Abstract
Synthesis of homogenous glycans in quantitative yields represents a major bottleneck to the production of molecular tools for glycoscience, such as glycan microarrays, affinity resins, and reference standards. Here, we describe a combined biological/enzymatic synthesis that is capable of efficiently converting microbially-derived precursor oligosaccharides into structurally uniform human-type N-glycans. Unlike starting material obtained by chemical synthesis or direct isolation from natural sources, which can be time consuming and costly to generate, our approach involves precursors derived from renewable sources including wild-type Saccharomyces cerevisiae glycoproteins and lipid-linked oligosaccharides from glycoengineered Escherichia coli. Following deglycosylation of these biosynthetic precursors, the resulting microbial oligosaccharides are subjected to a greatly simplified purification scheme followed by structural remodeling using commercially available and recombinantly produced glycosyltransferases including key N-acetylglucosaminyltransferases (e.g., GnTI, GnTII, and GnTIV) involved in early remodeling of glycans in the mammalian glycosylation pathway. Using this approach, preparative quantities of hybrid and complex-type N-glycans including asymmetric multi-antennary structures were generated and subsequently used to develop a glycan microarray for high-throughput, fluorescence-based screening of glycan-binding proteins. Taken together, these results confirm our combined synthesis strategy as a new, user-friendly route for supplying chemically defined human glycans simply by combining biosynthetically-derived precursors with enzymatic remodeling.
Similar content being viewed by others
Introduction
Carbohydrate chains, known as glycans, represent a vast source of biological diversity across all domains of life. They play a key role in almost every aspect of normal physiology, as well as in the etiology of nearly every major disease1. Approximately 1% of the human genome is dedicated to the biosynthesis and diversification of glycans, and the majority of human proteins are thought to be post-translationally modified by glycans, a process known as glycosylation2,3. Glycosylation can occur at several amino acid residues, most commonly asparagines (N-linked) and serines or threonines (O-linked). As conjugates to proteins, glycans add an additional information layer and have direct biological effects, ranging from stabilizing protein folds4,5,6 to signaling stem cell fate7,8,9. Glycans also feature prominently in disease. For example, in the context of cancer, tumor cells commonly express glycans at atypical levels or with altered structural attributes10,11,12 and there is growing interest in developing pharmaceutical agents that target these molecules13. However, while the functional importance of glycosylation is well established, methods for mass-producing glycans and glycoconjugates are lagging14,15. Such technological gaps are to be expected in light of the immense structural diversity and information density of glycans in nature16, which far exceeds that of DNA or proteins. Unfortunately, this complexity, coupled with a scarcity of materials, has hindered production of glycomolecules which in turn has limited the availability of chemically defined oligosaccharides for use as standards in glycan structure determination17,18, as probes to characterize glycan-binding proteins19,20, and as ligands in affinity resins21.
To furnish oligosaccharides for biological and structural studies, a variety of methods have been described that enable laboratory preparation of these important compounds. One such method involves the enzymatic or chemical deglycosylation of glycoproteins from natural sources (e.g., hen egg yolk) followed by a series of purification steps22,23. While natural sources are a rich supplier of glycans, the chromatography, desalting, and concentration steps used to purify released glycans are time consuming, expensive, and typically yield only small quantities of closely related structures that are difficult to fractionate. For these reasons, chemical or chemo-enzymatic production of synthetic carbohydrates has emerged as a preferred route for preparing libraries of pure oligosaccharides24,25,26,27,28,29,30. In the case of chemical synthesis, target oligosaccharides have been generated by either solution-phase or solid-phase methods. Solution-phase synthesis utilizes a one-pot strategy, wherein differences in the reactivity of glycosyl donors allow for ligation in the proper oligosaccharide sequence. In solid-phase synthesis, the reducing-end sugar is connected to a solid support and each additional sugar residue is added sequentially, after which the glycan is cleaved from the resin. While recent advances to these methods have greatly improved the number and diversity of glycans that can be produced chemically, including the addition of non-natural residues, this approach remains limited by the cost and scale of production31,32 and remains very time consuming, especially when highly complex structures are targeted24. Many of these issues can be circumvented by chemo-enzymatic methods, whereby a synthetic oligosaccharide precursor is modified by a range of glycosyltransferases (GTs) to supply more complex derivatives26,27,33. An illustrative example is the gram-scale production of 24 different mammalian blood group and tumor associated antigens based on poly-N-acetyllactosamine using six different GT enzymes and four nucleotide donors to elongate GlcNAc that was chemically synthesized with an azidoethyl spacer34. Chemo-enzymatic synthesis is not only more economical, but it enables efficient generation of extremely complex glycan structures, including asymmetrically branched N-glycans that have proven inaccessible to other existing synthesis technologies30.
These methods notwithstanding, new glycan synthesis technologies for glycobiology and glycomedicine that are low-cost, user-friendly, robust, and adaptable are in high demand. Along these lines, here we describe a combined biological/enzymatic route that enables efficient, low-cost production of structurally homogeneous human-type N-glycans. One of the main difficulties to chemo-enzymatic or enzymatic synthesis of glycan libraries is that generation of the starting material, typically by either chemical synthesis or direct isolation from natural sources, can be time consuming, costly, and difficult to scale. To bridge this technology gap, our synthesis strategy leverages precursor oligosaccharides that are derived in large quantities from two complementary microbial sources, either invertase glycoprotein from wild-type Saccharomyces cerevisiae or lipid-linked oligosaccharides (LLOs) from glycoengineered Escherichia coli 35. Following deglycosylation of these biosynthetic precursors, the microbially-derived oligosaccharides are subjected to a greatly simplified purification scheme followed by structural remodeling using GTs, including several from the mammalian glycosylation pathway that are not commercially available. The power of this methodology was demonstrated by supplying preparative quantities of 20 different eukaryotic N-glycans including asymmetric multi-antennary structures, several of which have not previously been synthesized by chemical or chemoenzymatic routes.
Results
Preparation of human-type oligomannose glycan precursors
To generate a renewable supply of precursor oligosaccharides (see Supplementary Table 1 for complete list with chemical structure), we investigated two complementary sources of N-glycans: hypermannoyslated invertase produced by recombinant expression in yeast; and undecaprenol pyrophosphate (Und-PP)-linked paucimannose glycans (e.g., mannose3-N-acetylglucosamine2, a.k.a. Man3GlcNAc2) produced by glycoengineered E. coli 35. Our initial focus was on invertase (Fig. 1a), which was chosen as a vehicle to isolate oligosaccharides because it has the highest glycan abundance of all known glycoproteins, and is produced at high yields when expressed recombinantly in wild-type yeast strains. Invertase carries high-mannose glycans comprised of eight or more mannose units23,36; however, at the core of these glycans is a common intermediate structure, Man5GlcNAc2, which is also present in mammalian N-glycans. Previous work has engineered yeast to eliminate hypermannosylation, producing an oligosaccharide that is sensitive to mammalian glycosylation enzymes37,38,39; however, to our knowledge, there has been no investigation of whether wild-type, hypermannosylated glycans can be converted to a glycoform that is sensitive to mammalian glycosylation enzymes. Thus, we investigated whether remodeling invertase glycans was a viable method for providing the starting material needed for production of a mammalian N-linked glycan library.
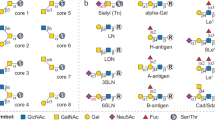
Glycan synthesis strategies. Precursor glycans (1–3) were derived from (a) yeast invertase or (b) lipid-linked oligosaccharides (LLOs) from glycoengineered E. coli cells carrying plasmid pYCG, and subsequently used to synthesize glycans 4–21. Enzymatic steps: (i) PNGase F (product shown is representative of high mannose yeast glycans); (ii) α1,2- and α1,6-mannosidase; (iii) jack bean α-mannosidase and α1,6-mannosidase; (iv) GnTI; (v) β1,4-galactosyltransferase; (vi) GnTIV; (vii) non-enzymatic hydrolysis of extracted LLOs; (viii) GnTII; (ix) GnTV; and (x) GnTIII.
The glycan profile of the oligomannose glycans released from Saccharomyces cerevisiae invertase by PNGase F treatment revealed a mixture of Man8GlcNAc2 to Man14GlcNAc2 oligomannose glycans (Supplementary Fig. 1a). To determine whether it was possible to trim these high-mannose glycans to the human-type Man5GlcNAc2, we subjected released glycan 1 to a series of mannosidases and determined that the additional mannose residues consisted of α(1,2)- and α(1,6)-mannose linkages. By addition of Xanthomonas manihotis α(1,6)-mannosidase and Aspergillus saitoi α(1,2)-mannosidase, hypermannosylated yeast glycans were trimmed to the human-type Man5GlcNAc2 glycoform 4 whereas treatment with Canavalia ensiformis (jack bean) α-mannosidase and the same α(1,6)-mannosidase resulted in glycoform 5. As discussed above, one of the drawbacks to isolation of glycans from natural sources is the difficulty in purification. Therefore, a point of emphasis was to develop a simplified purification method. After testing multiple separation techniques, we determined that an initial silica column followed by a charcoal/celite column with a single elution step, and a graphitized carbon column with a gradient elution was the simplest method. This purification method resulted in a product (glycan 4) having high purity as assessed by mass spectrometry (MS) and nuclear magnetic resonance (NMR) spectroscopy (Fig. 2a), as well as HPLC (Supplementary Fig. 2a). Isolation, mannose trimming, and purification of the oligosaccharides from 1 g of invertase resulted in a yield of 0.7 mg of 4 as determined by measuring dry weight and fluorophore-assisted carbohydrate electrophoresis (FACE) analysis40,41 (Supplementary Fig. 1b). A related procedure yielded glycan 5 in similar quantities (Supplementary Fig. 3a). Thus, we confirm that biosynthesis of S. cerevisiae invertase glycans provides a viable reservoir of human-type oligosaccharide precursors.
Biosynthesis of precursor oligosaccharides for enzymatic remodeling. MALDI-TOF MS analysis (top panels) and 600-MHz 1H NMR characterization (bottom panels) corresponding to: (a) Man5GlcNAc2 glycan synthesized by enzymatic deglycosylation of S. cerevisiae oligosaccharides; and (b) the Man3GlcNAc2 glycan synthesized by glycoengineered E. coli cells.
As an alternative biosynthetic route to precursor glycans, we investigated the use of glycoengineered E. coli as producers of the starting material for glycan diversification (Fig. 1b). Unlike all natural or engineered eukaryotic hosts, E. coli cells do not have native protein glycosylation pathways nor is glycosylation an essential mechanism in this host. As a result, E. coli provides an unformatted “operating system” that can be genetically reprogrammed to produce discrete, uniform glycans that are not attached to undesired targets and do not cause problems with cell physiology or morphology. Using natural pools of nucleotide sugars (e.g., UDP-GlcNAc, GDP-Man) as substrates, E. coli cells carrying a synthetic pathway of heterologous GTs, namely the β1,4-GlcNAc transferase Alg13/14, the β1,4-mannosyltransferase Alg1, and the bifunctional mannosyltransferase Alg2, are capable of synthesizing Und-PP-linked Man3GlcNAc2 as we showed previously35. This latter product was readily isolated by lipid extraction from E. coli followed by non-enzymatic removal of 2 from undecaprenol. Following this procedure, we reproducibly obtained ~750 μg of Man3GlcNAc2 glycan that was well resolved from contaminant peaks (Supplementary Fig. 2b) and had no detectable structural variability from batch-to-batch (Fig. 2b and Supplementary Fig. 4). It should be noted that addition of yeast mannosyltransferase Alg11 to the synthetic glycosylation pathway resulted in formation of Und-PP-linked 3 that could be similarly processed to generate nearly pure Man5GlcNAc2 (Supplementary Fig. 3b). Isolation of lipid-linked glycans from E. coli in this manner eliminates several difficulties associated with purification and heterogeneity. Moreover, the short time requirements and low costs associated with bacterial culture are advantageous for the synthesis of human-type N-glycan precursors.
Expression and purification of eukaryotic glycosyltransferases
A number of the glycosylation enzymes that are natively involved in the early stages of remodeling oligomannose glycan 4 are not commercially available, and methods to produce these enzymes have not been reported. Therefore, we undertook the development of methods for recombinant expression and purification of three N-acetylglucosaminyltransferases (GnTs), namely GnTI, GnTII, and GnTIV. We chose E. coli due to its proven track record as a protein expression host. To promote efficient folding in the cytoplasm, the genes encoding Nicotiana tabacum GnTI, Homo sapiens GnTII, and Bos taurus GnTIV were all fused to the C-terminus of the gene encoding E. coli maltose binding protein (MBP) lacking its native export signal. Both GnTI and GnTIV were solubly expressed over a wide range of temperatures (15–37 °C) and in a number of different E. coli strain backgrounds (e.g., MC4100, SHuffle T7 Express, and Origami2(DE3)). In contrast, GnTII was only expressed solubly at temperatures at or below 25 °C and required a host strain lacking both the thioredoxin reductase (trxB) and glutathione reductase (gor) genes (e.g., Origami2(DE3)), which greatly enhances disulfide bond formation in the E. coli cytoplasm42. All three enzymes were purified by amylose affinity chromatography (Supplementary Fig. 1c) and retained as fusions to MBP for enzymatic remodeling of yeast-derived glycan 4 and bacteria-derived glycan 2 as described below.
Generation of hybrid glycans from yeast-derived precursor
Hybrid oligosaccharide structures are inherently difficult to isolate from natural sources because they are transient glycans in the mammalian glycosylation pathway. Therefore, we attempted to produce four of these rare oligosaccharides using the yeast-derived Man5GlcNAc2 (glycan 4) as the starting material. First, MBP-GnTI was used to generate a hybrid GlcNAcMan5GlcNAc2 glycan 6 by conjugating a β1,2-linked GlcNAc residue to the α1,3 arm of 4. Production of this hybrid glycan was confirmed by MS and NMR analysis (Fig. 3a), as well as HPLC (Supplementary Fig. 2c). Importantly, the chemical shifts of 6 synthesized herein matched previous NMR characterization of this oligosaccharide structure (Supplementary Table 2)43. One advantage of enzymatic glycan synthesis is the ability to conduct multiple reactions in a single step. Synthesis of 7 is a good example of this, where both MBP-GnTI and MBP-GnTIV were added simultaneously to produce a glycan with β1,4-GlcNAc branching on the α1,3 mannose (Fig. 3b). Conducting multiple reactions in a single step eliminates purification of each intermediate, which in turn reduces loss of product associated with purification. As an example of further elaboration of these oligosaccharides, each glycan was treated with B. taurus β1,4-galactosyltransferase (GalT) to produce glycans 8 and 9. Addition of galactose residues was confirmed by MS, and by the appearance of a peak that corresponds to galactose in each NMR spectrum (Fig. 3c and d). In the case of glycan 9, the reaction must be conducted in a two-step process to eliminate the possibility of producing 8 as a side product. Importantly, these enzymatic conversions were all near 100% and final recoveries were ~75% for each with losses due to: (1) HPLC fractions that were deemed not clean enough thus excluded; and (2) desalting steps where fractional loss was expected. Taken together, these results confirm our combined biological/enzymatic synthesis strategy as a new route to human-type, hybrid oligosaccharides.
Generation of hybrid and complex glycans from bacteria-derived precursor
We next evaluated whether the E. coli-derived Man3GlcNAc2 glycan 2 could be used as the starting material for preparative-scale synthesis of hybrid- and complex-type glycans. An advantage of this precursor glycan is that it eliminates two mannosidase-catalyzed deglycosylation steps required to produce the hybrid GlcNAcMan3GlcNAc2 glycan 11. Indeed, following treatment with MBP-GnTI, efficient transfer of a GlcNAc residue directly to the α1,3 arm of 2 was observed (Fig. 4a and Supplementary Figs 2d and 5), despite the fact that this precursor glycan is not the native substrate for this enzyme. The chemical shifts of glycan 11 were in agreement with previous NMR spectroscopy of this oligosaccharide structure (Supplementary Table 2)22. Along similar lines, we synthesized the human-type complex GlcNAc2Man3GlcNAc2 glycan 14 using a one-pot strategy whereby both MBP-GnTI and MBP-GnTII were added simultaneously to 2 (Fig. 4b). This glycan was also characterized by NMR and matched previous characterization of this oligosaccharide structure (Supplementary Table 2)22. It is worth mentioning that the resulting complex glycan is a key intermediate in the mammalian N-glycosylation pathway, and in our glycan library synthesis efforts, because it acts as a substrate for all subsequent additions. For example, further GlcNAc addition was possible by treatment of 11 or 14 with MBP-GnTIV to produce glycans 12 and 17, respectively, with β1,4-GlcNAc branching (Fig. 4c and d, Supplementary Fig. 6 and Supplementary Table 2). Interestingly, when carrying out the synthesis of glycan 17, the reaction rate of GnTII was greatly decreased when acting on glycan 12 in the one-pot reaction. Therefore, glycan 17 was synthesized in a stepwise fashion whereby a first reaction with MBP-GnTI and MBP-GnTII was followed by a second reaction with MBP-GnTIV. Glycans with β1,6-GlcNAc branching were achieved by step-wise treatment of 14 or 17 with H. sapiens GnTV to yield 13 and 20, respectively (Fig. 4e and f, and Supplementary Table 2). Another interesting point is that two pairs of the GlcNAc-terminal glycans 12 and 14, 13 and 17 are structural isomers, which would be useful as standards for glycomics research and as affinity probes for determining lectin substrate specificity. As we observed with yeast precursor above, these enzymatic conversions were all near 100% and final recoveries were ~75% for each. Collectively, these results demonstrate that treatment of the bacteria-derived precursor glycan with different combinations of GnTI, GnTII, GnTIV, and GnTV is an efficient route to multiantennary, GlcNAc-terminal oligosaccharides.
Generation of hybrid and complex-type glycans using the bacteria-derived precursor. MALDI-TOF MS analysis of the following products: (a) glycan 11; (b) glycan 14; (c) glycan 12; (d) glycan 17; (e) glycan 13; (f) glycan 20; (g) glycan 18; (h) glycan 21; (i) glycan 10; (j) glycan 19; (k) glycan 15; (l) glycan 16.
Additional modification of select oligosaccharides described above was conducted to demonstrate the rapid increase in the number of glycan structures that can be produced by our combined biological/enzymatic method. For example, H. sapiens GnTIII adds a bisecting GlcNAc to the β1,4-mannose and is thought to be a natural regulator of branching, since attachment of this residue eliminates further elaboration by GnTs. To generate such bisecting glycan structures, glycans 14 and 17 were synthesized in a first reaction and then, for reasons described above, treated in a subsequent reaction with GnTIII to yield 18 and 21, respectively. Analysis of the reaction products revealed that GnTIII installed branching GlcNAc residues at a high conversion rate (Fig. 4g and h, and Supplementary Table 2). The resulting bisecting GlcNAc structures, which supply additional structural isomers to our panel, would be of particular interest for determining differences in lectin binding specificity. Finally, galactose residues were successfully added to 11, 12, 14, and 17 by treatment with B. taurus β1,4-GalT, yielding glycans 10, 15, 19, and 16, respectively (Fig. 4i–l, Supplementary Fig. 7 and Supplementary Table 2). Again, these enzymatic conversions were all near 100% with final recoveries of ~75% for each. While these latter reactions were only conducted on select glycan structures with two different GTs, one could easily apply these GTs to each glycan produced herein, resulting in an exponential increase in the number of oligosaccharides in the glycan library.
Construction and evaluation of an N-glycan microarray
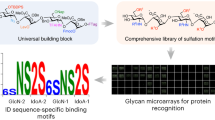
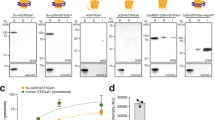
To demonstrate the functionality of the glycan library, we developed microarrays using a novel bifunctional fluorescent linker, 2-amino-N-(2-aminoethyl)-benzamide (AEAB)44. Specifically, glycans 2–21 were directly conjugated to AEAB through its arylamine group by reductive amination to form glycan-AEABs (GAEABs) and then purified by multidimensional HPLC. Following purification, GAEABs were covalently immobilized onto N-hydroxysuccinimide (NHS)-activated glass slides via their free alkylamine alongside the similarly immobilized reference compounds LNnT, LNT, and NA2 (Fig. 5a), which were prepared from natural glycans as described previously44. When probed with C. ensiformis Concanavalin A (ConA), binding of the biantennary N-glycan reference compound, NA2, was detected but not reference compounds LNT or LNnT, consistent with the specificity of ConA for primarily internal and non-reducing terminal α-mannose (Fig. 5b). Among the synthesized glycans, ConA bound most strongly to the oligomannose and hybrid glycans (2–12). It also bound complex glycans 13, 14, 17, 18, 20, and 21, albeit to a lesser extent, which was consistent with the established higher affinity of ConA for terminal versus internal α-linked mannose. In agreement with previous array data (http://www.functionalglycomics.org/glycomics/HServlet?operation=view&sideMenu=no&psId=primscreen_PA_v2_357_10312005), ConA bound efficiently to the complex glycan 19, which is structurally identical to reference compound NA2. In contrast, ConA showed virtually no binding to its hybrid isomer, glycan 15. Taken together, these results demonstrate the compatibility of our synthesized glycans with microarray development (i.e., chemical derivatization and immobilization steps) and fluorescence-based screening of glycan-binding proteins.
Binding of lectin ConA to microarray of synthesized human N-glycans. (a) Structures of reference compounds that were derivatized with AEAB and immobilized onto NHS-activated glass slides, alongside glycans 2–21. (b) Probing of immobilized glycans with biotinylated ConA (10 μg/ml). The amount of bound ConA was determined by streptavidin-Cy5 (5 μg/ml) fluorescence. Background subtracted mean fluorescence values are shown. Error bars represent the standard deviation of the mean. Representative structures for glycans 3, 12, 15 and 19 are shown. All glycans were printed at 100 μM in PBS in replicates of four. PBS and streptavidin-Cy5 spots served as controls. Biotin-LC-hydrazide spots printed on each subarray serve as an alignment feature to localize each subarray on the slide.
Discussion
As many have argued15, the development of simple and cost-effective methods to produce diverse glycomolecules is needed to meet the demand of, and grow, the glycoscience community. In this work, we addressed this need with the demonstration of an efficient new methodology that enables production of large quantities of hybrid- and complex-type human N-glycans. The novelty of this approach stems from first harnessing the power of microbial biosynthesis to furnish a renewable reservoir of starting material. Specifically, up to milligram amounts of precursor oligosaccharides were derived from either wild-type yeast that recombinantly express hypermannosylated invertase or glycoengineered E. coli that produce lipid-linked paucimannose glycans. In parallel, we developed simplified methods to purify the resulting oligosaccharides, as well as robust expression and purification methods for key glycan remodeling enzymes (e.g., eukaryotic GnTs) that are not commercially available. Armed with these and other commercially available GTs, we efficiently transformed precursor oligosaccharides into a library of hybrid and complex N-glycans including asymmetric multi-antennary structures. This latter point is particularly important because despite the urgent need for libraries of asymmetrically branched N-glycans, currently available methods are not capable of delivering such compounds, and previous synthetic efforts have focused predominantly on producing symmetrically branched compounds. Hence, a user-friendly synthetic strategy such as ours that can supply libraries of asymmetrically substituted glycans is not only novel but makes possible the creation of new probes for interrogating glycoconjugate synthesis pathways, new standards for the assignment of MS spectra, and new glycan microarrays for examining glycan-protein recognition.
Based on the yields obtained here, we expect that it should be possible to produce multiple milligrams of precursor glycan per week that can be easily and efficiently remodeled by GnTI-V treatment to furnish hybrid, tri-, and tetra-antennary structures much more easily than via chemical means reported in other glycan library synthesis efforts28,30. It is worth mentioning that while the purity of our synthesized glycans was consistently high, small impurities were observed in the MS and/or NMR spectra for many of the glycans. However, we believe these could be readily eliminated with more rigorous purification procedures. Beyond the structures reported here, one could also imagine readily generating fucosylated and sialylated glycoforms simply by the addition of commercial or recombinantly produced fucosyltransferases and sialyltransferases, respectively45,46. For example, by treating glycan 11 with the commercial human FUT8 fucosyltransferase we were able to generate a structurally uniform fucosylated glycan (Supplementary Fig. 3c). We anticipate that this technology will help to overcome many of the critical barriers to glycan production by offering a user-friendly synthesis methodology that can reliably deliver homogeneous glycans from a consistent source and flexibly adapt to production needs. Moreover, the libraries of glycans that can be supplied by our combined biological/enzymatic methodology presented here represent useful tools for understanding the biological role of glycans in health and disease as well as for defining and exploiting the human glycome.
Experimental Procedures
Strains and plasmids
E. coli strain MC4100 (F− araD139 Δ(argF-lac)U169 flbB5301 deoC1 ptsF25 relA1 rbsR22 rpsL150 thiA), SHuffle T7 Express (New England Biolabs), and Origami2(DE3) (Novagen) were used for heterologous expression of different eukaryotic GnTs. Origami2(DE3) gmd::kan ΔwaaL ΔnanA cells were used for producing lipid-linked Man3GlcNAc2 precursor. This strain was created by introducing sequential mutations using P1vir phage transduction47 with the respective strains from the Keio collection48 as donors. Briefly, to delete the nanA gene a lysate was generated from donor cells (JW31194-1) containing the ΔnanA735::kan mutation. Donor strains were obtained from the Coli Genetic Stock Center (CGSC). The resulting phage was used to infect Origami2(DE3) target cells and successful transductants were selected on LB plates supplemented with kanamycin (Kan). The Kan resistance cassette was subsequently removed by transforming the resulting strain with plasmid pCP20 as described49. This resulted in generation of strain Origami2(DE3) ΔnanA. Subsequent deletion of the waaL (rfaL) and gmd genes was performed in the same manner using donor strains JW3597-1 (ΔrfaL734::kan) and JW2038-1 (Δgmd751::kan). Yeast strain FY834 was used for homologous recombination using plasmid pMQ70 as described previously50. YPD growth medium was used to maintain yeast and synthetic defined uracil dropout medium was used to select and maintain yeast with plasmids. Plasmid pYCG was used for production of lipid-linked Man3GlcNAc2 and has been described previously35. A second plasmid containing the E. coli manB and manC genes for over-production of GDP-mannose was produced using homologous recombination in S. cerevisiae 50. Briefly, genes encoding phosphomannomutase (ManB) and mannose-1-phosphate guanylyltransferase (ManC) were PCR-amplified from E. coli genomic DNA using primer pairs that contained the priming region for the manB and manC genes, respectively, along with 40-bp overhangs of pMQ70. The resulting PCR products and the linearized pMQ70 plasmid were used to transform yeast strain FY834, yielding plasmid pManCB. Constructs assembled in yeast were electroporated into E. coli MC4100 for verification via PCR, restriction enzyme digestion, and/or sequencing. For expression of eukaryotic GnTs, N. tabacum GnTI, H. sapiens GnTII, B. taurus GnTIV were all cloned for expression as C-terminal fusions to E. coli MBP lacking its native export signal peptide. Briefly, genes encoding MBP and the respective GnT enzyme were amplified with primers containing regions of overlap to the fusion partner and pMQ70 plasmid. The PCR products were assembled with linearized pMQ70 by homologous recombination as described above, yielding plasmids pMBP-GnTI, pMBP-GnTII, and pMBP-GnTIV.
Synthesis of Man5GlcNAc2 precursor
5 g of S. cerevisiae invertase (Sigma-Aldrich) was denatured in the presence of sodium dodecyl sulfate and NP-40 as described in the New England Biolabs protocol for PNGase F treatment. The solution was allowed to cool to room temperature and 50,000 U of PNGase F (New England Biolabs) was added to the solution. The PNGase F reaction was incubated at 30 °C for 2 days to ensure complete removal of oligosaccharide from the protein. The glycan was precipitated using 80% (v/v) acetone, and resuspended in 60% methanol (v/v), where the remaining pellet was discarded. The glycan/protein mixture was then applied to a water-equilibrated charcoal/celite (1:1) column and eluted with 50% ethanol (v/v). The glycan was then dried by rotary evaporator, resuspended in a minimal amount of ethyl acetate:methanol:water (11:3:3), and applied to a 11:3:3 pre-equilibrated silica flash column (EMD Millipore Grade 62 60–200 Mesh). The column was washed with a ratio of 7:3:3 and eluted with a ratio of 3:3:3. The glycan was then resuspended in water and desalted by mixing with Ag 1-X8 and DOWEX 50WX8-400 resins. Both the characterization and purity were determined by MALDI-TOF MS, FACE, and 600-MHz 1D 1H NMR spectroscopy as described elsewhere22,35,40,41. The resulting high-mannose precursor oligosaccharides were specifically trimmed with both X. manihotis α1-6-mannosidase (ProZyme) and A. saitoi α1-2-mannosidase (New England Biolabs) to produce the human-type Man5GlcNAc2 glycan, since we empirically determined that there were no terminal α(1,3)-mannose residues. Alternatively, to generate the Man1GlcNAc2 glycan, which could serve as a reference standard or a substrate for GT- or glycosidase-based assays, C. ensiformis α-mannosidase (Sigma) and X. manihotis α(1,6)-mannosidase were used to trim invertase-derived glycans. To achieve this conversion, roughly 5 mg of purified precursor glycan was resuspended in mannosidase buffer (0.1 M sodium acetate, pH 5.0, ProZyme), to which 8 mU of α1-2-mannosidase and 80 U of α1-6-mannosidase were added. The reaction was allowed to proceed at 37 °C and monitored by MS until the reaction was complete. The glycan was then purified by a graphitized carbon column (Hypercarb, Thermo) using a 0–50% acetonitrile:water gradient. Characterization and purity of the glycan was assessed by 600-MHz NMR.
Synthesis of Man3GlcNAc2 precursor
Origami2(DE3) gmd::kan ΔnanA ΔwaaL cells were transformed with plasmids pYCG and pManCB by electroporation and used to inoculate a shake flask containing 200 mL LB containing 0.2% (v/v) glucose along with 100 µg/mL ampicillin (Amp) and 25 µg/mL chloramphenicol (Cam), and incubated overnight, with shaking at 30 °C. 50 mL of the 200-mL culture was then transferred to each of ten 1-L shake flasks containing LB supplemented with Amp and Cam and cultures were incubated overnight, with shaking at 30 °C. The cells were then pelleted by centrifugation at 4,236 × g, resuspended in 5 mL of methanol, and sonicated five times at 10-sec pulses. The sonicated lysate was then poured into a glass petri dish and heated at 60 °C until dry. The methods for extraction of LLOs and release of the glycan from the lipid were followed as described elsewhere40,41. Briefly, E. coli cell pellets were resuspended in 2:1 chloroform:methanol, sonicated, and the remaining solids collected by centrifugation. This pellet was sonicated in water and collected by centrifugation. The resulting pellet was sonicated in 10:10:3 chloroform:methanol:water to isolate the LLOs from the inner membrane. The LLOs were purified using acetate-converted DEAE anion exchange chromatography, as they bind to the anion exchange resin via the phosphates that link the lipid and glycan. The resulting compound was dried and treated by mild acid hydrolysis to release glycans from the lipids40,41. The released glycans were then separated from the lipid by a 1:1 butanol:water extraction, wherein the water layer contains the glycans. The glycans were then further purified with a graphitized carbon column using a 0–50% water:acetonitrile gradient. The Man3GlcNAc2 glycan was analyzed by MADLI-TOF MS, FACE, and 600-MHz NMR.
Expression and purification of eukaryotic GnTs
E. coli strains MC4100, SHuffle, or Origami2(DE3) were separately transformed with pMBP-GnTI, pMBP-GnTII, or pMBP-GnTIV by electroporation. Transformed cells were transferred to starter cultures containing 50 mL of LB with 0.2% (v/v) glucose and 100 µg/mL Amp and incubated overnight at 30 °C. The 50-mL cultures were then transferred to a shake flask containing 1 L LB supplemented with 100 µg/mL Amp and induced for 16 h with 0.2% arabinose when the absorbance (Abs600) reached 1.0. Cells were harvested by centrifugation at 4,320 × g, resuspended in a minimal amount of MBP binding buffer (20 mM Tris-HCl, 1 mM EDTA, 200 mM NaCl, pH 7.4), and sonicated 5 times at 30-sec pulses. The lysate was subjected to centrifugation at 12,000 × g for 20 min, and the cell debris was discarded. The clarified lysate was loaded onto an MBP binding buffer, pre-equilibrated amylose gravity flow column (New England Biolabs) and washed with the binding buffer. The protein was eluted with 10 mL MBP binding buffer supplemented with 10 mM maltose and concentrated to 1 mL by centrifugation (Vivaspin 20, GE Healthcare).
Synthesis of hybrid oligosaccharides from the yeast-derived precursor
To produce 6, a 500-µg sample of 4 was treated with an excess amount of GnTI and UDP-GlcNAc (Sigma-Aldrich) in GnT buffer (20 mM HEPES, 50 mM NaCl, 10 mM MnSO4, pH 7.2), and incubated at 37 °C until the reaction was complete, as determined by MALDI-TOF MS. To synthesize 7, a 500-µg sample of 4 was treated with an excess amount of GnTI and GnTIV along with an excess of UDP-GlcNAc and incubated at 37 °C until the reaction was complete. Each glycan was desalted as described above and purified by silica flash column chromatography as described above. The glycans were then characterized by MALDI-TOF MS and 1D 1H 600-MHz NMR. A 250-µg sample of 6 and 7 were treated with an excess amount of B. taurus β1,4-GalT (Sigma-Aldrich) and UDP-Gal (Sigma-Aldrich) to produce 8 and 9. The reaction was carried out at 37 °C until completion, as determined by MALDI-TOF MS. The glycan products were purified and analyzed as described above.
Synthesis of hybrid and complex glycans using the bacteria-derived precursor
A ~250-µg sample of E. coli-derived Man3GlcNAc2 was used for synthesis of each glycan produced. Both UDP-GlcNAc and UDP-Gal were added in 3-fold molar excess, and each enzyme was also added in excess as determined empirically. All synthesized glycans were desalted as described above and purified by graphitized carbon using a 0–50% acetonitrile:water gradient. The purified products were analyzed by MALDI-TOF MS, FACE, and 600-MHz NMR.
Synthesis of GlcNAc terminal glycans
To produce glycan 11, Man3GlcNAc2 was treated with GnTI and incubated overnight at 37 °C. To produce 14, both GnTI and GnTII were added simultaneously to 2 in GnT buffer and incubated overnight at 37 °C. To synthesize 12, GnTI and GnTIV were added simultaneously to 2 in GnT buffer and incubated overnight at 37 °C. For synthesis of 13, GnTI, GnTIV, and H. sapiens GnTV (R&D systems) were added simultaneously to 2 in GnTI buffer and incubated at 37 °C until the reaction was complete. To synthesize 17, GnTI and GnTII were added simultaneously to 2 in GnT buffer and incubated at 37 °C until the reaction was complete. GnTIV was then added to the solution and incubated at 37 °C until the reaction was complete. Similarly, 20 was synthesized by adding GnTI and GnTII to 2 in GnT buffer and incubated at 37 °C until the reaction was complete. GnTIV and GnTV were then added to the solution and incubated at 37 °C until the reaction was complete. The progress of each reaction was monitored by MALDI-TOF MS and determined to be complete when the substrate glycan was undetectable.
Synthesis of bisecting GlcNAc glycans
To produce 18 and 21, the glycans 14 and 17 were first synthesized as described above, and used as substrates for the addition of bisecting GlcNAc. H. sapiens GnTIII (R&D systems) was added to 14 and 17 separately in GnT buffer and incubated at 37 °C until the reaction was complete, as determined by MALDI-TOF MS.
Synthesis of Gal terminal glycans
For synthesis of 10, GnTI and GalT were added simultaneously to 2 in GnTI buffer and incubated at 37 °C until the reaction was complete. For synthesis of 15, 16, and 19, the GlcNAc terminal glycans (12, 14, 17) were first synthesized as described above before the addition of Gal to avoid production of 10 as a side product. GalT was added to 12, 14, and 17 in GnT buffer and incubated at 37 °C until the reactions were complete as determined by MS.
NMR analysis
Glycans 2, 3, 5, 10–21 (≥100 μg each, Complex Carbohydrate Research Center, University of Georgia, acquired spectra and performed analysis) were deuterium exchanged 3 times by suspending in D2O and subsequent lyophilization, were re-dissolved in 300 μl D2O (99.96%, Cambridge Isotopes) and placed in a 3-mm Shigemi tubes. Spectra were obtained at 25 °C on Varian Inova 600 MHz spectrometer equipped with cryoprobe using standard Varian pulse sequences. 1D 1H NMR was used to confirm the structures of glycans 5, 14, 15, 16, and 17; a combination of 1D 1H and 2D gCOSY for glycans 10, 11, 13, 18, 19, 20, and 21; a combination of 1D 1H, 2D gCOSY, and zTOCSY for glycans 2 and 3; and a combination of 1D 1H, 2D gCOSY, and 2D HSQC for glycan 12 (see Supplementary Figs 3–6 for representative 2D data). Glycans 4, and 6–9 (≥100 μg each) were prepared as above, except rotary evaporation was used for deuterium exchange (D2O, 99,9%, Sigma), and 1D 1H spectra were acquired at 25 °C using a Varian Inova 600-MHz spectrometer (Cornell University) with a pulse field gradient probe. Chemical shifts were measured relative to HOD peak (δH = 4.77 ppm).
Glycan-AEAB conjugation and purification
Free reducing glycans were conjugated with AEAB as described previously44. Briefly, 25 μg of glycan was mixed with 10 μl of AEAB hydrochloride salt solution freshly prepared at 0.5 M in DMSO/AcOH (7:3, v/v) followed by an equal volume of 1.0 M sodium cyanoborohydride solution freshly prepared in the same solvent. The mixture was vortexed briefly and incubated at 65 °C for 4 h. The mixture was chilled, and the products were precipitated upon the addition of 10 volumes of acetonitrile. After bringing the suspension to −20 °C and maintaining that temperature for 2 h, the products were separated by centrifugation at 10,000 × g for 3 min. The pellet was dried by centri-vap and stored at −20 °C for further purification. AEAB derivatives of LNnT, LNT, and the biantennary N-glycan, NA2, were prepared as reference standards.
Solid phase extraction by -NH2 column
The dried sample was reconstituted in 10 μl of water, loaded onto the -NH2 SPE column, which had been prewashed with acetonitrile, water and 85% acetonitrile in water. The -NH2 SPE column was then washed with 3 bed volumes of acetonitrile, followed by 3 bed volumes of 85% acetonitrile and the AEAB conjugate was eluted by 5% acetonitrile, 100 mM ammonium acetate. The combined eluents were dried under vacuum to remove the acetonitrile and lyophilized repeatedly to remove the ammonium acetate.
Printing, binding assay, and scanning
NHS-activated slides were purchased from Schott (Louisville, KY). Non-contact printing was performed using a Scienion printer. The printing, binding assay, and scanning conditions were essentially the same as described previously51. Briefly, all samples were printed at 100 μM in phosphate buffer (300 mM sodium phosphates, pH 8.5) in replicates of four. Before the assay, the slides were rehydrated for 5 min in TSM wash buffer (20 mM Tris-HCl, 150 mM sodium chloride (NaCl), 2 mM calcium chloride (CaCl2), and 2 mM magnesium chloride (MgCl2)). Samples were applied to the rehydrated slide in TSM buffer containing 1% BSA and 0.05% Tween 20 in a final volume of 100 μl and the slides were incubated in a humidified chamber at room temperature for 1 h. After incubation, sample was washed away by gently dipping the slides in buffer contained in Coplin jars. Biotinylated ConA (Vector Labs) lectin binding to glycans on the array was detected by a secondary incubation with streptavidin-cyanine 5 at 5 μg/ml (SA-Cy5). The slides were scanned with a Genepix fluorescence microarray scanner equipped with four lasers covering an excitation range from 488 to 637 nm. For Cy5 fluorescence, 633 nm (excitation) was used at laser power 70% and PMT 450. All images obtained from the scanner are in grayscale and colored for easy discrimination. The scanned images were analyzed and quantitated in relative fluorescence units (RFU) from each spot in an Excel spreadsheet, to determine the average and S.D. of the four replicates. The %CV (coefficient of variation expressed in percent) was calculated as 100 × S.D./mean.
Data availability
All datasets related to glycan array screening that were generated in this work are available through The Functional Glycomics Gateway (http://www.functionalglycomics.org/glycomics/publicdata/home.jsp), a comprehensive and free online resource provided by the CFG.
References
Varki, A. Biological roles of oligosaccharides: all of the theories are correct. Glycobiology 3, 97–130 (1993).
Varki, A. & Marth, J. Oligosaccharides in vertebrate development. Seminars in Developmental Biology 6, 127–138 (1995).
Apweiler, R., Hermjakob, H. & Sharon, N. On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim Biophys Acta 1473, 4–8 (1999).
Helenius, A. & Aebi, M. Intracellular functions of N-linked glycans. Science 291, 2364–9 (2001).
Imperiali, B. & O’Connor, S. E. Effect of N-linked glycosylation on glycopeptide and glycoprotein structure. Curr Opin Chem Biol 3, 643–9 (1999).
Hebert, D. N., Lamriben, L., Powers, E. T. & Kelly, J. W. The intrinsic and extrinsic effects of N-linked glycans on glycoproteostasis. Nat Chem Biol 10, 902–910 (2014).
Du, J. & Yarema, K. J. Carbohydrate engineered cells for regenerative medicine. Adv Drug Deliv Rev 62, 671–82 (2010).
Sampathkumar, S. G., Li, A. V., Jones, M. B., Sun, Z. & Yarema, K. J. Metabolic installation of thiols into sialic acid modulates adhesion and stem cell biology. Nat Chem Biol 2, 149–52 (2006).
Lanctot, P. M., Gage, F. H. & Varki, A. P. The glycans of stem cells. Curr Opin Chem Biol 11, 373–80 (2007).
Adamczyk, B., Tharmalingam, T. & Rudd, P. M. Glycans as cancer biomarkers. Biochim Biophys Acta 1820, 1347–53 (2012).
Hakomori, S. Aberrant glycosylation in cancer cell membranes as focused on glycolipids: overview and perspectives. Cancer Res 45, 2405–14 (1985).
Kim, Y. J. & Varki, A. Perspectives on the significance of altered glycosylation of glycoproteins in cancer. Glycoconj J 14, 569–76 (1997).
Fuster, M. M. & Esko, J. D. The sweet and sour of cancer: glycans as novel therapeutic targets. Nat Rev Cancer 5, 526–42 (2005).
Sheridan, C. Commercial interest grows in glycan analysis. Nat Biotechnol 25, 145–6 (2007).
Walt, D. et al. Transforming Glycoscience: A Roadmap for the Future, (The National Academies Press, 2012).
Werz, D. B. et al. Exploring the structural diversity of mammalian carbohydrates (“glycospace”) by statistical databank analysis. ACS Chem Biol 2, 685–91 (2007).
North, S. J., Hitchen, P. G., Haslam, S. M. & Dell, A. Mass spectrometry in the analysis of N-linked and O-linked glycans. Curr Opin Struct Biol 19, 498–506 (2009).
Marino, K., Bones, J., Kattla, J. J. & Rudd, P. M. A systematic approach to protein glycosylation analysis: a path through the maze. Nat Chem Biol 6, 713–23 (2010).
Rillahan, C. D. & Paulson, J. C. Glycan microarrays for decoding the glycome. Annu Rev Biochem 80, 797–823 (2011).
Oyelaran, O. & Gildersleeve, J. C. Glycan arrays: recent advances and future challenges. Curr Opin Chem Biol 13, 406–13 (2009).
Cummings, R. D. & Etzler, M. E. Antibodies and lectins in glycan analysis. In Essentials of Glycobiology (Cold Spring Harbor Laboratory Press, Cold Spring Harbor (NY), 2009).
Kajihara, Y. et al. Prompt chemoenzymatic synthesis of diverse complex-type oligosaccharides and its application to the solid-phase synthesis of a glycopeptide with Asn-linked sialyl-undeca- and asialo-nonasaccharides. Chemistry 10, 971–85 (2004).
Verostek, M. F., Lubowski, C. & Trimble, R. B. Selective organic precipitation/extraction of released N-glycans following large-scale enzymatic deglycosylation of glycoproteins. Anal Biochem 278, 111–22 (2000).
Boltje, T. J., Buskas, T. & Boons, G. J. Opportunities and challenges in synthetic oligosaccharide and glycoconjugate research. Nat Chem 1, 611–22 (2009).
Lepenies, B., Yin, J. & Seeberger, P. H. Applications of synthetic carbohydrates to chemical biology. Curr Opin Chem Biol 14, 404–11 (2010).
Palcic, M. M. Glycosyltransferases as biocatalysts. Curr Opin Chem Biol 15, 226–33 (2011).
Schmaltz, R. M., Hanson, S. R. & Wong, C. H. Enzymes in the synthesis of glycoconjugates. Chem Rev 111, 4259–307 (2011).
Li, L. et al. Efficient chemoenzymatic synthesis of an N-glycan isomer library. Chem Sci 6, 5652–5661 (2015).
Maki, Y., Okamoto, R., Izumi, M., Murase, T. & Kajihara, Y. Semisynthesis of Intact Complex-Type Triantennary Oligosaccharides from a Biantennary Oligosaccharide Isolated from a Natural Source by Selective Chemical and Enzymatic Glycosylation. J Am Chem Soc 138, 3461–8 (2016).
Wang, Z. et al. A general strategy for the chemoenzymatic synthesis of asymmetrically branched N-glycans. Science 341, 379–83 (2013).
Rich, J. R. & Withers, S. G. Emerging methods for the production of homogeneous human glycoproteins. Nat Chem Biol 5, 206–215 (2009).
Blow, N. Glycobiology: A spoonful of sugar. Nature 457, 617–620 (2009).
Blixt, O. & Razi, N. Chemoenzymatic synthesis of glycan libraries. Methods Enzymol 415, 137–53 (2006).
Vasiliu, D. et al. Large-scale chemoenzymatic synthesis of blood group and tumor-associated poly-N-acetyllactosamine antigens. Carbohydr Res 341, 1447–57 (2006).
Valderrama-Rincon, J. D. et al. An engineered eukaryotic protein glycosylation pathway in Escherichia coli. Nat Chem Biol 8, 434–6 (2012).
Trimble, R. B., Atkinson, P. H., Tschopp, J. F., Townsend, R. R. & Maley, F. Structure of oligosaccharides on Saccharomyces SUC2 invertase secreted by the methylotrophic yeast Pichia pastoris. J Biol Chem 266, 22807–17 (1991).
Chiba, Y. et al. Production of human compatible high mannose-type (Man5GlcNAc2) sugar chains in Saccharomyces cerevisiae. J Biol Chem 273, 26298–304 (1998).
Hamilton, S. R. et al. Humanization of yeast to produce complex terminally sialylated glycoproteins. Science 313, 1441–3 (2006).
Okbazghi, S. Z. et al. Production, Characterization, and Biological Evaluation of Well-Defined IgG1 Fc Glycoforms as a Model System for Biosimilarity Analysis. J Pharm Sci 105, 559–74 (2016).
Gao, N. Fluorophore-assisted carbohydrate electrophoresis: a sensitive and accurate method for the direct analysis of dolichol pyrophosphate-linked oligosaccharides in cell cultures and tissues. Methods 35, 323–7 (2005).
Gao, N. & Lehrman, M. A. Non-radioactive analysis of lipid-linked oligosaccharide compositions by fluorophore-assisted carbohydrate electrophoresis. Methods Enzymol 415, 3–20 (2006).
Bessette, P. H., Aslund, F., Beckwith, J. & Georgiou, G. Efficient folding of proteins with multiple disulfide bonds in the Escherichia coli cytoplasm. Proc Natl Acad Sci USA 96, 13703–8 (1999).
Chen, R., Pawlicki, M. A., Hamilton, B. S. & Tolbert, T. J. Enzyme-Catalyzed Synthesis of a Hybrid N-Linked Oligosaccharide using N-Acetylglucosaminyltransferase I. Advanced Synthesis & Catalysis 350, 1689–1695 (2008).
Song, X. et al. Novel fluorescent glycan microarray strategy reveals ligands for galectins. Chem Biol 16, 36–47 (2009).
Hug, I. et al. Exploiting bacterial glycosylation machineries for the synthesis of a Lewis antigen-containing glycoprotein. J Biol Chem 286, 37887–94 (2011).
Ortiz-Soto, M. E. & Seibel, J. Expression of Functional Human Sialyltransferases ST3Gal1 and ST6Gal1 in Escherichia coli. PLoS One 11, e0155410 (2016).
Thomason, L.C., Costantino, N. & Court, D. L. E. coli genome manipulation by P1 transduction. Curr Protoc Mol Biol Chapter 1, Unit1 17 (2007).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2(2006), 0008 (2006).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97, 6640–5 (2000).
Shanks, R. M., Caiazza, N. C., Hinsa, S. M., Toutain, C. M. & O’Toole, G. A. Saccharomyces cerevisiae-based molecular tool kit for manipulation of genes from gram-negative bacteria. Appl Environ Microbiol 72, 5027–36 (2006).
Song, X. et al. Shotgun glycomics: a microarray strategy for functional glycomics. Nat Methods 8, 85–90 (2011).
Acknowledgements
This research was supported by National Science Foundation Awards CBET-1605242, CBET-1159581, and MCB-1411715 (to M.P.D.), the New York State Office of Science, Technology and Academic Research Distinguished Faculty Award (to M.P.D.), the National Institutes of Health Small Business Innovation Research grants 1R43GM088905-01 and 2R44GM088905-02 (to J.H.M.), and the National Institutes of Health contract HHSN261201300046C (to J.H.M.). The NMR analysis was performed at the Complex Carbohydrate Research Center, supported in part by a grant from the NIH-funded Research Resource for Integrated Glycotechnology (5P41GM10339024 to P.A.). The glycan microarray analysis was performed at the National Center for Functional Glycomics (NCFG) at Beth Israel Deaconess Medical Center, supported in part by the National Center for Research Resources (P41GM103694 to R.D.C.). We also thank Anthony Condo and Ivan Keresztes of the Chemistry NMR Facility (Cornell University) for their assistance acquiring NMR spectra.
Author information
Authors and Affiliations
Contributions
B.H., J.W. and J.M. designed research, performed research, analyzed data, and wrote the paper. M.S., A.F., J.B., A.P., R.N., C.H., C.G., R.K., and J.H.-M. designed research, performed research, and analyzed data. P.A. and R.C. designed research and analyzed data. M.P.D. designed research, analyzed data, and wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
M.P.D. has a financial interest in Glycobia, Inc. M.P.D.’s interests are reviewed and managed by Cornell University in accordance with their conflict of interest policies.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hamilton, B.S., Wilson, J.D., Shumakovich, M.A. et al. A library of chemically defined human N-glycans synthesized from microbial oligosaccharide precursors. Sci Rep 7, 15907 (2017). https://doi.org/10.1038/s41598-017-15891-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15891-8
This article is cited by
-
Immobilized enzyme cascade for targeted glycosylation
Nature Chemical Biology (2024)
-
Successive remodeling of IgG glycans using a solid-phase enzymatic platform
Communications Biology (2022)
-
A universal glycoenzyme biosynthesis pipeline that enables efficient cell-free remodeling of glycans
Nature Communications (2022)
-
Streamlining the chemoenzymatic synthesis of complex N-glycans by a stop and go strategy
Nature Chemistry (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.