Abstract
Prion diseases are fatal neurodegenerative disorders with sporadic, genetic or acquired etiologies. The molecular alterations leading to the onset and the spreading of these diseases are still unknown. In a previous work we identified a five-gene signature able to distinguish intracranially BSE-infected macaques from healthy ones, with SERPINA3 showing the most prominent dysregulation. We analyzed 128 suitable frontal cortex samples, from prion-affected patients (variant Creutzfeldt-Jakob disease (vCJD) n = 20, iatrogenic CJD (iCJD) n = 11, sporadic CJD (sCJD) n = 23, familial CJD (gCJD) n = 17, fatal familial insomnia (FFI) n = 9, Gerstmann–Sträussler–Scheinker syndrome (GSS)) n = 4), patients with Alzheimer disease (AD, n = 14) and age-matched controls (n = 30). Real Time-quantitative PCR was performed for SERPINA3 transcript, and ACTB, RPL19, GAPDH and B2M were used as reference genes. We report SERPINA3 to be strongly up-regulated in the brain of all human prion diseases, with only a mild up-regulation in AD. We show that this striking up-regulation, both at the mRNA and at the protein level, is present in all types of human prion diseases analyzed, although to a different extent for each specific disorder. Our data suggest that SERPINA3 may be involved in the pathogenesis and the progression of prion diseases, representing a valid tool for distinguishing different forms of these disorders in humans.
Similar content being viewed by others
Introduction
Prion diseases are rare, fatal and rapidly progressive neurodegenerative disorders. In humans these diseases may be sporadic, genetic or acquired1. Acquired forms in humans include variant Creutzfeldt-Jakob disease (vCJD) – caused by exposure to bovine spongiform encephalopathy (BSE) infected material – whose incidence has been progressively declining due to preventive measures2. However, the recent report of the first vCJD patient heterozygous at codon 129 in the prion protein gene (PRNP) creates some concerns about a second wave of the disease in the next few years3. In addition, three major studies about prion accumulation in lymphoid tissue conducted in a large series of appendix specimens carried out to date all show the prevalence of abnormal prion protein in the UK population to be around 1 in 2,000–1 in 5,000 using immunohistochemical (IHC) testing, suggesting a high prevalence of asymptomatic carriers4,5,6. Although safety procedures were put in place in the past decades to prevent the iatrogenic transmission of CJD, new cases of iCJD continue to be identified with incubation periods of several decades7. Moreover, probable iatrogenic transmission of vCJD through blood transfusion has been reported for four cases, from 2003 until 2009, posing additional public health concerns8,9,10,11. However, sporadic CJD (sCJD) still accounts for the majority of prion disease cases, which overall affect 1–2 persons per million of population worldwide7. The pathological agent responsible for these diseases is an aberrant misfolded isoform of the cellular prion protein (PrPC) denoted as prion or PrPSc 12. The latter represents the only specific biomarker for the disease, which is readily detectable in post-mortem central nervous system (CNS) tissues by Western blotting after limited proteolytic treatment13. Other neuropathological alterations are a prominent reactive astrocytic gliosis, involving activation of astrocytes, assessed by detection of increased levels of glial fibrillary acidic protein (GFAP), vacuolation (also known as spongiform change) and neuronal loss in grey matter. Altogether, the type of abnormal deposition of PrPSc, astrocytic gliosis, and vacuolation in specific gray matter regions of the CNS are defining traits of different prion disorders13.
The prion protein is encoded by the PRNP gene located on chromosome 20. The gene contains 2 exons, with the entire coding region being contained in the last exon, thus excluding possible alternative splicing12. Human PrPC is a protein of 253 amino acids, prior to post-translational modifications, and in its mature form is a 208 amino acid polypeptide, which is glycosylphosphatidylinositol-anchored to the outer leaflet of the cellular membrane with a unique primary sequence14. Misfolding of PrPC into the pathogenic PrPSc occurs in different prion diseases, in which different forms of the protease-resistant core of PrPSc (sometimes referred as PrPres) can be identified. Different types of human prion diseases are characterized by diverse PrPSc isoforms and by the polymorphism present at PRNP codon 129. The two alleles of the gene can code for either methionine or valine in this position and homozygosity or heterozygosity can influence the pathological phenotypes in prion disorders. In addition, over 50 mutations in the PRNP open reading frame have been identified in genetic forms of prion diseases, in which the clinical and pathological features may also be influenced by the codon 129 polymorphism on the mutated and non-mutated alleles1.
In a recent analysis of the transcriptomic landscape in a model of prion infection using non-human primates, we identified a set of genes that are specifically dysregulated in the frontal cortex in the late stages of the disease15. In the present work we sought to validate whether up-regulated gene transcripts could be also dysregulated in human specimens of prion disease. We focused on SERPINA3, a gene encoding for a member of the serine protease inhibitor family of acute phase proteins16. Although serpins are predominantly produced in the liver, they are also synthesized in the brain, mainly by astrocytes and they have been reported to be involved in neuroinflammation and neurodegeneration16. Another member of the serpin superfamily named neuroserpin is predominantly expressed in neurons and its main function is to inhibit the enzyme tissue-type plasminogen activator. Point mutations in the neuroserpin protein determine a change in its conformation that leads to its abnormal cytoplasmic accumulation in neuroserpinopathies17. Concerning SERPINA3, it is a 56–66 kDa protein susceptible of heavy glycosylation at multiple sites18. Interestingly, it is the only serpin able to bind DNA, probably through lysine repeat motifs in regions 210–212 and 391–39619. Its dysregulation has been already linked to a number of different diseases, such as chronic obstructive pulmonary disease20, cystic fibrosis21, pancreas22, prostate23, lung24 and invasive breast cancer25 and Alzheimer disease (AD)26,27,28,29.
BSE infection in humans results in the acquired prion disease vCJD which has also been modelled experimentally in primates. However, given the limited numbers of reported vCJD patients (231 worldwide, including three from Italy), we decided to extend our analysis to sCJD patients. This would also allow us to detect differences in gene regulation between acquired and sporadic human prion disorders. We also included in our analysis cases of iCJD caused by treatment with prion-contaminated human growth hormone (hGH). Moreover, we included patients with genetic forms of prion disease resulting from different PRNP mutations, in particular FFI (with the D178N mutation), GSS (two with the P102L mutation, one with the P84S mutation and one with the Q212P mutation) and gCJD (11 with the E200K mutation, one with the E211Q mutation, one with the V210I mutation and three cases with 4 or 5 octarepeat region insertions). Regarding control samples, we used age- sex- and size-matched control groups for each disease analyzed. In order to enable the identification of prion-specific gene expression alterations, we decided to introduce in our study also some samples from patients with non-CJD neurodegenerative disorder (AD) as an additional control group. We collected over 200 samples of frontal cortex of post-mortem specimens from various forms of prion diseases, AD and age-matched controls, of which only 128 resulted of suitable quality for transcriptomic analysis. The different samples included vCJD (n = 20), sCJD (n = 23), iatrogenic CJD (iCJD, n = 11), familial CJD (gCJD, n = 17), fatal familial insomnia (FFI, n = 9), Gerstmann–Sträussler–Scheinker (GSS) syndrome (n = 4), AD (n = 14) and age-matched controls (n = 30).
Real Time-quantitative PCR (RT-qPCR) was performed for the SERPINA3 transcript, and four different housekeeping genes (ACTB, RPL19, GAPDH and B2M) were chosen as reference genes. We found strong up-regulation in the gene expression of SERPINA3 in all prion diseases considered. Limited up-regulation for this transcript was also observed in AD samples, although not comparable to that of prion-affected samples. In addition, we investigated GFAP and PRNP genes expression levels in all patients considered. As for GFAP, we found a similar dysregulation pattern as for SERPINA3 among all groups, up-regulation in all prion diseases together with a negligible increase in AD samples. Concerning PRNP transcriptional levels, after accurate analysis to rule out the influence of contaminating blood, a decrease in both iCJD and AD samples was observed. Nevertheless, these changes were limited when compared to SERPINA3 transcript. Immunohistochemistry and biochemical analyses for SERPINA3 protein were also performed, confirming the specific and differential up-regulation of the protein in sporadic, genetic or acquired forms of prion disease.
Results
qPCR and WB analysis
Two common issues with postmortem human brain samples are the stability of endogenous reference genes and the RNA integrity30. A number of different variables correlates to different extents with RNA integrity: for example, higher degradation is observed in samples subjected to prolonged thawing31 or storage32 as well as in samples of patients showing longer agony before death (probably due to brain acidosis)33. In studies involving postmortem human tissue is not possible to control all the variables that might impair RNA quality, and this inevitably leads to non-homogenous sample collections with high degree of biological variance. However, it is widely accepted that moderate degradation does not preclude reliable analyses of small amount of RNA. Indeed, it has been shown that gene expression profiles from partially degraded RNA samples with still visible ribosomal bands are highly similar to that of intact samples32, especially if the RT-qPCR amplified products are smaller than 250 bp34. This is not particularly surprising considering that reference genes transcripts most likely degrade gradually together with target ones30. For all these reasons, a RIN ≥ 4 was suggested as minimal threshold for human brain tissue35. In order to ensure a sufficient reliability of the samples, among the 225 samples collected, only 126 samples harbor RNA that showed RIN ≥ 4 or higher, and with the exception of few rare samples, were selected for further analysis, Regarding housekeeping genes, there is a lack of consensus about the optimal reference genes in postmortem brain tissues. Indeed, while GAPDH is the most widely used reference gene for transcriptomic studies, some issues have been reported about its involvement in human neurodegenerative processes36. However, ACTB, B2M and the ribosomal protein family seem to show good stability across different RIN values33 and brain samples36. Therefore, we decided to use GAPDH, B2M, ACTB and RPL19 as reference genes to normalize RT-qPCR data, in order to increase the robustness of our findings. The relative mRNA levels, expressed as the fold change (FC), were calculated using the 2−∆∆C T method and four reference genes, obtaining very similar results. Firstly, the four housekeeping genes were analyzed across diseased and control patients in order to assess their expression stability (Fig. 1S).
SERPINA3 was found to be strongly up-regulated in all prion diseases: sCJD (FC = 40.3), vCJD (FC = 38.8), FFI (FC = 52.7), gCJD (FC = 39.1) and GSS (FC = 12.17, not significant) cases (Table 1). This up-regulation is particularly pronounced in iCJD patients (FC = 347) while, in comparison, only a modest up-regulation was observed in AD cases (FC = 6.7) (Fig. 1). Regarding GSS, we have to stress that the lack of significance in this group is likely due to the sample size (n = 4) which is too small to allow reliable statistical analysis. Regarding AD patients, we have to mention that the mean age of this group was higher (68.7 years) than that of the oldest control group available in this study (63.3 years). In addition, we observed that SERPINA3 mRNA levels were increasing with age in non-neurodegenerative controls (CTRL group with mean 51yrs vs CTRL group with mean 29yrs FC = 3, CTRL group with mean 64yrs vs CTRL group with mean 29yrs FC = 7). Therefore, the limited up-regulation we detected in AD samples might be overestimated due to the lack of appropriate age-matched controls.
RT-qPCR analysis for SERPINA3 mRNA expression in brain normalized against GAPDH. gCJD (n = 17) *****p < 0.000000005, FFI (n = 9) **p < 0.0001, vCJD (n = 20) ***p < 0.000001, iCJD (n = 11) ****p < 0.00000005, sCJD (n = 23) ******p < 0.0000000005, AD (n = 14) *p < 0.005). Fold Change (FC) values are presented as average with CI.
Very similar results were obtained against all four reference genes (Figs 2S, 3S and 4S). As regards healthy controls, SERPINA3 mRNA was present at very low levels, in some cases barely detectable, with an average Ct of 31.2. The levels of SERPINA3 mRNA in controls was then used as zero value to normalize all the others groups.
Dealing with prion disease in which astrogliosis is a prominent feature, we assessed also the mRNA levels of GFAP in all our samples. As for SERPINA3, GFAP was up-regulated in all prion diseases, with the highest value observed in gCJD patients (FC = 5.2) and iCJD (FC = 5), sCJD (FC = 3), GSS (FC = 2.9, not significant) vCJD (FC = 2.8) and FFI (FC = 2.2). No changes were observed in AD samples (Fig. 2).
In addition, we investigated the gene expression pattern of PRNP. Also for this gene, the greatest change was observed in the iCJD group, where PRNP mRNA is strongly down-regulated. Similarly, sCJD patients displayed a significant but moderate down-regulation (Fig. 3).
Finally, we analyzed all the samples for expression of an erythrocyte specific marker, ALAS2, to evaluate blood contamination in the various samples. RT-qPCR analysis of ALAS2 revealed a small blood presence (CT < 35) in the majority of samples (Fig. 5S). However, when investigating the possible presence of SERPINA3, GFAP and PRNP mRNA in the blood, only a very low signal for PRNP was detected, confirming that the blood contamination did not influence our analysis (Fig. 6S).
After normalization against β-actin levels, Western blot analysis revealed the presence of higher level of SERPINA3 in all CJD brains (regardless the disease aetiology) if compared to that of AD and healthy controls. Interestingly, lower molecular weight bands – likely representing poorly glycosylated isoforms- are visible only in prion-infected samples. The presence of PrPSc was confirmed in all CJD brains (Fig. 4). According to densitometric analysis, the expression levels of SERPINA3 in CJD samples were six-fold higher than those of controls (Fig. 7S).
Western blotting analysis of brain samples from CJD, AD patients and healthy controls. The expression levels of SERPINA3 and PrPSc in brain samples of representative cases from each group were assessed by means of Western blotting, on two different gels run in parallel within the same experiment. β-actin was used as proteins loading control and to normalize the expression level of SERPINA3 for densitometric analysis. SERPINA3 and β-actin were developed on the same membrane, sequentially. SERPINA3 image has been cropped to improve clarity. Rec lane refers to recombinant SERPINA3 used as control. Molecular weight is represented on the left (kDa). Original full-length images are provided in Supplementary Material.
As for other tissues outside the brain, we also investigated the presence of SERPINA3 protein in the CSF where it is expressed, but with no significant differences found between AD and sCJD samples (Fig. 8S).
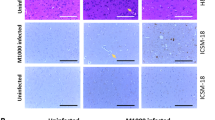
As for human brain samples, also RML-infected CD1 mice displayed a strong up-regulation of serpina3n at symptomatic stage, both at the mRNA and protein level (Figs 5, 6 and 9S). Even more importantly, this up-regulation was observed at the preclinical stage, suggesting the direct correlation between serpina3n and prion infection.
Western blotting analysis for serpina3n in the brain of RML-infected CD1 mice and related age-matched controls. The expression levels of serpina3n and PrPSc were assessed by means of Western blotting, on two different gels run in parallel within the same experiment. β-actin was used as proteins loading control and to normalize the expression level of serpina3n for densitometric analysis. serpina3n and β-actin were developed on the same membrane, sequentially. serpina3n image has been cropped to improve clarity. Molecular weights are shown on the left. Original full-length images are provided in Supplementary Material.
Neuropathological analysis
Immunohistochemical analysis revealed that in the frontal cortex of control subjects, immunostaining for SERPINA3 was present in isolated neurons and not in other cell types (Fig. 7A). On the contrary, in CJD brain samples, either sporadic, iatrogenic or variant, most of the residual cortical neurons and numerous reactive astrocytes were intensely immunostained by anti-serpina3 antibody. In neuronal cells, the immunodecoration was most evident in the perikarya and in the apical dendrites of pyramidal neurons (Fig. 7C–F). PrP focal deposits (perivacuolar, perineuronal or plaque-like) were not immunoreactive. A consistent number of SERPINA3-immunoreactive cortical neurons was found in AD samples where the amyloid of senile plaques and amyloid angiopathy (not shown) was also immunopositive.
Neuropathological analysis. Immunohistochemical analysis revealed strong SERPINA3 immunoreaction within residual neurons and reactive glial cells in the frontal cortex of CJD patients [variant (E,F), sporadic (D) and iatrogenic (C)] while it was scanty and limited to isolated neurons in control subjects (A). In AD patients, anti-SERPINA3 decorated some cortical neurons and the amyloid of senile plaques (B).
We also investigated the correlation between SERPINA3 and GFAP immunoreactivity. While a strong parallel between the two proteins was not observed, there is a clear diffuse and strong immunoreactivity in vCJD and iCJD samples, while in sCJD and in AD slices the reactivity was more confined to astrocytes and less pronounced (Fig. 10S)
Discussion
In this work we provide evidence for strong up-regulation of SERPINA3 mRNA in the CNS of patients deceased from different forms of prion diseases. A total of 128 samples were analysed from the frontal cortex of vCJD, sCJD, iCJD, gCJD, and FFI and GSS syndrome cases; as controls, AD and neurologically normal age-matched cases were used. Among the different prion diseases, there was striking up-regulation of SERPINA3 mRNA in iCJD specimens, up to about 350 fold. Nevertheless, other forms of prion disease show significant up-regulation of the SERPINA3 transcript, ranging from 30 to 40 folds in all groups considered. This up-regulation at the mRNA level is paralleled by enhanced SERPINA3 protein expression in astrocytes as well as in neurons. The discrepancy between mRNA and protein levels, concerning either the magnitude of up-regulation and the differences among the different groups, may depend on the half-life of the protein. While mRNA is continuously produced, the protein may be degraded quite readily, thus resulting the divergence seen between the mRNA and Western blotting results. In addition, we have to consider the different sensitivity of RT-qPCR and WB techniques, the latter being much lower in sensitivity, which could also help explain the discrepancies between mRNA and protein levels.
In addition, in an animal model of prion infection (CD1 mice intracranially inoculated with RML prions) the up-regulation of serpina3n and its protein product serpina3n follows the detection of prions as early as at the presymptomatic stage. Intriguingly, our data on mice inoculated with RML prions clearly show that the increase of serpina3n, particularly at the protein level but also its transcript, is concomitant with the appearance of PrPSc. All inoculated animals were sacrificed at 97 dpi; all animals with PrPSc detectable in the brain by Western blot also showed evidence of serpina3n mRNA upregulation. However, the single animal that did not display detectable brain PrPSc had no evidence of serpina3n upregulation, with a mRNA level comparable to that one of the age-matched controls (Fig. 5). These data strongly suggest a direct correlation between serpina3n and prion disease progression, as previously reported also in RML-infected C57BL/6 mice37. In a previous work, serpina3n mRNA, together with GFAP mRNA, was found up-regulated in the brain of Chandler-infected SWR/j mice, since week 15 post inoculation and progressively increased to the terminal stages of the disease38. Similar results were observed also in C506M3 scrapie-infected and, with a minor extent, in BSE infected C57BL/6 mice39.
The name SERPINA3, also known as alpha 1-antichymotrypsin, comes from serine protease inhibitor class A member 3 and it belongs to the serine protease inhibitor superfamily, which comprises several members in mammals40. The mature protein comprises about 398 amino acids after proteolytic removal of a 25 amino acid signal sequence. The mature protein has a molecular weight of about 56–66 kDa due to heavy glycosylation18. The glycosylation is characteristic for circulating plasma serpins but it is dispensable for their activity18. The protein inhibits serine proteases and several targets have been described, including cathepsins, chymases and elastases. The mechanism by which SERPINA3 inhibits its targets is through the binding to the protease and consequent irreversible conformational change in its structure41. SERPINA3 has been linked to neurodegeneration and in particular to AD pathology, since protein deposits have been found in association with Aβ-peptides in both non-congophilic amorphous and classical (pre)amyloid plaques42,43,44,45 but also in astrocytes, in few neurons and in neurofibrillary tangles46. More recently, transgenic mouse models overexpressing astroglial serpina3n together with mutant amyloid protein precursor (APP), showed an increased age-related amyloid plaque deposition if compared to mice expressing APP alone. The mechanism through which this occurs it has not been yet clarified, however it is suggested that serpina3n may promote the aggregation of Aβ and its deposition into the brain parenchyma or that it could interfere with the degradation and removal of Aβ aggregates47,48. Despite this evidence, controversial findings have not clarified the involvement of SERPINA3 in AD pathology and its role during the course of the disease16. Little is known about SERPINA3 and prion diseases. One report investigated the link between a polymorphism in the signal peptide sequence of SERPINA3 and incidence of CJD, but neither allele frequencies nor genotype distribution were found altered in CJD cases if compared to controls, and also the age at onset and disease duration were unaffected by the SERPINA3 genotype49. More recently, SERPINA3 has been found up-regulated in the occipital cortex of sCJD patients and the protein as well was found increased in urines37. How SERPINA3 is up-regulated in prion diseases remains to be established. Several intriguing hypotheses about the role of SERPINA3 in the prion replication and accumulation can be put forward. For example, SERPINA3 strong up-regulation can readily block target serine proteases and, in turn, the clearance of prions could be hampered, over time. Another captivating possibility is that increased expression of SERPINA3 could serve as a molecular chaperone which contributes to the conversion process of prion formation50. This hypothesis is supported by the finding of the dysregulation of PRNP gene, which is down-regulated in nearly all prion diseases analyzed, and again with particular relevance in iCJD samples. The exacerbation observed for SERPINA3 mRNA in iCJD samples may be explained by the younger age of these patients (29 years on average) if compared to the other acquired from of CJD studied (vCJD, 35 years on average) who might display a stronger reactivity to the infection. However, in order to exclude any age-effect, we normalized each disease group against a specific age-matched control group. In addition, we should take into consideration that iCJD patients have been repeatedly exposed to prion-contaminated hGH over time since early childhood, through intramuscular injections which represent an invasive and highly efficient transmission route. Moreover, we also observed a great difference between the different populations within the same disease. Indeed, the up-regulation of Italian-French vCJD cases was much higher (FC = 365) than the British one (FC = 27), suggesting a potential influence of different prion strain infectivity and/or hidden susceptibility factors (such as polymorphisms) that may differentially affect the dysregulation of SERPINA3.
Similarly, within the iatrogenic group, the up-regulation observed in French cases (FC = 702) was much higher when compared to that one of the British samples (FC = 248), reinforcing the latter hypothesis.
Concerning the correlation between SERPINA3 levels and PrPSc pathology, we performed IHC analyses on GFAP expression, which is a well-established marker of prion disorders. Even though we did not observe a perfect correlation, we still detected a similar trend between GFAP reactivity and SERPINA3 one. This, together with the fact that we detected an up-regulation of both gfap and serpina3n mRNA also at the preclinical stage in mice, strongly point toward a correlation between SERPINA3 and PrPSc pathology.
It is likely that the up-regulation of SERPINA3 is not exactly the same in all the different brain regions, indeed it is probably more prominent in regions were astrogliosis occurs, given its predominantly astrocytic expression. However, the up-regulation is widely spread across the different brain areas, given that both RNA and protein levels were strongly up-regulated also in mice were we analyzed whole brain tissue, both for RT-qPCR and WB.
As for other tissues outside the brain, we also investigated the presence of SERPINA3 protein in the CSF where it is well expressed but with no difference between AD and sCJD samples. One reason for this lack of difference between the two diseases might depend on a possible impairment of the blood-brain barrier in AD cases, being these patients much older and the disease having a longer course, which could lead to a higher presence of SERPINA3 in the CSF of AD patients.
Previous studies have already detected a down-regulation of the PRNP mRNA in the frontal cortex of MM1 sCJD brains, as well as the total PrP protein content. In contrast to the present study, reduced levels of PRNP mRNA have also been detected in AD patients51. One possible explanation for this discrepancy might reside in the different stages of the disease considered, which were early (Braak stage I-II) in our study and advanced (Braak stage V-VI) in their work. Even though a direct interaction between the two proteins is still to be clarified, we can envision a mechanism in which the up-regulation of a protease-inhibitors such as SERPINA3 might lead, in turn, to an inhibition of proteases that could facilitate prion conversion, replication and accumulation.
PRNP mRNA down-regulation could be the consequence of an increased content of total PrP due to PrPSc accumulation, which in turn up-regulates the production of protease-inhibitors such as SERPINA3 that would further sustain the prion accumulation and propagation. Interestingly, SERPINA3 – when present in a poorly glycosylated state - is also able to retrotranslocate in the nucleus52 and to regulate DNA synthesis by promoting chromatin condensation19. Therefore we could also hypothesize that the down-regulation of PRNP transcript might be a direct consequence of SERPINA3 localization within the nucleus. Besides this, it is known that also PrPSc can retrotranslocate from the cytoplasm to the nucleus where it concentrates in euchromatin areas53. We could then suggest that upon prions accumulation, PrPSc translocates in the nucleus where it could activate SERPINA3 transcription and subsequent SERPINA3 protein up-regulation, ultimately leading to decreased protease activity and consequent enhanced prion conversion and/or propagation. No effective treatment or cure is available for prion diseases54, and this lack of therapeutic approaches is further complicated by the heterogeneity and complexity of clinical presentation among different cases. With initial symptoms that span from personality changes, anxiety, depression, insomnia and impairment of autonomic functions, which worsen during the rapid course of the disease, it is difficult to determine a clinical diagnosis with certainty. Very recently, protein misfolding cyclic amplification (PMCA) technique has been successfully applied to human blood samples for the detection of minute amounts of vCJD prions. However, this approach still needs to be validated for its ability to monitor the disease at preclinical stages; indeed, only few samples from preclinical vCJD cases were available for these studies55,56. In addition, cerebrospinal fluid (CSF) and olfactory mucosa real-time quaking-induced conversion (RT-QuIC) has been successfully employed for detection of prions in the clinical phase of sCJD, providing virtually 100% diagnostic sensitivity and specificity57. In our work, we identified a potential additional biomarker highly specific for prion diseases, either sporadic, genetic of acquired. This specificity is further corroborated by GFAP mRNA expression pattern, which mirrors the one of SERPINA3. Indeed, GFAP is a well-known marker of astrogliosis, which is a hallmark of prion diseases: the fact that we find both GFAP and SERPINA3 -either mRNA and protein- strongly up-regulated in the brain tissue of patients with prion disease, reinforce the hypothesis of a strong correlation between prions and SERPINA3 and support our proposal of using SERPINA3 as a potential additional biomarker for human prion diseases. In blood SERPINA3 mRNA is absent (Fig. 6S), therefore this tissue is not suitable for diagnostic purposes. Additional studies in other readily accessible tissues such as CSF and urine may be needed to address the diagnostic potential of SERPINA3 mRNA detection in both preclinical and clinical stages of human prion diseases.
Conclusions
In light of the most recent epidemiologic data reported in the Appendix III study, it is plausible that we may face a new wave of prion cases in the near future, highlighting the importance of further investigation on additional genes modulating human prion diseases.
The elevated and specific up-regulation of SERPINA3 mRNA in human brain samples of prion diseases offers the possibility of using this transcript as a new tool to help distinguish different forms of these disorders. The ready detection of SERPINA3 mRNA and the corresponding protein product serpina3 in CNS samples and potentially in non-CNS samples may allow us to discriminate, in different stages of the disease, various forms of prion diseases.
Understanding the molecular mechanism for SERPINA3 up-regulation in prion diseases may shed light to the cellular events that regulates prion conversion, replication and accumulation.
Methods
Ethics statement
All samples were collected according to the ethical and safety regulations of the Countries in which samples were collected as follows.
Human biological samples and associated data were obtained from the Austrian CJD Surveillance and KIN-Biobank Medical University of Vienna (Ethics Committe approval: 396/2011). All tissue samples were obtained according to Austrian legislations. The human tissue examined in this study was provided by the MRC Edinburgh Brain Bank and its use was covered by ethical approval from the East of Scotland Research Ethics Service REC 1 (reference number 16/ES/0084). Informed consent for the research use of autopsy tissue was obtained from the relatives of the deceased whenever necessary.
Human brain tissue was obtained from the Institute of Neuropathology Brain Bank (HUB-ICO-IDIBELL Biobank, Barcelona, Spain) following the guidelines of Spanish legislation (Real Decreto 1716/2011) and the approval of the local ethics committee.
For the French cases, written informed consent for autopsy and research use was provided by patient’s relatives, according to the French regulation (L.1232–1 to L.1232-3, Code de la Santé Publique). The brain tissues with the corresponding written informed consent were referred for postmortem diagnosis and research to the French National Neuropathological Network for CJD (funded by the French Government) and to the French National Centre of Reference for prions (funded by the French Institute for Public Health Surveillance). PRNP gene was analyzed pre-mortem in all cases with the informed and signed consent of the patient’s family.
This study was approved by the institutional review board of Carlo Besta Neurological Institute and performed according to the guidelines approved by the ethics committee. Written informed consent for participation in research was done in accordance with the Declaration of Helsinki (1964-2008) and the Additional Protocol on the Convention of Human Rights and Biomedicine concerning Biomedical Research (2005).
Brain tissue samples were obtained at University hospital of Verona from patients with a clinical diagnosis of prion disorders for neuropathological studies. Written informed consent was obtained for the sample included in this study.
All experiments in mice were performed in accordance with European regulations [European community Council Directive, November 24, 1986 (86/609/EEC)] and were approved by the local authority veterinary service.
Mouse samples
Age- and sex-matched CD1 mice (Crl:CD1(ICR) strain code 022, charles river) were used for prion infection experiments. Rocky Mountain Laboratory (RML) prion infected brain homogenate was prepared at a 10% w/v in phosphate-buffered saline (PBS). 2.5 μl of 10% brain homogenate was stereotactically injected in the hippocampus of 2-months old outbred CD1 mice (n = 8). Not inoculated age- and sex-matched CD1 mice were used as controls (n = 8). Presymptomatic animals (n = 4) were sacrificed 97 days post inoculation, while terminal stage animals (n = 4) were monitored daily for clinical signs of prion disease and sacrificed at the end stage. Adult animals were sacrificed with CO2, brains were extracted, immediately frozen in liquid nitrogen and stored at −80 °C. Brains of prion-infected and age-matched controls were either homogenized for WB analysis or subjected to RNA extraction.
Patient samples
A total of 226 frontal cortex tissue samples from neurodegeneration-affected patients and healthy control subjects were collected. The samples included in our analysis (128) came from patients with sCJD (type 1 n = 15, type 2 n = 8), vCJD (n = 20), iCJD (n = 11), genetic prion diseases (FFI n = 9, gCJD n = 17, GSS n = 4), patients first diagnosed with AD as non-prion neurodegenerative surrogate controls (n = 14), and aged-matched non-neurodegenerative control cases (n = 30). All the prion disease cases have been confirmed by PrPSc detection with Western Blot or PRNP mutations. AD diagnosis was performed based on Braak scoring and neuropathologically confirmed. As for the controls, they died from unrelated conditions and all of them were all lacking any neurological sign. The full list of samples and patient details is reported in Table 1S.
Blood samples (n = 4) were obtained by Fondazione I.R.C.C.S Istituto Neurologico Carlo Besta.
Tissues and RNA extraction
Total RNA was isolated from about 100 mg of frontal cortex from frozen postmortem human brain tissue or from one hemisphere of mouse whole brain tissue. The tissue was homogenized with Stainless Steel Beads 5 mm in Tissue Lyser II (Qiagen), in TRIzol reagent (Invitrogen) following manufacturer’s instructions. RNA was extracted with PureLink RNA Mini Kit (Life Technologies) and on-column DNA digestion was performed using PureLink DNase Set (Life Technologies). Blood samples were collected in PAXgene Blood RNA Tubes (Qiagen) and the related RNA was extracted using PAXgene Blood RNA Kit (Qiagen). RNA was checked for concentration and purity on a NanoDrop 2000 spectrophotometer (Thermo Scientific) whereas RNA integrity was analyzed using 2100 Bioanalyzer (Agilent Technologies). Samples included in our gene expression analysis displayed RNA Integrity Number (RIN) ≥ 4, even if some RNA samples below this threshold were selected in order to increase the sample size of very rare prion diseases patients groups.
Reverse transcription and Real Time-quantitative PCR (RT-qPCR)
For human frontal cortex and mouse whole brain, cDNA was obtained starting from 3 µg of total RNA with 50 µM Oligo(dT)20, 10 mM dNTP mix, 5X First Strand Buffer, 0.1 M DTT, 40 U RNAse inhibitor and 200 U SuperScript® III Reverse Transcriptase (Life Technologies). cDNA from blood samples was obtained using 750 ng RNA, maintaining the same reaction conditions as above. For each sample a negative control was performed by omission of the reverse transcriptase (-RT control). qPCR primers were designed using the online tool Primer-Blast provided by NCBI and, possibly, spanning an exon-exon junction or flanking intron sequence/s, to prevent the amplification of genomic DNA.
The primers sequences for human targets were as follows: for GAPDH 5′-CCTGCACCACCAACTGCTTA-3′ and 5′-TCTTCTGGGTGGCAGTGATG-3′; for ACTB 5′-AGAGCTACGAGCTGCCTGAC-3′ and 5′-AGCACTGTGTTGGCGTACAG-3′; for RPL19 5′-CTAGTGTCCTCCGCTGTGG-3′ and 5′-AAGGTGTTTTTCCGGCATC-3′; B2M for 5′-AGATGAGTATGCCTGCCGTG-3′and 5’-TCATCCAATCCAAATGCGGC-3′; for SERPINA3 5′ TGCCAGCGCACTCTTCATC 3′ and 5′ TGTCGTTCAGGTTATAGTCCCTC 3′; for ALAS2 5′-TGTCCGTCTGGTGTAGTAATGA-3′ and 5′- GCTCAAGCTCCACATGAAACT-3′.
The primers sequences for mouse targets were as follows: Gapdh 5′-TTCACCACCATGGAGAAGGC-3′ and 5′-GGCATGGACTGTGGTCATGA-3′; for Actb 5′-CACACCCGCCACCAGTTC-3′ and 5′-CCCATTCCCACCATCACACC-3′; for Tubb3 5′-CGCCTTTGGACACCTATTC-3′ and 5′-TACTCCTCACGCACCTTG-3′; for Serpina3n 5′-ACCCTGAGGAAGTGGAAGAA-3′ and 5′-CCTGATGCCCAGCTTTGAAA-3′.
Gene expression assays were performed using iQTM SYBR® Green Supermix 2x (Bio-Rad Laboratories, Inc.), 400 nM final concentration of the corresponding forward and reverse primer (Sigma) and 10 ng/μl final concentration of cDNA samples. CFX96 TouchTM Real-Time PCR Detection System (Bio-Rad Laboratories, Inc.) was used to perform the amplification and the cycling conditions included an initial denaturation of 3 min at 95 °C then 45 cycles at 95 °C for 10 sec and 60 °C for 1 min. qPCR reaction was performed in technical triplicates for each primer pair and sample. -RT sample for each primer pairs was analyzed to evaluate possible genomic RNA contamination and non-template control (NTC) was added in each plate to exclude presence of contaminating DNA in the qPCR reaction mix. Melting curve analysis of amplicons was performed for each primer pair to verify that artificial products or primer dimers were not responsible for the obtained fluorescence signals.
RT-qPCR data analysis
Differential expression of human target genes was normalized to four reference genes (GAPDH, ACTB, RPL19 and B2M) expression. For mouse samples, Gapdh, Actb and Tubb3 were used as reference genes. The absolute expression value (CT) of a specific human erythrocyte marker, ALAS2, was used to account for blood contamination of brain samples. The relative expression ratio (fold change, FC) was calculated using 2−∆∆CT method58. ∆CT were calculated subtracting the CT of the housekeeping gene to the CT of the target one, both for “test” (disease affected patient) and “calibrator” (control). Then, ∆∆CT was calculated subtracting the average ∆CT of age-matched control groups to the average ∆CT of each disease groups. To select age-matched control for each prion diseases and AD group, the same sample size of non-neurodegenerative controls was chosen to match both median and average age of disease affected patient groups. Matching the sex was not possible due to a higher prevalence of males in the control group. However, it has been reported that no relevant sex variability for Serpina3 exists in different disease and tissues59,60, therefore we are reasonably confident that this sex mismatch would not bias our results. Fold change values smaller than 1 were converted using the equation -1/fold change, for representation.
Statistical analysis
Test-F was used to verify the homoscedasticity of variance between ∆CTs of disease-affected and the related age-matched control group (α = 0.05). Then, level of significance was calculated using unpaired student t-test (2 tails, p < 0.05) between ∆CTs of disease and control groups.
Immunohistochemical analysis
Brains were fixed in 10% formalin, dehydrated and embedded in para-plast. Seven-micrometer thick serial sections were stained with hematoxylin-eosin (H&E, Bioptica) or immunostained with polyclonal antibodies to serpina3 (Sigma), monoclonal antibodies to PrP (3F4, Dako) and polyclonal antibodies to Glial Fibrillary Acidic Protein (GFAP, Dako). Before PrP immunostaining formalin-fixed samples were treated with autoclave (120 °C, 10 min), 80% formic acid (20 min) and guanidine isothiocyanate (4 M, 1 hour). Before serpina3 immunostaining all sections were treated with 80% formic acid (20 min) at room temperature. Samples were finally incubated with the Envision Plus/Horseradish Peroxidase System for rabbit and mouse immunoglobulins (DakoCytomation). Immunoreactions were visualized using 3-3′-diaminobenzidine (DAB, Dako) as chromogen.
Production and purification of recombinant human SERPINA3
Plasmid pET-11° containing the sequence coding for HuSerA3 (GenScript) was transformed into Escherichia coli RosettaGami cells for the expression. Cells were grown in LB medium at 37 °C in presence of ampicillin (100 µg/mL) until OD = 600 nm, followed by induction with 0,5 mM IPTG overnight. Cells were collected by centrifugation and resuspended into Cell Lysis Buffer (25 mM Tris, 0,8% Triton X-100). Cells were disrupted using PANDA Homogenizer, and inclusion bodies containing the desired protein were subsequently isolated by centrifugation. Inclusion bodies were subjected to a series of washing steps, and then solubilized overnight at 37 °C in 8 M GdnHCl, 25 mM Tris. The purification process takes advantage of the His tag added at the C-terminal of the protein. The inclusion bodies were filtered and then applied onto a HisTrap column (GE Healthcare). The unspecific proteins were washed away with 5 column volumes of Binding Buffer, and then HuSerA3 was eluted with Elution Buffer, containing 500 mM imidazole. HuSerA3 fractions with higher purity were pooled together and refolded with serial dialysis steps to reduce the amount of denaturing agent. The protein was then dialysed against PBS and stored at −20 °C.
Immunoblot analysis
Frontal cortex of patients with sporadic (sCJD), genetic (gCJD, E200K), iatrogenic CJD (iCJD), Alzheimer’s disease (AD) and healthy controls were homogenized in lysis buffer (NaCl 100 mM, EDTA 10 mM, NP40 0.5%, Deoxycholic acid 0.5%, TRIS 10 mM pH 7.4) at 10% weight/volume. Samples were centrifuged (Eppendorf Centrifuge) for 1 minute at 4 °C (800 × g) to remove cellular debris. Two-hundred µg of proteins were loaded onto 12% Bis/Tris Acrylamide gels (NuPAGE, Invitrogen) and separated by SDS-Page, transferred to PVDF membrane and incubated with polyclonal anti-serpin3 rabbit antibody (1:500 Sigma-Aldrich), 3F4 antibody (1:10,000 Dako) and β-actin (1:10,000 Sigma-Aldrich). The reactions were visualized by chemiluminescence using Amersham ECL Prime (GE Healthcare Life Sciences). For PrP analysis, samples were digested with 50 µg/mL of proteinase-K (Life Technologies) at 37 °C for 1 hour. Densitometric analysis was carried out using UVIBand software.
For mouse samples the same procedure as above was followed. Detection of PrPSc was performed using W226 monoclonal antibody (1:1000) and a goat anti-mouse HRP secondary antibody. Actin beta was detected using monoclonal anti-β-actin−peroxidase antibody (1:10,000 Sigma), SerpinA3n detection was performed using Mouse Serpin A3N Antibody (1:500 R&D Systems) and a rabbit anti-goat HRP secondary antibody.
Data availability
All data generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Compliance with Ethical Standards
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. All human samples were anonymized. (for further details see Materials and Methods section).
All procedures performed in studies involving animals were in accordance with the ethical standards of the institution or practice at which the studies were conducted. Current animal husbandry and housing practices comply with the Council of Europe Convention ETS123 (European Convention for the Protection of Vertebrate Animals used for Experimental and Other Scientific Purposes; Strasbourg, 18.03.1986); Italian Legislative Decree 26/2014, Gazzetta Ufficiale della Repubblica Italiana, 26 July 2014; and with the 86/609/EEC (Council Directive of 24 November 1986 on the approximation of laws, regulations and administrative provisions of the Member States regarding the protection of animals used for experimental and other scientific purposes). Animal facility is licensed and inspected by the Italian Ministry of Health. Mice were housed in groups of 2–5 animals in individually ventilated cages, daily fed and water provided ad libitum. Lighting was on an automatic 12 hours basis. All surgery was performed under tribromoethanol anesthesia, and all efforts were made to minimize suffering and regular veterinary care was daily performed for assessment of animal health. The study, including its Ethics aspects, was approved by the Italian Ministry of Health (Permit Number: NP-02-14).
Informed consent
Informed written consent was obtained for all individual participants included in the study. (for details see Materials and Methods section).
References
Prusiner, S. B. Biology and genetics of prions causing neurodegeneration. Annu Rev Genet 47, 601–623, https://doi.org/10.1146/annurev-genet-110711-155524 (2013).
Collee, J. G., Bradley, R. & Liberski, P. P. Variant CJD (vCJD) and bovine spongiform encephalopathy (BSE): 10 and 20 years on: part 2. Folia Neuropathol 44, 102–110 (2006). doi:6212 [pii].
Mok, T. et al. Variant Creutzfeldt-Jakob Disease in a Patient with Heterozygosity at PRNP Codon 129. N Engl J Med 376, 292–294, https://doi.org/10.1056/NEJMc1610003 (2017).
Gill, O. N. et al. Prevalent abnormal prion protein in human appendixes after bovine spongiform encephalopathy epizootic: large scale survey. BMJ 347, f5675, https://doi.org/10.1136/bmj.f5675 (2013).
Advisory Committee on Dangerous Pathogens TSE Risk Assessment Subgroup (August 2016). “Appendix-III” position statement. Available from: ACDP TSE subgroup minutes, agendas and papers, https://app.box.com/s/hhhhg857fjpu2bnxhv6e (2016).
Hilton, D. A. et al. Prevalence of lymphoreticular prion protein accumulation in UK tissue samples. J Pathol 203, 733–739, https://doi.org/10.1002/path.1580 (2004).
Kim, M. O. & Geschwind, M. D. Clinical update of Jakob-Creutzfeldt disease. Current opinion in neurology 28, 302–310, https://doi.org/10.1097/WCO.0000000000000197 (2015).
Llewelyn, C. A. et al. Possible transmission of variant Creutzfeldt-Jakob disease by blood transfusion. Lancet 363, 417–421, https://doi.org/10.1016/S0140-6736(04)15486-X (2004).
Peden, A. H., Head, M. W., Ritchie, D. L., Bell, J. E. & Ironside, J. W. Preclinical vCJD after blood transfusion in a PRNP codon 129 heterozygous patient. Lancet 364, 527–529, https://doi.org/10.1016/S0140-6736(04)16811-6 (2004).
Wroe, S. J. et al. Clinical presentation and pre-mortem diagnosis of variant Creutzfeldt-Jakob disease associated with blood transfusion: a case report. Lancet 368, 2061–2067, https://doi.org/10.1016/S0140-6736(06)69835-8 (2006).
Peden, A. et al. Variant CJD infection in the spleen of a neurologically asymptomatic UK adult patient with haemophilia. Haemophilia 16, 296–304, https://doi.org/10.1111/j.1365-2516.2009.02181.x (2010).
Colby, D. W. & Prusiner, S. B. Prions. Cold Spring Harb Perspect Biol 3, a006833, doi:3/1/a006833 [pii] 10.1101/cshperspect.a006833 (2011).
DeArmond, S. J. & Bouzamondo, E. Fundamentals of prion biology and diseases. Toxicology 181–182, 9–16, doi:S0300483X02002494 [pii] (2002).
Gasperini, L. & Legname, G. Prion protein and aging. Front Cell Dev Biol 2, 44, https://doi.org/10.3389/fcell.2014.00044 (2014).
Barbisin, M. et al. Gene expression profiling of brains from bovine spongiform encephalopathy (BSE)-infected cynomolgus macaques. BMC Genomics 15, 434, https://doi.org/10.1186/1471-2164-15-434 (2014).
Baker, C., Belbin, O., Kalsheker, N. & Morgan, K. SERPINA3 (aka alpha-1-antichymotrypsin). Front Biosci 12, 2821–2835 (2007).
Miranda, E. & Lomas, D. A. Neuroserpin: a serpin to think about. Cell Mol Life Sci 63, 709–722, https://doi.org/10.1007/s00018-005-5077-4 (2006).
Hwang, S. R., Steineckert, B., Kohn, A., Palkovits, M. & Hook, V. Y. Molecular studies define the primary structure of alpha1-antichymotrypsin (ACT) protease inhibitor in Alzheimer’s disease brains. Comparison of act in hippocampus and liver. J Biol Chem 274, 1821–1827 (1999).
Naidoo, N., Cooperman, B. S., Wang, Z. M., Liu, X. Z. & Rubin, H. Identification of lysines within alpha 1-antichymotrypsin important for DNA binding. An unusual combination of DNA-binding elements. J Biol Chem 270, 14548–14555 (1995).
Elliott, P. R., Pei, X. Y., Dafforn, T. R. & Lomas, D. A. Topography of a 2.0 A structure of alpha1-antitrypsin reveals targets for rational drug design to prevent conformational disease. Protein Sci 9, 1274–1281, https://doi.org/10.1110/ps.9.7.1274 (2000).
Mahadeva, R. et al. Association of alpha(1)-antichymotrypsin deficiency with milder lung disease in patients with cystic fibrosis. Thorax 56, 53–58 (2001).
Koomen, J. M. et al. Plasma protein profiling for diagnosis of pancreatic cancer reveals the presence of host response proteins. Clin Cancer Res 11, 1110–1118 (2005).
Leinonen, J., Lovgren, T., Vornanen, T. & Stenman, U. H. Double-label time-resolved immunofluorometric assay of prostate-specific antigen and of its complex with alpha 1-antichymotrypsin. Clin Chem 39, 2098–2103 (1993).
Higashiyama, M. et al. Alpha-1-antichymotrypsin expression in lung adenocarcinoma and its possible association with tumor progression. Cancer 76, 1368–1376 (1995).
Yamamura, J. et al. mRNA expression level of estrogen-inducible gene, alpha 1-antichymotrypsin, is a predictor of early tumor recurrence in patients with invasive breast cancers. Cancer Sci 95, 887–892 (2004).
Abraham, C. R., Selkoe, D. J. & Potter, H. Immunochemical identification of the serine protease inhibitor alpha 1-antichymotrypsin in the brain amyloid deposits of Alzheimer’s disease. Cell 52, 487–501 (1988).
Fraser, P. E., Nguyen, J. T., McLachlan, D. R., Abraham, C. R. & Kirschner, D. A. Alpha 1-antichymotrypsin binding to Alzheimer A beta peptides is sequence specific and induces fibril disaggregation in vitro. J Neurochem 61, 298–305 (1993).
Janciauskiene, S., Rubin, H., Lukacs, C. M. & Wright, H. T. Alzheimer’s peptide Abeta1-42 binds to two beta-sheets of alpha1-antichymotrypsin and transforms it from inhibitor to substrate. J Biol Chem 273, 28360–28364 (1998).
Sun, Y. X., Wright, H. T. & Janciauskiene, S. Alpha1-antichymotrypsin/Alzheimer’s peptide Abeta(1-42) complex perturbs lipid metabolism and activates transcription factors PPARgamma and NFkappaB in human neuroblastoma (Kelly) cells. J Neurosci Res 67, 511–522, https://doi.org/10.1002/jnr.10144 (2002).
Wang, Q. et al. Stability of endogenous reference genes in postmortem human brains for normalization of quantitative real-time PCR data: comprehensive evaluation using geNorm, NormFinder, and BestKeeper. Int J Legal Med 126, 943–952, https://doi.org/10.1007/s00414-012-0774-7 (2012).
Stan, A. D. et al. Human postmortem tissue: what quality markers matter? Brain research 1123, 1–11, https://doi.org/10.1016/j.brainres.2006.09.025 (2006).
Schoor, O. et al. Moderate degradation does not preclude microarray analysis of small amounts of RNA. Biotechniques 35(1192–1196), 1198–1201 (2003).
Koppelkamm, A., Vennemann, B., Lutz-Bonengel, S., Fracasso, T. & Vennemann, M. RNA integrity in post-mortem samples: influencing parameters and implications on RT-qPCR assays. Int J Legal Med 125, 573–580, https://doi.org/10.1007/s00414-011-0578-1 (2011).
Fleige, S. & Pfaffl, M. W. RNA integrity and the effect on the real-time qRT-PCR performance. Mol Aspects Med 27, 126–139, https://doi.org/10.1016/j.mam.2005.12.003 (2006).
Weis, S. et al. Quality control for microarray analysis of human brain samples: The impact of postmortem factors, RNA characteristics, and histopathology. J Neurosci Methods 165, 198–209, https://doi.org/10.1016/j.jneumeth.2007.06.001 (2007).
Penna, I. et al. Selection of candidate housekeeping genes for normalization in human postmortem brain samples. Int J Mol Sci 12, 5461–5470, https://doi.org/10.3390/ijms12095461 (2011).
Miele, G. et al. Urinary alpha1-antichymotrypsin: a biomarker of prion infection. PLoS One 3, e3870, https://doi.org/10.1371/journal.pone.0003870 (2008).
Campbell, I. L., Eddleston, M., Kemper, P., Oldstone, M. B. & Hobbs, M. V. Activation of cerebral cytokine gene expression and its correlation with onset of reactive astrocyte and acute-phase response gene expression in scrapie. J Virol 68, 2383–2387 (1994).
Dandoy-Dron, F. et al. Enhanced levels of scrapie responsive gene mRNA in BSE-infected mouse brain. Brain Res Mol Brain Res 76, 173–179 (2000).
Silverman, G. A. et al. The serpins are an expanding superfamily of structurally similar but functionally diverse proteins. Evolution, mechanism of inhibition, novel functions, and a revised nomenclature. J Biol Chem 276, 33293–33296, https://doi.org/10.1074/jbc.R100016200 (2001).
Huntington, J. A., Read, R. J. & Carrell, R. W. Structure of a serpin-protease complex shows inhibition by deformation. Nature 407, 923–926, https://doi.org/10.1038/35038119 (2000).
Abraham, C. R., Shirahama, T. & Potter, H. Alpha 1-antichymotrypsin is associated solely with amyloid deposits containing the beta-protein. Amyloid and cell localization of alpha 1-antichymotrypsin. Neurobiol Aging 11, 123–129 (1990).
Abraham, C. R., Selkoe, D. J., Potter, H., Price, D. L. & Cork, L. C. Alpha 1-antichymotrypsin is present together with the beta-protein in monkey brain amyloid deposits. Neuroscience 32, 715–720 (1989).
Rozemuller, J. M. et al. Distribution pattern and functional state of alpha 1-antichymotrypsin in plaques and vascular amyloid in Alzheimer’s disease. A immunohistochemical study with monoclonal antibodies against native and inactivated alpha 1-antichymotrypsin. Acta Neuropathol 82, 200–207 (1991).
Verga, L. et al. Alzheimer patients and Down patients: cerebral preamyloid deposits differ ultrastructurally and histochemically from the amyloid of senile plaques. Neurosci Lett 105, 294–299 (1989).
Gollin, P. A., Kalaria, R. N., Eikelenboom, P., Rozemuller, A. & Perry, G. Alpha 1-antitrypsin and alpha 1-antichymotrypsin are in the lesions of Alzheimer’s disease. Neuroreport 3, 201–203 (1992).
Mucke, L. et al. Astroglial expression of human alpha(1)-antichymotrypsin enhances alzheimer-like pathology in amyloid protein precursor transgenic mice. Am J Pathol 157, 2003–2010 (2000).
Nilsson, L. N. et al. Alpha-1-antichymotrypsin promotes beta-sheet amyloid plaque deposition in a transgenic mouse model of Alzheimer’s disease. J Neurosci 21, 1444–1451 (2001).
Salvatore, M. et al. Alpha1 antichymotrypsin signal peptide polymorphism in sporadic Creutzfeldt-Jakob disease. Neurosci Lett 227, 140-142, doi:S030439409700308X [pii] (1997).
Zsila, F. Inhibition of heat- and chemical-induced aggregation of various proteins reveals chaperone-like activity of the acute-phase component and serine protease inhibitor human alpha(1)-antitrypsin. Biochem Biophys Res Commun 393, 242–247, https://doi.org/10.1016/j.bbrc.2010.01.110 (2010).
Llorens, F. et al. PrP mRNA and protein expression in brain and PrP(c) in CSF in Creutzfeldt-Jakob disease MM1 and VV2. Prion 7, 383–393, https://doi.org/10.4161/pri.26416 (2013).
Santamaria, M. et al. Nuclear alpha1-antichymotrypsin promotes chromatin condensation and inhibits proliferation of human hepatocellular carcinoma cells. Gastroenterology 144, 818–828 e814, https://doi.org/10.1053/j.gastro.2012.12.029 (2013).
Mange, A., Crozet, C., Lehmann, S. & Beranger, F. Scrapie-like prion protein is translocated to the nuclei of infected cells independently of proteasome inhibition and interacts with chromatin. J Cell Sci 117, 2411–2416, https://doi.org/10.1242/jcs.01094 (2004).
Bolognesi, M. L. & Legname, G. Approaches for discovering anti-prion compounds: lessons learned and challenges ahead. Expert opinion on drug discovery 10, 389–397, https://doi.org/10.1517/17460441.2015.1016498 (2015).
Concha-Marambio, L. et al. Detection of prions in blood from patients with variant Creutzfeldt-Jakob disease. Sci Transl Med 8, 370ra183, https://doi.org/10.1126/scitranslmed.aaf6188 (2016).
Bougard, D. et al. Detection of prions in the plasma of presymptomatic and symptomatic patients with variant Creutzfeldt-Jakob disease. Sci Transl Med 8, 370ra182, https://doi.org/10.1126/scitranslmed.aag1257 (2016).
Bongianni, M. et al. Diagnosis of Human Prion Disease Using Real-Time Quaking-Induced Conversion Testing of Olfactory Mucosa and Cerebrospinal Fluid Samples. JAMA Neurol 74, 155–162, https://doi.org/10.1001/jamaneurol.2016.4614 (2017).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408, https://doi.org/10.1006/meth.2001.1262 (2001).
Dimberg, J. et al. Expression of the serine protease inhibitor serpinA3 in human colorectal adenocarcinomas. Oncol Lett 2, 413–418, https://doi.org/10.3892/ol.2011.280 (2011).
Luo, D. et al. Serpin peptidase inhibitor, clade A member 3 (SERPINA3), is overexpressed in glioma and associated with poor prognosis in glioma patients. Onco Targets Ther 10, 2173–2181, https://doi.org/10.2147/OTT.S133022 (2017).
Acknowledgements
The authors wish to thank Helena Krmac for technical support. This work was also supported by a grant from the EC Joint Programme on Neurodegenerative Diseases (JPND-REfrAME) to GL.
Author information
Authors and Affiliations
Contributions
S.V. and G.L. conceived and designed the study. S.V., F.M., M.Z., E.B., E.D.C., M.R., G.G., F.T., S.H., J.P.D., G.Z., J.W.I., I.F., G.G.K. and G.L. contributed to the acquisition of samples and analysis of data. S.V., F.M., M.Z., E.B., M.R., G.G., G.L. contributed in drafting a significant portion of the manuscript or figures. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vanni, S., Moda, F., Zattoni, M. et al. Differential overexpression of SERPINA3 in human prion diseases. Sci Rep 7, 15637 (2017). https://doi.org/10.1038/s41598-017-15778-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-15778-8
This article is cited by
-
Integrated analysis of robust sex-biased gene signatures in human brain
Biology of Sex Differences (2023)
-
Gene expression and epigenetic markers of prion diseases
Cell and Tissue Research (2023)
-
Explaining a series of models by propagating Shapley values
Nature Communications (2022)
-
Novel Exosome Biomarker Candidates for Alzheimer’s Disease Unravelled Through Mass Spectrometry Analysis
Molecular Neurobiology (2022)
-
Serpin Signatures in Prion and Alzheimer’s Diseases
Molecular Neurobiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.