Abstract
The gene coding for the forkhead box protein P2 (FOXP2) is associated with human language disorders. Evolutionary changes in this gene are hypothesized to have contributed to the emergence of speech and language in the human lineage. Although FOXP2 is highly conserved across most mammals, humans differ at two functional amino acid substitutions from chimpanzees, bonobos and gorillas, with an additional fixed substitution found in orangutans. However, FOXP2 has been characterized in only a small number of apes and no publication to date has examined the degree of natural variation in large samples of unrelated great apes. Here, we analyzed the genetic variation in the FOXP2 coding sequence in 63 chimpanzees, 11 bonobos, 48 gorillas, 37 orangutans and 2 gibbons and observed undescribed variation in great apes. We identified two variable polyglutamine microsatellites in chimpanzees and orangutans and found three nonsynonymous single nucleotide polymorphisms, one in chimpanzees, one in gorillas and one in orangutans with derived allele frequencies of 0.01, 0.26 and 0.29, respectively. Structural and functional protein modeling indicate a biochemical effect of the substitution in orangutans, and because of its presence solely in the Sumatran orangutan species, the mutation may be associated with reported population differences in vocalizations.
Similar content being viewed by others
Introduction
Language is a defining feature of human uniqueness. Therefore, the cognitive, motor, and neural foundations that distinguish human speech and language from other animal communication systems have been a central focus of research in the social and biological sciences for more than 200 years1,2. To address the puzzle of human language origins, it is essential to examine the cognitive processes, neurobiology, and genetics underlying this unique form of communication in an evolutionary context, particularly in comparison to our species’ closest living relatives, the great apes (chimpanzees, bonobos, gorillas and orangutans)3.
Although it has been presumed that great apes are limited in their vocal capacity to produce the range of sounds in human speech4,5, they do show local variation in vocal calls that appear to be inherited across generations through social transmission (for review see6). Such vocal learning refers to the ability of an individual to modify the acoustic features or timing of existing species-typical calls, or to learn new calls altogether, expanding the vocal repertoire. Evidence for vocal learning by call modification has been documented in several primate species and recent studies also report some capacity for vocal invention, which typically encompasses voiceless vocalizations6,7,8,9,10,11. For example, in chimpanzees, the use of novel vocal signals, such as attention-getting sounds7,9 has been reported in captive populations, as a means of attracting the attention of an otherwise inattentive audience12. The capacity to produce these vocalizations is heritable and possibly socially learned, suggesting that these apes have voluntary control of both vocal signals and orofacial musculature8. Some apes can also acquire and use symbols during two-way interspecies communication through alternative and augmentative systems such as American Sign Language13 and visual-graphic symbols14,15. Furthermore, many parallels have been found in the gestures of apes compared to preverbal children, such as initiation and responding to pointing cues, intentional and referential signaling, and the elaboration and repair of failed communication16,17,18. Many nonverbal behaviors found in great apes, such as joint-attention19, have also been observed in preverbal children just prior to the onset of speech, which might serve as part of the cognitive foundation of language development20. Thus, in terms of understanding language evolution, great apes represent key reference species.
The genetic changes responsible for the human capacity for increased vocal learning likely occurred since our lineage split from chimpanzees and bonobos. One well-studied candidate is the gene coding for the transcription factor FOXP2 (forkhead box P2)21,22. FOXP2 is the first gene that was discovered to be associated with language disorders and fine orofacial motor control, as two functional copies are required for normal development of speech and language in humans21,22,23. Mutations affect primarily the coordination of orofacial movements required for speech24. Several recent studies compared the evolution of this gene in primates and other species21,25,26,27,28,29. Interestingly, the protein coding sequence is among the most highly conserved 5% of proteins in vertebrates, and its role in regulating vocal learning and communication appears to be shared across a range of animal species21,29,30,31,32, as is the expression of the gene in several key brain regions related to language and fine motor control28,33,34. More specifically, the gene is crucial for the development and function of brain circuits involving the neocortex, basal ganglia and cerebellum32,35,36,37. FOXP2 mRNA is expressed in these brain regions among mammals and avians, reinforcing the view that it plays a role in speech in humans and motor learning in other species, such as birds and mice28,38.
Although a number of nucleotide changes have accumulated in FOXP2 since primates diverged from the mouse lineage around 70 million years ago, only one of these changes resulted in an amino acid substitution (i.e. a nonsynonymous mutation)27. Strikingly, two more amino acid differences are found specifically on the human lineage, with an additional fixed lineage-specific difference in orangutans27. This indicates that modern humans have a uniquely derived version of FOXP2 that arose since the last common ancestor shared with the Pan lineage, only 4 to 6 million years ago3,27. In comparison to the highly conserved sequence, the rate of amino acid substitutions in this gene in modern humans is higher than expected by chance, indicating a signal of accelerated evolution27. Subsequent functional assessments of the human-specific changes to FOXP2 using mice engineered to express the human variant of the gene revealed changes in synaptic plasticity, axon and dendrite outgrowth, and physiological activity in medium spiny neurons of the striatum, supporting the idea that the human variant of FOXP2 causes alteration in brain development36,39. To date, however, it remains unclear when in the past these amino acid substitutions first occurred, as modern humans share them with both Neanderthals and Denisovans, indicating they originated at least ~400,000 years ago40. Recent findings suggest that other regulatory changes in the gene unique to modern humans lie at the base of the selection signal40, and experimental evidence shows that the human FOXP2 variant differentially regulates downstream targets compared to the ancestral version found in chimpanzees41.
Despite such tremendous interest in the FOXP2 gene among linguists, anthropologists, geneticists and neuroscientists, our understanding of within-species variation of this gene among nonhuman primates remains limited (but see42,43). Interestingly, the gene has been better studied in bats than primates, and despite the high conservation of the protein sequence in a majority of mammals, bats show remarkable coding variation in the FOXP2 gene which may be linked to differences in echolocation systems among species26. Thus, further examination of FOXP2 variation across primate species holds the potential to provide insight into the evolution of vocal control and communication systems within the human lineage.
The aim of this study was to assess FOXP2 coding variation in a relatively large sample of great apes. We also investigated potential within-species allelic length variation in the two polyglutamine (poly Q) tracts located in exons 5 and 6 of the gene. Poly Q tracts are encoded by a mixture of CAG and CAA codons repeated in tandem; these types of repeats typically have higher mutation rates and can serve as a functional modulator of eukaryotic transcription factors44. Poly Q tract variation has been shown to impact gene expression by regulating gradients of expression akin to a “tuning-knob” effect, as shown for example with the RUNX2 transcription factor45. As FOXP2 both upregulates and downregulates a large array of different target genes in human basal ganglia and inferior frontal cortex41, poly Q length variation could have important consequences for gene expression. Despite the nature of these repeats, length variation in the gene in humans is rare23,46 and is therefore commonly overlooked. Studies in nonhuman primates, in contrast, do report between-species length differences of these repeats27,43, but none of these studies to date have investigated within-species length variation of these poly Q tracts.
Results
FOXP2 coding variation
We identified 52 single nucleotide variants (SNV) among apes, of which 21 were fixed species-specific substitutions, and 31 were within-species single nucleotide polymorphisms (SNPs) (Fig. 1, Table S5). Out of 31 within-species SNPs found, three were nonsynonymous substitutions (Fig. 2). In chimpanzees, an A/T SNP in the first translated exon resulted in a Threonine to Serine substitution (Thr46Ser), present in just one individual in our sample (minor allele frequency = 0.008). In gorillas, a G/T SNP in exon 7 leads to an Alanine to Serine substitution (Ala326Ser). This SNP is only found in western lowland gorillas (Gorilla gorilla gorilla), with the ancestral G allele present at a higher frequency (0.74). Both allele and genotype frequencies were in Hardy Weinberg equilibrium (X 2 = 1.05, df = 1, p = 0.305), indicating that potential genotyping errors such as allelic dropout, did not pose a problem in this study. In orangutans, a C/A SNP in exon 16 causes a Proline to Threonine substitution (Pro626Thr). The latter SNP is found only in Sumatran orangutans (Pongo abelii), with the ancestral C allele again present at a higher frequency (0.71). Allele and genotype frequencies were in Hardy Weinberg equilibrium (X 2 = 2.20, df = 1, p = 0.138).
Alignment of the FOXP2 amino-acid sequences across apes. Both polyglutamine stretches (glutamine, denoted as Q, repeated 39–42 and 9–11 times in tandem see positions 151–210) are shaded. Dots indicate similarity to the consensus sequence. For species where within-species variation in amino acid substitutions were found, an individual with the ancestral (A) and derived (D) sequence are shown.
We also identified length variation in the poly Q tracts of both orangutans and chimpanzees (Table 1). In orangutans, two alleles were found for each poly Q tract, always differing in length by one 3 bp glutamine (Q) codon. In chimpanzees, two alleles were found for poly Q tract one (exon 5) and three alleles were distinguished in poly Q tract two (exon 6), again all differing by one Q codon. Allele and genotype frequencies were in Hardy Weinberg equilibrium for both loci in chimpanzees (Q1: X 2 = 0.47, df = 1, p = 0.491; Q2: X 2 = 0.50, df = 3, p = 0.919), and for Q2, but not Q1 in orangutans (Q1: X 2 = 14.88, df = 1, p = 0.0001; Q2: X 2 = 0.04, df = 1, p = 0.850).
Prediction of functional consequences and protein structure modeling for amino acid substitutions
SNAP2 prediction of the functional consequences of the chimpanzee Thr46Ser, gorilla Ala326Ser and orangutan Pro626Thr substitutions resulted in SNAP2 effect scores of −65 (expected accuracy 82%), −76 (expected accuracy 87%) and 64 (expected accuracy 80%), respectively (Fig. 3B).
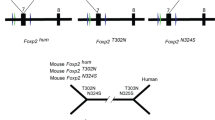
(A) Schematic representation of the FOXP2 gene. Boxes represent exons and lines represent introns. Translated exons included in this study are shaded in black. The domains coded for by the exons are shown above: two polyglutamine tracts (Q40 and Q10), a zinc-finger motif (ZnF), a leucine-zipper (LeuZ), the forkhead domain FOX, and an acidic C-terminus. CpG marks the site of a CpG island. (B) Heatmap showing predicted functional consequences of P626T mutation found in orangutans. The stronger the predicted effect, the redder; the stronger the predicted neutrality, the bluer. (C) Sequence alignment of the wild-type and Pro626Thr mutant FOXP2-CTR proteins. Secondary structure predictions are shown for each of the three algorithms used. (D) Comparison of the predicted tertiary structures of the wild-type and Pro626Thr FOXP2-CTR proteins; helix 2 of each structure are superimposed for comparison. Residues Pro626 and Thr626 are shown in cyan and as both sticks and spheres.
Next, the secondary and tertiary protein structural differences for ancestral (wild-type) and species-specific (mutant) FOXP2 regions containing the Ala326Ser and Pro626Thr substitutions were predicted. We refrained from structural modeling for Thr46Ser, because the mutation was only present in one chimpanzee and functional modeling predicted a neutral effect. The Ala326Ser and Pro626Thr substitutions were located in the region following the N-terminal Poly Q region (FOXP2-APQ, residues 210–339), and the FOXP2 C-terminal region (FOXP2-CTR, residues 583–715), respectively. Secondary structure prediction of both regions showed similar results when modeled as full-length FOXP2 sequences or as isolated regions. For both regions, two of the secondary prediction algorithms predicted two α-helical segments, while the other algorithm identified three. The helical segments were identified primarily within the first two-thirds of the region with the last 50 C-terminal amino acids predicted to be mainly unstructured. No differences in the secondary structure predictions were observed when the sequences included the Ala326Ser and Pro626Thr mutations. Both mutation sites were observed in regions predicted to be unstructured loops (Fig. 3C and Supplementary Fig. S1A). The Ala326Ser mutation is located in the last ~50 unstructured amino acids of the FOXP2-APQ region. For comparison, the secondary structure of the FOXP2 forkhead domain matched well with its known structure, consisting of four α-helices and two β-strands.
Tertiary structure predictions of the wild-type and Ala326Ser FOXP2-APQ showed that the region consists of three α-helices, one of which adopts a long central helix (Supplementary Fig. S1B). The α-helices encompass residues Ala210-Gln239, Pro262-Val272 and Met278-Gly286. As expected, Ala326 is in an unstructured region approximately 15 residues from the C-terminus. Analyses of all ten predicted structures of wild-type FOXP2-APQ revealed variation in the orientations of α-helices 2 and 3 and the location of the Ala326 residue. To identify the effect of the Ala326Ser mutation, the first and longest helix between the two sets of structures were superimposed. The structural comparison revealed variation in the orientations and locations of the Ser326 residue and α-helices 2 and 3, which were similar to the variation observed in the predicted structures of the wild-type FOXP2-APQ (Supplementary Fig. S1B). It is thus unclear whether the Ala326Ser mutation has a structural impact within this region. We then investigated the S321SVLNXRRDS region, in which X indicates the mutation (A or S), for phosphorylation signatures since serine amino acids are targeted by kinases. When X is alanine (wild-type), Ser322 and Ser330 would be sites for G-protein coupled receptor (GPCR) kinase and protein kinase C, respectively. When X is serine (mutation), the Ser326 introduces motifs for MAP kinases (SxxxpS326), PKA (S326RR), pyruvate dehydrogenase kinase (pS326xxDxx), glycogen synthase kinase (S322xxxpS326), and casein kinase (S326RRD), in which x = any amino acid (Supplementary Fig. S1C).
Next, we examined the Pro626Thr mutation within the FOXP2-CTR region. Given that Pro626 is observed in a loop region between α-helices 2 and 3, it is unclear how the Pro626Thr mutation might affect the structure and function of FOXP2. The predicted structures of the wild-type and Pro626Thr FOXP2-CTR contain three α-helices, found within the N-terminal half of the protein, while the C-terminal half consists of unstructured loops (Fig. 3D). The three α-helices encompass residues Pro591-Leu601, Gly604-Ala616 and Ala639-D649, respectively, held together by hydrophobic interaction. In contrast, the α-helices for the mutant FOXP2-CTR are 1–3 amino acids shorter. To identify the effect of the Pro626Thr mutation located in the loop between α-helices 2 and 3, the central and longest helix between the two structures was superimposed to reveal that α-helices 1 and 3 are oriented differently with respect to helix 2 (Fig. 3D). Likewise, the positions of Pro626 and Thr626, in their respective structures, are different.
Models of FOXP2 protein evolution (dN/dS sites analysis) show evidence of strong purifying selection across the polypeptide (Supplementary Fig. S2). Neither SLR (sitewise likelihood-ratio) nor codeml analyses detected evidence of positively selected sites, and likelihood ratio tests using both methods indicated a model of predominantly purifying selection with some sites experiencing neutral selection to be the most likely model. The three sites where we identified nonsynonymous single nucleotide polymorphisms had low dN/dS, with Thr46Ser and Pro626Ser yielding ratios of 0, and Ala326Ser yielding a ratio of 0.26.
Discussion
Within-species coding sequence variation of FOXP2 in the largest samples of great apes analyzed to date reveals likely functional variation potentially associated with communication skill and orofacial motor control. Length variation was found in both polyglutamine tracts in chimpanzees and orangutans. Additionally, three species-specific nonsynonymous SNPs were found, Thr46Ser in chimpanzees, Ala326Ser in gorillas and Pro626Thr in orangutans. Protein structure modeling did not indicate a clear effect for Ala326Ser, but it did reveal a potential impact of the Pro626Thr variant in orangutans: the mutation was predicted to alter the tertiary structure of a DNA binding interface in the C terminal region. Our results further confirmed interspecies differences in functional coding sequences between humans and great apes, as reported previously27,42,43.
The between-species comparison of the length of the two poly Q tracts is generally congruent with earlier reports. However, chimpanzees and orangutans show within-species variation in the length of both poly Q tracts, which is surprising given that reported variation in these tracts is rare in human studies46,47,48,49. One explanation is that orangutans and chimpanzees have much higher genetic diversity than humans, with populations having diverged millions of years ago3. Poly Q tract variation in transcription factors can potentially impact expression of downstream genes via a sort of tuning-knob effect50. Notably, the few studies that do report variation in these poly Q tracts show that they are likely functional, as higher frequencies of rare alleles are found in human individuals suffering from speech and/or language impairments46,48,49. Since the sequencing coverage at the position of these poly Q tracts was low in the whole-genome data, and we had an insufficient number of DNA samples for a large enough set of unrelated gorillas and bonobos, we were only able to examine poly Q length variation in orangutans and chimpanzees. Further sampling is required for the other species to determine whether potential within-species length variation exist in these tracts. Interestingly, chimpanzees show remarkable individual variation in the frequency and consistency of the use of vocal attention-getting sounds. The source of individual variation in the use of such vocalizations remains largely unknown, but since it is heritable8, it likely has a genetic component. Therefore, the individual genetic variation in both poly Q tracts presented in this study offers promising candidate loci for future research.
The Thr46Ser mutation identified in chimpanzees was present in just one individual, indicating that it is a rare variant. Therefore, it is difficult to investigate its potential phenotypic significance. Because of this, and the fact that functional SNAP2 prediction resulted in a likely neutral effect, we did not perform further structural modeling for this variant. In humans, many cases of rare de novo and familial nonsynonymous mutations and deletions in FOXP2 have been reported, and disruption of the gene typically results in severe motor speech disorders, or differences in cognitive and/or generalized motor skills22 but see51. It is unclear whether the Thr46Ser mutation affects this individual chimpanzee’s vocal, cognitive or motor skills, and it could potentially be due to a sequencing error. Selection modeling shows that the dN/dS ratio found at Ala326Ser was slightly higher than in other sites of the sequence, suggesting selection at this site may be somewhat more relaxed.
The second and third species-specific nonsynonymous variants, Ala326Ser and Pro626Thr, are present in western lowland gorillas and Sumatran orangutans, respectively. Both mutations had a relatively high frequency in the populations in which they were present, occurring in almost half of the genotyped individuals. Based on the three-dimensional homology search algorithm, DALI, the spatial orientation of the three α-helices in both the FOXP2-APQ and FOXP2-CTR structures revealed a helix-turn-helix motif commonly observed in DNA binding proteins52, such as transcription factors and repressor proteins. This is consistent with both regions located adjacent to the DNA-binding forkhead domain. The gorilla Ala326Ser variant is located in exon 7, which is the same exon that contains both previously reported fixed human nonsynonymous mutations27. The orangutan Pro626Thr mutation is located in exon 16, close to exons 12–14, which code for the DNA binding forkhead domain.
Although FOXP2 is under strong purifying selection and no sites showed significance evidence of directional selection, there are notable patterns of potential convergence across mammals. The exons of interest where variation was found in apes (exon 7, 16 and 17) also have the highest numbers of nonsynonymous fixed substitutions in bats26. This is interesting given that previous research suggests that the bat variants in these regions likely have adaptive significance, as opposed to being due to relaxed selection26.
For the Ala326Ser mutation found in gorillas, both functional and structural protein modeling revealed no clear differences between the wild-type and mutant FOXP-APQ region. However, since the mutation introduces a serine at position 326, and serine amino acids are commonly targeted by kinases, the mutation may still alter phosphorylation signatures at this position. The serine introduces more possibilities for phosphorylation by additional kinases that can affect how FOXP2 interacts with other proteins associated with gene expression. Although post-transcriptional modifications including sumoylation can have a functional impact on FOXP253, there is no empirical evidence of phosphorylation of the ancestral version of FOXP2 (but see27). While the Ala326Ser mutation may increase the chance of phosphorylation, it is unclear if this is actually the case at Ser326, so further experimental assays are needed to investigate the functionality of this SNP in gorillas. However, little evidence from field studies indicates differences in vocal repertoire, vocal learning or orofacial motor control, either within western lowland gorillas or between gorilla populations that could be attributable to the SNP found in this study54. Although there are reported differences in the production of “raspberry” vocalizations in wild mountain gorillas, but not in western lowland gorillas54, this observation is not easily reconciled with our finding that the SNP is present in some but not all western gorillas. Future studies of wild gorilla vocal behavior, nevertheless, might reveal if there is a relationship between this phenotypic variation and the genetic polymorphism we describe here.
For the Pro626Thr mutation found in orangutans, both functional and structural protein modeling showed a high likelihood that the mutation alters the native protein. Interestingly, compared to FOXP2-APQ, the FOXP2-CTR region does not contain a significant amount of lysine, arginine and aromatic amino acids, commonly involved in protein-nucleic acid interactions. Instead, it consists of many polar residues (serine, glutamine, asparagine, threonine) at the helix-turn-helix DNA binding interface that can form hydrogen bonds with the bases of DNA. The structure of the Pro626Thr mutant showed a complete rearrangement of these polar residues that may affect the DNA binding property of FOXP2-CTR. Further experimental evidence remains to be collected to confirm the exact functional effect of this mutation. However, it is interesting to note that the Pro626Thr mutation was found solely in Sumatran orangutans, which show remarkable differences in behavior and vocal skills compared to their Bornean sister species.
The Sumatran orangutans live in habitats with more stable food availability, they are more sociable and show lower frequencies of forced mating55. Notably, these populations also differ in their vocal repertoire and the pitch frequency of vocalizations56,57. For example, male long calls differ consistently between Sumatran and Bornean orangutans in number of pulses per call, call speed, call duration, bandwidth, pulse duration and dominant frequency57. Furthermore, male orangutans in Borneo are reportedly larger than Sumatran males and are therefore expected to have lower call pitches, yet exactly the opposite is found57. Since FOXP2 variation has been linked to differences in vocal learning and vocalization frequency differences in a variety of species28,32,38, the Pro626Thr mutation could be associated with reported vocal differences, not only between the two orangutan species, but also between individuals belonging to different populations within Sumatra, since the SNP is not present in all Sumatran individuals57. Results from captive studies also show that orangutans have the ability for vocal fold control10 and that they can more skillfully imitate human speech than any other apes11. This combined with evidence suggesting that Sumatran individuals are more avid oral tool users compared to the Bornean orangutans58, may be related to reported associations between FOXP2 mutations and levels of orofacial motor control25. These observations suggest that future studies of Pro626Thr may reveal salient individual or population differences in vocal behavior in Sumatran orangutans, although we cannot entirely rule out demographic history of these species as an additional factor influencing the genotype distribution pattern in this study59.
Overall, despite the relatively large number of great apes sampled in this study, the number of nonsynonymous substitutions found was low, indicating that FOXP2 is highly constrained and likely under purifying selection in great apes, and mammals in general26,27. Therefore, investigation of the impact of rare nonsynonymous variants found here in great apes could shed light on the proximate mechanisms shaping individual, population or even species level differences in vocal skills and orofacial motor control. Furthermore, although this study focused on coding variation because of its direct impact on protein structure and function, variation in regulatory and intronic regions, as well as tissue-specific alternative splice variants, also warrant further study47,60.
Methods
Genome sequencing, assembly and annotation
Genotyping was performed using publicly available whole genome data for chimpanzees (Pan troglodytes troglodytes N = 18, Pan troglodytes verus N = 12, Pan troglodytes schweinfurthii N = 16, Pan troglodytes ellioti N = 10, Pan troglodytes unknown subspecies N = 3), bonobos (Pan paniscus, N = 9), gorillas (Gorilla beringei beringei N = 7, Gorilla beringei graugeri N = 9, Gorilla gorilla diehl N = 1, Gorilla gorilla gorilla N = 27) and orangutans (Pongo pygmaeus N = 20, Pongo abelii N = 17)3,61,62. All genomes were mapped to human genome (version hg19) using BWA-MEM v0.7.5a-r405 (http://bio-bwa.sourceforge.net/bwa.shtml) with default parameters. After removing duplicates using PICARD v1.91 (https://sourceforge.net/projects/picard/files/picard-tools/1.91/), single nucleotide polymorphisms were called using GATK UnifiedGenotyper (https://software.broadinstitute.org/gatk/documentation/tooldocs/current/org_broadinstitute_gatk_tools_walkers_genotyper_UnifiedGenotyper.php). Gene consensus sequences were built for each individual using vcfconsensus (http://vcftools.sourceforge.net/man_latest.html). Intronic regions and non-protein coding exons were removed from the sequences and remaining exons were aligned to the human FOXP2 coding reference sequence (Ensembl: ENSG00000128573) using Geneious (version 6.0.6).
Sanger sequencing of exons
To maximize our sample size for this study, we included additional data from a preliminary study42,43, where genomic DNA was extracted from peripheral whole blood of four African-born chimpanzees (Pan troglodytes, including 1 P.t.verus, 1 P.t.troglodytes, 1 P.t.schweinfurthii, and 1 probable P.t.troglodytes/schweinfurthii), bonobos (Pan paniscus, N = 2), western lowland gorillas (Gorilla gorilla gorilla, N = 2), Sumatran orangutans (Pongo abelii, N = 2) and white-handed gibbons (Hylobates lar, N = 2). For details about the chimpanzees, subspecies ascertainment, DNA extraction, polymerase chain reaction (PCR) and DNA sequencing, see63. All coding exons (Fig. 3A) were amplified by PCR (primers and PCR conditions shown in Supplementary Table S1) and Sanger sequenced on a Li-cor 4200 DNA sequencer analyzer following manufacturer’s specifications. Multiple alignments of trimmed nucleotide and deduced amino acid sequences for all exons for each species were performed using Geneious (version 6.0.6). Resulting sequences are deposited in NCBI under accession numbers MG547712-MG547713, MG547714, MG547715, MG547716, MG547717, MG547718, MG547719, MG547720 and MG547721.
Microsatellite analysis of polyglutamine tracts
Since the sequencing coverage at the position of the poly Q tracts was low in the whole-genome data, we scored microsatellite genotypes for both tracts in available panels of chimpanzees (N = 54) and Sumatran orangutans (N = 32). For primer design, PCR conditions and fragment length analyses see supplementary information (Table S3). Individuals were genotyped using automated capillary electrophoresis on Applied Biosystems Genetic Analyzer platforms (DNA Analysis Facility at Yale University).
Function and structure prediction of coding variants
To infer the putative functional consequences of the identified coding variants, we first used SNAP2 to predict the effect of variants on protein function64. SNAP2 is a trained classifier that is based on a machine-learning device called “neural network”. It distinguishes between effect and neutral variants/non-synonymous SNPs by taking a variety of sequence and variant features into account. The effect of a variant is believed to be of importance to the native protein function if the SNAP2 score exceeds 50, neutral if the score is below −50 and unreliable when between 50 and −50.
The secondary structures of the full-length FOXP2 (residues 1–715) and the C-terminal region (residues 583–715, FOXP2-CTR) that follows the forkhead domain of the wild-type FOXP2 protein were predicted by PSIPRED65, Jpred66, and YASPIN67. A third prediction was performed for residues 210–339, that follows the N-terminal Poly Q region of the protein (FOXP2-APQ). The secondary structures of the FOXP2-APQ and FOXP2-CTR regions containing the Ala326Ser and Pro626Thr mutations were also predicted, respectively. As a control, the sequence encompassing the forkhead domain (residues 503–582), which has a known structure, was also submitted for secondary structure prediction.
The tertiary structures of the wild-type and mutant FOXP2-APQ and FOXP2-CTR were predicted using the ab initio modeling program QUARK68. While numerous ab initio modeling programs are available, QUARK has proven to be more accurate when predicting the structure of helical proteins68. For each of the wild-type and both mutant proteins, ten structures were predicted. Structures were ranked by their template modeling (TM) scores, the highest of which corresponds to the best structure, represented by the first model (Model 1). We also submitted two protein sequences for which the structures are known, including the forkhead domain of FOXP2. For these two proteins, the overall predicted and experimental structures have similar folds with root-mean-square deviation values of backbone atom superposition of 4.567 (forkhead, PDB accession code: 2A07) and 4.327 (UBR5 PABC domain, PDB accession code: 3NTW). In case the predicted structures of the wild-type and mutant FOXP2 regions did not yield apparent differences, further investigation was done to identify potential changes in phosphorylation motifs that could affect the function of the region using NetPhorest69.
To further identify potential functional implications of variants, we also investigated the evidence of selection on amino acid sites within FOXP2. We examined site-specific dN/dS ratios across a large alignment of 57 placental mammals (Supplementary Table S4), including 12 primates and five bats. A very low dN/dS ratio (i.e. close to zero) suggests that a site has been evolutionarily highly conserved, and thus is likely critical to protein structure/function. For details on dN/dS ratio calculations see Supplementary Methods 1.
Data Availability
The sequences generated during the current study are available at the NCBI repository.
Ethical statement
No animals were sacrificed or sedated for the purpose of this study. All aspects of this research adhered to the American Psychological Associations guidelines for the ethical treatment of animals in research.
References
Fitch, W. T. The evolution of speech: A comparative review. Trends Cogn. Sci. 4, 258–267 (2000).
Tallerman, M. & Gibson, K. G. The Oxford handbook of language evolution. (Oxford University Press, 2012).
Prado-Martinez, J. et al. Great ape genetic diversity and population history. Nature 499, 471–475 (2014).
Hayes, C. The ape in our house. (Harper and Brothers, 1951).
Kellogg, W. N. Communication and language in the home-raised chimpanzee. Science (80-.). 162, 423–427 (1968).
Lameira, A. R. Bidding evidence for primate vocal learning and the cultural substrates for speech evolution. Neurosci. Biobehav. Rev. (2017).
Russell, J. L., McIntyre, J., Hopkins, W. D. & Taglialatela, J. P. Vocal learning of a communicative signal in captive chimpanzees, Pan troglodytes. Brain Lang. 127, 520–525 (2013).
Taglialatela, J. P., Reamer, L., Schapiro, S. J. & Hopkins, W. D. Social learning of a communicative signal in captive chimpanzees. Biol. Lett. 8, 498–501 (2012).
Hopkins, W. D., Taglialatela, J. & Leavens, D. A. Chimpanzees differentially produce novel vocalizations to capture the attention of a human. Anim. Behav. 73, 281–286 (2007).
Lameira, A. R., Hardus, M. E., Mielke, A., Wich, S. A. & Shumaker, R. W. Vocal fold control beyond the species-specific repertoire in an orang-utan. Sci. Rep. 6, 30315 (2016).
Lameira, A. R. et al. Speech-like rhythm in a voiced and voiceless orangutan call. PLoS One 10, 1–12 (2015).
Leavens, D. A., Hostetter, A. B., Wesley, M. J. & Hopkins, W. D. Tactical use of unimodal and bimodal communication by chimpanzees, Pan troglodytes. Anim. Behav. 67, 467–476 (2004).
Miles, L. H. Language in primates: Perspectives and implications. (Springer, 1983).
Premack, D. Language in chimpanzees? Science (80-.). 172, 808–822 (1971).
Rumbaugh, D. M. Language learning by a chimpanzee: The Lana project. (Academic Press, 1977).
Liebal, K., Call, J. & Tomasello, M. Use of gesture sequences in chimpanzees. Am. J. Primatol. 64, 377–396 (2004).
Pika, S. Gestures of apes and pre-linguistic human children: Similar or different? First Lang. 28, 116–140 (2008).
Pika, S., Liebal, K. & Tomasello, M. Gestural communication in subadult bonobos (Pan paniscus): Repertoire and use. Am. J. Primatol. 65, 39–61 (2005).
Hopkins, W. D. et al. Poor receptive joint attention skills are associated with atypical gray matter asymmetry in the posterior superior temporal gyrus of chimpanzees (Pan troglodytes). Front. Psychol. 5, 1–8 (2014).
Leavens, D. A., Hopkins, W. D. & Thomas, R. K. Referential communication by chimpanzees (Pan troglodytes). J. Comp. Psychol. 118, 48–57 (2004).
Enard, W. FOXP2 and the role of cortico-basal ganglia circuits in speech and language evolution. Curr. Opin. Neurobiol. 21, 415–424 (2011).
Fisher, S. E. & Scharff, C. FOXP2 as a molecular window into speech and language. Trends Genet. 25, 166–177 (2009).
Lai, C. S. L., Fisher, S. E., Hurst, J. A., Vargha-Khadem, F. & Monaco, A. P. A forkhead-domain gene is mutated in a severe speech and language disorder. Nature 413, 519–523 (2001).
Watkins, K., Dronkers, N. & Vargha-Khadem, F. Behavioural analysis of an inherited speech and language disorder: comparison with acquired aphasia. Brain A J. Neurol. 125, 452–464 (2002).
Vargha-Khadem, F., Gadian, D. G., Copp, A. & Mishkin, M. FOXP2 and the neuroanatomy of speech and language. Nat. Rev. Neurosci. 6, 131–138 (2005).
Li, G., Wang, J., Rossiter, S. J., Jones, G. & Zhang, S. Accelerated FoxP2 evolution in echolocating bats. PLoS One 2 (2007).
Enard, W. et al. Molecular evolution of FOXP2, a gene involved in speech and language. Nature 418, 869–72 (2002).
Haesler, S. et al. FoxP2 Expression in Avian Vocal Learners and Non-Learners. J. Neurosci. 24, 3164–3175 (2004).
Webb, D. M. & Zhang, J. FoxP2 in song-learning birds and vocal-learning mammals. J. Hered. 96, 212–216 (2005).
Konopka, G. & Roberts, T. F. Insights into the Neural and Genetic Basis of Vocal Communication. Cell 164, 1269–1276 (2016).
Scharff, C. & Petri, J. Evo-devo, deep homology and FoxP2: implications for the evolution of speech and language. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 366, 2124–40 (2011).
Haesler, S. et al. Incomplete and inaccurate vocal imitation after knockdown of FoxP2 in songbird basal ganglia nucleus area X. PLoS Biol. 5, 2885–2897 (2007).
Kato, M. et al. Human speech- and reading-related genes display partially overlapping expression patterns in the marmoset brain. Brain Lang. 133, 26–38 (2014).
Teramitsu, I., Kudo, L. C., London, S. E., Geschwind, D. H. & White, S. A. Parallel FoxP1 and FoxP2 Expression in Songbird and Human Brain Predicts Functional Interaction. J. Neurosci. 24, 3152–3163 (2004).
French, C. A. et al. An aetiological Foxp2 mutation causes aberrant striatal activity and alters plasticity during skill learning. Mol. Psychiatry 17, 1077–85 (2012).
Groszer, M. et al. Impaired Synaptic Plasticity and Motor Learning in Mice with a Point Mutation Implicated in Human Speech Deficits. Curr. Biol. 18, 354–362 (2008).
Murugan, M., Harward, S., Scharff, C. & Mooney, R. Diminished FoxP2 levels affect dopaminergic modulation of corticostriatal signaling important to song variability. Neuron 80, 1464–1476 (2013).
Fujita, E. et al. Ultrasonic vocalization impairment of Foxp2 (R552H) knockin mice related to speech-language disorder and abnormality of Purkinje cells. Proc. Natl. Acad. Sci. USA 105, 3117–22 (2008).
Vernes, S. C. et al. FOXP2 regulates gene networks implicated in neurite outgrowth in the developing brain. PLoS Genet. 7 (2011).
Maricic, T. et al. A recent evolutionary change affects a regulatory element in the human FOXP2 gene. Mol. Biol. Evol. 30, 844–852 (2013).
Konopka, G. et al. Human-specific transcriptional regulation of CNS development genes by FOXP2. Nature 462, 213–217 (2009).
Ely, J. J. et al. Identical DNA sequences in FOXP2 language gene between humans, chimpanzees (Pan troglodytes) and bonobos (Pan paniscus). In The American Society of Human Genetics meeting (American Journal for Human Genetics, 2002).
Ely, J. J. et al. DNA sequence variation in FOXP2 language gene among humans and great apes. in American Society of Human Genetics meeting (American Journal for Human Genetics, 2002).
Gemayel, R. et al. Variable glutamine-rich repeats modulate transcription factor activity. Mol. Cell 59, 615–627 (2015).
Sears, K., Goswami, A., Flynn, J. & Niswander, L. The correlated evolution of Runx2 tandem repeats, transcriptional activity, and facial length in Carnivora. Evol Dev 9, 555–565 (2007).
Newbury, D. F. et al. FOXP2 is not a major susceptibility gene for autism or specific language impairment. Am. J. Hum. Genet. 70, 1318–1327 (2002).
Bruce, H. A. & Margolis, R. L. FOXP2: Novel exons, splice variants, and CAG repeat length stability. Hum. Genet. 111, 136–144 (2002).
Zhao, Y. et al. Association between FOXP2 gene and speech sound disorder in Chinese population. Psychiatry Clin. Neurosci. 64, 565–573 (2010).
Wassink, T. H. et al. Evaluation of FOXP2 as an autism susceptibility gene. Am. J. Med. Genet. - Neuropsychiatr. Genet. 114, 566–569 (2002).
Kashi, Y. & King, D. G. Simple sequence repeats as advantageous mutators in evolution. TRENDS Genet. 22, 253–259 (2006).
Mueller, K. L. et al. Common genetic variants in FOXP2 are not associated with individual differences in language development. PLoS One 11, 1–17 (2016).
Rohs, R. et al. Origins of specificity in protein-DNA recognition. Annu. Rev. Biochem. 79, 233–269 (2010).
Usui, N. et al. Sumoylation of FOXP2 regulates motor function and vocal communication through purkinje cell development. Biol. Psychiatry 1–11 (2015).
Robbins, M. M. et al. Behavioral variation in gorillas: Evidence of potential cultural traits. PLoS One 11, 1–18 (2016).
van Schaik, C. P., Marshall, A. J. & Wich, S. A. In Orangutans: Geographic Variation in Behavioral Ecology and Conservation (eds. Wich, S. A., Atmoko, S. S. U., Setia, T. M. & van Schaik, C. P.) (OUP Oxford, 2008).
Wich, S. A. et al. Call cultures in orang-utans? PLoS One 7 (2012).
Delgado, R. A. J. Geographic variation in the long calls of male orangutans (Pongo spp.). Ethology 113, 487–498 (2007).
van Schaik, C. P. et al. In Orangutans: Geographic Variation in Behavioral Ecology and Conservation (eds. Wich, S. A., Utami-Atmoko, S. S., Mitra Setia, T. & van Schaik, C. P.) (OUP Oxford, 2008).
Nater, A. et al. Reconstructing the demographic history of orang-utans using Approximate Bayesian Computation. Mol. Ecol. 24, 310–327 (2015).
Haygood, R., Babbitt, C. C., Fedrigo, O. & Wray, G. A. Contrasts between adaptive coding and noncoding changes during human evolution. Proc. Natl. Acad. Sci. 107, 7853–7857 (2010).
Xue, Y. et al. Mountain gorilla genomes reveal the impact of long-term population decline and inbreeding. Science (80-.). 348, 242–245 (2015).
Manuel, Mde et al. Chimpanzee genomic diversity reveals ancient admixture with bonobos. Science (80-.). 354, 477–481 (2016).
Ely, J. J. et al. Subspecies composition and founder contribution of the captive U.S. chimpanzee (Pan troglodytes) population. Am. J. Primatol. 67, 223–241 (2005).
Hecht, M., Bromberg, Y. & Rost, B. Better prediction of functional effects for sequence variants From VarI-SIG 2014: Identification and annotation of genetic variants in the context of structure, function and disease. BMC Genomics 16(Suppl 8), S1 (2015).
McGuffin, L. J., Bryson, K. & Jones, D. T. The PSIPRED protein structure prediction server. Bioinformatics 16, 404–405 (2000).
Cole, C., Barber, J. D. & Barton, G. J. The Jpred 3 secondary structure prediction server. Nucleic Acids Res. 36, W197–201 (2008).
Lin, K., Simossis, V. A., Taylor, W. R. & Heringa, J. A simple and fast secondary structure prediction method using hidden neural networks. Bioinformatics 21, 152–159 (2005).
Xu, D. & Zhang, Y. Ab initio protein structure assembly using continuous structure fragments and optimized knowledge-based force field. Proteins 80, 1715–1735 (2012).
Horn, H. et al. KinomeXplorer: an integrated platform for kinome biology studies. Nat. Methods 11, 603–604 (2014).
Acknowledgements
We thank collaborators for help with providing samples, specifically: the Yerkes National Primate Research Center and The Michale E. Keeling Center for Comparative Medicine and Research. We also thank Susmita Shrivastava, and all members of the Primate Genomics Lab and the Laboratory for Evolutionary Neuroscience at the George Washington University for helpful feedback on the project. This work was partially supported by NIA grant R43-AG17802 to JJE, by MINECO BFU2014-55090-P (FEDER), Howard Hughes International Early Career and Secretaria d’Universitats i Recerca del Departament d’Economia i Coneixement de la Generalitat de Catalunya to TMB, the James S. McDonnell Foundation (grant 220020293) and by NSF INSPIRE (SMA -1542848) for NS, CCS, WDH and BJB.
Author information
Authors and Affiliations
Contributions
B.J.B., C.C.S., W.D.H. and N.S. developed the study and designed the experimental set-up. Genomic/genotype data were provided by M.D.M., T.M.B., M.K. and J.J.E., and analyzed by N.S. Protein modelling was done by K.W. and M.M., and evolutionary selection modelling was done by E.E.G., N.S. also generated/analyzed the microsatellite data and wrote the manuscript with editing from all coauthors involved.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Staes, N., Sherwood, C.C., Wright, K. et al. FOXP2 variation in great ape populations offers insight into the evolution of communication skills. Sci Rep 7, 16866 (2017). https://doi.org/10.1038/s41598-017-16844-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16844-x
This article is cited by
-
Functional dissection of two amino acid substitutions unique to the human FOXP2 protein
Scientific Reports (2023)
-
Dissecting the cross-trait effects of the FOXP2 GWAS hit on clinical and brain phenotypes in adults with ADHD
European Archives of Psychiatry and Clinical Neuroscience (2023)
-
Sociality predicts orangutan vocal phenotype
Nature Ecology & Evolution (2022)
-
Orangutans show active voicing through a membranophone
Scientific Reports (2019)
-
Coupled whole-body rhythmic entrainment between two chimpanzees
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.