Abstract
Branch number is an important trait in plant architecture that can influence crop yield and quality in Brassica napus. Here, we detected the QTLs responsible for branch number in a DH population and its reconstructed F2 population over two years. Further, a GWAS research on branch number was performed using a panel of 327 accessions with 33186 genomic SNPs from the 60 K Brassica Illumina® Infinium SNP array. Through combining linkage analysis and association mapping, a new QTL was fine mapped onto C03. Subsequently, we tested the correlations between the SNP polymorphisms and mRNA expression levels of genes in the target interval to identify potential loci or genes that control branch number through expression. The results show that 4 SNP loci are associated with the corresponding gene expression levels, and one locus (BnaC03g63480D) exhibited a significant correlation between the phenotype variation and gene expression levels. Our results provide insights into the genetic basis for branching morphogenesis and may be valuable for optimizing architecture in rapeseed breeding.
Similar content being viewed by others
Introduction
The branch system is an important part of plant architecture1,2,3,4,5. The number of branches (BN) on the main stem is one of the most important properties in rapeseed architecture. A proper BN may aid in improving rapeseed yield and quality.
The shoot branching process generally involves two developmental stages, the formation of axillary meristems in the leaf axils and the growth of axillary buds6,7,8,9,10. This process is controlled by multiple factors and can be influenced by environmental conditions or developmental signals1,7,9,10,11,12,13,14,15,16,17,18,19,20,21,22. Three classical phytohormones, auxin, cytokinin, and strigolactone, and the genes associated with their homeostasis and signaling are likely largely responsible for regulating branching1,21,22,23. Moreover, other signals, such as sugars or molecular actors on plant phase transition, also strongly influence shoot branching11. To date, many genes involved in axillary meristem initiation and outgrowth have been reported in tomato, rice, Arabidopsis, maize, pea, petunia and barley24,25,26,27,28,29,30,31,32,33. These genes are mostly associated with homeostasis and signaling of phytohormones and growth regulators. In addition, certain transcription factors that play a role in regulating the transcription of other genes, such as REV 34, LAS 35 and RAX 1, 2, and 336, also control branching. Moreover, the genes encoding cytochrome P450, MAPKK7, arabinogalactan proteins, and other DNA-binding proteins have been identified that are involved in branch outgrowth37.
Linkage mapping is well-established and has been employed for mapping QTLs to determine quantitative traits in rapeseed38,39,40,41,42,43,44,45. However, fine mapping QTLs using a linkage analysis requires a population with thousands of individuals, and the limited polymorphic loci between the two parents will influence the mapping accuracy46. Recently, association mapping has been widely employed to detect quantitative trait loci47,48,49,50,51. It directly identifies associations between DNA markers and phenotypes in natural populations based on linkage disequilibrium (LD)52. Compared with linkage mapping, association mapping does not require constructing special mapping populations, and it uses high recombination in natural populations. Therefore, association mapping can complement linkage mapping and facilitate fine-scale QTL mapping52,53. However, in an association analysis, the population structure can produce a stronger LD between non-linked loci due to genetic drift, population stratification and natural selection54,55. Thus, combining association mapping and a linkage analysis not only avoids the false positives from associated loci due to high LD but also facilitates fine mapping of a target region with a large QTL interval56.
In rapeseed, certain QTLs that are related to branch number have been identified through linkage analyses in recent years57,58,59,60,61. Most QTLs were located on chromosomes A02, A06, A07, C03, C06 and C09 and were only detected in a single-year environment. Recently, Luo et al.62 reported six SNPs on A03 associated with rapeseed branch number using genome-wide association study (GWAS). However, despite the studies on rapeseed branching, the basis for genetically controlling branching morphogenesis has not been fully elucidated for rapeseed, and few genes have been predicted in the QTL intervals. Better understanding the genetic determinants for rapeseed branching is needed.
In the present study, the QTLs responsible for branch number were detected in a doubled haploid (DH) population and its reconstructed F2 (RC-F2) population. Further, a GWAS research on branch number was performed using a panel of 327 accessions with 33186 genomic SNPs from the 60 K Brassica Illumina® Infinium SNP array. Combined with the gene expression analysis, a candidate gene that controls the rapeseed branch number was identified.
Results
Phenotypic variation in DH, RC-F2 and association populations
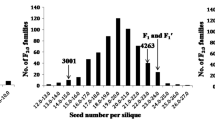
The average branch number of the parental line ‘SWU07’ is 7.92, whereas that of the other parental line ‘Express’ is 8.50. Extensive phenotypic variations for DH, RC-F2 and association mapping populations used in this study were observed (Fig. 1, Table 1). The frequency distribution of BN for the three populations appears to be an approximately normal distribution in all the separate experiments, which indicated that the branch number phenotype is governed by multiple genes (Fig. 1, Table S1). Transgressive segregation was observed in the DH and RC-F2 populations, which suggests that the loci control BN was harbored by different alleles of the two parents.
The results of ANOVA revealed significant differences among genotypes, environments and genotype-by-environment interactions for branch number in the three populations (P < 0.01) (Table 2). High broad-sense heritability was detected for BN with average of 49.19%, 67.02% and 79.17% in DH, RC-F2 and association mapping population, respectively (Table 2). The significant and positive correlation for BN was detected between the years in the three populations (r = 0.52, 0.28 and 0.59 in the DH, RC-F2 and association mapping populations, respectively; P < 0.01).
QTL analysis
Ten QTLs associated with BN are located on chromosomes A01, A03, A05, A07, C01 and C03 in the DH and RC-F2 populations over the two years, which explained 3.73–9.77% of the phenotypic variation (Fig. 2, Table 3). Two QTLs on C03 were detected in different years or in different populations. Finally, seven QTLs were generated after integrating the QTLs with overlapped confidence intervals in different experiments.
Population structure and relative kinship in association population
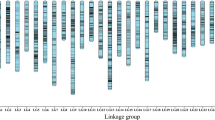
The population structure was estimated using 5700 SNP. We select 300 SNP in each chromosome of B. napus. The result showed that the most significant change of likelihood occurred when K increased from 3 to 4, and the highest Δk value was observed at k = 3 (Fig. 3a). The two parameters suggested that the 327 genotypes were assigned into three groups. The PCA based on the 33,186 genome-wide SNPs showed that the first two principal components explained 16.48 and 8.45% of the genetic variance, respectively (Fig. 3b). The 327 accessions were classified into three major groups, which are mainly composed of winter ecotype, spring ecotype, and semi-winter ecotype, respectively (Fig. 3b). It is largely in accordance with the growth habit of the accessions. We used 33,186 polymorphisms to estimate the relative kinship of the materials. The analysis showed that total of 54% of kinship coefficients between lines were equal to 0, and 69% kinship coefficients were less than 0.1(Fig. 3c). This pattern of genetic relatedness revealed that most lines have no or weak kinship in this association panel.
Genome-wide association analysis
Since the dataset used in the association study consists of three subgroups, winter, semi-winter and spring ecotype, the relative kinship is weak. However, it still has a certain impact for the GWAS analysis. To avoid the false negative associations, three mixed models, K model, Q + K model and PCA + K model were chosen to determine the statistical associations between phenotypes and genotypes to evaluate the effects of population structure (Q, PC) and relative kinship (K) on BN traits. According to the Q-Q plots of the three models, for the 2013 data, the observed P values of the Q + K model were closer to the expected P values than the PCA + K model and K modle (Fig. 4a), whereas for the 2014 data, the observed P values of the PCA + K model were closer to the expected P values than the other two modles (Fig. 4b). This indicated that the Q + K model and PCA + K model could effectively control false positive associations and avoid false negative associations for 2013 and 2014 data, respectively. Thus, we choose Q + K and PCA + K models to perform the association analysis for the 2013 and 2014 data, respectively.
We used 33,186 polymorphisms with a minor allele frequency (MAF) ≥ 0.05 for a GWAS. Notable positive associations were observed in the Manhattan plots (Fig. S1). The association analysis identified 50 SNP loci significantly associated with branch number at P < 3.01 × 10−5 over the two years (Table S2; Fig. S1). 44 and 15 SNPs were detected in the year of 2013 and 2014, respectively, and 9 were detected in the two-year experiments. According to the decay of LD, the physically closed SNPs were integrated to the single QTL interval. Totally, four GWAS-based QTL regions were generated in the two-year experiments (Table 4), which explain 8.65–20.03% of the phenotypic variance.
Candidate Gene Prediction using GWAS, QTL mapping and gene expression analyses
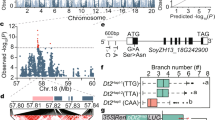
We compared the linkage mapping and association mapping results. Notably, one QTL interval on C03 was detected by the two mapping approaches. Four consecutive SNPs not only exhibited a significant correlation between DNA sequence polymorphisms and BN phenotypic variation in the GWAS analysis over the two years but were also located in the C03 QTL confidence interval by linkage mapping (Fig. 5a,b). This QTL were considered stably inherited. Therefore, we focused on the genes in this interval. The corresponding genomic sequences of the QTL region were extracted. An annotation analysis showed that chromosome C03 from 52. 18–53.69 Mb contains 138 genes with 125 SNP loci (Fig. 5c).
Identification of the target region controlling branch number by Linkage mapping and association mapping. (a) QTls associated with branch number detected on Chromosome C03; (b) Manhattanplots of association analysis for branch number. Each dot represents a SNP. The significance threshold −log10(p) = 4.52; (c) The target interval on Chromosome C03. The blue bar represents the gene which was identified to exhibit a significant correlation between phenotype variation of branch number and gene expression levels (at P < 0.05).
In the association mapping panel of this study, 31 inbred lines overlapped with the plant materials in a previous report by Bancroft et al.63. The RNA-seq data on the 31 inbred lines are available in an online resource63. Thus, we tested the correlations between the 125 SNP polymorphisms and mRNA expression levels of the 138 genes in the target interval to discern potential loci or genes that control branch number at the expression level. The results show that 4 of the 125 detected SNP loci, SNP46529, SNP50286, SNP50314, and SNP50315 are associated with the corresponding gene expression levels (Table S3). These 4 loci are positioned upstream region of 3 corresponding genes. SNP46529 and SNP50286 are positioned upstream of BnaC03g63340D and BnaC03g63530D, respectively. SNP50314 and SNP50315 are positioned upstream of BnaC03g63480D. The 4 SNP variation correlated (p < 0.01) with the corresponding gene expression levels at r = 0.5067 (SNP46529), r = −0.6142 (SNP50286), r = 0.5209 (SNP50314), and r = 0.5800 (SNP50315), respectively. Subsequently, we tested the correlations between phenotype variation and mRNA expression level of the three genes. Interestingly, among the three genes that showed a significant correlations between the SNP variation and gene expression levels, one gene (BnaC03g63480D) exhibited a significant correlation between the phenotype variation and gene expression level (at P < 0.05), while the other two have no correlation with the phenotype variation (P > 0.05). The gene expression levels of BnaC03g63480D correlated (p < 0.05) with the branch number at r = 0.1143 and r = 0.2679 in 2013 and 2014, respectively. These data strongly suggest that the gene BnaC03g63480D affects phenotypic variation via transcriptional regulation and it is a potential candidate gene for rapeseed branch number.
Discussion
Branch number is an important trait in plant architecture that can influence crop yield and quality1,2,3,4,5. Identifying the genetic loci for branch number would aid in understanding the heredity mechanism underlying branching morphogenesis and be valuable in optimizing the architecture for rapeseed breeding. Previous research shows that environmental factors can influence branching, such as planting density12,13, photoperiod9,10,14,15, and nutrient availability19,20. Thus, multi-environment testing is necessary to determine whether the effects of QTLs are due to different genes or environments. Here, we detected numerous distinct loci that influence branch number in a DH population and its reconstructed F2 population over two years. We also detected the association SNP loci via GWAS in the two-year environments. The QTL regions which detected by both linkage mapping and association mapping were considered stably inherited. Notably, most QTLs or SNP loci were only detected in a single-year environment, which suggests that environmental variation plays an important role in determining rapeseed branching morphogenesis. The concerted effects of genotype, environment and their interactions determine branching morphogenesis.
Along with the development of high-throughput SNP genotyping technology, genome-wide association studies (GWAS) have been broadly applied to unravel the genetic basis for complex traits in many crops, such as maize50, rice64 and sorghum65. Association mapping can be used to finely map linkage analysis results when a QTL region is large, and linkage mapping can exclude false positives from associated loci due to high linkage disequilibrium. The combination of association mapping and linkage analysis can increase mapping efficiency and accuracy56. In the present study, we preliminarily mapped the QTLs responsible for branch number through linkage mapping in rapeseed. Subsequently, a GWAS analysis was performed to quickly verify the QTL region and finely map the QTL within a 1.51 Mb interval (52. 18–53.69 Mb) containing 138 genes on C03. The GWAS analysis reduced the target region size and will aid in further identifying the candidate gene. Compared with previous studies for linkage mapping and GWAS analyses on rapeseed branch number57,58,59,60,61,62, the QTL we detected here is a new one.
Genetic mechanisms that regulate phenotypic variation can act not only on a gene structure level but also on a gene expression level. The transcript sequence data for Brassica napus would aid in analyzing the correlations between DNA sequence polymorphisms and mRNA expression levels. We tested the correlations between SNPs and mRNA expression levels for the genes in the target interval. We also examined the correlations between branch number phenotype and mRNA expression level of these genes. Finally, BnaC03g63480D was identified as a candidate gene according statistical correlations. The structure and function of the candidate gene were analyzed. BnaC03g63480D encodes a putative ubiquitin E3 ligase containing a RING domain. A previous study showed that the RING domain interacts with other proteins66. In Arabidopsis, BOI (AT4G19700), the BnaC03g63480D homologs, interact with DELLA proteins to inhibit GA responses by interacting with each other, binding to the same promoters of GA-responsive genes, and repressing these genes67. Gibberellins play an important role in internode elongation68, but their role in shoot branching process has not been clearly elucidated. Previous studies show that GA-deficient mutants had higher shoot branching than the wild type in Arabidopsis69, rice70, and pea71, and over expressing GA catabolism genes to reduce GA levels can increase a branching phenotype70,72. No direct evidence shows that the RING domain proteins are related to branch formation, but the RING domain can repress GA signals has been reported67. Our data confirm that the expression level of the RING domain gene BnaC03g63480D significantly correlates with branch number. We speculate that BnaC063480D controls branching morphogenesis by regulating the GA-responsive gene in rapeseed. Further studies should investigate and confirm the possible function of BnaC03g63480D in rapeseed.
Methods
Mapping population and phenotypic evaluation
The DH and RC-F2 populations used in the QTL analysis were previously described in Fu et al.73. The DH population consisting of 261 lines was developed by microspore culture, using a single F1 plant derived from a cross between the European winter oilseed rape cultivar ‘Express’ (female) and the Chinese semi-winter inbred line ‘SWU07’ (male). The RC-F2 population consisting of 233 lines was generated by two rounds of random crosses between DH lines. Each DH line was used once each round. The two populations and the parental lines were planted in the experimental field of Southwest University, Chongqing, China, in 2010 and 2011. A randomised complete block design with two replications was employed.
The association mapping panel used in the GWAS analysis was composed of 327 diverse inbred lines, including 71 winter ecotypes from Europe, 196 Chinese semi-winter accessions and 60 spring ecotype lines randomly selected from the ERANET-ASSYST B. napus diversity set. These lines were grown in the experimental field of Southwest University, Chongqing, China, in 2013 and 2014, and two replications were planted for each line in each year.
The branch number for ten representative B. napus plants in each line of the mapping populations were measured at maturity. Analysis of variance (ANOVA) was performed using the GLM procedure of SAS74.
QTL analysis
Development of molecular markers and construction of genetic linkage groups was described in a previous study73, where 293 markers were mapped to 19 linkage groups with a 1,188 cM map distance. Detection of QTLs and estimation of genetic parameters were performed with composite interval mapping (CIM) procedure of the software WinQTL Cartographer version 2.575. The QTLs were declared significant if the corresponding LR score was greater than 11.5 (equal to a 2.5 LOD score). The percent of phenotypic variance (PV) explained by a QTL (R2) was estimated at the highest probability peaks. A permutation test was performed 1,000 times at a significance level of p = 0.005 to minimize the experimental type-I error rate.
Genome-wide association analysis
The association mapping panel of 327 inbred lines was genotyped using the Brassica 60 K Illumina® Infinium SNP array by Emei Tongde Co. (Beijing) in accordance with the manufacturer’s protocol (http://www.illumina.com/technology/beadarray-technology/infinium-hd-assay.html). We excluded SNPs with either an AA or a BB frequency equal to zero, call frequency <0.9, or minor frequency <0.05.
Population structure was analyzed using the software package STRUCTURE v2.3.476. Five independent runs were performed with a K-value (the putative number of genetic groups) from 1 to 10, with 10,000 MCMC (Markov chain Monte Carlo) replications and 10,000 burn-ins. The optimal k-value was determined by the log probability of data [LnP(D)] and an ad hoc statistic Δk based on the rate of change of LnP(D) between successive k77. The software package SPAGeDi v1.4 was employed to calculate the relative kinship matrix comparing all pairs of the 327 accessions78. Negative values between two individuals were set to 079.
To avoid the false negative associations caused by relative kinship, the three mixed models controlling relative kinship, K model, controlling for K; Q + K model, controlling for both Q and K; and PCA + K model, controlling for both PC and K, were chosen to determine the statistical associations between phenotypes and genotypes to evaluate the effects of population structure (Q, PC) and relative kinship (K) on BN traits. These three models were performed with optimum compression and population parameters previously determined (P3D) by variance component estimation in TASSEL 5.079,80. Statistically significant loci were identified by comparing P values with the Bonferroni threshold (1/33186 = 3.01E-5)62.
Gene expression analysis
Thirty-one inbred lines of the association mapping panel in this study overlapped with the plant materials in a previous report by Bancroft et al.63. The RNA-seq data are available through an online resource. To determine whether the expression levels of genes in the target interval identified through QTL mapping and a GWAS analysis was associated with the BN observed in this study, the correlations between the gene expression levels and the SNP variations within or upstream of the corresponding genes were analyzed. The correlations between the BN phenotype variation and gene expression levels were also tested. Statistical analysis was completed using t test and correlation analysis.
Candidate gene predication
The QTL intervals were aligned to the B. napus reference genomes (http://www.genoscope.cns.fr/blat-server/cgi-bin/colza/webBlat) by BLAST the sequences of the SSR markers linked with BN QTLs. Based on the physical flanking marker positions of the QTLs, the corresponding genomic sequences of the QTL region were extracted. According to our previously study (Wei et al., data not published), the LD decay is 150 Kb in A subgenome and 750 kb in C subgenome. According to the decay of LD, the confident region of the GWAS-based QTL were determined. The QTL regions which identified by both linkage mapping and association mapping were selected and considered stably inherited. The corresponding genomic sequences of the region were extracted. The genes in the interval were used for a gene expression analysis. Finally, the correlations between the BN phenotype and the genes expression levels were examined to determine the candidate genes that control branch number.
References
Yaish, M. W. F., Guevara, D. R., El-Kereamy, A. & Rothstein, S. J. Axillary shoot branching in plants. In Plant developmental biology–biotechnological perspectives (ed. Pua E.C. & Davey M.R.) 37–52 (Springer, 2010).
Ledger, S. E., Janssen, B. J., Karunairetnam, S., Wang, T. & Snowden, K. C. Modified carotenoid cleavage dioxygenase8 expression correlates with altered branching in kiwifruit (Actinidia chinensis). New Phytol. 188, 803–813 (2010).
Cieslak, M., Seleznyova, A. N. & Hanan, J. A functional-structural kiwifruit vine model integrating architecture, carbon dynamics and effects of the environment. Ann. Bot. 107, 747–756 (2011).
Lauri, P. A. et al. Does knowledge on fruit tree architecture and its implications for orchard management improve horticultural sustainability? An overview of recent advances in the apple. Proc. 1st Is onHort In Europe (Ed. G. r. Dixon Acta Hort). 817, 243–249 (2009).
Upadyayula, N., Da Silva, H. S., Bohn, M. O. & Rocheford, T. R. Genetic and QTL analysis of maize tassel and ear inflorescence architecture. Theor. Appl. Genet. 112, 592–606 (2006).
Ward, S. P. & Leyser, O. Shoot branching. Curr. Opin. Plant Biol. 7, 73–78 (2004).
Shimizu-Sato, S. & Mori, H. Control of outgrowth and dormancy in axillary buds. Plant Physiol. 127, 1405–1413 (2001).
Hempel, F. D. & Feldman, L. J. Bi-directional inflorescence development in Arabidopsis thaliana: Acropetal initiation of flowers and basipetal iniation of paraclades. Planta. 192, 276–286 (1994).
Grbic, V. & Bleecker, A. B. Axillary meristem development in Arabidopsis thaliana. Plant J. 21, 215–223 (2000).
Stirnberg, P., Van de Sande, K. & Leyser, H. M. O. MAX1 and MAX2 control shoot lateral branching in Arabidopsis. Development. 129, 1131–1141 (2002).
Rameau, C. et al. Multiple pathways regulate shoot branching. Front. Plant Sci. https://doi.org/10.3389/fpls.2014.00741 (2015).
Casal, J. J., Sanchez, R. A. & Deregibus, V. A. The effect of plant density on tillering: The involvement of R/FR ratio and the proportion of radiation intercepted per plant. Environ. Exp. Bot. 26, 365–371 (1986).
Aguilar-Martínez, J. A., Poza-Carrión, C. & Cubas, P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell. 19, 458–472 (2007).
Arumingtyas, E. L., Floyd, R., Gregory, M. & Murfet, I. Branching in Pisum: inheritance and allelism tests with 17 ramosus mutants. Pisum Genet. 24, 17–31 (1992).
Napoli, C. A., Beveridge, C. A. & Snowden, K. C. Reevaluating concepts of apical dominance and the control of axillary bud outgrowth. Curr. Top. Dev. Biol. 44, 127–169 (1999).
Kebrom, T. H., Burson, B. L. & Finlayson, S. A. Phytochrome B represses teosinte branched1 expression and induces sorghum axillary bud outgrowth in response to light signals. Plant Physiol. 140, 1109–1117 (2006).
Snowden, K. C. & Napoli, C. A. A quantitative study of lateral branching in petunia. Funct. Plant Biol. 30, 987–994 (2003).
Cline, M. G. Exogenous auxin effects on lateral bud outgrowth in decapitated shoots. Ann. Bot. 78, 255–266 (1996).
Napoli, C. A. & Ruehle, J. New mutations affecting meristem growth and potential in Petunia hybrida Vilm. J. Hered. 87, 371–377 (1996).
Cline, M. G. Apical dominance. Bot. Rev. 57, 318–358 (1991).
Ongaro, V. & Leyser, O. Hormonal control of shoot branching. J. Exp. Bot. 59, 67–74 (2008).
Ferguson, B. J. & Beveridge, C. A. Roles for auxin, cytokinin, and strigolactone in regulating shoot branching. Plant Physiol. 149, 1929–1944 (2009).
Shimizu-Sato, S., Tanaka, M. & Mori, H. Auxin-cytokinin interactions in the control of shoot branching. Plant Mol Biol. 69, 429–435 (2009).
Groot, S. P. C., Keizer, L. C. P., Deruiter, W. & Dons, J. J. M. Seed and fruitset of the lateral suppressor mutant of tomato. Sci. Hortic-Amst. 59, 157–162 (1994).
Komatsu, K. et al. LAX and SPA: major regulators of shoot branching in rice. Proc. Natl. Acad. Sci. USA 100, 11765–11770 (2003).
Li, X. Y. et al. Control of tillering in rice. Nature. 422, 618–621 (2003).
Takeda, T. et al. The OsTB1 gene negatively regulates lateral branching in rice. Plant J. 33, 513–520 (2003).
Sorefan, K. et al. MAX4 and RMS1 are orthologous dioxygenase-like genes that regulate shoot branching in Arabidopsis and pea. Gene. Dev. 17, 1469–1474 (2003).
Doebley, J., Stec, A. & Hubbard, L. The evolution of apical dominance in maize. Nature. 386, 485–488 (1997).
Gallavotti, A. et al. The role of barren stalk1 in the architecture of maize. Nature. 432, 630–635 (2004).
Simons, J. L., Napoli, C. A., Janssen, B. J., Plummer, K. M. & Snowden, K. C. Analysis of the Decreased Apical Dominance genes of petunia in the control of axillary branching. Plant Physiol. 143, 697–706 (2007).
Dabbert, T., Okagaki, R. J., Cho, S., Boddu, J. & Muehlbauer, G. J. The genetics of barley low-tillering mutants: absent lower laterals (als). Theor. Appl. Genet. 118, 1351–1360 (2009).
Dabbert, T. et al. The genetics of barley low-tillering mutants: low number of tillers-1 (lnt1). Theor. Appl. Genet. 121, 705–715 (2010).
Otsuga, D., De Guzman, B., Prigge, M. J., Drews, G. N. & Clark, S. E. Revoluta regulates meristem initiation at lateral positions. Plant J. 25, 223–236 (2001).
Greb, T. et al. Molecular analysis of the LATERAL SUPPRESSOR gene in Arabidopsis reveals a conserved control mechanism for axillary meristem formation. Gene. Dev. 17, 1175–1187 (2003).
Muller, D., Schmitz, G. & Theres, K. Blind homologous R2R3 Myb genes control the pattern of lateral meristem initiation in. Arabidopsis. Plant Cell. 18, 586–597 (2006).
Nambeesan, S. U. et al. Association mapping in sunflower (Helianthus annuus L.) reveals independent control of apical vs. basal branching. BMC Plant Biology. 15, 84 (2015).
Zhao, J., Becker, H., Zhang, D., Zhang, Y. & Ecke, W. Conditional QTL mapping of oil content in rapeseed with respect toprotein content and traits related toplant development and grain yield. Theor. Appl. Genet. 113, 33–38 (2006).
Uzunova, M., Ecke, W., Weissleder, K. & Robbelen, G. Mapping the genome of rapeseed (Brassica napus L.). I. Construction of an RFLP linkagemap andlocalization of QTLs for seedglucosinolate content. Theor. Appl. Genet. 90, 194–204 (1995).
Zhao, J., Dimov, Z., Becker, H., Ecke, W. & Mollers, C. Mapping QTL controlling fatty acid composition in a doubled haploid rapeseed population segregating for oil content. Mol. Breed. 21, 115–125 (2008).
Long, Y. et al. Flowering time quantitative trait loci analysis of oilseed Brassica in multiple environments and genomewide alignment with Arabidopsis. Genetics. 177, 2433–2444 (2007).
Shi, J. et al. Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics. 182, 851–861 (2009).
Radoev, M., Becker, H. & Ecke, W. Genetic analysis of heterosis for yield and yield components in rapeseed (Brassica napus L.) by quantitative trait locus mapping. Genetics. 179, 1547–1558 (2008).
Yang, P. et al. Identification of a major QTL for silique length and seed weight in oilseed rape (Brassica napus L.). Theor. Appl. Genet. 125, 285–296 (2012).
Zhang, L., Li, S., Chen, L. & Yang, G. Identification and mapping of a major dominant quantitative trait locus controlling seeds per silique as a single Mendelian factor in Brassica napus L. Theor. Appl. Genet. 125, 695–705 (2012).
Holland, J. B. Genetic architecture of complex traits in plants. Curr. Opin. Plant Biol. 10, 156–161 (2007).
Zhao, K. et al. Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat. Commun. https://doi.org/10.1038/ncomms1467 (2011).
Tian, F. et al. Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat. Genet. 43, 159–162 (2011).
Huang, X. et al. Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat. Genet. 44, 32–39 (2012).
Li, H. et al. Genome-wide association study dissects the genetic architecture of oil biosynthesis inmaize kernels. Nat. Genet. 45, 43–50 (2013).
Atwell, S. et al. Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature. 465, 627–631 (2010).
Nordborg, M. & Weigel, D. Next-generation genetics in plants. Nature. 456, 720–723 (2008).
Nordborg, M. & Tavare, S. Linkage disequilibrium: what history has to tell us. Trends Genet. 18, 83–90 (2002).
Lander, E. & Kruglyak, L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat. Genet. 11, 241–247 (1995).
Cardon, L. R. & Palmer, J. L. Population stratification and spurious allelic association. Lancet. 361, 598–604 (2003).
Hu, G. et al. Fine mapping of the awn gene on chromosome 4 in rice by association and linkage analyses. Sci. Bull. 56, 835–839 (2011).
Li, Y. et al. QTL analysis of yield-related traits and their association with functional markers in Brassica napus L. Aust. J. Agr. Res. 58, 759–766 (2007).
Zhao, Z. K., Wu, L. K., & Nian, F. Z. Dissecting quantitative trait loci for boron efficiency across multiple environment in Brassica napus. Plos One. https://doi.org/10.1371/journal.pone.0045215 (2012).
Shi, J. Q. et al. Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics. 182, 851–861 (2009).
Ding, G. D. et al. Quantitative trait loci for seed yield and yield-related traits, and their responses to reduced phosphorus supply in Brassica napus. Ann. Bot. 109, 747–759 (2012).
Chen, W. et al. Detection of QTL for six yield-related traits in oilseed rape (Brassica napus) using DH and immortalized F2 populations. Theor. Appl. Genet. 115, 849–858 (2007).
Luo, X. et al. Unravelling the complex trait of harvest index in rapeseed (Brassica napus L.) with association mapping. BMC Genomics. 16, 379 (2015).
Bancroft, I. et al. Dissecting the genome of the polyploid crop oilseed rape by transcriptome sequencing. Nat. biotechnol. 29, 762–768 (2011).
Huang, X. et al. Genome-wide association studies of 14 agronomic traits in rice landraces. Nat. Genet. 42, 961–967 (2010).
Morris, G. P. et al. Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc. Natl. Acad. Sci. USA 110, 453–458 (2013).
Houvras, Y. et al. BRCA1 physically and functionally interacts with ATF1. J. Biol. Chem. 275, 36230–36237 (2000).
Park, J., Nguyen, K. T., Park, E., Jeon, J. & Choi, G. DELLA proteins and their iInteracting RING finger proteins repress gibberellin responses by binding to the promoters of a subset of gibberellin-responsive genes in. Arabidopsis. The Plant Cell. 25, 927–943 (2013).
Davies, J. P. The plant hormones: their nature, occurrence, and functions. Ithaca, NY: Springer (2007).
Silverstone, A. L., Mak, P. Y., Martinez, E. C. & Sun, T. P. The new RGA locus encodes a negative regulator of gibberellin response in Arabidopsis thaliana. Genetics. 146, 1087–1099 (1997).
Lo, S. F. et al. A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell. 20, 2603–2618 (2008).
Murfet, I. C. & Reid, J. B. Developmental mutants. In Peas: Genetics, Molecular Biology and Biotechnology (eds Casey R. & Davies D. R.) 165–216 (CAB, 1993).
Agharkar, M. et al. Stable expression of AtGA2ox1 in a low-input turfgrass (Paspalum notatum Flugge) reduces bioactive gibberellin levels and improves turf quality under field conditions. Plant Biotechnol. J. 5, 791–801 (2007).
Fu, Y. et al. Development of genic cleavage markers in association with seed glucosinolate content in canola. Theor. Appl. Genet. 128, 1029–1037 (2015).
SAS Institute, SAS/STAT user’s guide, version 8. SAS Institute, Cary (2000).
Wang, S., Basten, C. J., Basten, C. J. & Zeng, Z. B. Windows QTL cartographer version 2.5. Statistical genetics, North Carolina State University, Raleigh (2005).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics. 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol. 14, 2611–2620 (2005).
Hardy, O. & Vekemans, X. SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol. Ecol. Notes. 2, 618–620 (2002).
Yu, J. et al. A unified mixedmodel method for association mapping that accounts for multiple levels of relatedness. Nat Genet. 38, 203–208 (2006).
Yang, X., Yan, J. & Shah, T. Genetic analysis and characterization of a new maize association mapping panel for quantitative trait loci dissection. Theor Appl Genet. 121, 417–431 (2010).
Acknowledgements
This program was financially supported in part by the National Key Research and Development Program of China (2016YFD0100202), the National Basic Research Program of China (973 Program, 2015CB150201), National Natural Science Foundation of China (31671729), Chongqing Research Program of Basic Research and Frontier Technology (cstc2017jcyjAX0391), and the Fundamental Research Funds for the Central Universities (XDJK2016B031).
Author information
Authors and Affiliations
Contributions
W.Q. designed research, Y.H., D.W., D.W., Y.F., Y.C., H.D. and C.T. performed research, Y.H. and D.W. analyzed data and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, Y., Wu, D., Wei, D. et al. GWAS, QTL mapping and gene expression analyses in Brassica napus reveal genetic control of branching morphogenesis. Sci Rep 7, 15971 (2017). https://doi.org/10.1038/s41598-017-15976-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15976-4
This article is cited by
-
Identification of environment-insensitive genes for oil content by combination of transcriptome and genome-wide association analysis in rapeseed
Biotechnology for Biofuels and Bioproducts (2024)
-
Genotypic variation of tocopherol content in a representative genetic population and genome-wide association study on tocopherol in rapeseed (Brassica napus)
Molecular Breeding (2023)
-
Linkage and association mapping of ovule number per ovary (ON) in oilseed rape (Brassica napus L.)
Molecular Breeding (2023)
-
The current scenario and future perspectives of transgenic oilseed mustard by CRISPR-Cas9
Molecular Biology Reports (2023)
-
Multi-omics analysis dissects the genetic architecture of seed coat content in Brassica napus
Genome Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.