Abstract
In this work, performance and microbial structure of a digestion (food waste-only) and a co-digestion process (mixture of cow manure and food waste) were studied at mesophilic (37 °C) and thermophilic (55 °C) temperatures. The highest methane yield (480 mL/g VS) was observed in the mesophilic digester (MDi) fed with food waste alone. The mesophilic co-digestion of food waste and manure (McoDi) yielded 26% more methane than the sum of individual digestions of manure and food waste. The main volatile fatty acid (VFA) in the mesophilic systems was acetate, averaging 93 and 172 mg/L for McoDi and MDi, respectively. Acetate (2150 mg/L) and propionate (833 mg/L) were the main VFAs in the thermophilic digester (TDi), while propionate (163 mg/L) was the major VFA in the thermophilic co-digester (TcoDi). The dominant bacteria in MDi was Chloroflexi (54%), while Firmicutes was dominant in McoDi (60%). For the mesophilic reactors, the dominant archaea was Methanosaeta in MDi, while Methanobacterium and Methanosaeta had similar abundance in McoDi. In the thermophilic systems, the dominant bacteria were Thermotogae, Firmicutes and Synergistetes in both digesters, however, the relative abundance of these phyla were different. For archaea, the genus Methanothermobacter were entirely dominant in both TDi and TcoDi.
Similar content being viewed by others
Introduction
Anaerobic digestion process has widely been employed for treatment of various organic wastes because the process can be used for production of value-added products such as an energy-rich gas and bio-fertilizer. This process is carried out by a complex microbial community which degrade various organic compounds into final products such as methane and carbon dioxide, collectively called biogas.
There are presently many research efforts worldwide on anaerobic digestion of food waste to improve process efficiency, stability and economic competitiveness. Studies of co-digestion of food waste generally found that inclusion of food waste was beneficial for methane yield1,2,3, while digestion processes with food waste as the sole substrate were often found to be unstable3,4,5. Several researchers have reported the benefits of using mixed feedstocks, including increased biogas production, enhanced degradation rates and higher digester capacity1,6,7. The beneficial effects of co-digestion are mostly related to a balanced availability of macro- and micronutrient required by the microbial community, optimal moisture content, buffer capacity and dilution of inhibitory or toxic compounds. Additionally, co-digestion may improve the process kinetics rather than the bioavailability of the feedstock. Ebner et al.8 measured hydrolysis rates using bio-methane potential assays, and found that co-digestion increased hydrolysis rates when food waste and manure was co-digested compared to mono-digestion in BMP assays. The synergistic effect was attributed to dilution of inhibitory compounds and improved nutrient balance due to co-digestion8,9. The enzymes involved in hydrogenotrophic methanogenesis and syntrophic acetate oxidation requires trace elements such as selenium (Se), molybdenum (Mo) tungsten (W), cobolt (Co), nickel (Ni) and iron (Fe)4. Lack of these trace elements can limit the syntrophic acetate oxidation as well as formate oxidation3,4,5. The resulting accumulation of formate may again inhibit propionic acid oxidation. This will result in an overall acid accumulation, which eventually can cause the pH in the digester to drop, severely affecting or completely stopping the methanogenesis. Notably, the toxicity of intermediate compounds also increases with increasing temperature10 and thermophilic digesters are commonly considered to be more prone to process inhibition than mesophilic digesters7. Moreover, anaerobic digestion processes operating at high-temperature often selects for a less diverse microbial community, which is more vulnerable to stress and operational changes11,12. Most studies in the literature have focused on enhancing functionality and operation of anaerobic co-digesters using food waste and other feedstocks. Several studies have compared performance of mesophilic and thermophilic digesters2,3, but comparison of community structures and diversity in anaerobic digesters and anaerobic co-digesters at these different temperatures are rare in the literature.
Accordingly, the aim of this study was to investigate the microbial structure of co-digestion of food waste and cow manure under mesophilic (37 °C) and thermophilic (55 °C) conditions. Additionally, we compared the microbial structure of the co-digestion process to that of food waste digestion alone to determine how the co-digestion process influences the microbial communities. Also, performance parameters were studied under the various conditions and it was attempted to explain performance efficiencies using microbial data.
Results and Discussion
Performance of the biogas reactors
The average pH values for MDi, McoDi, TDi and TcoDi were 7.7 ± 0.1, 7.9 ± 0.1, 7.8 ± 0.2 and 8.2 ± 0.1, respectively. As illustrated in Fig. 1, the average pH of McoDi and MDi was comparable, although slightly higher in McoDi. Notably, while the pH of TDi was similar to mesophilic processes, an elevated pH was clearly seen for TcoDi. This agrees with the higher ammonia concentrations in the co-digestion systems (McoDi and TcoDi), which were 16.5% and 13.7% higher than the MDi and TDi digesters, respectively (Fig. 2). Additionally, as shown in Fig. 1, alkalinity was also higher in the co-digesters than the digestion systems. This is most likely due to the addition of manure, as manure typically has high content of nitrogen-bearing material8 that are released as ammonia during the fermentation process and acts as a buffering system.
The degradation of organic material in all the digesters was measured in terms of TCOD removal (Fig. 3). Regardless of the operational temperature, the removal efficiencies were higher for the digestion systems (MDi: 73.0% and TDi: 66.4%) than the co-digesters (McoDi: 61.4% and TcoDi: 56.7%). This was expected due to a general high degradability of food waste13. MDi had the highest methane yield of all four digesters with 479.5 ± 33.9 ml CH4/g VSfeed, which was 11.5%, 7.0% and 31.6% higher than the McoDi, TDi and TcoDi, respectively (Fig. 3). These results are in agreement with earlier studies reported elsewhere8,14. Additionally, lower methane production in the thermophilic reactors may be related to the presence of higher free ammonia concentrations that was, on average, 198 mg/L for TDi and 431 mg/L in TcoDi (Fig. 2), potentially causing inhibition of the methanogenesis process15.
It has been observed that co-digestion of food waste and manure may enhance biogas production, and lead to more stable digestion processes7,16,17,18. We also observed higher methane production when we compared the methane yield of McoDi fed with the mixture of food waste and manure with that of manure-only fed mesophilic (37 °C) digester. The methane yields of manure-fed digester and McoDi were 133 ± 18 and 430 ± 28 mL CH4/g VSfeed, respectively. Based on the measured specific methane yield from MDi and the manure-only reactor, the expected methane yield for McoDi without any synergistic effects would be 341 mL CH4/g VSfeed. However, our results showed that the observed methane yield of McoDi was 430 mL CH4/g VSfeed, meaning that the co-digestion of food waste and manure (McoDi) resulted in 26% higher methane production than the sum of digestions of individual substrates.
The solubilization values estimated for MDi, McoDi, TDi and TcoDi were 56, 55, 63 and 48%, respectively (see Equation 1). The highest solubilization extent was observed in the TDi (63%), although less methane was generated in this system as compared to MDi and McoDi (discussed above). It was also noticed that the SCOD fraction of the extent of solubilization in TDi was quite high (10%) and thus less solubilized compounds were converted into the final product methane. This accumulation of soluble COD was likely prompted by the higher degradability of the food waste as compared to manure. The SCOD accounted for 1, 2 and 6% of the solubilization extent in MDi, McoDi and TcoDi, respectively. Moreover, the TcoDi showed the lowest solubilization extent (48%), indicating less efficient solubilization of the substrate mixture. Additionally, when the SCOD results were compared between the digester sets (i.e., MDi vs. TDi and McoDi vs. TcoDi), it revealed that the SCOD was 10 and 4 times higher in the effluent of the TDi and TcoDi than those of MDi and McoDi. However, the effluent SCOD concentrations were statistically comparable in the mesophilic digester (932 ± 151) and co-digester (1537 ± 511) (pvalue = 0.05).
Overall, both mesophilic digesters had low concentrations of volatile fatty acids (VFAs). Analysis of the VFAs measurements (Fig. 4) revealed acetate as the main VFA in the mesophilic digesters MDi and McoDi, which was, on average, 172 ± 61 and 93 ± 54 mg/L, respectively. The remaining VFAs detected in the mesophilic digesters, propionic, iso-butyric and butyric acids, were all below 50 mg/L (Fig. 4). Thus, it appeared that the fermenting and methanogenic processes were in balance preventing accumulation of intermediate products in MDi and McoDi. Low VFAs concentrations were reported for an anaerobic digester operated under mesophilic condition for food waste treatment19. The analysis of the VFA profiles (Fig. 4) revealed a totally different behavior in the thermophilic digester TDi and co-digester TcoDi. Acetate and propionate accumulated in TDi, averaging 2150 ± 208 and 833 ± 282 mg/L, respectively. These results were consistent with previous works reported in literature that under thermophilic condition increased concentrations of VFAs were observed, while mesophilic digesters were capable of achieving lower VFA concentrations19,20. As can be seen from Fig. 4, the concentrations of total VFAs in TcoDi were less than 300 mg/L, in which propionate was the main VFA with an average concentration of 163 ± 27 mg/L. This difference in the VFA profile in TDi and TcoDi might be due to an improved synergistic performance of acetogens and methanogens in TcoDi that prevented the accumulation of the intermediate products and resulted in significantly lower concentrations of VFAs in TcoDi. The slow degradation of the manure which constituted 40% (on VS basis) of the feedstock in the co-digesters may also explain this difference.
Microbial composition of the mesophilic reactors
Statistical analysis demonstrated that the anaerobic co-digestion process resulted in a significantly (pvalue < 0.005) higher microbial richness compared to the digesters fed with food waste alone (see Supplementary Fig. S1). The major bacteria in both mesophilic digesters included Firmicutes, Chloroflexi, Bacteroidetes and Actinobacteria (Fig. 4). However, the distribution of these major bacteria in the digesters was different. Chloroflexi, which in the final phase constituted 54% of the sequences, was the dominant phylum in MDi, followed by 25% Firmicutes and 15% Bacteroidetes. Firmicutes (60% of the sequences in the final phase) was the dominant phylum in McoDi, while the relative abundance of Chloroflexi (22%) and Bacteroidetes (8%) was noticeably lower in McoDi than MDi. Additionally, the candidate phylum WWE1 was identified in McoDi and accounted for 5% of the relative abundance. Limam et al.21 investigated the metabolic function of WWE1 members and suggested that the members of this division were involved in hydrolysis of cellulosic materials. WWE1 was also found in mesophilic co-digestion studies of manure with various agricultural residues22,23. Thus, the addition of cow manure to the co-digestion system seems to spur the growth of WWE1 members, probably involved in decomposition of cellulose content of the manure. It should be noted that WWE1 was not detected in the cow manure in the current study. The dominance of Chloroflexi in MDi (Fig. 5), which was mainly made up of the T78 group of family Anaerolinacea, was probably due to the presence of fermentable carbohydrates in the preprocessed food waste used (pasteurized at 70 °C). Anaerolinacea are mostly saccharolytic anaerobes and use a number of carbohydrates for growth24,25. Use of manure in the feedstock of the co-digestion systems resulted in a different relative abundance of bacterial communities in McoDi and prompted the prevalence of Firmicutes, which include members with very versatile metabolic characteristics and more potential to degrade the recalcitrant manure26,27. Firmicutes has been reported as one of the major microbial contributors in several studies carried out on anaerobic digesters, indicating that the phylum is common in both mesophilic and thermophilic processes28,29. Additionally, Firmicutes dominance has also been linked to better reactor performance20. The higher relative abundance of Bacteroidetes in MDi, which was fed with the preprocessed food waste, probably indicates involvement of their members in degradation of intermediate degradation products of carbohydrates and proteins.
Notably, the relative abundance of Fimicutes increased in the final phase of McoDi compared to MDi. This could be due to the addition of manure which is a potential source of Firmicutes, as organisms belonging to this phylum dominated the microbial profile of the manure feedstock with 78% of all sequences (Fig. 5). To evaluate this, the genus level distribution of the sequences was investigated and a high diversity within the Firmicutes-phylum was noticed (Fig. 6A). An unclassified genus of the family Tissierellaceae accounted for 32% of the sequences assigned to the phylum Firmicutes in the final phase of MDi, while this value was much lower in McoDi (11% of phylum) where the main genus was Clostridium (42% of phylum). In compliance, three OTUs assigned to Clostridium were significantly more abundant in McoDi compared to MDi (Supplementary Fig. S2, see also Fig. 7A). Thus, it would appear that Firmicutes in general and Clostridium in particular played an important role in McoDi system. This genus was also represented in the cow manure samples, accounting for 9% of the Firmicutes-related sequences. A principle component analysis (PCA) was used to investigate possible links between microbiome and performance. Based on this analysis an association of Clostridium to the concentration of n-Butyrate was observed, although only low levels of butyrate were measured in both mesophilic digesters (Fig. 7B). Notably, a correlation was observed between the abundance of Clostridium and the cow manure used in the feedstock mixture of the co-digestion system. It is therefore reasonable to believe that the increase in relative abundance of Clostridium in the co-digestion system was originated from the cow manure as a feedstock. It should nevertheless be mentioned that some Clostridium species can form endospores that enable them to tolerate moist heat30 and pasteurization pretreatment applied on the food waste collected from the processing center. The food waste can therefore not be eliminated as a source of Clostridium. However, a carry-over from the cow manure used seems more likely due to the abovementioned increase of Clostridium in McoDi. This was further supported by the correlation of higher numbers of Clostridium with the addition of cow manure.
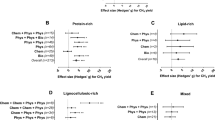
Phylogenetic distribution of the genera within (A) phylum Firmicutes and (B) phylum Euryarchaeota. Although all genera (>0.005% of total sequences) are included in the pie chart for Firmicutes, only the genera representing ≥1% of the sequences in at least one of the samples is included in the legend to reduce size. The most dominant genera are highlighted in bold type to ease the visual interpretation.
Relative abundance of the OTUs, classified at genus level or highest possible ranked taxonomic level (A), and their association with operational conditions and process variables in the mesophilic (B) and the thermophilic (C) digesters assessed through principal component analysis (PCA). The chemical variables included in the PCA plots are the values of NH3, CH4 (ml/week), and VFAs (propionate; “Pro”, Acetate; “AC”, n-Butyrate; “n-Bu”, i-Butyrate; “i-Bu”, i-Valerate; “i-Va”). Only the most abundant taxa are annotated in the barchart to reduce the complexity. A comprehensive OTU table is supplied in the Electronical supplementary material, Table S1.
Archaea belonging to the phylum Euryarchaeota were dominated by Methanobacterium and Methanosaeta in both MDi and McoDi (Fig. 6B). The genus distribution within this phylum was similar for initial and final phase in MDi, where Methanosaeta was dominant, constituting 53% and 62% (in initial and final phase, respectively) of the archaeal sequences. Methanosaeta was also prominent in the McoDi (43% and 36% in initial and final phase, respectively), yet significantly lower compared to MDi (Supplementary Fig. S2). In addition to Methanobacterium and Methanosaeta, a noticeable portion of the methanogenic population was also assigned to the genus Methanobrevibacter in McoDi, with increasing relative abundance over time (from 7% of the archaeal sequences in the initial phase, to 22% in the final phase). Methanobrevibacter most likely originated from the manure used in this study as the analysis of manure samples showed Methanobrevibacter as the only dominant archaea (Fig. 6B). While Methanosaeta is known as an acetate-utilizing methanogen, Methanobacterium and Methanobrevibacter both contain H2 utilizing methanogens31, suggesting a mixed pathway for methane production in the mesophilic co-digestion system. The reason for the presence of high hydrogen utilizers might partly be due to slightly higher free ammonia observed in McoDi (81 mg/L vs. 50 mg/L in MDi) and partly due to the continual addition of Methanobrevibacter through manure. This agrees with previous studies indicating the dominance of hydrogen utilizing methanogens in manure-fed digesters32,33.
Microbial composition in the thermophilic digesters
The bacterial communities in TDi and TcoDi were mainly composed of the four phyla (relative abundancy >1%) Thermotogae, Firmicutes, Synergistetes and Bacteroidetes (Fig. 5). However, the relative abundance of these phyla was remarkably different within TDi and TcoDi, except for Bacteroidetes that accounted for 1–2% of the total sequences in both digesters. The profound effect of co-digestion on the bacterial community in the final phase of TcoDi was reflected by increased relative abundance of Firmicutes (62% of all sequences) and a decreased relative abundance of Thermotogae (15% of all sequences). In comparison, the relative abundance of Firmicutes and Thermotogae was 35% and 40% in the TDi, respectively. Notably, the distribution of Synergistetes, mainly represented by genus Anaerobaculum, differed significantly in the thermophilic digesters (Supplementary Fig. S2), accounting 11% and 4% in the final phase of TDi and TcoDi, respectively. The members of this taxon fermentatively convert polypeptides and organic acids to acetate, H2 and CO2 34. Compared to the initial phase, the relative abundance of Thermotogae increased over time in both thermophilic reactors, while the relative abundance of Firmicutes decreased only in TDi (Fig. 5). Similar to the mesophilic digesters, co-digestion of manure and food waste seemingly spurred the growth and dominance of Firmicutes in the TcoDi, while the digestion of food waste alone induced more even-distribution of Thermotoga and Firmicutes and supported the development of more Synergistetes in TDi.
While phylotypes assigned to genus Coprothermobacter accounted for more than 50% of the sequences assigned to Firmicutes in the initial samples of both TDi and TcoDi, this portion was largely reduced in the final phase (15% and 8% of Firmicutes, in TDi and TcoDi respectively). Instead, a more even distribution of several genera was observed in the final phase (Fig. 6A), with prominence of Syntrophomonas (16% and 11% of Firmicutes, in TDi and TcoDi respectively), Thermactogenium (9% and 10% in TDi and TcoDi respectively) and unclassified phylotypes assigned to the candidate divisions SHA-98 (17% and 9% in TDi and TcoDi respectively) and MBA08 (8% and 28% in TDi and TcoDi respectively). Coprothermobacter is a proteolytic bacterium involved in the syntrophic fermentation of polypeptides, and the high dominance in the initial phase was most likely reflected by a strong dominance of the Coprothermobacter population in the seed culture from the FREVAR biogas plant, as reported in a previous study35. The OTUs assigned to genus Thermacetogenium were in all probability affiliated to Thermacetogenium phaeum 35, a bacterium able to oxidize acetate syntrophically and grow acetogenically on organic acids and alcohols36,37.
The phylotype affiliated to candidate order MBA08 was noticeably higher in the TcoDi compared to TDi. This probably suggests the role of the members of the candidate group MBA08 in the co-digestion of food waste and manure. Li38 also reported the candidate order MBA08 as one of the major bacterial groups in the thermophilic reactor of a staged system used for the co-digestion of whey and manure. However, none of the Firmicutes-associated sequences obtained from cow manure were related to MBA08 in the current study. On the contrary, the relative abundance of the candidate order SHA-98 was greater in the food waste-fed digester (i.e., TDi). There is almost no knowledge regarding the function of the members of the unclassified order SHA-98 and no genera could be assigned in this group.
Most of the phylotypes that could be assigned to a known genus in the order Clostridiales were probably involved in the degradation of polysaccharides, fermentable carbohydrates and syntrophic oxidation of saturated fatty acids26. Members of Syntrophomonas are believed to oxidize anaerobically C4-C18 saturated fatty acids39. Clostridium consists of bacteria that display metabolic versatility26,40. Ruminococccaceae, Caldicoprobacteriaceae and Lachnospiraceae were less dominant families. Caldicoprobacter, which exclusively was represented by Caldicoprobacteriaceae in both digesters, ferments xylan and simple sugars to lactate, acetate, H2 and CO2 41. Interestingly, the family Lachnospiracea, although less abundant, differed in the genus and was mainly composed of Butyrivibrio in TcoDi and of Coprococcus in TDi. Members of the Butyrivibrio and Coprococcus both use fermentable carbohydrates, however, the Butyrivibrio members are also involved in degradation of plant materials and are a major component of rumen microbiota41. The Butyrivibrio members probably came from cow manure and were able to retain their activities and growth in TcoDi.
Unlike the large diversity observed within Firmicutes, the second most dominant phylum, Thermotogae (Fig. 5), demonstrated very low diversity as almost all sequences were assigned to the candidate division Thermotoga S1. Notably, a clear difference was found in the relative abundance of this phylum in the final phase of TDi (40% of all sequences) and TcoDi (15% of all sequences). As described earlier42, the members of Thermotoga are capable to grow on the various simple (e.g., glucose) and complex (e.g., xylan and starch) polysaccharides. The lower abundance of Thermotoga species may explain the lower methane yield in the TcoDi (23% less) than the TDi, especially considering the higher amount of particulate COD (44% higher) that left the TcoDi as compared to that of the TDi. Presence of the higher particulate COD might be due to an inefficient conversion of the complex carbohydrates in the feed, in particular in the recalcitrant manure. Additionally, principle component analysis (Fig. 7C) showed a correlation between the relative abundance of S1 and the concentrations of VFAs, indicating that the elevated concentrations of VFAs in TDi could be a cause-effect of an enhanced degradation of polysaccharides by Thermotoga S1. Furthermore, the free ammonia measured was 2.2 times greater in TcoDi (431 mg/L) than TDi (198 mg/L), suggesting that ammonium may possibly influence the abundance of the Thermotoga species43.
Overall, it would appear that the detection of higher relative abundance of 16S rRNA genes assigned to the genera Anaerobaculum, Coprothermobacter, Thermotoga and Syntrophomonas in TDi might indirectly imply an enhanced hydrolysis and acidogenesis of the food waste as compared to the co-digestion of food waste and manure. This might further be supported by the presence of significantly greater amount of VFAs (acetate, propionate and butyrate) in TDi than TcoDi (Fig. 4).
Analysis of the archaeal sequences (Fig. 6) showed that the process configuration (digestion vs. co-digestion) had little influence on the composition of methanogens and that the genus Methanothermobacter, which contains hydrogen utilizers, was almost entirely predominant in TDi and TcoDi. A correlation between Methanothermobacter and Coprothermobacter was observed, as well as an association with methane production (Fig. 7C). Such co-existence has frequently been reported in literature, drawing a scenario of a synergic relationship where Coprothermobacter supply Methanothermobacter with hydrogen44. The dominance of Methanothermobacter agrees with a previous study on community structure in a thermophilic biogas plant (FREVAR), from where the inoculum was taken for the start-up of the thermophilic digesters used in this study35. The lack of Methanosaetaceae species reflected an unfavorable environment (e.g., high free ammonia content) for their activities in TDi and TcoDi, suggesting the prevalence of the hydrogenotrophic methanogensis pathway in both digesters. In addition to an unfavorable environment, the prevalence of Methanothermobacter members might be due to the improved hydrolysis and fermentation at the elevated temperature that required syntrophic reactions to efficiently convert intermediates such as H2 and carboxylic organic acids.
Conclusion
The anaerobic digesters fed solely food waste performed better than the co-digesters (food waste and cow manure), most probably due to the addition of a more recalcitrant material in the form of cow manure in the co-digesters. Nevertheless, co-digestion resulted in a higher microbial diversity at both temperatures, compared to anaerobic digestion of food waste as sole substrate. This could be a reflection of the increased complexity of feedstocks in co-digestion, selecting for a richer microbial community. Although similar in the initial phase, the microbial community compositions diverged when cow manure was added at both temperatures. Based on our observations, we speculate that this variation is mostly explained by cow manure providing trace minerals and a balanced C/N ratio, rather than carry-over of microorganisms from the cow manure. However, the increased population Clostridium in both McoDi and TcoDi indicates that the establishment of this population is a direct result of microbiome transmission from the cow manure. Carry-over of methanogens from the cow manure, represented by Methanobrevibacter was also suggested for the mesophilic co-digestion system (McoDi), while only to a minor extent in the thermophilic co-digestion system (TcoDi). As higher microbial diversity often is associated with a microbiome that is more resilient to environmental changes and stress, co-digestion could potentially enhance the robustness of the anaerobic digestion process. Additionally, co-digestion at mesophilic temperature clearly showed a synergistic effect, yielded more methane than the digestion of manure-alone.
Materials and Methods
Feedstock
Food waste (FW) was shipped from Norsk Matretur AS (Norwegian food recycling, Finstadjordet, Norway), which is a central food waste pre-treatment plant which reduces particle size to <7 mm and sanitizes the waste at 70 °C for 1 hour, as required by Norwegian health regulations. Dairy cow manure was collected at the farm of the Norwegian University of Life Sciences (Ås, Norway). Both manure and FW were stored at 4 °C and diluted using tap water to achieve the targeted organic loading rate. The waste batch were characterized on a weekly basis to ensure a constant organic loading. The average characteristics of the substrates are shown in Table 1.
Set up and operation of digesters
Four completely mixed reactors (Belach Bioteknik, Sweden) with a working volume of 6 L were used in this study. Two of the digesters were only fed with food waste, where one of them was kept at mesophilic temperature (37 ± 0.1 °C; MDi), while the other one was kept at thermophilic temperature (55 ± 0.1 °C; TDi). The Co-digestion systems consisted of one mesophilic co-digester (McoDi) and one thermophilic co-digester (TcoDi), which both were fed with a mixture of food waste and cow manure in a ratio of 60:40 on volatile solid (VS) basis. The start-up of the reactors was performed as previously described by Estevez et al.45. Seed sludge for the mesophilic reactors came from the Oslo EGE biogas plant (Nes, Norway); a full scale mesophilic anaerobic digester with food waste as its sole substrate. Seed sludge for the thermophilic reactors came from the FREVAR biogas plant (Fredrikstad, Norway); a full scale thermophilic reactor with sludge and food waste as its substrates. The feeding of the experimental reactors was done manually once per day, 6 days a week. Hydraulic retention time (HRT) and organic loading rate (OLR) for all the digesters were 20 days and 3 g VS/L/d, respectively. Temperature, pH, biogas volume and stirrer speed (set at 100 rpm) of the digesters were monitored and recorded online using BIOPHANTOM software (Belach Bioteknik, Sweden). Additionally, samples of the effluent were regularly taken to monitor the performance of the digesters and co-digesters.
Analytical methods
Total solids (TS), volatile solids (VS), total chemical oxygen demand (TCOD) and soluble chemical oxygen demand (SCOD) were measured following Standard Methods46. Chemical oxygen demand was measured using Merck Spectroquant® COD Cell test with measuring range 0.5–10 g/L. The extent of solubilization was calculated using Equation (1).
where CODCH4 is the COD equivalent of the CH4 produced; SCOD is effluent soluble COD; SCODin is influent soluble COD and PCODin is the influent particulate COD.
Ammonium (NH4 +) was measured with an ammonium ion selective electrode according to the company’s manual (Orion 93; Thermoscientific, USA). Biogas composition was analyzed online for methane and carbon dioxide as previously described Zamanzadeh et al.15, with the use of an SRI gas chromatograph (Model 8610 C) equipped with a thermal conductivity detector (TCD, RCH3100, USA) and a 2 m Haysep-D column. Volatile fatty acids (VFAs) were determined by high-pressure liquid chromatography (HPLC; Dionex 3100, USA) with a Zorbax Eclipse Plus C18 column (150 × 2.1 mm column; 3.5 µm particles; Agilent, USA) as previously described by Zamanzadeh et al.15. The samples were centrifuged at 14000 rpm for 5 min, adjusted to pH < 2.5 using 95–98% H2SO4 and then filtered through a 0.45 µm cellulose acetate syringe filter.
DNA extraction
Samples were collected for 16S rRNA gene sequencing during the initial stable phase of the anaerobic digestion (after 68 days) and in the final phase (after 152 days) from reactors MDi, McoDi, TDi and TcoDi, in addition to the cow manure used for co-digestion. Food waste was also sampled for the same purpose, but genomic DNA was not successfully recovered, likely due to the sanitization treatment (70 °C for 1 hour). All samples were frozen immediately after sampling, and stored at −20 °C until DNA extraction. For DNA extraction, thawed samples were centrifuged at 18 800 x g for 7 min. to remove the liquid. The pellet was then re-suspended in S.T.A.R buffer (Roche Diagnostics Corporation, USA) to stabilize nucleic acid and prevent bacterial growth. The suspension was vortexed followed by a subsequently slow spin to dissociate cells from large particles. The cell-containing suspension was transferred to FastPrep24 tubes with acidic washed glass beads for mechanical lysis. DNA was extracted using a commercial DNA extraction kit (LGC Genomics, UK), and DNA concentration measured with Qubit™ fluorometer and Quant-iT™ dsDNA BR Assay Kit (Invitrogen, USA). The DNA samples were stored at −20 °C until sequencing preparation.
16 S rRNA gene sequencing
The 16S rRNA gene amplicons were prepared for the Illumina MiSeq system (Illumina Inc.) as described in Zamanzadeh et al.15. In brief, 16S rRNA gene PCR amplification was carried out using the Pro341F/Pro805R primer set selected from Takahsahi et al.47 modified with an Illumina adapter overhang in, and iProof HF DNA polymerase (BioRad, USA). A second PCR was carried out to attach unique 8-bp indices (Nextera XT Index Kit) to the Illumina sequencing adaptors to allow multiplexing of samples. A clean-up step (Agencourt AMPure XP beads, Beckman Coulter, USA) was preformed after each PCR. The amplicons were quantified (Quant-iT™ dsDNA HS Assay Kit and Qubit™ fluorometer, Invitrogen, USA), normalized and pooled to equimolar concentration, and then spiked with 30% PhiX control. A final concentration of 8 pm denaturated DNA was sequenced on an Illumina MiSeq instrument using the MiSeq reagent kit V3.
Sequencing analysis
All 16S rRNA gene sequences were processed using the QIIME version 1.8.0 software package48. Single-ends were trimmed to 200 bp and quality filtered as follows: only three sequential low-quality (Phred quality score <20) bases were allowed per sequence before truncating, and sequences with <75% (of total length) consecutive high-quality base calls were discarded. No N characters or barcodes were allowed in the sequence. Chimeric sequences were removed from the dataset using UCHIME incorporated in USEARH49 and a threshold of 3% dissimilarity between 16S rRNA gene sequences was used to cluster sequences into de novo operational taxonomic units (OTUs)49. Taxonomy (up to rank ‘genus’) was assigned to each OTU using the uclust-based consensus taxonomy assigner implemented in QIIME with default parameters. Alpha rarefaction plot of the phylogenetic diversity was generated using the script alpha_rarefaction.py with default parameters in QIIME. Low abundant OTUs (those with a total count less than 0.005%) and singletons were filtered out. Statistical analysis and visualization was carried out using Calypso version 8.2050,51. The alpha diversity of the microbial communities was measured by OTU richness in addition to the Shannon index. The diversity were further compared with ANOVA to evaluate the significance between the different subgroups (digestion vs. co-digestion, mesophilic vs. thermophilic). ANOVA was also used to compare taxa abundance across the different digesters. Finally, associations between reactor performance and the community composition were assessed. For this, a principle component analysis (PCA) was applied to examine how much of the variance in the 16S rRNA gene sequencing dataset could explain the process variables (day-specific concentration of VFA and NH3, and weekly measurements of CH4 production).
Data Availability
Sequence data are available at NCBI Sequence Read Archive under accession number SRP123045.
References
Aichinger, P. et al. Synergistic co-digestion of solid-organic-waste and municipal-sewage-sludge: 1 plus 1 equals more than 2 in terms of biogas production and solids reduction. Water Res. 87, 416–423 (2015).
Marañón, E. et al. Co-digestion of cattle manure with food waste and sludge to increase biogas production. Waste Manag. 32, 1821–1825 (2012).
Zhang, L., Lee, Y. W. & Jahng, D. Anaerobic co-digestion of food waste and piggery wastewater: Focusing on the role of trace elements. Bioresour. Technol. 102, 5048–5059 (2011).
Banks, C. J., Zhang, Y., Jiang, Y. & Heaven, S. Trace element requirements for stable food waste digestion at elevated ammonia concentrations. Bioresour. Technol. 104, 127–135 (2012).
Zhang, W., Zhang, L. & Li, A. Enhanced anaerobic digestion of food waste by trace metal elements supplementation and reduced metals dosage by green chelating agent [S, S]-EDDS via improving metals bioavailability. Water Res. 84, 266–277 (2015).
Edelmann, W., Engeli, H. & Gradenecker, M. Co-digestion of organic solid waste and sludge from sewage treatment. Water Science and Technology 41(3), 213–221 (2000).
Labatut, R. A., Angenent, L. T. & Scott, N. R. Conventional mesophilic vs. thermophilic anaerobic digestion: a trade-off between performance and stability? Water research 53, 249–258 (2014).
Ebner, J. H., Labatut, R. A., Lodge, J. S., Williamson, A. A. & Trabold, T. A. Anaerobic co-digestion of commercial food waste and dairy manure: Characterizing biochemical parameters and synergistic effects. Waste Management 52, 286–294 (2016).
Astals, S., Batstone, D., Mata-Alvarez, J. & Jensen, P. Identification of synergistic impacts during anaerobic co-digestion of organic wastes. Bioresource technology 169, 421–427 (2014).
Angelidaki, I. & Ahring, B. K. Thermophilic anaerobic digestion of livestock waste: the effect of ammonia. Applied Microbiology and Biotechnology 38(4), 560–564 (1993).
Karakashev, D., Batstone, D. J. & Angelidaki, I. Influence of environmental conditions on methanogenic compositions in anaerobic biogas reactors. Applied and Environmental Microbiology 71(1), 331–338 (2005).
Levén, L., Eriksson, A. R. B. & Schnürer, A. Effect of process temperature on bacterial and archaeal communities in two methanogenic bioreactors treating organic household waste. FEMS microbiology ecology 59(3), 683–693 (2007).
Fisgativa, H., Tremier, A. & Dabert, P. Characterizing the variability of food waste quality: a need for efficient valorisation through anaerobic digestion. Waste Management 50, 264–274 (2016).
El-Mashad, H. M. & Zhang, R. Biogas production from co-digestion of dairy manure and food waste. Bioresource technology 101(11), 4021–4028 (2010).
Zamanzadeh, M., Hagen, L. H., Svensson, K., Linjordet, R. & Horn, S. J. Anaerobic digestion of food waste-effect of recirculation and temperature on performance and microbiology. Water Research 96, 246–254 (2016).
Zhang, C., Xiao, G., Peng, L., Su, H. & Tan, T. The anaerobic co-digestion of food waste and cattle manure. Bioresource technology 129, 170–176 (2013).
Agyeman, F. O. & Tao, W. Anaerobic co-digestion of food waste and dairy manure: Effects of food waste particle size and organic loading rate. Journal of environmental management 133, 268–274 (2014).
Pagés-Díaz, J., Pereda-Reyes, I., Taherzadeh, M. J., Sárvári-Horváth, I. & Lundin, M. Anaerobic co-digestion of solid slaughterhouse wastes with agro-residues: synergistic and antagonistic interactions determined in batch digestion assays. Chemical Engineering Journal 245, 89–98 (2014).
Guo, C. L. et al. Effects of temperature and organic loading rate on the performance and microbial community of anaerobic co-digestion of waste activated sludge and food waste. Chemosphere 105(3), 146–151 (2014b).
Wang, P., Wang, H., Qiu, Y., Ren, L., & Jiang, B. Microbial characteristics in anaerobic digestion process of food waste for methane production–A review. Bioresource Technology, In Press (2017).
Limam, R. D. et al. Members of the uncultured bacterial candidate division WWE1 are implicated in anaerobic digestion of cellulose. MicrobiologyOpen 3(2), 157–167 (2014).
Sun, L., Pope, P. B., Eijsink, V. G. & Schnürer, A. Characterization of microbial community structure during continuous anaerobic digestion of straw and cow manure. Microbial biotechnology 8(5), 815–827 (2015).
Stolze, Y. et al. Identification and genome reconstruction of abundant distinct taxa in microbiomes from one thermophilic and three mesophilic production-scale biogas plants. Biotechnology for Biofuels 9(1), 156 (2016).
Yamada, T. et al. Anaerolineathermolimosa sp. nov., Levilineasaccharolytica gen. nov., sp. nov. and Leptolineatardivitalis gen. nov., sp. nov., novel filamentous anaerobes, and description of the new classes Anaerolineae classis nov.andCaldilineae classis nov.in the bacterial phylum Chloroflexi. Int. J. Syst. Evol. Microbiol. 56(6), 1331–1340 (2006).
Sekiguchi, Y. et al. Anaerolineathermophila gen. nov., sp. nov. and Caldilineaaerophila gen. nov., sp. nov., novel filamentous thermophiles that represent a previously uncultured lineage of the domain Bacteria at the subphylum level. Int. J. Syst. Evol. Microbiol. 53(6), 1843–1851 (2003).
Riviere, D. et al. Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. The ISME journal 3(6), 700–714 (2009).
Vanwonterghem, I., Jensen, P. D., Ho, D. P., Batstone, D. J. & Tyson, G. W. Linking microbial community structure, interactions and function in anaerobic digesters using new molecular techniques. Curr. Opin. Biotechnol. 27, 55–64 (2014).
Sundberg, C. et al. 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS microbiology ecology 85(3), 612–626 (2013).
Guo, X., Wang, C., Sun, F., Zhu, W. & Wu, W. A comparison of microbial characteristics between the thermophilic and mesophilic anaerobic digesters exposed to elevated food waste loadings. Bioresource technology 152, 420–428 (2014a).
Dürre, P. Physiology and sporulation in Clostridium. In The Bacterial Spore: from Molecules to Systems (pp. 315–329). American Society of Microbiology (2016).
Zinder, S. H. Physiological ecology of methanogens. In Methanogenesis (pp. 128–206). Springer US (1993).
Demirel, B. & Scherer, P. The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Reviews in Environmental Science and Bio/Technology 7(2), 173–190 (2008).
Karakashev, D., Batstone, D. J., Trably, E. & Angelidaki, I. Acetate oxidation is the dominant methanogenic pathway from acetate in the absence of Methanosaetaceae. Applied and environmental microbiology 72(7), 5138–5141 (2006).
Menes, R. J. & Muxí, L. Anaerobaculum mobile sp. nov., a novel anaerobic, moderately thermophilic, peptide-fermenting bacterium that uses crotonate as an electron acceptor, and emended description of the genus Anaerobaculum. International journal of systematic and evolutionary microbiology 52(1), 157–164 (2002).
Hagen, L. H. et al. Quantitative metaproteomics highlight the metabolic contributions of uncultured phylotypes in a thermophilic anaerobic digester. Applied and Environmental Microbiology 83(2), e01955–16 (2016).
Etchebehere, C., Pavan, M. E., Zorzopulos, J., Soubes, M. & Muxi, L. Coprothermobacter platensis sp. nov., a new anaerobic proteolytic thermophilic bacterium isolated from an anaerobic mesophilic sludge. International journal of systematic bacteriology 48(4), 1297–1304 (1998).
Hattori, S., Kamagata, Y., Hanada, S. & Shoun, H. Thermacetogenium phaeum gen. nov., sp. nov., a strictly anaerobic, thermophilic, syntrophic acetate-oxidizing bacterium. International Journal of Systematic and Evolutionary Microbiology 50(4), 1601–1609 (2000).
Li, Y. F. An integrated study on microbial community in anaerobic digestion systems (Doctoral dissertation, The Ohio State University) (2013).
Zhao, H., Yang, D., Woese, C. R. & Bryant, M. P. Assignment of fatty acid-β-oxidizing syntrophic bacteria to Syntrophomonadaceae fam. nov.on the basis of 16S rRNA sequence analyses. International journal of systematic bacteriology 43(2), 278–286 (1993).
Hippe, H., Andreesen, J. & Gottschalk, G. The genus Clostridium—nonmedical. The prokaryotes 2, 1800–1866 (1992).
Yokoyama, H., Wagner, I. D. & Wiegel, J. Caldicoprobacter oshimai gen. nov., sp. nov., an anaerobic, xylanolytic, extremely thermophilic bacterium isolated from sheep faeces, and proposal of Caldicoprobacteraceae fam. nov. International journal of systematic and evolutionary microbiology 60(1), 67–71 (2010).
De Vos, P. et al. Bergey’s Manual of Systematic Bacteriology Volume 3: The Firmicutes. 2nd Edition. Springer, New York (2009).
Conners, S. B. et al. Microbial biochemistry, physiology, and biotechnology of hyperthermophilic Thermotoga species. FEMS microbiology reviews 30(6), 872–905 (2006).
Pervin, H. M. et al. Drivers of microbial community composition in mesophilic and thermophilic temperature-phased anaerobic digestion pre-treatment reactors. Water research 47(19), 7098–7108 (2013).
Sasaki, K. et al. Syntrophic degradation of proteinaceous materials by the thermophilic strains Coprothermobacter proteolyticus and Methanothermobacter thermautotrophicus. Journal of Bioscience and Bioengineering 112(5), 469–472 (2011).
Estevez, M. M., Sapci, Z., Linjordet, R., Schnürer, A. & Morken, J. Semi-continuous anaerobic co-digestion of cow manure and steam-exploded Salix with recirculation of liquid digestate. J. Environ. Manage. 136, 9–15 (2014).
APHA, AWWA & WEF. Standard Methods for the Examination of Water and Wastewater, twentiethed. American Public Health Association, Washington, DC., USA (1998).
Takahashi, S., Tomita, J., Nishioka, K., Hisada, T. & Nishijima, M. Development of a Prokaryotic Universal Primer for Simultaneous Analysis of Bacteria and Archaea Using Next-Generation Sequencing. PloS one 9(8), e105592 (2014).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature methods 7(5), 335–336 (2010).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27(16), 2194–2200 (2011).
Zakrzewski, M. et al. Calypso: a user-friendly web-server for mining and visualizing microbiome–environment interactions. Bioinformatics 33(5), 782–783 (2016).
Acknowledgements
This work was financially supported by the ENERGIX-program of the Research Council of Norway, grant no. 228747 (BiogasFuel).
Author information
Authors and Affiliations
Contributions
R.L., S.J.H. and M.Z. participated in the design of the study. M.Z. and K.S. carried out the reactor digestion experiments. L.H.H. carried out the microbial analyses and the bioinformatics. M.Z. wrote the original manuscript. All authors read, revised and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zamanzadeh, M., Hagen, L.H., Svensson, K. et al. Biogas production from food waste via co-digestion and digestion- effects on performance and microbial ecology. Sci Rep 7, 17664 (2017). https://doi.org/10.1038/s41598-017-15784-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15784-w
This article is cited by
-
Impact of tumbling on production of biomethane from household waste
Environmental Science and Pollution Research (2024)
-
Research Progress on the Mechanism and Control Strategy of Anaerobic Digestion Imbalance of Food Waste
Waste and Biomass Valorization (2024)
-
Evaluation of biomethane potential of codigested sheep manure and kitchen refuse
Biomass Conversion and Biorefinery (2023)
-
Anaerobic co-digestion of kitchen waste and animal manure: a review of operating parameters, inhibiting factors, and pretreatment with their impact on process performance
Biomass Conversion and Biorefinery (2023)
-
Biotechnological interventions in food waste treatment for obtaining value-added compounds to combat pollution
Environmental Science and Pollution Research (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.