the standard Illumina paired-end protocol, and 99,070,380 pairs of 120 bp reads were produced on the Illumina Genome Analyzer platform (Illumina, San Diego, CA), providing 88 fold coverage of the genome. The shotgun library of 300–800 bp fragments was prepared from 5ug of DNA using the standard GS FLX shotgun library protocol, and a total of 1,641,411 reads with an average length of 388 bp were produced by Roche 454 GS FLX, providing 2.3 fold coverage of the A. cerana genome. The 3 kb mate-pair library was constructed combing the GS FLX and Illumina mate-pair protocol, with an adaptor sequence inserted between the mate-pair reads. A total of 46,124,432 mate-pair reads of 120 bp was produced on the Illumina Genome Analyzer platform, providing 32 fold sequencing coverage.
The Illumina pair-end reads and Roche 454 reads were assembled using Velvet V1.2.0361, which produced 21,784 contigs with an average length of 9,601 bp. Then mate-pair reads were mapped to the contigs to construct a scaffold of the A. cerana genome. Finally, we obtained 879 scaffolds with a total size of 228,791,026 bp, with a maximum length of 4,477,204 bp.
cDNA sequencing
The honeybee samples were taken from an A. cerana cerana colony at the Huajiachi campus of Zhejiang University, Hangzhou, China. The five samples of newly emerging workers, nurse workers, guard workers, dancing workers and foraging workers were collected from the same colony and each sample included thirty individuals. Newly emerging workers were identified as bees emerging from the cells. Nurses were caught when they entered into the cells and were nursing the larvae62. Guard workers were identified as bees stinging at the black lint ball which was jumping before the entrance of the hive63. Foragers were identified as bees returning to a colony loaded with pollen on their corbicula. Dancers were identified as pollen foragers performing waggle dances on the combs64. Each individual was put into liquid nitrogen with a pair of forceps immediately after collection and stored at −80 °C until head dissection.
To prevent mRNA degradation, brains were dissected in a solution of 0.4 M guanidine thiocyanate on an ice bag. The brain tissue was transferred into TRIzol reagent (Invitrogen) immediately and total RNA was prepared according to the manufacturer’s protocols. mRNA [poly(A) RNA] was then purified from total RNA using the Micropoly(A)PuristTM mRNA purification kit (Ambion, Cat.No.1919, Foster, CA, USA).
Gene Prediction
Four gene prediction sets were independently generated using Augustus65, Genemark66, Gnomon67, and SNAP68. Augustus and Genemark predicted using an A. mellifera model. Gnomon predicted based on ESTs. For the first step, the longest gene was selected, and if the gene was predicted at the same start or same end locus by two or more software; the second, the gene was selected which had no overlap with other genes; the third, gene was selected if it could be mapped by EST sequences or had homolog with A. mellifera genes; the fourth, a gene was abandoned if it had a stop codon in the middle of the sequences. After above steps, we manually checked all of the genes. Finally, a total of 10,182 genes were predicted.
Bioinformatics analysis
Based on the deduced amino acid sequences, the annotation was performed through BLASTP against the Swiss-Prot database and non-redundant peptide database (NR), with parameters set at E-value 1e-3. Gene ontology analysis was performed using Blast2GO69 through BLASTP against the Swiss-Prot database with a parameter of 1e-3. Protein motif and KOG (the assignment was predicted through RPS‐BLAST with the Conserved Domain Database (CDD) with E-value 1e-1070. The metabolic pathway was constructed based on the KEGG database by BBH method71. Differently expressed genes were identified by the DEGseq package using the MARS (MA-plot-based method with Random Sampling model)72 method, with the reads number of each gene transformed into RPKM (Reads Per Kilo bases per Million reads)73.
Candidate species for comparison with A. cerana
Seven species were chosen for comparison analysis with A. cerana, including A. mellifera, Camponotus floridanus, Harpegnathos saltator, Bombyx mori, Nasonia vitripennis, Drosophila melanogaster (release 5) and Anopheles gambiae. The set of orthologs were defined by pair-wise comparison of the proteomes using BLASTP with E-value ≤ 1e-50, identity ≥50% and alignment length covering at least 40% of the shorter protein. In addition, proteins matching the same KOG were also taken as orthologs.
Mite infection experiment
For both A. cerana and A. mellifera, newly emerged workers were marked on their thorax with non-toxic paint and returned to their original colony consisting of a normal age-structured worker population. Twelve days later, when they were at the age of performing duties within the hive, they were collected and subjected to behavioral tests regarding their behavioral responses to the presence of Varroa mites. The cages used to house the bees in the test were plastic containers (diameter = 5.0 cm; height = 5.0 cm), each with a single hole (diameter = 1.0 cm) on the top to allow the introduction of bees and mites. After the introduction, the hole was covered with mesh to prevent bees escaping. Single bees were introduced into each cage. The bees were given ten minutes without any disturbance to calm them down. Using a fine brush, a mite, collected by opening capped brood cells, the mites removed and kept in a petri dish, were transferred onto the abdomen of the bee. Following mite introduction, the bee was observed for 2 minutes. During the observation, two different levels of behavior by individual bees were noticed in response to the presence of the mites. In the first case,, bees groomed themselves using their legs, twisting or pivoting their abdomens while grooming. In the second case, bees immediately moved their bodies rapidly in a side-to-side movement, shaking their bodies repetitively. These two cases were assigned as self-grooming group, and body-shaking group, respectively. As A. mellifera did not show significantly triggered behaviors in response to the presence of Varroa, only A. cerana with behavior responses were used for subsequent molecular analysis. Five bees of each typical behavior of A. cerana were sampled. Bees without exposed to Varroa mite were used as negative controls. Bees with obscure behaviors were discarded. Five heads of bees in each group were used to extract RNA.
Caste determination experiment
The newly laid eggs in a comb were used to reveal the mechanism of the caste determination of A. cerana by RNA-seq. After two days, 30 larvae of A. cerana were moved into artificial queen cells with royal jelly which was collected from the same hive. The remaining larvae were still kept in their own cells. After two days, the larvae were collected into EP tubes and stored immediately in nitrogen for future use. After another two days, the fourth-day larvae were collected from the royal jelly feed cells and the original cells into EP tubes and stored immediately in liquid nitrogen for future RNA extraction.
Accession Numbers
A. cerana genome assembly contigs and scaffolds have been deposited in GenBank under accession number JPYL00000000. Sequences and functional annotations of A. cerana protein-encoding genes are available from the NCBI.
References
Arias, M. C. & Sheppard, W. S. Phylogenetic relationships of honey bees (Hymenoptera:Apinae:Apini) inferred from nuclear and mitochondrial DNA sequence data. Mol Phylogenet Evol 37, 25–35, https://doi.org/10.1016/j.ympev.2005.02.017 (2005).
Akratanakul, P. Beekeeping in Asia. (Food & Agriculture Org., 1986).
Hepburn, H. R. & Radloff, S. E. Honeybees of Asia. (Springer, 2011).
Gallai, N., Salles, J.-M., Settele, J. & Vaissière, B. E. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol Econ 68, 810–821 (2009).
Lever, J. J., van Nes, E. H., Scheffer, M. & Bascompte, J. The sudden collapse of pollinator communities. Ecol Lett 17, 350–359, https://doi.org/10.1111/ele.12236 (2014).
Potts, S. G. et al. Global pollinator declines: trends, impacts and drivers. Trends Ecol Evol 25, 345–353, https://doi.org/10.1016/j.tree.2010.01.007 (2010).
Henry, M. et al. A common pesticide decreases foraging success and survival in honey bees. Science 336, 348–350, https://doi.org/10.1126/science.1215039 (2012).
Doublet, V., Labarussias, M., de Miranda, J. R., Moritz, R. F. & Paxton, R. J. Bees under stress: sublethal doses of a neonicotinoid pesticide and pathogens interact to elevate honey bee mortality across the life cycle. Environ Microbiol. https://doi.org/10.1111/1462-2920.12426 (2014).
He, X. L. X. Y. Factors of Apis cerana decline in China. Apiculture of China 62, 21–23 (2011).
Cox-Foster, D. L. et al. A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318, 283–287, https://doi.org/10.1126/science.1146498 (2007).
Consortium, H. G. S. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949 (2006).
Radloff, S. E. et al. Population structure and classification of Apis cerana. Apidologie 41, 589–601 (2010).
Hepburn, H. R., Smith, D. R., Radloff, S. E. & Otis, G. W. Infraspecific categories of Apis cerana: morphometric, allozymal and mtDNA diversity. Apidologie 32, 3–24 (2001).
Abrol, D. P. Asiatic honey bee Apis cerana: biodiversity conservation and agricultural production Springer Dordrecht Heidelberg London New York, 761–793 (2013).
Evans, J. D. & Spivak, M. Socialized medicine: individual and communal disease barriers in honey bees. J Invertebr Pathol 103(Suppl 1), S62–72, https://doi.org/10.1016/j.jip.2009.06.019 (2010).
Anderson, D. L. & Trueman, J. W. Varroa jacobsoni (Acari: Varroidae) is more than one species. Exp Appl Acarol 24, 165–189 (2000).
Peng, Y.-S., Fang, Y., Xu, S. & Ge, L. The resistance mechanism of the Asian honey bee, Apis cerana Fabr., to an ectoparasitic mite, Varroa jacobsoni Oudemans. J Invertebr Pathol 49, 54–60 (1987).
Tewarson, N., Singh, A. & Engels, W. Reproduction of Varroa jacobsoni in colonies of Apis cerana indica under natural and experimental conditions. Apidologie 23, 161–171 (1992).
Yang, G.-H. Harm of introducing the western honeybee Apis mellifera L. to the Chinese honeybee Apis cerana F. and its ecological impact. Acta Entomologica Sinica 3, 015 (2005).
Oldroyd, B. P. & Nanork, P. Conservation of Asian honey bees. Apidologie 40, 296–312 (2009).
Moritz, R. F., Härtel, S. & Neumann, P. Global invasions of the western honeybee (Apis mellifera) and the consequences for biodiversity. Ecoscience 12, 289–301 (2005).
Li, J. et al. The prevalence of parasites and pathogens in Asian honeybees Apis cerana in China. PLoS One 7, e47955, https://doi.org/10.1371/journal.pone.0047955 (2012).
Menzel, R., Leboulle, G. & Eisenhardt, D. Small brains, bright minds. Cell 124, 237–239, https://doi.org/10.1016/j.cell.2006.01.011 (2006).
Shi, Y. Y. et al. A SNP based high-density linkage map of Apis cerana reveals a high recombination rate similar to Apis mellifera. PLoS One 8, e76459, https://doi.org/10.1371/journal.pone.0076459 (2013).
Evans, J. D. et al. Immune pathways and defence mechanisms in honey bees Apis mellifera. Insect Molecular Biology 15, 645–656, https://doi.org/10.1111/j.1365-2583.2006.00682.x (2006).
Harpur, B. A. & Zayed, A. Accelerated evolution of innate immunity proteins in social insects: adaptive evolution or relaxed constraint? Molecular biology and evolution 30, 1665–1674, https://doi.org/10.1093/molbev/mst061 (2013).
Sadd, B. M. et al. The genomes of two key bumblebee species with primitive eusocial organization. Genome Biology16:76. https://doi.org/10.1186/s13059-015-0623-3 (2015).
Barribeau, S. M. et al. A depauperate immune repertoire precedes evolution of sociality in bees. Genome Biology 16, 83, https://doi.org/10.1186/s13059-015-0628-y (2015).
Straub, T. & Becker, P. B. Dosage compensation: the beginning and end of generalization. Nat Rev Genet 8, 47–57 (2007).
Morra, R., Smith, E. R., Yokoyama, R. & Lucchesi, J. C. The MLE subunit of the Drosophila MSL complex uses its ATPase activity for dosage compensation and its helicase activity for targeting. Mol Cell Biol 28, 958–966, https://doi.org/10.1128/mcb.00995-07 (2008).
Gu, W., Wei, X., Pannuti, A. & Lucchesi, J. C. Targeting the chromatin-remodeling MSL complex of Drosophila to its sites of action on the X chromosome requires both acetyl transferase and ATPase activities. The EMBO Journal 19, 5202–5211, https://doi.org/10.1093/emboj/19.19.5202 (2000).
Hilfiker, A., Hilfiker‐Kleiner, D., Pannuti, A. & Lucchesi, J. C. mof, a putative acetyl transferase gene related to the Tip60 and MOZ human genes and to the SAS genes of yeast, is required for dosage compensation in Drosophila. The EMBO Journal 16, 2054–2060 (1997).
Kozmik, Z. Pax genes in eye development and evolution. Curr Opin Genet Dev 15, 430–438, https://doi.org/10.1016/j.gde.2005.05.001 (2005).
Jang, C. C. et al. Two Pax genes, eye gone and eyeless, act cooperatively in promoting Drosophila eye development. Development 130, 2939–2951 (2003).
Dominguez, M., Ferres-Marco, D., Gutierrez-Avino, F. J., Speicher, S. A. & Beneyto, M. Growth and specification of the eye are controlled independently by Eyegone and Eyeless in Drosophila melanogaster. Nat Genet 36, 31–39, https://doi.org/10.1038/ng1281 (2004).
Yao, J. G. et al. Differential requirements for the Pax6(5a) genes eyegone and twin of eyegone during eye development in Drosophila. Dev Biol 315, 535–551, https://doi.org/10.1016/j.ydbio.2007.12.037 (2008).
Czerny, T. et al. twin of eyeless, a second Pax-6 gene of Drosophila, acts upstream of eyeless in the control of eye development. Molecular cell 3, 297–307 (1999).
Hider, R. C. Honeybee venom: a rich source of pharmacologically active peptides. Endeavour 12, 60–65 (1988).
Grunwald, T. et al. Molecular cloning and expression in insect cells of honeybee venom allergen acid phosphatase (Api m 3). J Allergy Clin Immunol 117, 848–854, https://doi.org/10.1016/j.jaci.2005.12.1331 (2006).
Winningham, K. M., Fitch, C. D., Schmidt, M. & Hoffman, D. R. Hymenoptera venom protease allergens. J Allergy Clin Immunol 114, 928–933, https://doi.org/10.1016/j.jaci.2004.07.043 (2004).
Blank, S. et al. Identification, recombinant expression, and characterization of the 100 kDa high molecular weight Hymenoptera venom allergens Api m 5 and Ves v 3. Journal of immunology 184, 5403–5413, https://doi.org/10.4049/jimmunol.0803709 (2010).
Schmidt, J. O. Toxinology of venoms from the honeybee genus Apis. Toxicon: official journal of the International Society on Toxinology 33, 917–927 (1995).
Peiren, N., de Graaf, D. C., Evans, J. D. & Jacobs, F. J. Genomic and transcriptional analysis of protein heterogeneity of the honeybee venom allergen Api m 6. Insect molecular biology 15, 577–581, https://doi.org/10.1111/j.1365-2583.2006.00669.x (2006).
Balfanz, S. et al. Functional characterization of transmembrane adenylyl cyclases from the honeybee brain. Insect Biochem Mol Biol 42, 435–445, https://doi.org/10.1016/j.ibmb.2012.02.005 (2012).
Heisenberg, M. Mushroom body memoir: from maps to models. Nat Rev Neurosci 4, 266–275, https://doi.org/10.1038/nrn1074 (2003).
Pollard, T. D. & Beltzner, C. C. Structure and function of the Arp2/3 complex. Curr Opin Struct Biol 2, 244–249, https://doi.org/10.1016/S0959-440X(02)00396-2 (1999).
Rotty, J. D., Wu, C. & Bear, J. E. New insights into the regulation and cellular functions of the ARP2/3 complex. Nat Rev Mol Cell Biol 14, 7–12, https://doi.org/10.1038/nrm3492 (2013).
Suraneni, P. et al. The Arp2/3 complex is required for lamellipodia extension and directional fibroblast cell migration. J Cell Biol 197, 239–251, https://doi.org/10.1083/jcb.201112113 (2012).
Xu, P., Shi, M. & Chen, X. X. Antimicrobial peptide evolution in the Asiatic honey bee Apis cerana. PLoS One 4, e4239, https://doi.org/10.1371/journal.pone.0004239 (2009).
Guzmán-Novoa, M. E. A.-Va. E. Relative effect of four characteristics that restrain the population growth of the mite Varroa destructor in honey bee (Apis mellifera) colonies. Apidologie 32, 157–174 (2001).
Abdullah Ibrahim, M. S. The relationship between hygienic behavior and suppres-sion of mite reproduction as honey bee (Apis mellifera) mechanisms of resistance to Varroa destructor. Apidologie 37, 31–40 (2006).
Otto Boeckinga, M. S. Behavioral defenses of honey bees against Varroa jacobsoni Oud. Apidologie 30, 141–158 (1999).
Chow, K. L., Hall, D. H. & Emmons, S. W. The mab-21 gene of Caenorhabditis elegans encodes a novel protein required for choice of alternate cell fates. Development 121, 3615–3626 (1995).
Arechavaleta-Velasco, M. E., Alcala-Escamilla, K., Robles-Rios, C., Tsuruda, J. M. & Hunt, G. J. Fine-scale linkage mapping reveals a small set of candidate genes influencing honey bee grooming behavior in response to Varroa mites. PLoS One 7, e47269, https://doi.org/10.1371/journal.pone.0047269 (2012).
Qin, Q. H., He, X. J., Tian, L. Q., Zhang, S. W. & Zeng, Z. J. Comparison of learning and memory of Apis cerana and Apis mellifera. Journal of comparative physiology. A, Neuroethology, sensory, neural, and behavioral physiology 198, 777–786, https://doi.org/10.1007/s00359-012-0747-9 (2012).
Barron, A. B., Sovik, E. & Cornish, J. L. The roles of dopamine and related compounds in reward-seeking behavior across animal phyla. Front Behav Neurosci 4, 163, https://doi.org/10.3389/fnbeh.2010.00163 (2010).
Honeybee Genome Sequencing, C. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 443, 931–949 (2006).
Johnston, M. V., Alemi, L. & Harum, K. H. Learning, memory, and transcription factors. Pediatric research 53, 369–374, https://doi.org/10.1203/01.PDR.0000049517.47493.E9 (2003).
Gade, G., Hoffmann, K. H. & Spring, J. H. Hormonal regulation in insects: facts, gaps, and future directions. Physiological reviews 77, 963–1032 (1997).
Park, D. et al. Uncovering the novel characteristics of Asian honey bee, Apis cerana, by whole genome sequencing. BMC Genomics 16, 1, https://doi.org/10.1186/1471-2164-16-1 (2015).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome research 18, 821–829, https://doi.org/10.1101/gr.074492.107 (2008).
Liu, F. et al. High-abundance mRNAs in Apis mellifera: comparison between nurses and foragers. J Insect Physiol 57, 274–279, https://doi.org/10.1016/j.jinsphys.2010.11.015 (2011).
Arechavaleta-Velasco, M. E., Hunt, G. J. & Emore, C. Quantitative trait loci that influence the expression of guarding and stinging behaviors of individual honey bees. Behavior genetics 33, 357–364 (2003).
Li, L. et al. Differences in microRNAs and their expressions between foraging and dancing honey bees, Apis mellifera L. J Insect Physiol 58, 1438–1443, https://doi.org/10.1016/j.jinsphys.2012.08.008 (2012).
Nayak, A. et al. Cricket paralysis virus antagonizes Argonaute 2 to modulate antiviral defense in Drosophila. Nature structural & molecular biology 17, 547–554 (2010).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic Acids Res 33, 6494–6506, https://doi.org/10.1093/nar/gki937 (2005).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res 40, D13–25, https://doi.org/10.1093/nar/gkr1184 (2012).
Bromberg, Y. & Rost, B. SNAP: predict effect of non-synonymous polymorphisms on function. Nucleic Acids Res 35, 3823–3835, https://doi.org/10.1093/nar/gkm238 (2007).
Conesa, A. & Gotz, S. Blast2GO: A comprehensive suite for functional analysis in plant genomics. International journal of plant genomics 2008, 619832, https://doi.org/10.1155/2008/619832 (2008).
Marchler-Bauer, A. et al. CDD: a Conserved Domain Database for protein classification. Nucleic Acids Research 33, D192 (2005).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28, 27–30 (2000).
Wang, L., Feng, Z., Wang, X. & Zhang, X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26, 136–138, https://doi.org/10.1093/bioinformatics/btp612 (2010).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature methods 5, 621–628, https://doi.org/10.1038/nmeth.1226 (2008).
Acknowledgements
This research was supported by earmarked funds for the Modern Agro-industry Technology Research System (No. CARS-45), the National Natural Science Foundation of China (No. 31340061 and No. 31072095), and the Shanghai Municipal Commission for Science and Technology (15DZ2291400).
Author information
Authors and Affiliations
Contributions
J.W., F.H., S.S., Z.Z., Y.P.C., and S.W. coordinated the project; Q.D., L.S., Z.L., F.L., L.Z., L.C. and C.D. collected the A.cerana samples and extracted DNA and RNA; H.Z., Y.Z., Y.W., Y.S., Z.Z. and S.W. performed the genome and transcriptome sequencing; H.Z., Y.W., S.W., Y.S., L.S., and Z.Z. assembled the genome and EST sequence data; Q.D., L.S., H.Z., H.Q.Z., S.X., Y.S., Y.W., F.M., L.C., Q.S., M.Y. and S.S. performed gene prediction, annotation, and genomic analysis; S.C., R.C., S.Z., W.F.L., J.D.E., Q.H., J.L. and F.H. commented on the genome sequencing and analysis; Y.P.C., H.Z., S.X., H.Q.Z., Q.D., L.S., J.D.E., S.W., Z.Z., S.S., F.H. and J.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Diao, Q., Sun, L., Zheng, H. et al. Genomic and transcriptomic analysis of the Asian honeybee Apis cerana provides novel insights into honeybee biology. Sci Rep 8, 822 (2018). https://doi.org/10.1038/s41598-017-17338-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-17338-6
This article is cited by
-
A key gene for the climatic adaptation of Apis cerana populations in China according to selective sweep analysis
BMC Genomics (2023)
-
Varroa resistance in Apis cerana: a review
Apidologie (2023)
-
Minus-C subfamily has diverged from Classic odorant-binding proteins in honeybees
Apidologie (2023)
-
Geometric morphology and population genomics provide insights into the adaptive evolution of Apis cerana in Changbai Mountain
BMC Genomics (2022)
-
Organ-specific transcriptome analysis reveals differential gene expression in different castes under natural conditions in Apis cerana
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.