Abstract
Oestrogen controls Foxp3 expression in regulatory T cells (Treg cells) via a mechanism thought to involve oestrogen receptor alpha (ERα), but the molecular basis and functional impact of ERα signalling in Treg cells remain unclear. We report that ERα ligand oestradiol (E2) is significantly increased in human cervical cancer (CxCa) tissues and tumour-infiltrating Treg cells (CD4+CD25hiCD127low), whereas blocking ERα with the antagonist ICI 182,780 abolishes FOXP3 expression and impairs the function of CxCa infiltrating Treg cells. Using a novel approach of co-immunoprecipitation with antibodies to E2 for capture, we identified binding of E2:ERα complexes to FOXP3 protein in CxCa-derived Treg cells. Chromatin immunoprecipitation analyses of male blood Treg cells revealed ERα occupancy at the FOXP3 promoter and conserved non-coding DNA elements 2 and 3. Accordingly, computational analyses of the enriched regions uncovered eight putative oestrogen response elements predicted to form a loop that can activate the FOXP3 promoter. Together, these data suggest that E2-mediated ERα signalling is critical for the sustenance of FOXP3 expression and Treg cell function in human CxCa via direct interaction of ERα with FOXP3 promoter. Overall, our work gives a molecular insight into ERα signalling and highlights a fundamental role of E2 in controlling human Treg cell physiology.
Similar content being viewed by others
Introduction
Regulatory T cells (Treg cells) expressing forkhead box P3 (FOXP3) critically control immune responses by maintaining tolerance of self-antigens and restricting inflammation. Treg cells are indispensable for host protection against autoimmunity, allergy and inflammatory tissue damage, but these cells can also exert detrimental effects by mediating inappropriate tolerance of infections and tumours1. FOXP3 is a lineage-specific transcription factor that governs Treg development, differentiation, maintenance and function2. While a major proportion of Treg cells differentiates in the thymus (tTreg cells), a small percentage arises in the periphery (pTreg cells), with both subsets being evenly distributed between the lymphoid and non-lymphoid organs, as well as in the tissue-resident Treg cell pool3. Signals in the local microenvironment stimulate tissue-resident Treg cells to undergo phenotypic and functional specialization tailored to specific anatomic sites and organs3. These environment-specific cues include cytokines such as IL-2 and TGFβ, hormones such as oestradiol (E2) and 1,25-Dihyroxyvitamin D3, vitamin metabolites such as retinoic acid, and even microbial products such as short-chain fatty acids4,5. Indeed, sex steroid hormone E2 has previously been shown to drive Treg cell expansion and induce/potentiate their suppressive functions by signalling through oestrogen receptor alpha (ERα), thereby enhancing the phosphorylation of Akt and/or increasing cell surface expression of the inhibitory receptor PD-16,7,8,9,10,11,12. While these data have convincingly demonstrated that oestradiol can potently modulate Treg cell function, the molecular basis of E2 signalling in Treg cells and how this impacts on immune surveillance in the human female genital tract remains largely unknown largely because in ERα (Esr1) knockout mice Foxp3 expression was retained6.
In the present study, we demonstrate that human cervical cancer (CxCa) and tumour-infiltrating Treg cells (CD4+CD25hiCD127low cells) contain elevated levels of hormone E2 which is in complex with ERα in the latter. Using the ER-specific antagonist ICI 182,780 (ICI), we further show that ERα modulates FOXP3 expression and suppressive function of Treg cells isolated from CxCa tumour tissues. Using a novel approach of immunoblotting of E2-bound proteins revealed that ERα can form complexes with FOXP3 protein. Further, analysis in male blood Treg cells by chromatin immunoprecipitation (ChIP)-coupled quantitative PCR (qPCR) demonstrated ERα occupancy of the FOXP3 promoter and multiple intronic enhancers, consistent with an ability of ERα to directly modulate FOXP3 gene expression. Accordingly, computational analyses of the enriched regions of the FOXP3 locus identified eight putative oestrogen responsive elements (ERE) predicted to form a loop that may be capable of activating the FOXP3 promoter. Taken together, these data reveal a novel role of E2-mediated ERα signalling in the transcriptional regulation of FOXP3 and control of human Treg cell function.
Results
Human cervical tumours display accumulation of sex steroid hormone oestradiol
The hormone oestradiol has been strongly implicated in the pathogenesis of human cervical cancer, but the exact role that E2 plays in tumor formation is currently unclear. In order to clarify how E2 promotes tumorigenesis in the human female genital tract, we first assessed levels of 17β-oestradiol in blood and tissue samples obtained from patients with squamous cell carcinoma (SCC) of the cervix. There was a significant difference in average concentrations of circulating hormone between patients and controls, however the levels were very low in both the groups (mean 26 pg/ml vs.39 pg/ml respectively; P < 0.002; (Fig. 1A.i). These data are consistent with previous reports that blood levels of oestrogen, although difficult to measure accurately at low concentrations, are known to be modulated in female cancers13. E2 concentrations in SCC tissue samples (mean 691 pg/100 mg, n = 30) were ~3 to 4-fold higher than those detected in tissue samples of normal cervix (172 pg/100 mg, n = 30; P< 0.0001; Fig. 1A.ii) or healthy tissue sampled from sites adjacent to the tumours (240 pg/100 mg, n = 30; P < 0.0001) irrespective of patient’s age or menopausal status (13 of 30 study volunteers were post-menopausal women).
Cervical tumours are enriched in oestradiol (E2) and express oestrogen receptor α. (A) (i) Concentrations of 17β-oestradiol as determined by ELISA in blood plasma from healthy donors (Pl HD) or patients with CxCa (Pl CxCa) as well as in (ii) tissue samples of cervical tumours (CxCa), areas adjacent to the tumours (CxCa adj), and healthy cervices (Normal Cx). Graph shows mean values ± SEM of n = 30 per group. (B) Staining distribution of 17β oestradiol, oestrogen receptor α, and aromatase in a representative tissue section of SCC cervix. Upper left image (i) shows haematoxylin and eosin staining of a tumour section; upper right image (ii) shows E2 staining which was predominantly cytoplasmic in the tumor and both nuclear and cytoplasmic in the stroma and infiltrating cells; lower left image (iii) shows the nuclear staining of ERα in the stromal cells only; lower right image (iv) shows aromatase expression detected in the cytoplasm of the tumour, stroma and infiltrating cells. Inset: normal rabbit serum negative control. Symbol T indicates tumour location in each picture; *Indicates stroma. Images are representative of n = 30.
Having confirmed that E2 concentrations are increased in SCC tissues, we next investigated the cellular localization of the hormone using immunohistochemistry (IHC). For all cases of SCC tested (n = 30), IHC of tissue sections revealed marked E2 staining, which ranged in intensity from mild to moderate and was primarily located in the cytoplasm of tumour cells (30–80% stained E2 positive; (Fig. 1B.ii). Among tumour-infiltrating cell types, E2 staining varied from mild to strong and was more evenly distributed between the nucleus and cytoplasm. Nuclear E2 staining was detected in >80% of infiltrating inflammatory cells, which were comprised primarily of lymphocytes and fibroblasts (based on morphological criteria). Similar data were obtained when assessing tumour tissue sections for expression of the E2 biosynthetic enzyme aromatase, although enzyme staining was detected only in the cytoplasm (Fig. 1B.iv). Together, these data suggested that ongoing E2 synthesis in human cervical tumours may contribute to high local levels of this hormone despite the low concentrations present in blood plasma.
Tumour-infiltrating Treg cells express oestrogen receptor α and exhibit elevated levels of intracellular E2
Having confirmed E2 enrichment in cervical SCC tumours, we next assessed which of the infiltrating cell types expressed the corresponding oestrogen receptor - ERα and might be sensitive to modulation by E2 exposure. Sections of tumour tissue exhibited moderate ERα expression throughout the stroma and inflammatory cell infiltrate (30–50% staining in any given positive field), but staining was strongest in the nuclei of putative lymphocytes and fibroblasts which were distributed unevenly across the tumours (Fig. 1B.iii). In order to identify these ERα-expressing cell types, we next isolated the infiltrating lymphocytes from tumour tissues/peripheral blood by magnetic/flow-sorting and subjected these to reverse transcription PCR (primers listed in supplementary Table 1) to determine expression of ERα mRNA. Using this approach, we detected ERα expression in both CD8+ and CD4+ (CD4+CD25int) subsets of effector T-cells as well as in infiltrating Treg cells (CD4+CD25hiCD127lo cells) (Figs. 2A and S1a,b), suggesting that E2 sensitivity is prominent in antigen-experienced/regulatory cell types. Consistent with these data, we also detected marked ERα expression in peripheral blood Treg cells whether obtained from female CxCa patients or healthy male volunteers (Figs. 2A and S1a,b).
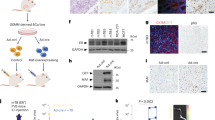
Primary human Treg cells contain high levels of oestradiol hormone complexed with ERα. (A) Reverse-transcription PCR assessment of ERα and FOXP3 expression in Treg cells (TR), Teffs (TE) and Naïve T cells (TN) isolated from cervical tumours (t) or blood (b) obtained from CxCa patients (p) or healthy male donors (d). The ERα-expressing breast cancer cell line MCF7 was included as a positive control. Data are representative of 20 independent experiments. Gels have been run under the same experimental conditions. Full-length gels are presented in Supplementary Figs. S1 to S3. (B) E2 concentrations as quantified by ELISA in different T-cell subsets, MACS-purified and flow-sorted from tumour tissue of SCC cervix; Treg cells - TRt (CD4 + CD25hiCD127lo), Teffs − TEt (CD4+CD25int), cytotoxic T lymphocytes (CD3+CD8+cells) and peripheral blood naïve T cells – NT PB (CD4+CD25−). A total of 105 cells per subset were used for quantification. Graphs show mean ± SEM of n = 20 per subset (PB = peripheral blood; t = tumour). (C) Oestradiol complexes present in MACS/sort-purified tumour Treg cells as detected by immunoprecipitation of oestradiol complexes using anti-E2 antibodies followed by immunoblotting with anti-ERα antibodies (i) or isotype-matched control antibodies (ii). β-actin immunoblots were carried out using lysates pre-immunoprecipitated with anti β-actin antibodies. Data are representative of 6 independent experiments. Gels have been run under the same experimental conditions. Please see Figs. S4 and S5 for original blot pictures.
We next assessed whether ERα expression by tumour-infiltrating lymphocytes was associated with hormone accumulation and potential E2 signalling in these populations. To do this, we isolated various T-cell subsets from CxCa tissues by magnetic/flow-based separation and quantified E2 levels in the cell lysates by competitive ELISA. We observed that mean concentration of E2 was ~2 to 3-fold higher in tumour-derived Treg cells (294.4 pg/105 cells) when compared with CD4+ effector T-cells (Teff cells) (97.3 pg/105 cells), CD8+ effector T-cells (138 pg/105 cells), or peripheral blood naïve T cells (94.9 pg/105 cells) (n = 20, P < 0.0001; Fig. 2B). These data indicated that human Treg cells are enriched in intracellular E2, consistent with their high expression of ERα, suggesting that E2 hormone may have a significant role to play in the biology of regulatory T-cell subsets. Accordingly, we also detected high levels of E2 in Treg cells separated from the peripheral blood of healthy male donors and in Treg cells isolated from the draining lymph nodes (LNs) of patients with CxCa (data not shown). Together, these findings indicate that high concentrations of intracellular E2 are characteristic of human Treg cell populations not only in CxCa tissues, but also in tumour-draining LNs and in the systemic circulation.
In addition, when we used anti-E2 antibodies to immunoprecipitate the hormone from cell lysates of CxCa-derived Treg cells, subsequent immunoblotting with anti-ERα antibodies revealed a 66 kDa band consistent with receptor binding (Figs. 2C and S4 and S5). This finding confirmed that the E2 content of human CxCa-derived Treg cells is at least partially complexed with the corresponding receptor ERα and may therefore exert an influence on the suppressive function of these cells.
ERα regulates FOXP3 expression in human Treg cells
Pioneering experiments in mice have demonstrated that E2 can enhance the suppressive activity of Treg cells by increasing Foxp3 expression via a mechanism thought to depend on ERα6,14: however, Esr1 knockout surprisingly did not result in complete disappearance of Foxp3 6. We therefore sought to determine whether E2 signalling through ERα is also capable of modulating FOXP3 expression and suppressive function of human tumour-infiltrating Treg cells in patients with CxCa. First, we analysed FOXP3 mRNA in different T-cell subsets in the circulation and tumour mass, and we observed prominent FOXP3 expression in Treg cells in the peripheral blood and CxCa infiltrate, minimal expression in tumour-derived effector T-cells, and a complete lack of expression in naïve T-cell populations from either patients or donors (Figs. 2A and S2a,b). We therefore proceeded to test whether expression of FOXP3 by Treg cells could be decreased by exposure to the ER-specific pure antagonist ICI, which is already being used for therapeutic applications under the trade name Fulvestrant15. Using this approach, we observed that treatment with pharmacologically relevant concentrations of ICI9,16 abolished the expression of both ERα and FOXP3 transcripts in both CxCa-derived Treg cells (Figs. 3A and S6–S8), and in peripheral blood Treg cells from healthy male volunteers (Fig. 3B). The loss of expression of both markers in male peripheral blood Treg cells was also confirmed in experiments with another ER antagonist and Selective Estrogen Receptor Disruptor (SERD) - RU58,668 (Fig. 3B). We were also able to confirm the drug-induced loss of ERα protein in human Treg cells by using anti-ERα antibodies to perform immunoprecipitation and immunoblotting assays (Figs. 3C and S9 and S10) as well as analysis by flow cytometry (Fig. S11). Consistent with the concept that E2:ERα signalling exerts a direct influence on human Treg cell function, we also observed that ICI treatment ablated FOXP3 protein expression as revealed by immunoprecipitation and immunoblotting using anti-FOXP3 antibodies (Figs. 3D and S12 and S13). Furthermore, supplementing these Treg cells with physiological concentrations of E2 was unable to rescue gene expression of either ERα or FOXP3 (Figs. 3A–D; S6 to S13). Together, these results indicated that ERα critically regulates the expression of FOXP3 in human Treg cells.
ERα control of FOXP3 expression in tumour-derived and peripheral blood Treg cells. (A) ERα and FOXP3 mRNA detection in CxCa Treg cells or MCF7 cells after treatment or not with the ERα antagonist ICI 182,780 (ICI; 1.8 µM) for 72 h in the presence or absence of exogenous oestradiol (E2;3.6 nM). Data are representative of 6 independent experiments. Gels have been run under the same experimental conditions. Original gel images are shown in Figs. S6–S8. (B) Relative expression levels of ERα and FOXP3 in peripheral blood Treg cells subjected to the same treatment as described in Fig. 3A (n = 6 healthy male donors). (C) Loss of ERα protein expression in CxCa Treg cells after 72 h treatment with ICI even in the presence of E2 supplementation throughout. Cellular extracts were immunoprecipitated using anti-ERα antibodies followed by immunoblotting with antibodies against ERα. The ERα-expressing breast cancer cell line MCF7 served as a positive control. β-actin immunoblots were carried out using lysates pre-immunoprecipitated with anti β-actin antibodies. Data are representative of 6 independent experiments. Gels have been run under the same experimental conditions. Please see Figs. S9 and S10 for original blot pictures (see also Fig. S11). (D) FOXP3 protein expression in CxCa Treg cells after treatment or not with 1.8 µM ICI in the presence or absence of E2 for 72 h. Cell lysates were immunoprecipitated and immunoblotted using anti-FOXP3 antibodies. β-actin immunoblots were carried out using lysates pre-immunoprecipitated with anti β-actin antibodies. Data are representative of 6 independent experiments. Gels have been run under the same experimental conditions. Please see Figs.S12 and S13 for original blot pictures.
Intracellular oestradiol-liganded oestrogen receptor regulates CxCa Treg cell function
We have previously demonstrated that in co-cultures of CxCa-derived Teff and Treg cells, effector cell secretion of IL4 and IFNγ is efficiently suppressed by regulatory cells that produce TGFβ and IL1017. We therefore sought to investigate whether E2:ERα signalling effects on Treg cell function might contribute to the restriction of anti-tumour effector T-cell responses in human CxCa. In co-cultures of CxCa-derived Treg and Teff cells, we observed that ICI induced a significant drop in the expression of suppressive cytokines TGFβ and IL10 (~10 and 4-fold decrease respectively, P < 0.01), and increased production of effector T-cell cytokines IL4 and IFNγ (~7-fold rise in both cytokines, P < 0.01) perhaps indicating restoration of effector potential upon inhibition of tumour-derived Treg cells (Fig. 4). We also observed a dose-dependent decrease in TGFβ production by CxCa-derived Treg cells when exposed to ICI either in the presence or absence of autologous Teff cells (Fig. S14). The significant reduction in TGFβ output in the presence of ICI was accompanied by a progressive decrease in the suppressive function of CxCa Treg cells, which displayed a dose-dependent decline in their ability to restrict the proliferation of autologous Teff cells (P < 0.001; Figs. 5A,B and S15). Annexin V staining demonstrated that ICI-treated Treg cells maintained high levels of cell viability throughout culture (~95% at 1.8 µM ICI; Supplementary Fig. S16), hence the loss of cytokine secretion and suppressive potential of CxCa-derived Treg cells could not be attributed to cell death. Indeed, E2 supplementation in co-cultures of Teff cells and ICI-treated Treg cells partially restored secretion of suppressive cytokines (P < 0.01) and restriction of CD4+ effector T-cell proliferation (P < 0.001; Figs. 4, 5A,B and S15; P < 0.001).
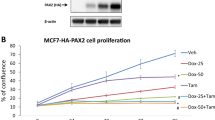
ERα and E2 modulate human Treg cell function. Equal numbers of tumour-dervied Treg and Teff cells (1 × 105 cells per subset) were co-cultured with or without ICI and exogenous E2 for 5 days and cytokine concentrations in the supernatant were determined by ELISA. Graphs show mean ± SEM of n = 6 experiments (see also Fig. S14).
ERα and E2 modulate human Treg cell proliferation. (A) Effect of ICI and E2 treatment on Treg cell’s suppressive function during 5 days co-culture with CFSE-labelled Teff cells. a: Lymphocyte gating based on light scatter; b: Percentage of Teff cells proliferating in response to stimulation with anti-human CD3/CD28. c: Treg cell inhibition of Teff cell proliferation during co-culture (1:1). d: Teff cell proliferation in co-culture with ICI-treated Treg cells (1:1). e: rescue of Treg cells’ suppressive function in co-cultures with Teff cells (1:1) treated with 1.8 µM ICI and 3.6 nM E2. M1: Percentage of non-proliferating T-cells; M2: Percentage of proliferating T-cells. Shown is a representative example of the FACS analysis (See also Figs. 5B and S15 and S16). (B) Percentage proliferating Teff cells in co-culture with Treg cells after treatment or not with ICI and exogenous E2. Mean ± error bars of 6 independent experiments performed in triplicate. (See also Figs. S15 and S16).
Together these data indicated that E2:ERα signalling effects on FOXP3 expression in human CxCa Treg cells alter cytokine secretion by these cells to modulate their regulatory function. However, since we have previously established that only ~50% of CxCa-infiltrating Treg cells (CD4+CD25hiCD127lo cells) express FOXP3 as measured by flow-cytometry17, we cannot exclude additional effects of the hormone on FOXP3-negative Treg cells or rule out the possibility that E2 may also act via non-ERα signalling pathways in CxCa Treg cells. Nonetheless, these data provide compelling evidence that intracellular complexes of E2:ERα play a vital role in the functional regulation of human FOXP3+ Treg cells that infiltrate human CxCa tumours.
Oestrogen receptor alpha binds the FOXP3 locus and expressed protein
Since oestrogen receptor is known to modulate the expression of multiple genes by binding directly to target promoters, we next sought to clarify whether ERα effects on FOXP3 expression in Treg cells were mediated by binding to the corresponding promoter. To do this, we used chromatin immunoprecipitation-coupled quantitative PCR (ChIP-qPCR) to map potential ERα binding sites at the FOXP3 promoter of primary human Treg cells. Since only limited numbers of Treg cells can be isolated from CxCa tissues, and these cells are not uniformly FOXP3+17, we performed these analyses using CD4+CD25hiCD127lo Treg cells separated from the peripheral blood of healthy male donors.
Using primer pairs targeting different regions of the FOXP3 promoter and enhancer regions (Supplementary Table 2), we identified ERα binding at multiple sites along the locus, including regions both upstream and within the core promoter, as well conserved non-coding DNA sequence elements (CNS)-2 and CNS3, but not CNS1 (Fig. 6A). These data likely reflected genuine ERα interaction with the FOXP3 locus, since we also detected receptor binding to known oestrogen response elements (EREs) at the loci BCL11B and pS2 in primary human Treg and MCF7 cells, but not to the SETB1 and FKBP6 genes respectively which lack the ERα target sequence (Fig. 6B and C). Together, these data indicated that ERα interaction with the FOXP3 locus occurs primarily at sites both upstream and within the core promoter, as well as at the regions CNS2 and CNS3.
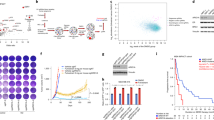
ERα-enriched regions of the FOXP3 locus. ChIP-qPCR analysis of the FOXP3 promoter and enhancer regions in chromatin fragments pulled down by anti-ERα antibodies (filled bars) or control IgG (open bars) in the cell lysates of blood Treg cells obtained from healthy male volunteers (A and C) or MCF7 cells (B). Data shown are mean ± SEM of 6 independent experiments performed in triplicate.
When we searched for ER-responsive elements of the FOXP3 promoter/enhancer using in-silico analysis of the enriched regions, we detected eight putative ERα binding sites distributed between regions upstream of the core promoter, adjacent to the TSS, and at CNS2 and CNS3 (P < 0.01)18. The majority of these putative ERα binding sites exhibited sequences characteristic of ‘ERE half-sites’, which have been reported to exhibit strong affinity for ERα binding19. Many of these EREs were also found to be conserved between the human, mouse and rat genomes (Figs. S17–S20; Supplementary Table 3). The enriched region upstream of the core promoter incorporated two ERE half-sites at positions −943 and −846 (Fig. S17), whereas the core promoter itself included conserved binding motifs at +39 and +114 bp downstream to the TSS (Fig. S18). CNS2b harboured a conserved ERE half-site at +4030 and was strongly enriched in ERα binding relative to CNS2a, which displayed a full ERE at +3895, suggesting that the former region may be critical for receptor binding to the FOXP3 locus (Fig. 6A and S19). Given the marked ERα enrichment at CNS3b (Fig. 6A and S20) and evolutionary conservation of both putative ERE half-sites, it seems likely that one or both of these sites is crucial for ERα binding to CNS3.
Since FOXP3 is known to auto-regulate its own promoter by binding to CNS2 region, we next sought to probe the physical interaction of E2 hormone and ERα with FOXP3 in primary human Treg cells isolated from CxCa tissues. To do this, we conducted co-IP experiments using the total cell lysates of Treg cells magnetically/flow sorted from CxCa tissues. We detected both ERα and FOXP3 in the immunoprecipitates obtained using anti-E2 antibodies (Figs. 7 and S21 to S23). Similarly FOXP3 was successfully detected in the anti-ERα immunoprecipitates (Fig. S24), thus indicating that ERα exists in a form complexed with FOXP3 in CxCa-derived Treg cells. Importantly, antibodies against E2 or ERα were unable to immunoprecipitate FOXP3 protein from the cell lysates of Treg cells that had been treated with ICI (Figs. 7 and S21 to S24). Together, these in silico and experimental data suggest that ERα may form part of an auto-regulatory loop in which the interaction of FOXP3 protein and E2:ERα complexes with the FOXP3 locus contribute to the maintenance of transcription factor expression20, and potentially influence the clinical course of human cervical cancers.
ERα protein interacts with FOXP3 protein in human tumour-derived Treg cells. CxCa Treg cells were treated or not with ICI in the presence or absence of exogenous E2 for 72 h before lysis. The cell lysates were then immunoprecipitated using anti-E2 antibodies and the protein complexes obtained were immunoblotted using antibodies against ERα and FOXP3. β-actin immunoblots were carried out using lysates pre-immunoprecipitated with anti β-actin antibodies. Data are representative of 6 independent experiments. Gels have been run under the same experimental conditions. Please see Figs. S21 to S23 for original blot pictures. See also Fig. S24.
Discussion
In the current report, we provide evidence that E2:ERα complexes drive FOXP3 expression and suppressive function of primary human Treg cells in both healthy male individuals and in tumours from patients with CxCa. Our data suggest that ERα critically regulates FOXP3 expression by binding to multiple elements along the locus as well as by directly interacting with FOXP3 protein intracellularly. Collectively, these findings uncover a pivotal role for ERα in Treg cell biology that may have important implications for the therapeutic targeting of these cells in patients with various types of cancer including CxCa.
It has been widely reported that various solid tumours exhibit oestradiol accumulation despite normal levels of this hormone being present in the circulation21. Oestradiol up-regulation in tumours is due to increased uptake and retention of circulating oestrogens E1 and/or E2, as well as enhanced hormone biosynthesis via the action of aromatase enzyme locally in the tumours22. Indeed, we detected marked aromatase expression in the tumour tissues analysed here, consistent with an earlier report that enzyme levels are increased in human CxCa23. Accordingly, the hormone products of the aromatase pathway have been strongly implicated in the pathogenesis of CxCa in both an animal model and in human patients24,25,26. In-line with these earlier findings, we detected high levels of E2 in human CxCa tumour tissue, whereas hormone levels in blood plasma remained low in both cancer patients and healthy volunteers. More surprising was our novel finding that primary human Treg cells are also characterized by high levels of intracellular E2, both in the peripheral circulation and CxCa tumours, suggesting that this hormone may have an important role to play in suppression of host anti-tumour immunity.
The cellular response to E2 hormone is predominantly mediated by the intracellular receptors ERα and ERβ, as well as the membrane G protein-coupled Oestrogen Receptor. In-line with the findings of a previous report27, our immunohistochemical analyses of CxCa tissues confirmed that both E2 hormone and ERα were primarily localized in the nuclei of stromal fibroblasts and tumour-infiltrating lymphocytes (TILs), and analysis of purified cell subsets identified strong expression of E2 and ERα in isolated Treg cells. In partial agreement with these data, a recent tissue microarray analysis of human CxCa tumours identified ERα expression by stromal fibroblasts, but failed to detect receptor expression by CD45+ hematopoietic cells28, perhaps due to the non-uniform distribution of FOXP3+cells within the tumours29. Previous studies in mice and human blood have shown that Treg cells express both ERα and ERβ, but population expansion triggered by E2 exposure is primarily mediated via ERα signalling6,8,30. Accordingly, when we used a novel approach of using anti-E2 antibodies to capture proteins that were complexed with this hormone in blood and tumour-derived Treg cells, we observed that intracellular E2 was at least partially complexed with ERα, and might therefore be capable of modulating Treg cells function via this pathway.
The hypothesis that E2 could induce FOXP3 expression was first proposed over a decade ago6. Direct evidence of this mechanism and its relevance in humans have until now remained obscure particularly since studies in Esr1 knockout mice did not abolish FOXP3 expression6. In the current report, we demonstrated that treatment with a SERD was able to abolish FOXP3 expression in Treg cells isolated from tumour tissues or peripheral blood obtained from CxCa patients or healthy volunteers, suggesting a direct influence of the classical ERα signalling pathway on sustenance of FOXP3 expression and Treg cell function. This is in contrast to studies in mice wherein basal expression of FOXP3 was retained in Esr1 knockout animals6. We could argue that this baseline expression could have had its origin in cord blood Treg cells where FOXP3 expression is progesterone dependent31. Hence these experiments unambiguously indicated that ER signaling through the classical pathway controlled FOXP3 expression in Treg cells. Loss of FOXP3 expression has previously been shown to result in a corresponding loss/dysfunction of Treg cells in mice and humans respectively32,33,34. E2:ERα signalling effects on FOXP3 expression are therefore likely to exert a significant influence on Treg cells function in human CxCa. However, therapeutic targeting of this pathway is likely to be complicated by reports from our lab and others that the Treg cell population of solid tumours can include both FOXP3+ and FOXP3− cells17,35. Accordingly, we observed that supplementation with E2 induced only a partial revival of suppressive function in ICI-treated Treg cells, suggesting additional effects of the hormone that do not depend on ERα and/or FOXP311,12. Our data suggest that ERα ranks among a select group of cellular factors, including TGF-β and NR4A2 that can exert major effects on Treg cell’s expression of FOXP3 36. Despite the known ability of TGFβ to induce Foxp3 expression in pTreg cells, this cytokine is unable to increase Foxp3 levels in tTreg cells37, and failed to sustain FOXP3 expression by CxCa-derived Treg cells upon ERα inhibition in the current study. These data suggest that FOXP3+ Treg cells residing in CxCa tissues are tTreg cells rather than pTreg cells3, which may have implications for the therapeutic manipulation of tumour Treg cell function in human patients.
Our ChIP-qPCR analyses of the FOXP3 locus revealed ERα occupancy of four distinct regions; upstream of the core promoter, within the core promoter, and at CNS2 and CNS3. By combining data from the literature with TRANSFAC and/or Jasper analysis, we identified Oestrogen Responsive Elements that not only appeared to be conserved across the mammalian species, but were also located in the vicinity of other known transcription factor binding sites proposed to influence ERα binding, FOXP3 expression, and/or Treg cell function20,38,39,40,41,42,43. In particular, the CNS2 region features binding sites for numerous transcription factors known to interact with ERα44,45, and may form short distance loops to interact with the core promoter and induce FOXP3 gene expression46,47. Less clear are the roles of the putative binding sites at CNS3 and in regions upstream of the core promoter, although future studies should be able to determine whether these are involved in stabilizing DNA looping and possibly facilitate interactions between other critical regions. Global analyses of receptor occupancy and expression profiling may also uncover additional gene targets of ERα modulation in human Treg cells with potential for novel clinical applications.
Earlier studies have demonstrated that complexes of Foxp3 protein with Runx1 and Cbf-β bind to the de-methylated CNS2 region of the FOXP3 locus to ‘auto-induce’ gene expression20,48, and proteomic studies have identified at least 361 interacting partners of Foxp3 in Treg cells49. In our study, we observed that in addition to ERα binding to the FOXP3 locus in human Treg cells, E2:ERα complexes also physically interacted with FOXP3 protein in these cells. Other known co-activators of ERα include the histone acetyl transferases (HATs) p300/CBP50, which act on the CNS2 region of the Foxp3 locus to maintain Treg cell stability51. We are therefore tempted to speculate that the FOXP3 locus in Treg cells is maintained in an active state by HAT-mediated recruitment of ERα to FOXP3-RUNX1-CBFβ complexes at CNS2. In light of these data, as well as earlier reports, we propose a model in which multiple ERα binding sites flanking the FOXP3 promoter regulate gene activity through a combination of direct DNA binding and indirect tethering/looping in a manner similar to that reported for ER-regulated genes in MCF-7 breast cancer cells (Fig. 8)44,52.
Diagrammatic representation of proposed ERα interactions with the FOXP3 locus in human Treg cells: RUNX1(R), CBFβ (C), Oestrogen Receptor (ER), CREB (CR), NFAT (N), NFKB (NK) and FOXP3 (F). Transcription factors that have previously been suggested to be involved in the looping of relatively distant DNA elements near the core promoter are marked with ‘*’. See also Figs. S17 to S20.
Cervical cancer remains a major burden globally especially in developing countries. Currently, chemo-radiation is the mainstay of treatment; however, five year survival rates are low as most of the patients present with advanced disease. Immunotherapy is a useful alternative to treat the disease. However, simultaneous to potentiation of effector responses in the tumor microenvironment, cancer immunotherapies must also overcome host immune suppression mediated by Tregs cells. Accordingly, we have presented evidence that E2:ERα interactions with FOXP3 locus exert potent effects on gene expression and can modulate the suppressive function of primary human tumor infiltrating Treg cells in patients with CxCa. These data not only advance our current understanding of basic Treg cell biology, but will also inform attempts to target ERα for therapeutic benefit in CxCa as well as other solid tumours (unpublished data from our lab has revealed elevated concentrations of the hormone E2 in various types of solid tumors in males), autoimmune diseases, and inflammatory disorders in which Treg cells play a central role. Also, in support of this proposed line of action are recent studies including ours showing paracrine ERα signaling in Cancer Associated Fibroblasts (CAFs) to be relevant in human CxCa28,53. Hence, we envisage that it may be worth exploring targeted therapy with ER antagonist in the management of CxCa: interfering with both Treg cell function and paracrine signaling through fibroblast ERα may help in checking the growth of cancer cells. Although we did not assess the effect of ICI on tumor Teff function which also expressed ERα, we envisage that ER antagonists would have an additive effect of boosting tumor infiltrating Th1 subset as well54. We therefore believe that the potentially far-reaching implications of our study have great translational value.
Materials and Methods
Study Design
Subjects
Fresh cervical tumour tissue (FIGO stage IB2), draining lymph nodes, and whole blood were obtained from patients undergoing radical hysterectomy for invasive cervical cancer. Normal cervices were obtained from women undergoing hysterectomy for benign conditions. Additional whole blood samples were collected from healthy male donors and used for isolation of Treg cells. Written informed consent was obtained from all study participants. The study was approved by Institutional Medical Ethics Committee (Kidwai Memorial Institute of Oncology, Bangalore, India). All experiments were performed in accordance with relevant guidelines and regulations.
Replicates: All experiments were performed in triplicates.
Oestradiol quantification
Oestradiol was extracted and quantified according to the procedure described by Rao et al.55. Briefly, 100 mg tumour tissue or 105 sorted cells were homogenized in 1 ml digestion buffer (0.1 M Phosphate Buffered Saline [PBS], pH 7.4, containing 100 mM EDTA, 100 μg/ml proteinase K [No. P2308, Sigma, St. Louis, MO], and 250 μg/ml DNase [No. DN5025, Sigma]) for 30 mins at 37oC before being subjected to chemical-based Oestradiol extraction. A total of 3 ml di-ethyl ether was added to the lysate and the mixture was snap-frozen in liquid nitrogen for 10 min. Upon thawing, the supernatant was collected and left at room temperature overnight to allow ether evaporation. The hormone-containing pellet was re-suspended in 1 ml PBS containing 10% gelatin and then used for Oestradiol quantification using Chemiluminescence Immuno Assays (no. 03000079190, Elecsys Oestradiol II, COBAS, USA) or Oestradiol competitive ELISA kits (no: 74070, Equipar, Saronno VA; Italy).
Immunohistochemical (IHC) staining of Oestradiol, ERα, and Aromatase
IHC staining of tissue sections of human cervical SCC was performed using the following antibodies (Abs): rabbit anti-human oestradiol (no. AR038-5R) and rabbit anti-human oestrogen receptor α (no. AM272-2M) from Biogenex, CA, USA; and rabbit anti-human aromatase (no. Ab35604), from Abcam UK. Secondary Abs (no. BA-1000) and ABC-Peroxidase kit VECTASTAIN® (no. PK-4001) were from Vector Laboratories Inc., CA, USA. Immunohistochemistry was performed as previously described17. Images of the stained sections were captured using an Olympus Bx 500 microscope and staining intensity/distribution in the tumour and stroma were scored by a histopathologist using Image Pro Plus software (Media Cybernatics, Rockville, MD, USA).
Cell Isolation, Purification and Analysis
Tumour-infiltrating lymphocytes (TILs) were isolated from enzymatically digested tumour tissues as detailed in our earlier report17. Additional procedures adopted for the isolation of specific cell types are described below:
Treg cells (CD4+CD25hi CD127lo cells)
Cell suspensions prepared from homogenized tumour tissues or peripheral blood samples were initially enriched for T cells by passing through nylon wool columns. Putative Treg cells were then isolated from the total T cell pool using a MACS CD4+CD25+ Regulatory T Cell Isolation Kit (no. 130-093-337, Miltenyi Biotec, GmBH, Germany). MACS-enriched CD4+CD25hiCD127lo Treg cells isolated from peripheral blood were used without further purification, whereas tumour-derived Treg cells were additionally subjected to flow sorting to obtain high purity CD4+CD25hi cells. We have previously determined the purity of MACS-enriched CD4+CD25hiCD127lo cervical tumour Treg cells to be 96% pure as assessed by flow cytometry17.
Teff cells (CD4+CD25int)
Nylon wool-enriched TILs were flow-sorted to obtain CD4+CD25int cells without prior MACS enrichment. Uniform sort gates were applied to all samples and cell purity was confirmed by re-acquisition of the sorted populations17.
Naïve T-cells (CD4+CD25−), B cells and CD8+ T-cells
CD4+CD25− naïve T cells, B cells, and CD8+ T cells were isolated from blood samples and/or CxCa tumour tissues using MACS kits (Miltenyi nos. 130-094-131, 130-091-151 and 130-096-495 respectively) as per the manufacturer’s instructions. Naïve T cells were additionally flow-sorted to obtain a highly pure CD4+CD25− population.
Flow cytometry
Lymphocyte subset analysis was performed using a FACS caliburTM flow-cytometer (Becton Dickinson, San Jose, USA). Cell sorting was conducted using a FACS Vantage SE apparatus (Beckton Dickinson, San Jose, USA.).
Treg cell suppression assays
CD4+CD25int Teff cells isolated from CxCa tissues were used as responder cells in Treg cells suppression assays. Teff (5 × 104) cells were cultured with or without an equal number of Treg cells and 5 × 103 autologous B cells to serve as antigen presenting cells. In some experiments, Treg cells were incubated with different concentrations of ICI 182,780 (36.7 nM, 367 nM, 1.8μM; No.I4409, Sigma) in the absence or presence of E2 hormone (3.6 nM; kind gift from Prof. A J Rao) for 30 min at 37 °C, 5%CO2 before being washed and co-cultured with Teff that had been pre-labelled with CFSE (1 μg/ml; 21888, Sigma). The cultures were then incubated for 5 days in indicator-free RPMI-1640 medium (No. R8755, Sigma), containing 10% charcoal-stripped Foetal Calf Serum (FCS; F2442, Sigma), and 300U/ml recombinant human IL2, and stimulated with plate bound anti-CD3ε (1 μg/ml functional grade purified clone OKT3) and anti-CD28 (0.5 μg/ml functional grade purified clone CD28.2, eBiosciences Inc. San Diego, CA, USA). Responder cell proliferation was assessed by CFSE dilution measured using a MoFlo flow cytometer (DAKO, USA). Cell divisions were quantitated using summit V4 software (DAKO cytomation, USA) and the results expressed as mean percentage proliferating cells ± SEM of triplicate cultures. RU 58,668 (1 μM; 3224, Tocris Bioscience) was used to confirm the results obtained with ICI56. Stock solutions of ICI, RU and oestradiol were prepared in ethanol and serially diluted in culture media to obtain the specified concentrations.
Cytokine quantitation in cell culture supernatants
After 5 days incubation, the culture supernatants were aspirated and assayed for concentrations of the cytokines IFNγ, IL4 and IL10, TGFβ1 (latent form) using the following ELISA kits; Human IFNγ (no. 88–731); Human IL4 (no. 88–7046); Human IL10 (no. 88–7106); and Human LAP (TGF β1) (no. 88–50390), (all from eBiosciences), as per the manufacturer’s instructions. Mean values of triplicate measurements were tabulated.
PCR determination of FOXP3 and ERα mRNA expression
Total cellular RNA was extracted using TRIzol reagent (no.15596-026, Ambion, New York, USA). Reverse transcription PCR determination of FOXP3 and ERα mRNA levels was performed using oligo dT and Moloney Murine Leukaemia Virus reverse transcriptase (no. 210210; Qiagen). All samples were run in triplicate. MCF-7 cell line which expresses ER α and Human FOXP3 plasmids pCMV-Tag2-human FoxP3, which generates full-length human FoxP3 by EcoRI and SalI, and retroviral pMCsIg (IRES-GFP)-human FoxP3 (kind donation from Dr. Ono M) were used as positive controls. Expression of β actin gene served as internal control. Primer sequences are provided in Supplementary Table 1 57,58,59.
Real-Time quantitative Reverse Transcription PCR
Messenger RNA levels of FOXP3 and ERα were determined by real-time PCR using a Step One Plus thermal cycler (Applied Biosystems). Briefly, 250 ng of total RNA was reverse transcribed, 10-fold diluted in sterile nuclease free water, and 2 μl used per 10 μl reaction volume with 1X Power SYBR Master Mix, and 100 nM of forward and reverse primers (listed in Supplementary Table 1). Cycling conditions were: 1 min at 95 °C, followed by 40 cycles of 15 s at 95 °C denaturation, 60 s at 60 °C annealing. Samples were run in duplicate and expression levels were determined using the 2ΔΔCt method with GAPDH as the internal control60,61.
Co-immunoprecipitation assays
Capture antibodies (directed against epitopes at/near the N termini of the respective target proteins); Rabbit anti-E2 (AR038-5R, BioGenex); mouse anti-human FOXP3 (clone 259D/C7, BD Pharmingen, NJ, USA); and rabbit anti-human ERα clone 1D5 from Biogenex (NU509-UCE) or Dako Cytomation, CA, USA (M7047).
Probing antibodies (directed against epitopes at/near the C termini of the respective target proteins); mouse anti-human ERα (clone F10, Santa Cruz Biotechnology Inc, USA); and rabbit anti-human FOXP3 (ab10563; Abcam, against epitope aa 400).
Detection antibodies: goat anti-mouse IgG HRP and goat anti-rabbit IgG HRP (31436 and 31466 respectively, from Pierce Biotechnology Inc., USA). TrueBlot® anti-rabbit and anti-mouse Ig IP Beads (nos. 008800 and 00881125 respectively) were from eBiosciences.
For immunoprecipitation assays, total cellular protein extracts were first obtained using various methods. To assess whether E2 was complexed to ERα, primary tumour-derived Treg cells were purified by MACS/FACS and then lysed. To assess the extent of ERα co-localization with FOXP3, tumour-derived Treg cells were incubated in the presence of different concentrations of ICI and/or E2 hormone (or ethanol-only control) prior to stimulation for 72 h with plate bound anti-CD3 (1 μg/ml), anti-CD28 (0.5 μg/ml) and recombinant human IL2 (100 IU/ml) prior to lysis. The protein extracts were then processed for immunoprecipitation using antibody-coated IP beads according to the manufacturer’s instructions. Immunoprecipitated material/protein extracts were separated on 10% SDS-PAGE gels and transferred onto Biodyne C nylon membranes (no. 60315, Pall Life Sciences, New York, USA). The membranes were probed with antibodies and visualized using the super signal® west pico chemiluminescent substrate (no. 34079, Thermo Scientific, Rockford, USA). The MCF-7 breast cancer cell line was used as a positive control both for FOXP3 and ERα detection. SDS sample buffer was used as the negative control.
Cell viability staining
Propidium iodide (PI) and annexin V–FITC apoptosis detection kits (no. K101; BioVision Inc, California, USA) were used to determine the viability of ICI treated Treg cells isolated by MACS/FACS sorting from tumour tissues. Cell staining was determined by flow cytometry.
ERα staining using flow cytometry
For assessment of ERα expression by flow cytometry, a total of 2 × 105 Treg cells were cultured for 72 h with various concentrations of ICI 182, 780 (36.7 nM, 367 nM, or 1.8μM), then surface staining with anti-human CD4-PE (clone OKT4, eBiosciences), followed by permeabilization and intracellular staining with rabbit anti-human ERα antibodies (no. ab37438, Abcam). After 30 min incubation, the cells were stained for a further 30 min with goat anti-rabbit IgG-FITC conjugated antibodies (No. F6005, Sigma) and washed in 1x PBS containing 0.1% BSA to remove excess/unbound antibodies. A total of 20,000 cells were acquired using a BD FACS Calibur apparatus and lymphocyte staining for ERα was determined relative to lymphocytes treated with normal rabbit/goat sera.
Chromatin Immunoprecipitation (ChIP)
CD4+CD25hiCD127lo Treg cells were isolated from the peripheral blood buffy coats of healthy donors using the MACS Treg cell isolation kit (no. 130-093-337, Miltenyi). The ERα-expressing breast cancer cell line MCF7 was used as a positive control. Chromatin was sheared in a Bioruptor (Diagenode, Belgium) using 12 pulses of 30 s duration with an amplitude of 90% and 90 s pause between pulses. ChIP on the fragmented DNA was performed using the low cell # ChIP kit (no. kch-maglow-016, Diagenode, Belgium) and ChIP-grade monoclonal antibodies against ERα (no.Mab-009-050, Diagenode) or a control Ig (provided in the kit). The relative abundance of regions of interest in the precipitated DNA was quantitated by qPCR using SYBR Select PCR master mix (no. 4472903, Life Technologies, Carlsbad, CA, USA).
Primers for ChIP qPCR
Bcl11b is a molecular partner of Foxp3 and has been shown to be essential for Treg cell’s suppressive function and maintenance of optimal Foxp3 gene expression49,62,63. An in silico search revealed that the BCL11B promoter incorporates an ERE, hence we were able to design qPCR primers for this region to use as a known positive control gene for ChIP-qPCR in primary human Treg cells. In contrast, our searches indicated that the SETB1 promoter lacked any ERE, hence primers targeted against this gene served as a negative control (Supplementary Table 2).
FOXP3 gene
Based on our in silico analyses and published literature describing the FOXP3 gene22,42,64,65,66,67,68, we designed quantitative PCR primers against: (a) Upstream region (−3888 to −3793 and −3665 to −3565); (b) tiling primers covering the region upstream of the core promoter (−943 to −664); (c) tiling primers covering the core promoter body (−456 to +104); (d) tiling primers covering CNS1 (+1593 to +1924); (e) tiling primers covering CNS2 (+3820 to +4051); (f) tiling primers covering CNS3 (+6443 to +6615). We then carried out pilot ChIP-qPCR analyses on blood Treg cells obtained from two healthy donors, followed by validation of the ER-enriched regions detected using ChIP-qPCR analyses of Treg cells on a further cohort of six healthy donors. Primer sequences are listed in Supplementary Table 2.
MCF7 cells
The ERα-expressing breast cancer cell line MCF7 was used as a positive control in the ERα ChIP-qPCR experiments. Primers to the ERE-containing gene pS2, and no ERE- containing FKBP6 on chromosome 7 were used as positive and negative controls respectively (Supplementary Table 2)69.
In silico identification of binding sites for ERα and interacting proteins
Position Specific Weight Matrices (PSWM) for the following Transcription Factor Binding Sites (TFBSs): ETS1, NFkB, RUNX1 and SMAD; and for complete ERE and ERE half sites were obtained using the Transfac tool (http://www.biobase-international.com/product/transcription-factor-binding-sites). The human promoter sequence for the FOXP3 gene was obtained from the UCSC genome browser70. The TFBSs within regions of interest on the human FOXP3 locus were identified using the matrices derived from Transfac and the Matrix-scan tool71 and the p-values assigned by the tool were recorded.
Evolutionary conservation of FOXP3 promoter regions
Regions of interest around the FOXP3 gene in human, mouse and rat species were analysed using both the Evolutionary Conserved Regions (ECR) browser72 and the MULAN tool73 to identify similar TFBS patterns between species. The latter tool revealed the conserved regions (sequences shown in Figs. S8–S11). The conserved TFBSs were manually identified and marked in the output file. The ECR browser provided a graphical summary of the sequences studied and the corresponding human genome (top portions in Figs. S8–S11). Evolutionary conserved regions were identified based on sequence similarity between species (assessed in ECR browser). Sequences were aligned by optimizing alignment identity scores and a threshold of 50% sequence homology per 100 base block was used to define conservation (performed using the MULAN tool in the ECR browser).
Statistics
All tests were performed in triplicate. Significant differences between groups were determined using either student’s t-test or paired t-tests as appropriate (Graph Pad Prism 5 Software, San Diego, CA, USA). P-values < 0.05 were considered significant (*P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001).
Study approval
Written informed consent was obtained from all study participants prior to inclusion in the study. The study was approved by Institutional Medical Ethics Committee of Kidwai Memorial Institute of Oncology (vide letter no. PER/CAB-I/D-1-13/2006).
References
Sakaguchi, S. et al. Foxp3+CD25+CD4+ natural regulatory T cells in dominant self-tolerance and autoimmune disease. Immunol. Rev. 212, 8–27 (2006).
Ochs, H. D., Ziegler, S. F. & Torgerson, T. R. FOXP3 acts as a rheostat of the immune response. Immunol. Rev. 203, 156–164 (2005).
Burzyn, D., Benoist, C. & Mathis, D. Regulatory T cells in nonlymphoid tissues. Nat. Immunol. 14, 1007–1013 (2013).
Yang, X. F. Factors regulating apoptosis and homeostasis of CD4+CD25 (high) FOXP3+ regulatory T cells are new therapeutic targets. Front. Biosci. 13, 1472–1499 (2008).
Hoeppli, R. E., Wu, D., Cook, L. & Levings, M. K. The environment of regulatory T cell biology: cytokines, metabolites, and the microbiome. Front. Immunol. 6, 61–74 (2015).
Polanczyk, M. J. et al. Cutting edge: Estrogen drives expansion of the CD4+CD25+ regulatory T cell compartment. J. Immunol. 173, 2227–2230 (2004).
Polanczyk, M. J., Hopke, C., Vandenbark, A. A. & Offner, H. Estrogen-mediated immunomodulation involves reduced activation of effector T cells, potentiation of Treg cells, and enhanced expression of the PD-1 costimulatory pathway. J. Neurosci. Res. 84, 370–378 (2006).
Prieto, G. A. & Rosenstein, Y. Oestradiol potentiates the suppressive function of human CD4+CD25+ regulatory T cells by promoting their proliferation. Immunology 118, 58–65 (2006).
Tai, P. et al. Induction of regulatory T cells by physiological level estrogen. J. Cell. Physiol. 214, 456–464 (2008).
Valor, L. et al. Estradiol-dependent perforin expression by human regulatory T-cells. Eur. J. Clin. Invest. 41, 357–364 (2011).
Yates, M. A., Li, Y., Chlebeck, P. J. & Offner, H. GPR30, but not estrogen receptor-alpha, is crucial in the treatment of experimental autoimmune encephalomyelitis by oral ethinyl estradiol. BMC Immunol. 11, 20–24 (2010).
Luo, C. Y., Wang, L., Sun, C. & Li, D. J. Estrogen enhances the functions of CD4(+)CD25(+)Foxp3(+) regulatory T cells that suppress osteoclast differentiation and bone resorption in vitro. Cell. Mol. Immunol. 8, 50–58 (2011).
Dowsett, M. & Folkerd, E. Deficits in plasma oestradiol measurement in studies and management of breast cancer. Breast Cancer Res. 7, 1–4 (2005).
Polanczyk, M. J., Hopke, C., Huan, J., Vandenbark, A. A. & Offner, H. Enhanced FoxP3 expression and Treg function in pregnant and estrogen-treated mice. J. Neuroimmunol. 170, 85–92 (2005).
Osborne, C. K., Wakeling, A. & Nicholson, R. I. Fulvestrant: an oestrogen receptor antagonist with a novel mechanism of action. Br. J. Cancer. 90(Suppl 1), S2–S6 (2004).
Robertson, J. F. et al. Pharmacokinetic profile of intramuscular fulvestrant in advanced breast cancer. Clin. Pharmacokinet. 43, 529–538 (2004).
Adurthi, S. et al. Functional tumor infiltrating TH1 and TH2 effectors in large early-stage cervical cancer are suppressed by regulatory T cells. Int. J. Gynecol. Cancer. 22, 1130–1137 (2012).
Mason, C. E. et al. Location analysis for the estrogen receptor-alpha reveals binding to diverse ERE sequences and widespread binding within repetitive DNA elements. Nucleic Acids Res. 38, 2355–2368 (2010).
Anderson, I. & Gorski, J. Estrogen receptor alpha interaction with estrogen response element half-sites from the rat prolactin gene. Biochemistry. 39, 3842–3847 (2000).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature. 463, 808–812 (2010).
Ito, K. Hormone replacement therapy and cancers: the biological roles of estrogen and progestin in tumorigenesis are different between the endometrium and breast. Tohoku J. Exp. Med. 212, 1–12 (2007).
Lønning, P. E. et al. Exploring breast cancer estrogen disposition: the basis for endocrine manipulation. Clin. Cancer Res. 17, 4948–4958 (2011).
Nair, H. B. et al. Induction of aromatase expression in cervical carcinomas: effects of endogenous estrogen on cervical cancer cell proliferation. Cancer Res. 65, 11164–11173 (2005).
Moodley, M., Moodley, J., Chetty, R. & Herrington, C. S. The role of steroid contraceptive hormones in the pathogenesis of invasive cervical cancer: a review. Int. J. Gynecol. Cancer. 13, 103–110 (2003).
Arbeit, J. M., Münger, K., Howley, P. M. & Hanahan, D. Progressive squamous epithelial neoplasia in K14-human papillomavirus type 16 transgenic mice. J. Virol. 68, 4358–4368 (1994).
Chung, S. H., Shin, M. K., Korach, K. S. & Lambert, P. F. Requirement for stromal estrogen receptor alpha in cervical neoplasia. Horm. Cancer. 4, 50–59 (2013).
Kwasniewska, A. et al. Estrogen and progesterone receptor expression in HPV-positive and HPV-negative cervical carcinomas. Oncol. Rep. Oncol. Rep. 26, 153–160 (2011).
den Boon, J. A. et al. Molecular transitions from papillomavirus infection to cervical precancer and cancer: Role of stromal estrogen receptor signaling. Proc. Natl. Acad. Sci. USA 112, E3255–3264 (2015).
Adurthi, S. et al. Regulatory T Cells in a Spectrum of HPV-Induced Cervical Lesions: Cervicitis, Cervical Intraepithelial Neoplasia and Squamous Cell Carcinoma. Am. J. Reprod. Immunol. 60, 55–65 (2008).
Aristimuño, C. et al. Sex-hormone receptors pattern on regulatory T-cells: clinical implications for multiple sclerosis. Clin. Exp. Med. 12, 247–55 (2012).
Lee, J. H., Ulrich, B., Cho, J., Park, J. & Kim, C. H. Progesterone promotes differentiation of human cord blood fetal T cells into T regulatory cells but suppresses their differentiation into Th17 cells. J Immunol. 187, 1778–87 (2011).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nature Immunol. 4, 330–336 (2003).
Bacchetta, R. et al. Defective regulatory and effector T cell functions in patients with FOXP3 mutations. J. Clin. Invest. 116, 1713–1722 (2006).
Filaci, G. et al. CD8+CD28− T regulatory lymphocytes inhibiting T cell proliferative and cytotoxic functions infiltrate human cancers. J. Immunol. 179, 4323–4334 (2007).
Sekiya, T. et al. The nuclear orphan receptor Nr4a2 induces Foxp3 and regulates differentiation of CD4+ T cells. Nat. Commun. 2, 269–280 (2011).
Okada, M., Hibino, S., Someya, K. & Yoshmura, A. Regulation of regulatory T cells: epigenetics and plasticity. Adv. Immunol. 124, 249–273 (2014).
Ruan, Q. et al. Development of Foxp3 (+) regulatory t cells is driven by the c-Rel enhanceosome. Immunity 31, 932–940 (2009).
Polansky, J. K. et al. Methylation matters: binding of Ets-1 to the demethylated Foxp3 gene contributes to the stabilization of Foxp3 expression in regulatory T cells. J. Mol. Med. (Berl). 88, 1029–1040 (2010).
Ogawa, C. et al. TGF-β-mediated Foxp3 gene expression is cooperatively regulated by Stat5, Creb, and AP-1 through CNS2. J. Immunol. 192, 475–483 (2014).
Long, M., Park, S. G., Strickland, I., Hayden, M. S. & Ghosh, S. Nuclear factor-kappaB modulates regulatory T cell development by directly regulating expression of Foxp3 transcription factor. Immunity. 31, 921–931 (2009).
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nat. Immunol. 11, 618–627 (2010).
Morelli, C. et al. Akt2 inhibition enables the forkhead transcription factor FoxO3a to have a repressive role in estrogen receptor alpha transcriptional activity in breast cancer cells. Mol. Cell. Biol. 30, 857–870 (2010).
Stender, J. D. et al. Genome-wide analysis of estrogen receptor alpha DNA binding and tethering mechanisms identifies Runx1 as a novel tethering factor in receptor-mediated transcriptional activation. Mol. Cell. Biol. 30, 3943–3955 (2010).
Hirano, S., Furutama, D. & Hanafusa, T. Physiologically high concentrations of 17 beta-estradiol enhance NF-kappaB activity in human T cells. Am. J. Physiol. Regul. Integr. Comp. Physiol. 292, R1465–1471 (2007).
Feng, Y. et al. Control of the inheritance of regulatory T cell identity by a cis element in the Foxp3 locus. Cell 158, 749–763 (2014).
Li, X., Liang, Y., LeBlanc, M., Benner, C. & Zheng, Y. Function of a Foxp3 cis-element in protecting regulatory T cell identity. Cell 158, 734–748 (2014).
Gavin, M. A. et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature 445, 771–775 (2007).
Rudra, D. et al. Transcription factor Foxp3 and its protein partners form a complex regulatory network. Nat. Immunol. 13, 1010–1019 (2012).
Acevedo, M. L. & Kraus, W. L. Mediator and p300/CBP-steroid receptor co-activator complexes have distinct roles, but function synergistically, during estrogen receptor alpha-dependent transcription with chromatin templates. Mol. Cell. Biol. 23, 335–348 (2003).
Liu, Y. et al. Two Histone/Protein Acetyl transferases, CBP and p300, are indispensable for Foxp3+ T-Regulatory Cell Development and Function. Mol. Cell. Biol. 34, 3993–4007 (2014).
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009).
Kumar, M. M. et al. Role of estrogen receptor alpha in human cervical cancer-associated fibroblasts: a transcriptomic study. Tumour Biol. 37, 4409–4420 (2016).
Polese, B. et al. The Endocrine Milieu and CD4 T Lymphocyte Polarization during Pregnancy. Front Endocrinol (Lausanne). 5, 106–117 (2014).
Rao, A. J., Kotagi, S. G. & Moudgal, N. R. Serum concentrations of chorionic gonadotrophin, oestradiol-17 beta and progesterone during early pregnancy in the south Indian bonnet monkey (Macaca radiata). J. Reprod. Fertil. 70, 449–455 (1984).
Kocanova, S., Mazaheri, M., Subra, S. C. & Bystricky, K. Ligands specify estrogen receptor alpha nuclear localization and degradation. BMC Cell Biology 11, 98 (2010).
Ishikawa, H. et al. High aromatase expression in uterine leiomyoma tissues of African-American women. J Clin Endocrinol Metab 94, 1752–1756 (2009).
Dulkys, Y. et al. Detection of mRNA for eotaxin-2 and eotaxin-3 in human dermal fibroblasts and their distinct activation profile on human eosinophils. J Invest Dermatol. 116, 498–505 (2001).
Morgan, M. E. et al. Expression of FOXP3 mRNA is not confined to CD4+CD25+ T regulatory cells in humans. Hum Immunol. 66, 13–20 (2005).
Matsuzaki, S. et al. Expression of estrogen receptor alpha and beta in peritoneal and ovarian endometriosis. Fertil Steril. 75, 1198–205 (2001).
Zhang, Y. et al. PGC-1alpha induces apoptosis in human epithelial ovarian cancer cells through a PPARgamma-dependent pathway. Cell Res. 17, 363–73 (2007).
Vanvalkenburgh, J. et al. Critical role of Bcl11b in suppressor function of T regulatory cells and prevention of inflammatory bowel disease. J. Exp. Med. 208, 2069–2081 (2011).
Fu, W. et al. A multiply redundant genetic switch ‘locks in’ the transcriptional signature of regulatory T cells. Nat. Immunol. 13, 972–980 (2012).
Mantel, P. Y. et al. Molecular mechanisms underlying FOXP3 induction in human T cells. J. Immunol. 176, 3593–3602 (2006).
Kim, H. P. & Leonard, W. J. CREB/ATF-dependent T cell receptor induced FoxP3 gene expression: a role for DNA methylation. J. Exp. Med. 204, 1543–1551 (2007).
Burchill, M. A., Yang, J., Vogtenhuber, C., Blazar, B. R. & Farrar, M. A. IL-2 receptor beta-dependent STAT5 activation is required for the development of Foxp3+ regulatory T cells. J. Immunol. 178, 280–290 (2007).
Klunker, S. et al. Transcription factors RUNX1 and RUNX3 in the induction and suppressive function of Foxp3+ inducible regulatory T cells. J. Exp. Med. 206, 2701–2715 (2009).
Barbarulo, A. et al. Notch3 and canonical NF-kappaB signaling pathways cooperatively regulate Foxp3 transcription. J. Immunol. 186, 6199–6206 (2011).
Wang, C., Yu, J. & Kallen, C. B. Two estrogen response element sequences near the PCNA gene are not responsible for its estrogen enhanced expression in MCF7 cells. PLoS One. 3, e3523–3532 (2008).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Turatsinze, J. V., Thomas-Chollier, M., Defrance, M. & van Helden, J. Using RSAT to scan genome sequences for transcription factor binding sites and cis-regulatory modules. Nat. Protoc. 3, 1578–1588 (2008).
Ovcharenko, I., Nobrega, M. A., Loots, G. G. & Stubbs, L. ECR Browser: a tool for visualizing and accessing data from comparisons of multiple vertebrate genomes. Nucleic Acids Res. 32(Web Server Issue), W280–W286 (2004).
Ovcharenko, I. et al. Multiple-sequence local alignment and visualization for studying function and evolution. Genome Res. 15, 184–194 (2005).
Acknowledgements
We thank A.J. Rao, Professor Emeritus, Indian Institute of Science, Bangalore for his help in guiding SA with oestradiol extraction and assay and for providing pure oestradiol powder; Dr. Ono M., Department of Experimental Pathology, Institute for Frontier Medical Sciences, Kyoto University, Japan for providing human FOXP3 plasmids; Dr. Bhuvaneswari Adurthi for helping SA in ChIP; Dr. Uday Kumar Ranga, Professor, Jawaharlal Nehru Center for Advanced Scientific Research, Bangalore, India and Dr. Girdhari Lal, Scientist, National Centre for Cell Science, Pune, India, for critically reviewing our work and providing very useful suggestions on the manuscript; Dr. Neil McCarthy of Insight Editing London for providing writing assistance; Dr. Anita Mahadevan, Additional Prof. Department of Neuropathology, NIMHANS for microphotography; Alaguraj D., Project Assistant FACS Facility, IISc, Bangalore for help with flow sorting; Dr. Harsha Gowda, Faculty Scientist, Institute of Bioinformatics, Bangalore, for useful discussions for improving the manuscript and Dr. Remya Raja, Research Scientist, Institute of Bioinformatics for her support at a crucial time in the experiments. This work was supported by grants from the Department of Biotechnology, Government of India (to RSJ) (no. BT/PR7014/MED/14/927/2005); SA received a Senior Research fellowship from the Indian Council of Medical Research - (no. 3/2/2/72/2005/NCD-III); MMK was funded by the Council for Scientific and Industrial Research - Senior Research fellowship (no. 09/999 (0001)/2009-EMR-I). RSJ had full access to all the data in the study and takes responsibility for the integrity of the data and the accuracy of the data analysis.
Author information
Authors and Affiliations
Contributions
S.A. did oestradiol extraction and estimation by ELISA, IHC, RT PCR, IP/IB, functional experiments on tumor Tregs: cytokine ELISA; suppression assays, analyzed data with the direction and supervision of R.S.J. and contributed to the discussions; M.M.K. did experiments on peripheral blood Tregs from male donors, designed ChIP qPCR primers for various regions of FOXP3 locus with the direction and supervision of R.S.J., did real time PCR and ChIP qPCR, analyzed the data and contributed to the discussions; V.H.S. and A.K.K. did mapping of TFBS on FOXP3 locus with help from M.M.K. and R.S.J.; Model hypothesis was done by V.H.S. with inputs from A.K.K., R.S.J. and M.M.K.; G.M. grossed and sampled fresh tissues of uterine cervical specimens, scored IHC slides; A.B. did buffy coat separation from voluntary blood donor bags; U.D.B. and U.D.K. did surgical work up of cases for the study. S.K. and H.K. helped S.A. in sorting cells from tumors; A.P. and A.M. did the additional experiment with RU 58668, Real Time PCR, RTPCR and agarose gel electrophoresis with the direction and supervision of R.S.J.; R.S.J. planned, directed and supervised the study, wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
V.H.S., A.K.K.: The research and development division of a commercial organization (Shodhaka LS Pvt. Ltd.) was involved in the research work, but the participation was independent of the commercial interests of the company. There are no financial competing interests in conducting or reporting this work. There are no competing interests for any of the other authors.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Adurthi, S., Kumar, M.M., Vinodkumar, H.S. et al. Oestrogen Receptor-α binds the FOXP3 promoter and modulates regulatory T-cell function in human cervical cancer. Sci Rep 7, 17289 (2017). https://doi.org/10.1038/s41598-017-17102-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-17102-w
This article is cited by
-
An increase in tumor-infiltrating lymphocytes after treatment is significantly associated with a poor response to neoadjuvant endocrine therapy for estrogen receptor-positive/HER2-negative breast cancers
Breast Cancer (2023)
-
Revisiting steroidogenesis and its role in immune regulation with the advanced tools and technologies
Genes & Immunity (2021)
-
The impact of 17β-estradiol and progesterone therapy on peripheral blood mononuclear cells of asthmatic patients
Molecular Biology Reports (2021)
-
Increased β-cell proliferation before immune cell invasion prevents progression of type 1 diabetes
Nature Metabolism (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.