Abstract
Glycated haemoglobin (HbA1c) is widely used as a biomarker for the diagnosis of diabetes, for population-level screening, and for monitoring the glycaemic status during medical treatment. Although the heritability of HbA1c has been estimated at ~55–75%, a much smaller proportion of phenotypic variance is explained by the HbA1c-associated variants identified so far. To search for novel loci influencing the HbA1c levels, we conducted a genome-wide meta-analysis of 2 non-diabetic Japanese populations (n = 7,704 subjects in total). We identified 2 novel loci that achieved genome-wide significance: TMC6–TMC8 (P = 5.3 × 10−20) and SIX3–SIX2 (P = 8.6 × 10−9). Data from the largest-scale European GWAS conducted for HbA1c supported an association between the novel TMC6–TMC8 locus and HbA1c (P = 2.7 × 10−3). The association analysis with glycated albumin and glycation gap conducted using our Japanese population indicated that the TMC6–TMC8 and SIX3–SIX2 loci may influence the HbA1c level through non-glycaemic and glycaemic pathways, respectively. In addition, the pathway-based analysis suggested that the linoleic acid metabolic and 14-3-3-mediated signalling pathways were associated with HbA1c. These findings provide novel insights into the molecular mechanisms that modulate the HbA1c level in non-diabetic subjects.
Similar content being viewed by others
Introduction
The glycated haemoglobin (HbA1c) level represents the percentage of haemoglobin proteins bound by glucose. The glycation of haemoglobin is a non-enzymatic and predominantly irreversible reaction; therefore, the HbA1c level reflects the average blood glucose level over approximately 3 months prior to the measurement1. Measuring HbA1c is more convenient than measuring the fasting plasma glucose (FPG) level because HbA1c does not need to be measured in the fasting state, has greater pre-analytical stability and is not subject to intra-individual day-to-day variability2. Moreover, the HbA1c level, which represents long-term hyperglycaemia, is an independent risk factor for cardiovascular events3. Thus, HbA1c is widely used as a biomarker for diagnosing diabetes, for population-level screening, and for monitoring the glycaemic status during medical treatment4.
Data from twin and familial studies have shown that the HbA1c level is a heritable trait, with a heritability of approximately 55% to 75%5,6,7. Genome-wide association studies (GWASs) have revealed ~20 HbA1c-associated genetic loci8,9,10,11,12,13,14,15. Previous GWASs have indicated that genetic effects on the HbA1c level may involve both glycaemic and non-glycaemic pathways10,14. HbA1c-associated variants located at the ANK1, CDKAL1, G6PC2/ABCB11, GCK, MTNR1B, SLC30A8, and TCF7L2 loci confer an increased risk for type 2 diabetes (T2D) and/or are associated with 1 or more glycaemic traits, including FPG, 2-hour glucose, and fasting proinsulin16. In addition, non-glycaemic variants have been identified that are associated with HbA1c but not with glycaemic traits and the T2D risk. Of these non-glycaemic variants, those at the HFE and TMPRSS6 loci have been associated with red blood cell parameters17.
The largest-scale GWAS conducted for HbA1c to date was a meta-analysis of non-diabetic European-ancestry subjects (n = ~46,000 subjects)10. The second-largest GWAS was a meta-analysis of non-diabetic East Asian populations (n = ~21,000 subjects)14. Arab, Malay and South Asian populations have also been analysed12,15. However, the HbA1c-associated variants identified to date explain a much smaller proportion of the phenotypic variance than the heritability estimates from twin and familial studies10,14. Accordingly, the genetic factors that influence the HbA1c level have not been fully determined. To search for novel HbA1c-associated loci and to elucidate the molecular pathways involved in HbA1c biology, we conducted a genome-wide meta-analysis of HbA1c in 2 Japanese populations of non-diabetic subjects (n = 7,704 subjects).
Methods
Study subjects
Over 80,000 apparently healthy adults living in the Iwate and Miyagi Prefectures (residing along the Pacific coast of the Tohoku region of Japan) were recruited from May 2013 to March 2016 for the Tohoku Medical Megabank (TMM) Project. The study design and recruitment methods were previously described18. Briefly, the participants were aged from 20 to 75 years and completed questionnaires covering a wide range of topics, including sociodemographic factors, lifestyle habits, and medical history. Blood and urine tests were conducted at the baseline survey. In addition, blood samples were stored at our biobank. Participants living in the Iwate and Miyagi Prefectures were recruited by Iwate Medical University and Tohoku University, respectively. We obtained approval from the relevant ethics committees at both facilities. All participants gave written, informed consent at the time of study enrolment. This study was conducted according to the principles expressed in the Declaration of Helsinki.
The HbA1c levels were measured using National Glycohemoglobin Standardization Program (NGSP)-certified methods. A high-performance liquid chromatography (HPLC) method was used for the Iwate subjects, and a latex agglutination method was used for the Miyagi subjects. We excluded diabetic participants defined based on self-reported diabetes, self-reported diabetes treatment, or HbA1c ≥ 6.5%. For the Iwate subjects, FPG was measured using a hexokinase method, and glycated albumin (GA) was assayed with an enzymatic method. Glycation gaps (GGs) were calculated as the difference between the measured and GA-based predicted HbA1c levels as previously described19,20. The plasma creatinine and cystatin C levels were measured using enzymatic and latex-coagulating nephelometry methods, respectively. The creatinine- and cystatin C-based estimated glomerular filtration rates (eGFRcrea and eGFRcys, respectively) were estimated using the Japanese equation for the eGFR calculation21,22. The red blood cell (RBC) counts, haemoglobin (Hb) concentrations, and haematocrit (HCT) values were measured with flow cytometry, sodium lauryl sulphate, and sheath flow detection methods, respectively. The mean corpuscular volume (MCV), mean corpuscular haemoglobin (MCH), and mean corpuscular haemoglobin concentration (MCHC) values were calculated from the RBC, Hb, and HCT values.
Genotyping, quality control, and genotype imputation
A total of 9,966 participants enrolled in 2013 were genotyped using the HumanOmniExpressExome BeadChip Array (Illumina Inc., San Diego, CA, USA). Of these participants, 8,678 were non-diabetic with body mass index (BMI) data available. Sex was inferred using the PLINK software (version 1.90b3.45)23,24. Subjects in whom the inferred sex was ambiguous (n = 77) or inconsistent with the sex recorded in the questionnaire (n = 29) were excluded. In addition, subjects with a low call rate (<0.99; n = 8) and an estimated non-Japanese ancestry (n = 13; Supplementary Fig. 1) were excluded. Ancestry was estimated based on principal component analysis25,26. We found 1,173 close relationship pairs using the identity-by-descent method implemented in the PLINK software (PI_HAT >0.1875). We randomly excluded one of the closely related subjects for each pair; thus, 847 subjects were excluded. Single-nucleotide polymorphisms (SNPs) with low call rates (<0.95), low Hardy–Weinberg equilibrium exact test P-values (<1 × 10−6) or low minor allele frequencies (MAFs; <0.01) were filtered out. These quality-control filters resulted in the inclusion of 3,664 Iwate and 4,040 Miyagi subjects and 596,877 autosomal SNPs.
Genotype imputation was performed using the SHAPEIT (version 2.r790)27 and Minimac3 (version 1.0.11)28 software packages with the 1000 Genomes reference panel (phase 3)29,30. After genotype imputation, variants with a low imputation quality (R 2 < 0.8) and a low MAF (<0.01) were excluded, and 7,135,436 variants were retained for further analysis.
Estimation of variance explained by common variants
The variance in HbA1c explained by common variants was estimated based on a linear mixed model31 (LMM) implemented in the GCTA software (version 1.24.2)32. For the estimation, we additionally excluded directly genotyped SNPs with moderately low Hardy–Weinberg equilibrium exact test P-values (P < 0.05). A genetic relationship matrix (GRM) was calculated from the remaining 534,808 SNPs. Then, narrow-sense heritability was estimated with adjustments for age, sex, BMI, and recruitment site. We combined the Iwate and Miyagi subjects for this analysis.
Association with HbA1c
The association between each variant and the HbA1c level was tested using an LMM association method implemented in the GCTA software32. We modified the software to accept genotype dosage data as an input for the association test33. For each of the Iwate and Miyagi populations, we tested all 7,135,436 imputed variants with adjustments for age, sex, and BMI. The same GRM used in the heritability estimation was used in this analysis. Based on the summary association statistics from both populations, we performed a meta-analysis of the association between all imputed variants and the HbA1c level using a fixed-effect model and the inverse-variance weighting method with the METAL software (version 2011-03-25)34. Variants with an association P-value less than the genome-wide significance (GWS; P < 5 × 10−8) were considered HbA1c-associated variants. All GWS variants within 500 kb were grouped into a single locus, and we determined a lead variant for each HbA1c-associated locus by choosing the variant with the lowest P-value at that locus.
Expression quantitative trait locus (eQTL) analysis
Whole-genome and transcriptome data from 105 Japanese subjects registered in a multi-omics database (iMETHYL) were analysed to search for significant cis-eQTL variant-gene pairs. Pairs of a novel lead variant and neighbouring genes within ±100 kb were tested. Adaptor trimming, mapping, quality control filtering, base calling, and gene expression profiling of the iMETHYL data were previously described35. Briefly, genotype calling was performed with the same filtering procedures used in the 1KJPN Japanese population reference panel, including single-nucleotide variant (SNV) filtering according to read coverage, software-derived biases, departures from the Hardy–Weinberg equilibrium, and complexities of genomic regions around variants36. For the gene expression profiling, fragments per kb of exon per million mapped fragments (FPKM) values were calculated and normalised across subjects using the cuffquant and cuffnorm programs in the Cufflinks (version 2.2.1) software package37. The eQTL association was tested using linear regression and additive genetic models, i.e., log10 (FPKM + 1) was used as a target variable, and the genotype data (coded as 0, 1, or 2) was used as an explanatory variable. No adjustment variable was included. P-values < 0.05 were considered significant.
Pathway analysis
Based on the GWAS summary data (chromosomal position and P-value) for the directly genotyped SNPs, gene- and pathway-based analyses were conducted using the MAGMA software (version 1.06)38. Variants were mapped onto protein-coding genes based on gene annotations downloaded from the NCBI Gene database (https://www.ncbi.nlm.nih.gov/gene). Then, gene-based P-values were calculated by aggregating variant-based P-values after accounting for the linkage-disequilibrium (LD) structure. The LD information was based on the East Asian population of the 1000 Genomes Project29,30. Pathway-based P-values were calculated by aggregating the gene-based P-values.
Data availability
The datasets analysed in the current study are not publicly available for ethical reasons but are available upon request after approval from the Ethical Committee of Iwate Medical University, the Ethical Committee of Tohoku University, and the Materials and Information Distribution Review Committee of the TMM Project.
Results
Genome-wide meta-analysis in Japanese populations
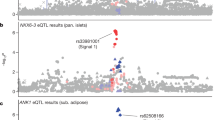
The demographic characteristics of the study subjects are shown in Table 1 and Supplementary Table S1. In the combined Iwate and Miyagi subjects, the variance explained by common variants (MAF ≥ 0.01) was estimated to be 32.1% (standard error [SE] = 4.1%). Genome-wide association tests were conducted for each of the Iwate (n = 3,664) and Miyagi (n = 4,040) populations, and a meta-analysis was performed for the association evidence obtained from the 2 populations. The inflation factor (λ) was 1.008 (95% confidence interval [CI]: 1.006–1.009) for the Iwate population, 1.002 (95% CI: 1.000–1.003) for the Miyagi population, and 1.023 (95% CI: 1.021–1.025; Supplementary Fig. S2) for the meta-analysis, indicating that the population stratification was well-controlled. The meta-analysis showed that 4 independent loci achieved GWS (P < 5 × 10−8), as shown in Fig. 1 and Table 2. Of the 4 loci, the TMC6–TMC8 locus (lead variant: rs2748427; Fig. 2a) on chromosome 17 and the SIX3–SIX2 locus (lead variant: rs10168523; Fig. 2b) on chromosome 2 have not been reported by previous GWASs for HbA1c; therefore, these 2 loci were novel findings. The remaining 2 loci (FN3KRP–FN3K and SMG5) were previously reported10,14. No heterogeneity on the effect of the SIX3–SIX2 lead variant rs10168523 was observed (I 2 = 0), but a large heterogeneous effect was observed for the TMC6–TMC8 lead variant rs2748427 (I 2 = 96.8). The conditional analyses did not find additional independent signals that achieved GWS for the 2 novel loci (Supplementary Tables S2 and S3).
Association signals around novel lead variants. The x-axis represents chromosomal positions and the y-axis represents −log10 P-values. The lead variant is shown in purple. Colours represent the degree of LD (r 2) between each variant and the lead variant. The LD (r 2) was calculated based on the combined dataset of Iwate and Miyagi subjects. (a) The TMC6–TMC8 locus. Lead variant was rs2748427. (b) The SIX3–SIX2 locus. Lead variant was rs10168523.
To investigate the association of the novel loci with HbA1c in Europeans, we looked up the summary statistics data from the largest-scale HbA1c GWAS10. For the TMC6–TMC8 locus, although the lead variant rs2748427 was not found in the summary data, a proxy variant rs429216 was included (the LD r 2 between rs2748427 and rs429216 was 0.494 in East Asians and 0.615 in Europeans according to the 1000 Genomes Project data30 and the LDlink server39). The rs429216 reached GWS in our meta-analysis (Supplementary Table S2), and the association between rs429216 and HbA1c was significant in the European population (P = 0.0027). The effect size was 0.0414% (SE = 0.0138%) in the European population. For the SIX3–SIX2 locus, the proxy variant rs4953155 (the LD r 2 between rs10168523 and rs4953155 was 0.919 in East Asians and 0.913 in Europeans), which achieved GWS in our meta-analysis (Supplementary Table S3), was not significantly associated with HbA1c in Europeans (P = 0.74).
To elucidate whether the 2 novel loci influenced the HbA1c level through glycaemic or non-glycaemic pathways, we examined the association of the novel lead variants with GA and GG using the Iwate population. For the TMC6–TMC8 locus, the lead variant rs2748427 was strongly associated with GG (P = 5.3 × 10−23) but not with GA (P = 0.65) (Fig. 3 and Supplementary Table S4), indicating that the TMC6–TMC8 variants were non-glycaemic. For the SIX3–SIX2 locus, the lead variant rs10168523 was associated with GA (P = 2.5 × 10−4; Fig. 3 and Supplementary Table S4) but not with GG (P = 0.65), suggesting that the SIX3–SIX2 variants were glycaemic. We also examined the association of the genetic variants with the BMI, FPG, eGFRcrea, eGFRcys, and erythrocyte-related traits (RBC, Hb, HCT, MCV, MCH and MCHC) using the Iwate population. Neither novel variant was significantly associated with those variables (Supplementary Tables S4 and S5).
Association of genetic variants with glycated albumin and glycation gap. For novel and previously-reported HbA1c-associated variants, the association with glycated albumin (GA) and glycation gap (GG) was tested using the Iwate population by a linear regression model adjusted for age and sex. The x- and y-axes represent −log10 P-value of association with GA and GG, respectively. Novel lead variants were shown in red, whereas other variants were shown in brown.
To examine the relationship between the novel locus and glycaemic traits, we looked up the summary statistics data available from the DIAGRAM40,41, MAGIC42, and CKDGen43,44 consortia. For the TMC6–TMC8 locus, trans-ethnic T2D GWAS data40 from Europeans, East Asians, South Asians, Mexicans, and Mexican-Americans showed an association between the novel locus and the T2D risk (P = 7.0 × 10−4), but later European T2D GWAS data41 did not support this association (P = 0.25) (Supplementary Table S6). We did not find an appropriate proxy for rs2748427 when looked up the MAGIC data42 (the maximum LD r 2 in East Asians was 0.144; Supplementary Table S7). The CKDGen data43,44 weakly supported an association between the TMC6–TMC8 locus and the urinary albumin-to-creatinine ratio (UACR) in nondiabetic subjects (P = 0.030) but did not support an association with the chronic kidney disease (CKD) risk (P = 0.92), eGFRcrea (P = 0.39), eGFRcys (P = 0.72), and microalbuminuria (MA; P = 0.81) (Supplementary Tables S8 and S9). For the SIX3–SIX2 locus, previous GWASs of FPG showed that the locus achieved GWS in East Asian populations, although the association was not significant in Europeans45. Consistent with this lack of association with FPG in Europeans, variants at the SIX3–SIX2 locus were not associated with the T2D risk, FPG, fasting insulin, beta-cell functions, CKD risk, eGFRcrea, eGFRcys, UACR, or microalbuminuria in Europeans (P > 0.05) (Supplementary Tables S6–S9).
Replication analysis of previously reported variants
We examined the association of 21 previously reported lead variants8,9,10,11,12,13,14,15 based on our meta-analysis results. Of the lead variants, 4 were monoallelic or had a very low MAF (<0.01); therefore, these variants were excluded from this analysis. Most of the remaining 17 variants were nominally significantly associated with the HbA1c level (P < 0.05), with the exceptions of rs7998202 (at the ATP11A/TUBGCP3 locus), rs12603404 (C17orf53), and rs11667818 (MYO9B) (Table 3). The effect direction for the 14 significant variants was perfectly consistent between the Japanese and previously reported East Asian populations. Previous reports showed that the rs6474359 (ANK1) C allele was associated with an increased HbA1c level in East Asians14 but with a decreased level in Europeans10. In the Japanese population, the C allele was associated with an increased HbA1c level, which was consistent with the findings from the East Asian population.
Of the 14 replicated variants in the Iwate population, 5 (rs6684514 at the TMEM79 locus, rs9399137 at the HBS1L/MYB locus, rs4737009 at the ANK1 locus, rs9933309 at the CYBA locus, and rs1046896 at the FN3K locus) were associated with GG, and 6 (rs3755157 at the G6PC2/ABCB11 locus, rs7772603 at the CDKAL1 locus, rs730497 at the GCK locus, rs13266634 at the SLC30A8 locus, rs1387153 at the MTNR1B locus, and rs9933309 at the CYBA locus) were associated with GA (Fig. 3 and Supplementary Table S4). All 5 variants associated with GG were associated with 1 or more erythrocyte-related traits (P < 0.05) in the Iwate population, with the exception of rs1046896 at the FN3K locus (Supplementary Table S5). For the 6 variants associated with GA, 5 were associated with the T2D risk and/or FPG in the European GWAS summary data40,41,42 (Supplementary Tables S6 and S7). The lone exception was rs9933309 at the CYBA locus, which was also associated with GG.
Variant functions at novel loci
For the TMC6–TMC8 locus, we found 17 variants that met GWS (Supplementary Table S2). Most of the variants with GWS showed moderate LD with the lead variant rs2748427 (Supplementary Table S2). Two variants caused missense alterations in the TMC6 amino acid sequence (Supplementary Table S10). Based on bioinformatics analysis using SIFT46 and PolyPhen47, 1 amino acid change (rs2748427, W125R) was predicted to be a tolerated and benign variant, and 1 substitution (rs12449858, L153F) was predicted to be a deleterious and possibly damaging variant.
To interrogate the effects of the novel variants with GWS on the expression of neighbouring genes, we accessed the GTEx database48,49, which contains significant eQTL variant-gene pairs from 44 tissues. The alleles associated with an increased HbA1c level were also associated with decreased expression levels of the TMC6 and TNRC6C-AS1 genes in the heart, artery, and thyroid and were associated with increased TMC8 gene expression level in whole blood (Supplementary Table S11). Two variants were associated with increased TMC6 gene expression level in whole blood, although 1 variant had the opposite effect.
Furthermore, we performed a cis-eQTL analysis using the Japanese multi-omics iMETHYL database35. Transcriptome data were available for purified CD4+ T cells and monocytes. The results showed that the lead variant rs2748427 was not associated with TMC6 and TMC8 gene expression in the 2 purified cell types but was associated with arylformamidase (AFMID) gene expression (Supplementary Table S12). The transcription start site of the AFMID gene is located ~62 kb from the lead variant. The rs2748427 G allele, which was associated with an increased HbA1c level, was associated with decreased AFMID gene expression level.
For the SIX3–SIX2 locus, 6 variants located in the intergenic regions between SIX3 and SIX2 achieved GWS (Supplementary Table S3). Accordingly, these 6 variants did not change the amino acid sequence of any protein-coding gene (Supplementary Table S13). No significant cis-eQTL was found for the SIX3–SIX2 locus in the GTEx and iMETHYL databases (Supplementary Tables S14 and S15).
HbA1c-associated molecular pathways
Variants with weak genetic effects may be clustered on certain genes even after accounting for LD, and these genes may be overrepresented in certain molecular pathways. Accordingly, we searched for molecular pathways that were collectively associated with HbA1c by combining our GWAS data and prior knowledge in pathway databases. Based on the KEGG pathway database50, which consists of 168 pathways, the linoleic acid (LA) metabolic pathway was significantly associated after multiple testing correction (P < 0.05/168) (Supplementary Table S16). The pathway is composed of 34 genes, of which 12 genes were nominally associated with HbA1c (P < 0.05). The 12 significant genes included fatty acid desaturase genes (FADS1, FADS2, and FADS3), cytochrome P450 enzyme genes (CYP1A2, CYP2C8, CYP2C18, CYP2C19, and CYP2E1), phospholipase A2 genes (PLA2G2E and PLA2G2F), an aldo-keto reductase gene (AKR1B10), and a hydroxy-delta-5-steroid dehydrogenase gene (HSD3B7) (Supplementary Table S17).
Based on the Ingenuity Pathway Database (http://www.ingenuity.com/index.html) analysis, the 14-3-3-mediated signalling pathway was significantly associated with HbA1c (P < 0.05/92) (Supplementary Table S18). Of the 23 genes in the pathway, BAD, CDKN1B, PDCD6IP, and VIM were nominally associated (P < 0.05) (Supplementary Table S19).
We also examined the PANTHER51 and GO term52 classifications, but no pathway or gene set was associated with HbA1c after Bonferroni correction (Supplementary Tables S20–S23).
Furthermore, we investigated the association of the HbA1c-associated KEGG LA and Ingenuity 14-3-3-mediated signalling pathways with GA, GG, FPG, eGFRcrea, eGFRcys, and erythrocyte-related traits in the Iwate population. The results showed that neither pathway was associated with any traits, with the exception of a weak association between the KEGG LA pathway and Hb (P = 0.045) (Supplementary Tables S24 and S25).
Discussion
A genome-wide meta-analysis of 2 Japanese populations revealed 2 novel HbA1c-associated loci (TMC6–TMC8 and SIX3–SIX2). The association between the TMC6–TMC8 locus and HbA1c was replicated in European populations, and the association between the SIX3–SIX2 locus and FPG was previously reported in East Asian populations45. Thus, we successfully identified these 2 loci as new genetic factors influencing the HbA1c levels in non-diabetic subjects. The association analysis with GA and GG indicated that the TMC6–TMC8 locus may be involved in a non-glycaemic pathway, whereas the SIX3–SIX2 locus may be involved in a glycaemic pathway.
GG, indicating the discordance between HbA1c and other measures of glycaemic control (e.g., GA and fructosamine), has been associated with renal impairment and diabetic nephropathy19,53,54,55. GG was shown previously to be a heritable trait56,57. The novel HbA1c-associated lead variant rs2748427 at the TMC6–TMC8 locus was strongly associated with GG in our Japanese population. This result indicated that the TMC6–TMC8 variants may influence the HbA1c level through a non-glycaemic pathway. Recent findings have indicated that the erythrocyte lifespan and glucose gradient across the erythrocyte membrane, i.e., the intracellular versus extracellular glucose concentration, may account for GG57,58. A previous genetic study showed an association between rs2748427 and MCV in their discovery populations with a suggestive significance (P = 1.6 × 10−5), but the association was not replicated in their replication populations59. In our Japanese population, the novel lead variant was not associated with any erythrocyte-related parameters, including MCV. Accordingly, the association between the TMC6–TMC8 locus and MCV was inconclusive. Taken together, it is hypothesized that the TMC6–TMC8 variants may affect GG through the erythrocyte life span, iron handling, glucose distribution across the erythrocyte membrane or an as-yet-undiscovered mechanism60.
Contrary to this hypothesis, look up of a trans-ethnic T2D GWAS showed an association between variants at the TMC6–TMC8 locus and T2D that did not reach GWS, which could be interpreted as suggesting that the effects of the variants on HbA1c may be mediated through their effects on glycaemia. However, the association was not significant in a European GWAS for T2D. In addition, we found weak genetic evidence that variants at this locus were associated with UACR in Europeans. The significance of these findings is unclear at this time.
In the eQTL analyses, we showed that variants at the TMC6–TMC8 locus affected gene expression levels of 3 protein-coding genes, i.e., TMC6, TMC8 and AFMID. TMC6 and TMC8 play central roles in anti-human papillomavirus (HPV) barrier. Rare loss-of-function genetic variants in either gene can lead to epidermodysplasia verruciformis (EV; OMIM 226400), which is characterized by abnormal susceptibility to specific HPVs and is associated with a high risk of skin carcinoma61,62. AFMID encodes an enzyme that converts N-formyl-L-kynurenine to L-kynurenine (KYN)63. In turn, KYN and several of its metabolites have an impact on insulin secretion and sensitivity64,65. However, it is difficult to interpret these data in the light of the associations between this locus and GG, which suggests that the effect of variants at this locus on HbA1c may relate to non-glycemic determinants of HbA1c.
For the SIX3−SIX2 locus, our data showing that the locus was associated with GA and evidence from a previous East Asian GWAS showing that the locus was associated with FPG consistently indicated that the locus may influence HbA1c through a glycaemic pathway. Although the association between the SIX3–SIX2 locus and FPG was not significant in our Japanese population, the association analysis may lack sufficient statistical power due to the limited number of subjects with available FPG data (n = 604). Previous studies showed that the effect size of the SIX3−SIX2 variants on FPG was not heterogeneous among East Asian populations, whereas the association between the locus and FPG was not significant among European populations45. Our data also showed that the effect size of the locus on HbA1c was not heterogeneous among 2 Japanese populations. The eQTL analysis did not identify genes with expression levels that were significantly affected by this locus. The mechanisms by which the SIX3–SIX2 locus affects FPG and HbA1c should be investigated in future studies.
A total of 7,704 non-diabetic subjects were included in our meta-analysis, making our sample smaller than the samples included in previous European10 (up to 46,368 subjects) and East Asian14 (n = 21,026) meta-analyses. A recent trans-ethnic genome-wide meta-analysis on HbA1c identified 42 novel HbA1c-associated loci from an analysis of up to ~160,000 non-diabetic individuals of European, African, East Asian or South Asian ancestry66. The recent analysis independently found HbA1c-associated variants at the TMC6–TMC8 locus with relatively small effect sizes (β = 0.013 [SE = 0.033; P = 1.3 × 10−4; n = 41,300] for Europeans, and β = 0.019 [SE = 0.059; P = 1.2 × 10−3; n = 9,477] for East Asians) compared to those observed in our 2 Japanese populations (Table 2). In previous meta-analyses and recent trans-ethnic studies, only one-third of the subjects were available for analysis of the TMC6–TMC8 locus, possibly because less dense SNP arrays (e.g., Illumina 300 K) were included in their datasets10,66. A combination of effect size heterogeneity and SNP array coverage would explain why previous meta-analyses10,14 did not identify variants at the TMC6–TMC8 locus. The recent trans-ethnic analysis did not identify variants at the SIX3–SIX2 locus, possibly because majority of the subjects in their datasets had European ancestry, and the association between variants at the SIX3–SIX2 locus and FPG/HbA1c was not significant in Europeans10,45. The effect size heterogeneity among ethnicities would explain why the previous European meta-analysis10 did not detect variants at the locus.
By combining our GWAS data and pathway knowledge, we provided genetic evidence that the LA metabolic and 14-3-3-mediated signalling pathways were collectively associated with HbA1c. LA is abundant in vegetable oils and is a major dietary source of ω-6 polyunsaturated fatty acids (PUFAs). Arachidonic acid (AA), which is a 20-carbon ω-6 PUFA synthesized from LA, is a dominant substrate for ω-6 eicosanoids, which have pro-inflammatory activities67,68. Observational and interventional studies have demonstrated beneficial health outcomes of long-chain ω-3 PUFAs69, which competitively inhibit the synthesis of ω-6 eicosanoids from AA67,68. The 14-3-3 proteins integrate multiple signalling cues by recognizing post-transcriptional phosphorylation of cellular proteins and coordinating their subcellular localization70. The 14-3-3 proteins have been shown to protect pancreatic β-cells from pro-inflammatory cytokines by mediating pro-survival signals71. Knockout of the 14-3-3ζ isoform resulted in glucose intolerance and insulin resistance72. These data provided genetic evidence that the genetic risk for elevated HbA1c levels was attributable to the LA metabolic and 14-3-3-mediated signalling pathways.
A limitation of this study is that we analysed only Japanese populations and studied GWAS summary data only from individuals with European ancestry. Although the novel TMC6–TMC8 locus showed significance in both our Japanese and previous European meta-analyses, the effect size showed a large degree of heterogeneity. According to the 1000 Genomes Project data (phase 3)29,30, the rs2748427 G allele frequency is 0.216 in Japanese populations, which is in agreement with the value of 0.179 observed in our Japanese populations. The G allele frequency was 0.291 in East Asians, which was higher than the value observed in Europeans (0.213) and Americans (0.220) and lower than the value observed in South Asians (0.333) and Africans (0.523). The association between the novel locus and HbA1c is testable in various populations, because the novel variant is common in several ethnic groups. Future studies are needed to uncover the genetic effect of the novel locus in other ethnic groups. In addition, the HbA1c measurement method differed between the Iwate and Miyagi populations. HbA1c levels measured using immunoassay-based methods tend to be slightly lower than those measured using HPLC-based methods73. Indeed, the average HbA1c level in the Miyagi subjects was lower than that in the Iwate subjects (Table 1). However, we separately analysed the association between genetic variants and HbA1c for each of the Iwate and Miyagi populations. Then, we performed a meta-analysis of the association evidence from the 2 Japanese populations. Accordingly, the effects of the differences in measurement methods were minimized.
In summary, we identified the TMC6–TMC8 and SIX3–SIX2 loci as novel genetic factors associated with HbA1c. The TMC6–TMC8 locus may influence the HbA1c level through a non-glycaemic pathway, whereas the SIX3–SIX2 locus may affect the HbA1c level via a glycaemic pathway. In addition, we provided genetic evidence that the LA metabolic and 14-3-3-mediated signalling pathways may modulate the HbA1c level. Genetic evidence from this study provides insights into the molecular mechanisms that modulate the HbA1c level in non-diabetic subjects.
References
Nathan, D. M. et al. Translating the A1C assay into estimated average glucose values. Diabetes Care 31, 1473–1478, https://doi.org/10.2337/dc08-0545 (2008).
American Diabetes Association. (2) Classification and diagnosis of diabetes. Diabetes Care 38Suppl, S8–S16, https://doi.org/10.2337/dc15-S005 (2015).
Selvin, E. et al. Glycated hemoglobin, diabetes, and cardiovascular risk in nondiabetic adults. N Engl J Med 362, 800–811, https://doi.org/10.1056/NEJMoa0908359 (2010).
Sherwani, S. I., Khan, H. A., Ekhzaimy, A., Masood, A. & Sakharkar, M. K. Significance of HbA1c Test in Diagnosis and Prognosis of Diabetic Patients. Biomark Insights 11, 95–104, https://doi.org/10.4137/BMI.S38440 (2016).
Snieder, H. et al. HbA(1c) levels are genetically determined even in type 1 diabetes: evidence from healthy and diabetic twins. Diabetes 50, 2858–2863 (2001).
Mills, G. W. et al. Heritability estimates for beta cell function and features of the insulin resistance syndrome in UK families with an increased susceptibility to type 2 diabetes. Diabetologia 47, 732–738, https://doi.org/10.1007/s00125-004-1338-2 (2004).
Simonis-Bik, A. M. et al. The heritability of HbA1c and fasting blood glucose in different measurement settings. Twin Res Hum Genet 11, 597–602, https://doi.org/10.1375/twin.11.6.597 (2008).
Pare, G. et al. Novel association of HK1 with glycated hemoglobin in a non-diabetic population: a genome-wide evaluation of 14,618 participants in the Women’s Genome Health Study. PLoS Genet 4, e1000312, https://doi.org/10.1371/journal.pgen.1000312 (2008).
Franklin, C. S. et al. The TCF7L2 diabetes risk variant is associated with HbA(1)(C) levels: a genome-wide association meta-analysis. Ann Hum Genet 74, 471–478, https://doi.org/10.1111/j.1469-1809.2010.00607.x (2010).
Soranzo, N. et al. Common variants at 10 genomic loci influence hemoglobin A(1)(C) levels via glycemic and nonglycemic pathways. Diabetes 59, 3229–3239, https://doi.org/10.2337/db10-0502 (2010).
Ryu, J. & Lee, C. Association of glycosylated hemoglobin with the gene encoding CDKAL1 in the Korean Association Resource (KARE) study. Hum Mutat 33, 655–659, https://doi.org/10.1002/humu.22040 (2012).
Chen, P. et al. A study assessing the association of glycated hemoglobin A1C (HbA1C) associated variants with HbA1C, chronic kidney disease and diabetic retinopathy in populations of Asian ancestry. PloS one 8, e79767, https://doi.org/10.1371/journal.pone.0079767 (2013).
An, P. et al. Genome-wide association study identifies common loci influencing circulating glycated hemoglobin (HbA1c) levels in non-diabetic subjects: the Long Life Family Study (LLFS). Metabolism 63, 461–468, https://doi.org/10.1016/j.metabol.2013.11.018 (2014).
Chen, P. et al. Multiple nonglycemic genomic loci are newly associated with blood level of glycated hemoglobin in East Asians. Diabetes 63, 2551–2562, https://doi.org/10.2337/db13-1815 (2014).
Hebbar, P. et al. Genetic risk variants for metabolic traits in Arab populations. Sci Rep 7, 40988, https://doi.org/10.1038/srep40988 (2017).
Leong, A. & Meigs, J. B. Type 2 Diabetes Prevention: Implications of Hemoglobin A1c Genetics. Rev Diabet Stud 12, 351–362, https://doi.org/10.1900/RDS.2015.12.351 (2015).
Soranzo, N. Genetic determinants of variability in glycated hemoglobin (HbA(1c)) in humans: review of recent progress and prospects for use in diabetes care. Curr Diab Rep 11, 562–569, https://doi.org/10.1007/s11892-011-0232-9 (2011).
Kuriyama, S. et al. The Tohoku Medical Megabank Project: Design and Mission. J Epidemiol 26, 493–511, https://doi.org/10.2188/jea.JE20150268 (2016).
Kim, M. K., Yun, K. J., Kwon, H. S., Baek, K. H. & Song, K. H. Discordance in the levels of hemoglobin A1C and glycated albumin: Calculation of the glycation gap based on glycated albumin level. J Diabetes Complications 30, 477–481, https://doi.org/10.1016/j.jdiacomp.2015.12.022 (2016).
Kim, M. K., Jeong, J. S., Kwon, H. S., Baek, K. H. & Song, K. H. Concordance the hemoglobin glycation index with glycation gap using glycated albumin in patients with type 2 diabetes. J Diabetes Complications, https://doi.org/10.1016/j.jdiacomp.2017.04.015 (2017).
Matsuo, S. et al. Revised equations for estimated GFR from serum creatinine in Japan. Am J Kidney Dis 53, 982–992, https://doi.org/10.1053/j.ajkd.2008.12.034 (2009).
Horio, M. et al. GFR estimation using standardized serum cystatin C in Japan. Am J Kidney Dis 61, 197–203, https://doi.org/10.1053/j.ajkd.2012.07.007 (2013).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81, 559–575, https://doi.org/10.1086/519795 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7, https://doi.org/10.1186/s13742-015-0047-8 (2015).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet 38, 904–909, https://doi.org/10.1038/ng1847 (2006).
Yamaguchi-Kabata, Y. et al. Japanese population structure, based on SNP genotypes from 7003 individuals compared to other ethnic groups: effects on population-based association studies. Am J Hum Genet 83, 445–456, https://doi.org/10.1016/j.ajhg.2008.08.019 (2008).
Delaneau, O., Marchini, J. & Zagury, J. F. A linear complexity phasing method for thousands of genomes. Nat Methods 9, 179–181, https://doi.org/10.1038/nmeth.1785 (2011).
Das, S. et al. Next-generation genotype imputation service and methods. Nat Genet 48, 1284–1287, https://doi.org/10.1038/ng.3656 (2016).
1000 Genomes Project Consortium et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65, https://doi.org/10.1038/nature11632 (2012).
1000 Genomes Project Consortium et al. A global reference for human genetic variation. Nature 526, 6–74, https://doi.org/10.1038/nature15393 (2015).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat Genet 42, 565–569, https://doi.org/10.1038/ng.608 (2010).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 88, 76–82, https://doi.org/10.1016/j.ajhg.2010.11.011 (2011).
Hachiya, T. et al. Genetic Predisposition to Ischemic Stroke: A Polygenic Risk Score. Stroke 48, 253–258, https://doi.org/10.1161/strokeaha.116.014506 (2017).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics (Oxford, England) 26, 2190–2191, https://doi.org/10.1093/bioinformatics/btq340 (2010).
Hachiya, T. et al. Genome-wide identification of inter-individually variable DNA methylation sites improves the efficacy of epigenetic association studies. NPJ Genom Med. 2, 11, https://doi.org/10.1038/s41525-017-0016-5 (2017).
Nagasaki, M. et al. Rare variant discovery by deep whole-genome sequencing of 1,070 Japanese individuals. Nature communications 6, 8018, https://doi.org/10.1038/ncomms9018 (2015).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics (Oxford, England) 25, 1105–1111, https://doi.org/10.1093/bioinformatics/btp120 (2009).
de Leeuw, C. A., Mooij, J. M., Heskes, T. & Posthuma, D. MAGMA: generalized gene-set analysis of GWAS data. PLoS computational biology 11, e1004219, https://doi.org/10.1371/journal.pcbi.1004219 (2015).
Machiela, M. J. & Chanock, S. J. LDlink: a web-based application for exploring population-specific haplotype structure and linking correlated alleles of possible functional variants. Bioinformatics (Oxford, England) 31, 3555–3557, https://doi.org/10.1093/bioinformatics/btv402 (2015).
DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet 46, 234–244, https://doi.org/10.1038/ng.2897 (2014).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47, https://doi.org/10.1038/nature18642 (2016).
Dupuis, J. et al. New genetic loci implicated in fasting glucose homeostasis and their impact on type 2 diabetes risk. Nat Genet 42, 105–116, https://doi.org/10.1038/ng.520 (2010).
Teumer, A. et al. Genome-wide Association Studies Identify Genetic Loci Associated With Albuminuria in Diabetes. Diabetes 65, 803–817, https://doi.org/10.2337/db15-1313 (2016).
Pattaro, C. et al. Genetic associations at 53 loci highlight cell types and biological pathways relevant for kidney function. Nature communications 7, 10023, https://doi.org/10.1038/ncomms10023 (2016).
Kim, Y. J. et al. Large-scale genome-wide association studies in East Asians identify new genetic loci influencing metabolic traits. Nat Genet 43, 990–995, https://doi.org/10.1038/ng.939 (2011).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nature protocols 4, 1073–1081, https://doi.org/10.1038/nprot.2009.86 (2009).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat Methods 7, 248–249, https://doi.org/10.1038/nmeth0410-248 (2010).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet 45, 580–585, https://doi.org/10.1038/ng.2653 (2013).
GTEx Consortium. Human genomics. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660, https://doi.org/10.1126/science.1262110 (2015).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic acids research 45, D353–d361, https://doi.org/10.1093/nar/gkw1092 (2017).
Mi, H., Muruganujan, A., Casagrande, J. T. & Thomas, P. D. Large-scale gene function analysis with the PANTHER classification system. Nature protocols 8, 1551–1566, https://doi.org/10.1038/nprot.2013.092 (2013).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29, https://doi.org/10.1038/75556 (2000).
Cohen, R. M., Holmes, Y. R., Chenier, T. C. & Joiner, C. H. Discordance between HbA1c and fructosamine: evidence for a glycosylation gap and its relation to diabetic nephropathy. Diabetes Care 26, 163–167 (2003).
Rodriguez-Segade, S., Rodriguez, J., Cabezas-Agricola, J. M., Casanueva, F. F. & Camina, F. Progression of nephropathy in type 2 diabetes: the glycation gap is a significant predictor after adjustment for glycohemoglobin (Hb A1c). Clin Chem 57, 264–271, https://doi.org/10.1373/clinchem.2010.144949 (2011).
Cosson, E. et al. Glycation gap is associated with macroproteinuria but not with other complications in patients with type 2 diabetes. Diabetes Care 36, 2070–2076, https://doi.org/10.2337/dc12-1780 (2013).
Cohen, R. M. et al. Evidence for independent heritability of the glycation gap (glycosylation gap) fraction of HbA1c in nondiabetic twins. Diabetes Care 29, 1739–1743, https://doi.org/10.2337/dc06-0286 (2006).
Leslie, R. D. & Cohen, R. M. Biologic variability in plasma glucose, hemoglobin A1c, and advanced glycation end products associated with diabetes complications. J Diabetes Sci Technol 3, 635–643, https://doi.org/10.1177/193229680900300403 (2009).
Herman, W. H. & Cohen, R. M. Hemoglobin A1c: teaching a new dog old tricks. Ann Intern Med 152, 815–817, https://doi.org/10.7326/0003-4819-152-12-201006150-00011 (2010).
Pankratz, N. et al. Meta-analysis of rare and common exome chip variants identifies S1PR4 and other loci influencing blood cell traits. Nat Genet 48, 867–876, https://doi.org/10.1038/ng.3607 (2016).
Cohen, R. M. & Lindsell, C. J. When the blood glucose and the HbA(1c) don’t match: turning uncertainty into opportunity. Diabetes Care 35, 2421–2423, https://doi.org/10.2337/dc12-1479 (2012).
Ramoz, N. et al. Mutations in two adjacent novel genes are associated with epidermodysplasia verruciformis. Nat Genet 32, 579–581, https://doi.org/10.1038/ng1044 (2002).
Lazarczyk, M. et al. Regulation of cellular zinc balance as a potential mechanism of EVER-mediated protection against pathogenesis by cutaneous oncogenic human papillomaviruses. The Journal of experimental medicine 205, 35–42, https://doi.org/10.1084/jem.20071311 (2008).
Dobrovolsky, V. N. et al. Effect of arylformamidase (kynurenine formamidase) gene inactivation in mice on enzymatic activity, kynurenine pathway metabolites and phenotype. Biochimica et biophysica acta 1724, 163–172, https://doi.org/10.1016/j.bbagen.2005.03.010 (2005).
Stone, T. W. & Darlington, L. G. Endogenous kynurenines as targets for drug discovery and development. Nature reviews. Drug discovery 1, 609–620, https://doi.org/10.1038/nrd870 (2002).
Hugill, A. J. et al. Loss of arylformamidase with reduced thymidine kinase expression leads to impaired glucose tolerance. Biology open 4, 1367–1375, https://doi.org/10.1242/bio.013342 (2015).
Wheeler, E. et al. Impact of common genetic determinants of Hemoglobin A1c on type 2 diabetes risk and diagnosis in ancestrally diverse populations: A transethnic genome-wide meta-analysis. PLoS Med 14, e1002383, https://doi.org/10.1371/journal.pmed.1002383 (2017).
James, M. J., Gibson, R. A. & Cleland, L. G. Dietary polyunsaturated fatty acids and inflammatory mediator production. The American journal of clinical nutrition 71, 343s–348s (2000).
Calder, P. C. Polyunsaturated fatty acids and inflammation. Biochemical Society transactions 33, 423–427, https://doi.org/10.1042/bst0330423 (2005).
Ruxton, C. H., Reed, S. C., Simpson, M. J. & Millington, K. J. The health benefits of omega-3 polyunsaturated fatty acids: a review of the evidence. Journal of human nutrition and dietetics: the official journal of the British Dietetic Association 17, 449–459, https://doi.org/10.1111/j.1365-277X.2004.00552.x (2004).
Morrison, D. K. The 14-3-3 proteins: integrators of diverse signaling cues that impact cell fate and cancer development. Trends in cell biology 19, 16–23, https://doi.org/10.1016/j.tcb.2008.10.003 (2009).
Lim, G. E., Piske, M. & Johnson, J. D. 14-3-3 proteins are essential signalling hubs for beta cell survival. Diabetologia 56, 825–837, https://doi.org/10.1007/s00125-012-2820-x (2013).
Lim, G. E. et al. 14-3-3zeta coordinates adipogenesis of visceral fat. Nature communications 6, 7671, https://doi.org/10.1038/ncomms8671 (2015).
Manley, S. E. et al. Comparison of IFCC-calibrated HbA(1c) from laboratory and point of care testing systems. Diabetes Res Clin Pract 105, 364–372, https://doi.org/10.1016/j.diabres.2014.05.003 (2014).
Acknowledgements
This work was supported by the Tohoku Medical Megabank Project (Special Account for Reconstruction from the Great East Japan Earthquake) from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) and the Japan Agency for Medical Research and Development (AMED). The authors thank the members of the Iwate Tohoku Medical Megabank Organization of Iwate Medical University (IMM) and the Tohoku Medical Megabank Organization of Tohoku University (ToMMo) for their encouragement and support. We are grateful to the Tohoku Medical Megabank Project participants. GWAS summary statistics for T2D risk, glycemic traits, and kidney-related traits have been contributed by DIAGRAM, MAGIC and CKDGen investigators, respectively, and have been downloaded from http://www.diagram-consortium.org (DIAGRAM), http://www.magicinvestigators.org (MAGIC) and http://ckdgen.imbi.uni-freiburg.de (CKDGen).
Author information
Authors and Affiliations
Contributions
T.H., S.K., Y.H., H.O., Y.I. and A.S. wrote the manuscript. K.T. and A.H. were in charge of cohort data management. G.Y. prepared for the genotype dataset for GWAS. T.H. performed statistical analysis, GWAS and replication analyses. M.Y., K.O., M.N., J.H., Y.I., M.S. and A.S. supervised the work. T.H. and A.S. designed and coordinated the project. All authors commented on and approved the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hachiya, T., Komaki, S., Hasegawa, Y. et al. Genome-wide meta-analysis in Japanese populations identifies novel variants at the TMC6–TMC8 and SIX3–SIX2 loci associated with HbA1c . Sci Rep 7, 16147 (2017). https://doi.org/10.1038/s41598-017-16493-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-16493-0
This article is cited by
-
scParser: sparse representation learning for scalable single-cell RNA sequencing data analysis
Genome Biology (2024)
-
Impact of diet and host genetics on the murine intestinal mycobiome
Nature Communications (2023)
-
Development and validation of genome-wide polygenic risk scores for predicting breast cancer incidence in Japanese females: a population-based case-cohort study
Breast Cancer Research and Treatment (2023)
-
Comparison of the loci associated with HbA1c and blood glucose levels identified by a genome-wide association study in the Japanese population
Diabetology International (2023)
-
CRISPR-based genome editing in primary human pancreatic islet cells
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.