Abstract
Homologous Recombination (HR) repair is essential for repairing DNA double strand breaks (DSB) in dividing cells and preventing tumorigenesis. BRCA2 plays an important role in HR by recruiting the DNA recombinase RAD51 to the DSB. Despite being a popular model organism in genetic and cancer research, knowledge on the conservation of the HR pathway and function of zebrafish Brca2 is limited. To evaluate this, we developed a Rad51 foci assay in zebrafish embryos. We identified the zebrafish embryonic intestinal tissue as an ideal target for Rad51 immunostaining. After inducing DSB through irradiation, Rad51 foci were present in irradiated embryos but not in unirradiated controls. We present a method for accurate quantification of HR. Both morpholino-induced knockdown and knockout of Brca2 lead to almost complete absence of Rad51 foci in irradiated embryos. These findings indicate conserved function of Brca2 in zebrafish. Interestingly, a statistically significant decrease in Rad51 foci was observed in Brca2 heterozygous carriers compared to wild types, indicative of haploinsufficiency, a hypothesised cause of some tumours in patients with a germline BRCA2 mutation. In conclusion, we demonstrated the suitability of zebrafish as an excellent in vivo model system for studying the HR pathway and its functionality.
Similar content being viewed by others
Introduction
In every cell, the genomic material is constantly being damaged by exogenous or endogenous mechanisms1. Double Strand Breaks (DSB) are especially pathogenic, as there is no complementary base template available for accurate repair. These DSB can be repaired through two pathways: Non-Homologous End Joining (NHEJ) and Homologous Recombination (HR)1,2. NHEJ ligates the DNA ends, but does so in an error-prone fashion3. Misrepair can lead to gross chromosomal translocations, amplifications of proto-oncogenes, or deletions of tumour suppressor genes, all of which can result in tumorigenesis and ultimately carcinogenesis1,4. HR uses the sister chromatid as a template to repair in an error-free fashion. Therefore, HR can only occur during the late S-/G2-phase of the cell cycle, when the DNA has been duplicated prior to cell division3,5,6. Two key players in the HR pathway are BRCA1 and BRCA2, which play a crucial role in recruiting the DNA recombinase RAD51 to the vicinity of the DSB. Localisation of RAD51 to the site of the DSB is essential for strand invasion of the sister chromatid and can be visualised through a RAD51 foci immunostaining. Knockdown of BRCA1 or BRCA2 leads to loss of these RAD51 foci6,7. Germline mutations in BRCA1 or BRCA2 are associated with a highly increased risk for breast and ovarian cancer6.
Zebrafish (Danio Rerio) contains orthologues for many genes involved in DNA repair pathways that are available in higher eukaryotes, including the brca2 gene8,9,10. Despite its poor overall homology with human BRCA2 (22%), the functional N-terminal transcription activation domain, C-terminal DNA binding domain and RAD51 binding BRCA repeats are conserved in the zebrafish Brca2 protein9,10. It was shown that zebrafish brca2 acts as a caretaker gene and is involved in maintaining genomic integrity through HR9,10. Brca2 deficient zebrafish models were shown to be sensitive to mitomycin C (MMC), a DNA crosslinker for which repair through HR is essential, and develop an all-male phenotype due to failure of meiotic recombination during the sex determination period9,10. Human BRCA2 −/− patients develop Fanconi Anaemia, which is characterised by genomic instability and sensitivity to MMC11. Liu et al. provided evidence that zebrafish Rad51 is essential for HR in zebrafish, as morpholino mediated knockdown of this protein resulted in decreased HR12.
So far, up to 20 Fanconi Anaemia (FA) genes have been identified (FANCA, B, C, D1 (BRCA2), D2, E, F, G (XRCC9), I, J (BRIP1), L, M, N (PALB2), O (RAD51C), P (SLX4), Q (ERRC4), R (RAD51), S (BRCA1), T (UBE2T) and U (XRCC2))13. With the notable exception of BRCA1, zebrafish has the complete set of FA genes at its disposal14. The apparent absence of zebrafish brca1, a gene with a plethora of important functions, suggests that some modifications between the human and zebrafish HR repair pathway have occurred during evolution14,15.
Despite the importance of HR in maintaining genomic stability, not much is known about this pathway in zebrafish8. As zebrafish is an increasingly important model organism for studying cancer (pathway discovery, compound screening, drug discovery, …) it is crucial to unravel how this repair pathway functions in zebrafish16,17. Therefore, we developed the Rad51 foci assay in zebrafish, enabling the accurate measurements of in vivo HR activity under a wide range of experimental conditions.
Results
Determination of proliferative tissues in 72 hpf zebrafish embryos
As HR can only occur in dividing cells, it is essential to target a tissue that has high proliferative capabilities. Sagittal and transversal sections of 72 hours post fertilisation (hpf) wild type embryos were made and stained for Proliferating Cell Nuclear Antigen (PCNA), which stains cells in G1-, S-, G2- and M-phase of the cell cycle and acts as a general marker for dividing cells. PCNA is most noticeable in the tectum and throughout the intestinal tract (Fig. 1a,b). All nuclei in the intestinal tract show strong PCNA staining, which indicates the high proliferative capacity of these cells at 72 hpf. Due to the prominent staining and large morphology of the nuclei, we focused on the intestinal tract for further stainings.
Stainings for PCNA (a,b), BrdU (c) and geminin (d-f). All stainings were performed on 72 hpf embryos. (a) Sagittal section stained for PCNA. Strong staining is found in the tectum (blue arrow) and gastro-intestinal tract (black arrows). (b) Transversal section stained for PCNA. PCNA stains positive in the gut tube, swim bladder and liver. (c) Transversal section stained for BrdU. BrdU intake is prominent in intestinal cells. (d–f) Separate images of nuclear staining (DAPI) (d), mCherry (zGem signal) (e) and overlay (f). (f) mCherry signal is present in a large proportion of intestinal cells.
To further confirm the proliferative capacity of the intestinal tract, embryos were administered Bromodeoxyuridine (BrdU) at 72 hpf for 2 hours and subsequently fixed and stained with an anti-BrdU antibody. BrdU is an analogue of thymine and will be incorporated into the DNA of cells during the S-phase. Transversal sections reveal that throughout the gastro-intestinal tract, cells incorporate BrdU (Fig. 1c). Quantification of the number of BrdU + cells show that up to 40% of the intestinal epithelial cells have incorporated BrdU, indicating that these cells are actively going through S-phase.
In a third experiment, we examined the intestinal tract of 72 hpf transgenic Tg(EF1a: mCherry-zGem)oki011 zebrafish, which expresses an mCherry-fused 100-amino acid peptide of geminin under the control of the EF1α promotor and allows for fluorescent visualisation of the nuclei of cells that are in late S-/G2-phase of the cell cycle18,19. Since the mCherry signal is destroyed during the fixation process, an immunostaining with an anti-mCherry antibody was performed. Geminin positive cells were clearly visible in and around the intestinal tract (Fig. 1d–f). Quantification of intestinal cells in three embryos shows that on average 40–60% of the nuclei in the intestinal tract were positive for geminin.
The three different markers (PCNA; BrdU; geminin), therefore, manifest the proliferative nature of the intestinal tissue in zebrafish embryos. Moreover, a large proportion of the cells in the intestinal tract are in late S-/G2-phase of the cell cycle, which are the phases where HR is active. As such, this tissue was chosen for optimisation of the Rad51 foci assay.
Wild type embryos show Rad51 foci upon irradiation
To optimise conditions for visualisation of Rad51 foci, a time kinetics experiment was performed. Embryos at 72 hpf were irradiated with 10 or 20 Gy and fixed at 1, 3, 5, 7 and 9 hours post irradiation (hpi). Unirradiated controls at 5 and 9 hours were also included. Transversal sections of the intestinal tract were made. To show that irradiation with 10 and 20 Gy induces DSB, immunostaining for γH2AX, a DSB marker, was conducted. An abundance of nuclear γH2AX foci was detected in irradiated embryos, but not in unirradiated controls (Supplementary Fig. S1). Staining for Rad51 showed formation of Rad51 foci in irradiated embryos while unirradiated controls contain almost no Rad51 foci (Fig. 2a,b).
Representative images of Rad51 staining. Rad51 foci are absent in unirradiated sample (a), while irradiated sample displays clear Rad51 foci (b) (green fluorescence). (c) Schematic drawing of an unirradiated (left) and irradiated (right) gut tube section. Cells in blue are in late S-/G2-phase and thus capable of HR. Cells in white are not capable of HR and will never show Rad51 foci. Red dots indicate Rad51 foci. Quantifying all cells would lead to a serious underestimation of your signal, due to the large proportion of non-HR capable cells. Instead, quantifying only cells containing foci will give a more true value for irradiated conditions. To estimate the amount of HR capable cells in unirradiated samples, one can extrapolate the percentage of cells that contain Rad51 foci from the irradiated samples.
Considering all intestinal cells to determine the mean number of foci/cell would result in an underestimation of Rad51 foci, as geminin staining showed that a substantial proportion (~40–60%) of these cells are not in late S-/G2-phase. Instead the following method was developed: as the embryos were irradiated with high doses, all cells have DSB (as observed with the γH2AX foci assay). Thus, we hypothesise that all cells that were in late S- or G2-phase at the moment of irradiation should show Rad51 foci and we restricted the quantification to cells containing foci.
This approach cannot be followed for unirradiated embryos (no DSB induced). Instead, we took the total number of foci and divided it by the expected number of cells that are capable of HR in these controls. This number was calculated by multiplying the percentage of Rad51 positive cells from irradiated conditions in the same experiment (typically 40–60%) with the total number of intestinal cells in the controls.
For convenience purposes, the values obtained in irradiated and unirradiated embryos will always be referred to as ‘foci/cell’. A visual representation of this quantification approach can be found in Fig. 2c. All irradiated embryos display distinct nuclear foci in intestinal cells while controls are almost completely devoid of Rad51 foci (Fig. 3; Supplementary Table S5). Two-way ANOVA did not indicate a difference in foci/cell between the different time points. However, the foci observed at 1 hour were more subtle, hampering adequate quantification, compared to later time points. This indicates that 1 hpi, Rad51 is still being recruited to the site of the DSB. For all subsequent experiments, embryos of 72 hpf were irradiated with 20 Gy, fixated at 5 hpi and sectioned.
Time kinetics experiment. Graph displaying the number of foci/cell in function of different time points. Three embryos per condition were irradiated with 10 or 20 Gy and fixed at different time points, ranging from 1–9 hours. Unirradiated embryos were also fixed at 5 and 9 hours. Error bars display 95% CI.
Of note, in cells of the tectum and the retina, other highly proliferative regions of the zebrafish embryos 72 hpf, Rad51 foci are also visible. However, visualisation and adequate quantification of the number of Rad51 foci in these regions is challenging due to the smaller size of the cells, the close proximity of the cells to one another and the difficulty for sectioning the correct region.
The quantification method used in the time kinetics experiment was based on the assumption that all cells in late S-/G2-phase repair a number of DSB by means of HR and thus show Rad51 foci after irradiation. To confirm that this quantification is reliable, three transgenic Tg (EF1a : mCherry-zGem)oki011 embryos were irradiated with 20 Gy, fixed at 5 hpi and stained for both Rad51 and mCherry (zGem) (Fig. 4). Unirradiated embryos were also included.
Quantification was conducted in two ways. First, all cells containing foci were included (disregarding if cells stained positive for geminin or not) and the mean number of foci/cell was calculated using formulas (1) and (2). In Supplementary Fig. S2 and Supplementary Table S6 these “hypothesised values” are shown. Second, only the geminin positive cells were included and the number of foci divided by the amount of geminin positive cells was calculated. These data are indicated in Supplementary Fig. S2 and Supplementary Table S6 as“true values”. For both the irradiated and unirradiated embryos, the “hypothesised value” correlates very well with the “true value” (Irradiated: 6.14 versus 6.06 foci/cell respectively; unirradiated: 0.67 vs 0.47 foci/cell respectively, Supplementary Table S6), confirming the validity of our quantification method.
Knockout/knockdown of Brca2 results in absence of Rad51 foci
A brca2 zebrafish knockdown model was obtained by injecting 1-cell stage wild type embryos with a splice-blocking morpholino targeting the exon7/intron7 splice site of brca2. At 72 hpf, embryos were irradiated with 20 Gy, followed by fixation at 5 hpi. Three uninjected embryos (MO−) were also irradiated. RT-PCR was performed to check the functionality of the morpholino. Control embryos show normal amplification of a region between exon 4 and exon 8 (250 bp). Injection of the morpholino causes a band shift to about 1600 bp, corresponding to an intron 7 retention (Fig. 5a), and multiple stop codons are predicted within the first 50 bp by this intron retention.
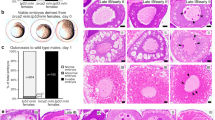
Results of morpholino experiment. (a) Digital gel of RT-PCR (obtained by Labchip®GX software) of uninjected (MO−) and injected (MO+) embryos. Injecting the morpholino results in retention of intron 7, resulting in a transcript containing a premature stop codon. This is visualised as a longer RT-PCR fragment, using primers located in exon 4 and 8 (expected size: 250 bp; in case of intron 7 retention: 1600 bp). In the morphants, there is a clear band of 1600bp, indicative for intron retention. (b) Quantification of Rad51 foci in three morphants (MO+) and uninjected (MO−) embryos. Almost no Rad51 foci are present in the morphants. Error bars display 95% CI. (c) γH2AX foci are present in both irradiated wild types (upper left) and morphants (lower left). Wild types display clear Rad51 foci (upper right), while morphants display no Rad51 foci.
In morphants, a severe loss of the number of Rad51 foci/cell was noticed when compared to controls (6.3 versus 0.2 foci/cell; p = 10−4) (Fig. 5b). To confirm that DSB were induced in both groups, the γH2AX foci assay was performed. Both wild types and morphants show an abundance of γH2AX foci compared to unirradiated embryos (Fig. 5c).
In addition to the brca2 MO-mediated knockdown model, three different knockout lines for brca2 were tested with the Rad51 foci assay: brca2 sa22682, brc2 hg5 and brca2 cmg35 (in-house developed). As brca2 −/− fish develop to infertile males, most likely due to apoptosis of oocytes during the critical period of sex determination of zebrafish, these homozygotes cannot be used for propagation9,10. This ‘all-male’ phenotype was confirmed in our in-house developed brca2 cmg35 line, where 16 adult brca2 −/− fish developed exclusively as males (chi², p = 6.10−5). In addition, using offspring from brca2 −/− fish is discouraged as these fish would have a dysfunctional HR pathway and may present with genomic instability. This predisposition could be transmitted to their offspring and affect results. Therefore, in this experiment, heterozygous carriers of each knock-out line were in-crossed and resulting embryos were irradiated with 20 Gy and fixed at 5 hpi. At least three embryos per condition were quantified. For all lines, Rad51 foci were present in both brca2 +/+ and brca2 +/− embryos, whilst being absent in brca2 −/− embryos (Fig. 6; Supplementary Table S7). Interestingly, a statistically significant difference was also observed between brca2 +/+ and brca2 +/− in the brca2 cmg35+/− line, with less foci being present in the brca2 +/− condition (p = 0.02). The two other lines showed a similar trend but results were not significant. When combining the data from all three lines, the difference between brca2 +/+ and brca2 +/− was statistically significant (p = 0.02).
Rad51 foci quantification of brca2 sa22682/+, brca2 hg5/+ and brca2 cmg35/+ in-cross. At least three embryos per genotype were stained. All brca2 −/− lines show a complete absence of Rad51 foci. brca2 +/+ embryos display more Rad51 foci than brca2 +/−. This difference is statistically significant in brca2 cmg35 or when combining data from all lines. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
Discussion
Zebrafish is an increasingly popular model system in genetic and cancer research because of its relative small size, ease of housing, ex utero development and initial transparency. However, current knowledge on the HR repair pathway of this model organism is limited8,20.
In this work, we aimed to develop an assay to accurately quantify HR activity in zebrafish through means of Rad51 foci quantification. Furthermore, using this method, we wanted to determine if zebrafish Brca2 conserved its function in recruitment of Rad51 to the site of DSB.
To be able to study HR in zebrafish embryos, we first needed to identify a suitable tissue showing high HR activity. Through PCNA staining, we identified that intestinal cells 72 hpf are particularly highly proliferative. This confirms previous findings where it has been reported that between 26 hpf and 72 hpf the entire intestinal endoderm shows high proliferative capacity21,22. BrdU stainings performed on 72 hpf embryos indicated that about 40% of cells in the intestinal tract are in S-phase. In contrast, Wallace et al. found about 20% S-phase in intestinal cells in 72 hpf embryos. This might be attributed by differences between protocols, as they performed a 1 hour incubation while we applied 2 hours23. Finally, geminin stainings determined that about 40–60% of intestinal cells are in late S-/G2-phase at 72 hpf. The high number of cells in late S-/G2-phase makes intestinal tissue highly interesting for studies on HR because during these phases sufficient genetic material has been duplicated allowing the cell to repair DSB through HR24. Another advantage of intestinal tissue is the large size of the cells and their nuclei. This allows for adequate visualisation of discrete nuclei as well as Rad51 foci in these nuclei. Finally, the gut tube spans a large part of the embryo allowing the sectioning of intestinal cells to be straightforward. In conclusion, zebrafish embryonic intestinal tissue is an ideal target tissue when assessing HR functionality in this organism.
To activate the HR pathway, DSB were induced through ionising irradiation. A time kinetics experiment was performed, where 10 and 20 Gy were administered to 72 hpf embryos. A dose of 20 Gy and assessment of HR activity at 5 hpi was identified as optimal. Also, other studies reporting on radiation experiments in zebrafish, such as screening of radiation modifiers25 and engraftment experiments26, use up to 20 Gy. Studies where the RAD51 foci assay has been performed in human tumour biopsies and mammalian cell lines also use rather high doses (ranging from 5–12 Gy)7,27,28. Although a majority of the DSB are repaired through NHEJ, a sub-fraction of radiation-induced DSB are specifically repaired by HR. This is due to the complexity of the DSB repair resulting from increasing doses29,30. The Rad51 foci assay, therefore, requires high doses.
HR activity was quantified as the mean number of Rad51 foci/cell as described in the results section. This quantification was validated using the transgenic line Tg(EF1a: mCherry-zGem)oki011. Geminin is known to be active in the late S-/G2-phase and was previously shown to be an excellent HR marker in combination with RAD51 staining in human tumour tissues7,31. Of note, at any given dose and time point γH2AX foci are more prominent than Rad51 foci. This suggests that besides HR, other DSB repair pathways are also active. Liu et al. previously showed that NHEJ is the most dominant DSB repair pathway in zebrafish embryos12.
After optimizing the zebrafish Rad51 foci assay on wild type embryos, we determined if deficiency of Brca2 leads to absence of Rad51 foci recruitment to the DSB. Using a brca2 MO-mediated knockdown, a complete loss of Rad51 foci was observed. However, as morpholinos may produce off-targeted effects32, we validated our results in three stable brca2 mutant lines, harbouring different inactivating mutations in exon 8 (brca2 cmg35) or 11 (brca2 sa22682, brca2 hg5). For all stable lines, Rad51 foci were absent in mutants, while being present in both wild types and heterozygous carriers. We conclude that Brca2 has conserved its main function in recruiting Rad51 and that our assay can accurately quantify HR functionality. Morpholinos have the advantage that every injected embryo is deficient for Brca2, while only 25% of offspring from heterozygous carriers will be mutant. This makes the use of stable lines more labour intensive. However, morpholino-mediated knockdown is only temporary. The choice should be made in function of the data and experiments required.
Interestingly, for the brca2 cmg35 line we observed a borderline statistically significant difference in the number of Rad51 foci/cell between brca2 +/+ and brca2 +/− embryos. This suggests that HR is less efficient in brca2 +/− cells compared to brca2 +/+ cells. In general, tumours from patients with a germline BRCA2 mutation contain an inactivating mutation of the second allele. However, in a considerable number of human BRCA1/2 associated breast tumours, loss of heterozygosity (LOH) of the wild type allele is not observed33,34,35. It has been suggested that BRCA1/2 haploinsufficiency may cause genomic instability, initiating tumorigenesis6,33,35. Recently, a study on 160 BRCA1 and BRCA2 germline mutation-associated breast and ovarian cancer found absence of LOH in a subset of patients, in which they postulate haploinsufficiency as a possible mechanism for some of these cases36. Our data obtained in the zebrafish brca2 +/− embryos indeed support reduced repair capacity through HR. The observed difference, however, is small (13% reduction). Nevertheless, a small reduction in HR could hypothetically lead to additional accumulation of mutations and eventually tumorigenesis in some cases. More studies are warranted to evaluate if the observed difference has an impact on repair capacity and how this can be translated to a tumorigenic process.
In conclusion, we have identified zebrafish embryonic intestinal cells as an ideal target to study the functionality of the HR pathway due to their highly proliferative activity and good morphology. Our study shows that zebrafish Brca2, like human BRCA2, plays a role in HR through recruitment of Rad51; furthermore, disabling Brca2 results in absence of Rad51 foci after irradiation. These results were observed in both morpholino injected embryos and in mutant zebrafish lines. We demonstrate here that zebrafish is a suitable model organism for studying HR functionality. Interestingly, the outcome of our experiments also suggests less efficient HR capacity in brca2 +/− embryos. Further studies are warranted to evaluate if this leads to increased genomic instability due to haploinsufficiency.
We are convinced that, in the future, our Rad51 foci assay can be applied in combination with rescue experiments to study the pathogenic effect of BRCA2 variants of unknown clinical significance37. Related to this, PARP inhibitors are a novel type of drug to selectively target HR deficient tumours through synthetic lethality38; our assay may be useful for testing the efficiency of new inhibitors. Another application may be related to evaluate the efficiency of Crispr-Cas9 mediated genome editing through HR in zebrafish.
Material and Methods
Zebrafish maintenance and lines
Zebrafish lines were housed in a Zebtec semi-closed recirculation housing system at a constant temperature (27–28 °C), pH (~7.5), conductivity (~550 µS) and light/day cycle (14/10). Fish were fed twice a day with dry food (Gemma Micro, Skretting) and once with artemia (Ocean Nutrition). The mutant brca2 sa22682 line was purchased from the European Zebrafish Resource Center (EZRC, Karlsruhe institute, Germany); brca2 cmg35 was generated using Crispr-Cas9 mutagenesis (cfr below); brca2 hg5 was a kind gift from Dr. Heather Shive (North Carolina State University)9; Tg(EF1a: mCherry-zGem)oki011 was a kind gift from Dr. Ichiro Masai (Okinawa Institute of Science and Technology)18.
Ethic statement
This study was approved by the local animal ethics committee (Ghent University hospital, Ghent, Belgium) application numbers: ECD 14/70 and 16/67 K. All methods were carried out in accordance with the approved guidelines.
Crispr-Cas9 mutagenesis
Three sgRNA sequences targeting brca2 were designed using CRISPRdirect39. From these sequences, synthetic dsDNA consisting of a 5′ random sequence, T7 promotor, target-specific sequence (minus PAM) and a constant region were designed according to Boel et al.40. These dsDNA sequences were ordered as G-blocks at Integrated DNA Technologies (IDT). G-blocks were dissolved in 20 µl nuclease-free water. PCR conditions can be found in Supplementary data (Supplementary Table 2–4).
PCR product was purified with the Qiaquick PCR purification kit (Qiagen) following general guidelines of the manufacturer. In vitro transcription was performed using the MEGAshortscript™ Kit (Thermo Fisher). The MEGAclear™ Kit (Thermo Fisher) was used to purify the transcription reaction. RNA quality was verified on the Experion (Bio-Rad). 25 pg sgRNA was co-injected with 250 pg Cas9 protein (Cas9 wild type nuclease protein with NLS, ToolGen) during the 1-cell stage. Phenol red 6% was used to visualise injection. At 24 hpf, 10 embryos were harvested for DNA extraction, using the KAPA Express Extract DNA Extraction kit. PCR was performed according to manufacturer’s protocol using primers flanking the crispr region. Samples were deep sequenced on the MiSeq and somatic indel frequency was analysed using BATCH-GE40. If a high out-of-frame indel percentage was identified, mosaic fish (F0) were grown to adulthood. These were out-crossed and offspring (F1) was tested for the presence of the mutation. If present, offspring was grown to adulthood and genotyped for the mutation.
Morpholino injection
To knockdown zebrafish Brca2, a splice-blocking morpholino oligonucleotide (MO; Genetools) was designed targeting the exon7/intron7 boundary (5′-ATTGCgtatgtatgaatggtcttgc-3′) of brca2. At the 1-cell stage, wild type embryos were injected with 10 ng of the morpholino. The morpholino was dissolved in a 2% Phenol red solution for visualisation of injection.
RNA extraction and RT-PCR
Ten embryos per condition were dechorionated and rinsed with PBS. 175 µl TRIZOL (Invitrogen) was added and embryos were homogenised. After 5 min incubation 70 µl of chloroform was added and samples were centrifuged for 15 minutes at 11000 rpm at 4 °C. The upper phase was transferred to a new tube and 100% ethanol was added. This mixture was brought onto a RNeasy mini-spin column (Qiagen) and centrifuged at 9600 rcf for 30 seconds. Samples were treated with 350 µl RW1 and briefly centrifuged. 35 µl RDD buffer and 5 µl DNaseI were added on each column and after 15 min incubation 350 µl RW1 was added, followed by a brief centrifugation. Samples were subsequently treated twice with 500 µl RPE buffer after which the RNA was finally eluted in 30 µl RNAse free water. cDNA synthesis was performed using the IScript™ cDNA Synthesis Kit (Bio-Rad). Samples underwent RT-PCR with primers surrounding the target area of the morpholino (F: 5′-CCTTGTTTGTTTGGCTCAGC-3′; R: 5′-CAGATGCAACCCGGTCC-3′). Sample sizing was conducted on the Labchip® GX capillary electrophoresis system.
BrdU labelling
At 72 hpf, embryos were treated with 10 mM BrdU (Sigma-Aldrich) in E3 medium for 2 hours. BrdU was washed away prior to fixation.
Irradiation and fixation
At 72 hpf, embryos were irradiated with the Small Animal Radiation Research Platform (SARRP, Infinity Lab, Ghent University). Embryos underwent irradiation with doses of 10 or 20 Gy X-rays (220 kV, dose rate 2.77 Gy.min−1). Embryos were euthanised and fixed for 1 hour in 4% PFA after different time points post irradiation (hpi) ranging between 1–9 hpi. Following fixation, embryos were placed in increasing alcohol concentrations and embedded in paraffin.
Immunostainings
Paraffin embedded embryos were sectioned in 5 µm sections and placed on slides. The embryos were then deparaffinised and underwent 20 minutes of steam antigen retrieval in citric buffer (pH 6.1). For immunohistochemical stainings, slides were treated with 10% H2O2. Blocking serum was added to the samples to prevent aspecific binding. Slides were incubated with mouse anti-BrdU (1/100; Dako; M0744) or mouse anti-PCNA (1/100; Novo Castra; NCL-PCNA) for 2 hours. This was followed by incubation with biotinylated RAM (1/200, Dako) and Strep-HRP (1/200, Dako) antibodies for 30 minutes and 3,3′-diaminobenzidine tetrahydrochloride (DAB, Sigma-Aldrich) exposure for 10 minutes. Slides were counterstained with Mayer’s Haematoxylin (Merck).
For immunofluorescent stainings, slides were treated with anti-γH2AX (1/1000; Bethyl Laboratories; IHC-00059), anti-RAD51 (1/2000; Santa-Cruz, H-92) or anti-mCherry (Abcam; ab125096; 1C51) antibodies overnight at 4 °C. Slides were then incubated with Goat-anti-Rabbit Dylight 488 antibody (1/1000, Sigma-Aldrich) or Goat-anti-Mouse Dylight 594 (1/1000 Sigma-Aldrich). Finally slides were treated with DAPI + fluoromount (Sigma-Aldrich). For PCNA staining, no antigen retrieval or counterstaining was performed. Unless mentioned otherwise, three embryos per condition were used.
Visualisation
Slides were scanned on a Zeiss Axio Observer.Z1 inverted microscope, using the Zen pro 2012 software. γH2AX, Rad51 and geminin images were captured using fluorescent microscopy. To allow for foci analysis (γH2AX, Rad51), a 100x enlargement was used and Z-stacks were made (0.22 µm thickness). Images were deconvoluted (fast iterative) and an orthogonal projection of 10 slices was performed. For the quantification of %BrdU positive cells, 236 intestinal cells were counted. For the quantification of %geminin positive cells, 1024 intestinal cells were counted. Quantification of Rad51 foci is explained in the results section.
Genotyping
During sectioning of paraffin embedded embryos, some sections were reserved for genotyping. DNA was extracted with the Kapa Express Extract kit (Kapa Biosystems) and PCR according to the manufacturer’s protocol was performed. Inactivating mutations and primers used for genotyping brca2 hg5, brca2 sa22682 and brca2 cmg35 can be found in Supplementary Table S1. PCR products were Sanger sequenced to determine the genotype.
Statistical analysis
Data was analysed using SPSS statistics 24 (IBM). Unless mentioned otherwise, groups were compared using the student’s t-test (two-sided, α = 0.05). Levene’s test for equality of variances was performed. Graphs were made with Sigmaplot (Systat Software Inc.).
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Khanna, K. K. & Jackson, S. P. DNA double-strand breaks: signaling, repair and the cancer connection. Nat. Publ. Gr. 27, 247–254 (2001).
Ohnishi, T., Mori, E. & Takahashi, A. DNA double-strand breaks: Their production, recognition, and repair in eukaryotes. Mutat. Res. Mol. Mech. Mutagen. 669, 8–12 (2009).
Lieber, M. R. The Mechanism of Human Nonhomologous DNA End Joining. J. Biol. Chem. 283, 1–5 (2008).
Tsai, A. G. & Lieber, M. R. Mechanisms of chromosomal rearrangement in the human genome. BMC Genomics 11(Suppl 1), S1 (2010).
Thompson, L. H. & Schild, D. Homologous recombinational repair of DNA ensures mammalian chromosome stability. Mutat. Res. 477, 131–153 (2001).
Roy, R., Chun, J. & Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nat. Rev. Cancer 12, 68–78 (2012).
Naipal, K. A. T. et al. Functional ex vivo assay to select homologous recombination-deficient breast tumors for PARP inhibitor treatment. Clin. Cancer Res. 20, 4816–4826 (2014).
Pei, D.-S. & Strauss, P. R. Zebrafish as a model system to study DNA damage and repair. Mutat. Res. 743–744, 151–159 (2013).
Shive, H. R. et al. Brca2 in Zebrafish Ovarian Development, Spermatogenesis, and Tumorigenesis. Proc. Natl. Acad. Sci. USA 107, 19350–19355 (2010).
Rodríguez-Marí, A. et al. Roles of brca2 (fancd1) in oocyte nuclear architecture, gametogenesis, gonad tumors, and genome stability in zebrafish. PLoS Genet. 7 (2011).
Moldovan, G.-L. & D’Andrea, A. D. How the fanconi anemia pathway guards the genome. Annu. Rev. Genet. 43, 223–249 (2009).
Liu, J. et al. Development of novel visual-plus quantitative analysis systems for studying DNA double-strand break repairs in zebrafish. J. Genet. Genomics 39, 489–502 (2012).
Bogliolo, M. & Surrallés, J. Fanconi anemia: a model disease for studies on human genetics and advanced therapeutics. Curr. Opin. Genet. Dev. 33, 32–40 (2015).
Dong, H. et al. Update of the human and mouse Fanconi anemia genes. Hum. Genomics 9, 32 (2015).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Mione, M. C. & Trede, N. S. The zebrafish as a model for cancer. Dis Model Mech 3, 517–523 (2010).
Tat, J., Liu, M. & Wen, X. Y. Zebrafish cancer and metastasis models for in vivo drug discovery. Drug Discov. Today Technol. 10, e83–e89 (2013).
Mochizuki, T., Suzuki, S. & Masai, I. Spatial pattern of cell geometry and cell-division orientation in zebrafish lens epithelium. Biol. Open 3, 982–994 (2014).
McGarry, T. J. & Kirschner, M. W. Geminin, an inhibitor of DNA replication, is degraded during mitosis. Cell 93, 1043–1053 (1998).
Amatruda, J. F. & Patton, E. E. Genetic Models of Cancer in Zebrafish. Int. Rev. Cell Mol. Biol. 271, 1–34 (2008).
Ng, A. N. Y. et al. Formation of the digestive system in zebrafish: III. Intestinal epithelium morphogenesis. Dev. Biol. 286, 114–135 (2005).
Wullimann, M. F. & Knipp, S. Proliferation pattern changes in the zebrafish brain from embryonic through early postembryonic stages. Anat. Embryol. (Berl). 202, 385–400 (2000).
Wallace, K. N., Akhter, S., Smith, E. M., Lorent, K. & Pack, M. Intestinal growth and differentiation in zebrafish. Mech. Dev. 122, 157–173 (2005).
Rothkamm, K., Krüger, I., Thompson, L. H., Kru, I. & Lo, M. Pathways of DNA Double-Strand Break Repair during the Mammalian Cell Cycle Pathways of DNA Double-Strand Break Repair during the Mammalian Cell Cycle. Mol. Cell. Biol. 23, 5706–5715 (2003).
Lu, B., Hwang, M., Yong, C., Moretti, L. & Lu, B. Zebrafish as a Model System to Screen Radiation Modifiers. Curr. Genomics 8, 360–369 (2007).
Glass, T. J., Hui, S. K., Blazar, B. R. & Lund, T. C. Effect of Radiation Dose-Rate on Hematopoietic Cell Engraftment in Adult Zebrafish. PLoS ONE 8 (2013).
Vaclova, T. et al. DNA repair capacity is impaired in healthy BRCA1 heterozygous mutation carriers. Breast Cancer Res. Treat. 152, 271–282 (2015).
Godthelp, B. C. et al. Inducibility of nuclear Rad51 foci after DNA damage distinguishes all Fanconi anemia complementation groups from D1/BRCA2. Mutat. Res. 594, 39–48 (2006).
Jeggo, P. A., Geuting, V. & Löbrich, M. The role of homologous recombination in radiation-induced double-strand break repair. Radiother. Oncol. 101, 7–12 (2011).
Johnson, R. D. & Jasin, M. Double-strand-break-induced homologous recombination in mammalian cells. Biochem. Soc. Trans. 29 (2001).
Graeser, M. et al. A marker of homologous recombination predicts pathologic complete response to neoadjuvant chemotherapy in primary breast cancer. Clin. Cancer Res. 16, 6159–6168 (2010).
Kok, F. O. et al. Reverse Genetic Screening Reveals Poor Correlation between Morpholino-Induced and Mutant Phenotypes in Zebrafish. Dev. Cell 32, 1–12 (2014).
King, T. A. et al. Heterogenic loss of the wild-type BRCA allele in human breast tumorigenesis. Ann. Surg. Oncol. 14, 2510–2518 (2007).
Clarke, C. L. et al. Mapping loss of heterozygosity in normal human breast cells from BRCA1/2 carriers. Br. J. Cancer 95, 515–519 (2006).
Tung, N. et al. Prevalence and predictors of loss of wild type BRCA1 in estrogen receptor positive and negative BRCA1-associated breast cancers. Breast Cancer Res. 12, R95 (2010).
Maxwell, K. N. et al. BRCA locus-specific loss of heterozygosity in germline BRCA1 and BRCA2 carriers. Nat. Commun. 8 (2017).
Guidugli, L. et al. Functional assays for analysis of variants of uncertain significance in BRCA2. Hum. Mutat. 35, 151–164 (2014).
Ledermann, J. A. & El-Khouly, F. PARP inhibitors in ovarian cancer: Clinical evidence for informed treatment decisions. Br. J. Cancer 113, S10–S16 (2015).
Naito, Y., Hino, K., Bono, H. & Ui-Tei, K. CRISPRdirect: Software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 31, 1120–1123 (2015).
Boel, A. et al. BATCH-GE: Batch analysis of Next-Generation Sequencing data for genome editing assessment. Sci. Rep. 6, 30330 (2016).
Acknowledgements
We wish to thank Dr. Heather Shive (North Carolina State University) and Dr. Ichiro Masai (Okinawa Institute of Science and Technology) for donating the brca2 hg5 line and Tg(EF1a: mCherry-zGem)oki011 line respectively. We thank Leen Pieters for her technical assistance. This study was funded by ‘Bijzonder Onderzoeksfonds UGent’, grant number 01J15713, and ‘Stichting Tegen Kanker’, grant number C/2014/385.
Author information
Authors and Affiliations
Contributions
J.V., A.W., A.V. and K.C. conceived the experiments. P.C., A.V. and K.C. provided the facilities for performing the experiments. J.V. performed the experiments and analysed the data. A.W., P.V., A.V. and K.C. provided conceptual advice. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vierstraete, J., Willaert, A., Vermassen, P. et al. Accurate quantification of homologous recombination in zebrafish: brca2 deficiency as a paradigm. Sci Rep 7, 16518 (2017). https://doi.org/10.1038/s41598-017-16725-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16725-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.